Early definitions of personalised nutrition were gene focused; however, in recent times, the definition has been extended and now incorporates the concept of levels( Reference Ronteltap, van Trijp and Berezowska 1 ). This reworked definition of personalised nutrition now includes the following: level 1 personalised advice, which involves delivering personalised advice based on dietary intake; level 2 personalised advice, which involves personalised advice based on diet and phenotypic markers such as blood markers and BMI; and level 3 personalised advice, which builds on the previous levels and includes diet, phenotype and genotype information( Reference Gibney and Walsh 2 ). Although such definitions focus on personalised advice delivered at an individual level, there is an emerging concept that has gained momentum in recent years, where dietary advice can be tailored to specific groups of individuals and is referred to as targeted nutrition( Reference Brennan 3 – Reference O’Donovan, Walsh and Gibney 5 ).
These groups of individuals have similar characteristics and are referred to as metabotypes( Reference Morris, O’Grada and Ryan 6 ). There are numerous examples of metabotyping in the medical literature where it has been utilised to sub-group patients with diseases with differential symptomology( Reference Erro, Vitale and Amboni 7 – Reference Castaldi, Dy and Ross 10 ). For example, several studies have used cluster analysis to identify sub-groups of patients with characteristic phenotypes of asthma, a disease that is very heterogeneous in nature( Reference Haldar, Pavord and Shaw 11 – Reference Park, Song and Kim 14 ). Metabotyping has also been used to identify groups of individuals with differing responses to drug treatments( Reference Clayton, Lindon and Cloarec 15 – Reference van Bochove, van Schalkwijk and Parnell 17 ) and dietary interventions( Reference O’Sullivan, Gibney and Connor 18 – Reference Vazquez-Fresno, Llorach and Perera 20 ).
However, although there are many examples of identifying groups of similar individuals in the population( Reference Erro, Vitale and Amboni 7 – Reference Viniol, Jegan and Hirsch 9 , Reference Botelho, Fioratti and Abdalla 21 ), the evidence base for developing tailored health solutions for these groups is weak. Previous work from our group demonstrated a framework for the delivery of targeted nutrition advice to metabolically similar groups or metabotypes in the population( Reference O’Donovan, Walsh and Nugent 22 ). In this study, three distinctly different metabotypes were identified on the basis of four routinely measured markers of metabolic health including blood TAG, total cholesterol, HDL-cholesterol and glucose (n 896). Using a decision tree approach, targeted dietary advice messages were developed on the basis of the characteristics of each cluster. Good agreement was observed between the targeted dietary advice method and an individualised method without the need for collection of detailed dietary data( Reference O’Donovan, Walsh and Nugent 22 ). Overall, this previous work demonstrated the potential of the metabotyping approach to deliver appropriate tailored dietary advice at a group level with minimal data collection required.
In the current study, this concept is further advanced using data from the Food4Me study, a personalised nutrition intervention study( Reference Celis-Morales, Livingstone and Marsaux 23 ). In Food4Me, participants received personalised advice based on the three levels of personalisation, delivered by trained nutritionists, and thus provides a valuable resource for testing the targeted nutrition approach( Reference Celis-Morales, Livingstone and Marsaux 23 ). Therefore, the objectives of this study were to: (1) identify metabotypes in a European population group and (2) develop targeted dietary advice solutions for these metabotypes and compare them with personalised dietary advice given within the Food4Me study.
Methods
Study design and ethical approval
As part of the Food4Me project (CinicalTrials.gov no.: NCT01530139, https://clinicaltrials.gov/ct2/show/NCT01530139), a proof-of-principle study was conducted, which compared the effectiveness of personalised nutrition advice, based on the three levels of personalisation, on health-related outcomes, compared with generic healthy eating advice. This was an internet-based study, designed to emulate a personalised nutrition service, and was conducted in seven research centres across Europe (Ireland, UK, the Netherlands, Spain, Germany, Poland, Greece). Ethical approval was obtained from the Research Ethics Committees at each university or research centre. This study was conducted according to the guidelines laid down in the Declaration of Helsinki, and all procedures involving human subjects were approved by the Research Ethics Committees at each university or research centre. Participants (>18 years) (n 1607) were randomised into one of four groups: control group received general European-based healthy eating guidelines; level 1 participants received personalised advice based on their dietary intake; level 2 participants received personalised advice based on their diet and phenotype; and level 3 participants received advice on their diet, phenotype and genotype. More details on the overall study design and the main outcomes of the personalised nutrition randomised controlled trial can be found elsewhere( Reference Celis-Morales, Livingstone and Marsaux 23 , Reference Celis-Morales, Livingstone and Marsaux 24 ). Written informed consent was obtained from all subjects.
Data collection and personalised feedback
All data were self-collected by participants using detailed instructions provided by researchers and online video demonstrations. A more detailed description of the data collection methods is reported elsewhere( Reference Celis-Morales, Livingstone and Marsaux 23 ). In brief, habitual dietary intake was assessed using the online Food4Me FFQ, which was previously developed and validated for the purposes of the study( Reference Forster, Fallaize and Gallagher 25 , Reference Fallaize, Forster and Macready 26 ). The foods included in the FFQ were aggregated to form thirty-two food groups. The list of the foods contributing to each of the food groups is found in the online Supplementary Table S1. Participants were provided with a measuring tape to perform anthropometric measures including weight (kg), height (m) and circumferences including waist (cm), hip (cm) and thigh (cm); all were collected according to standard previously published protocols( Reference Celis-Morales, Livingstone and Marsaux 23 ). A validation study was conducted to assess the accuracy of these measurements and strong correlation coefficients (0·983 for BMI and 0·993 for weight) were observed between the self-reported measurements and measurements performed face-to-face by researchers( Reference Celis-Morales, Livingstone and Woolhead 27 ).
Metabolic markers were measured by finger-prick blood samples collected by participants using a collection pack provided by Vitas Ltd. Participants were asked to fast 8 h before collection in the morning and to fill two dry blood spot (DBS) cards (five drops of blood or 150 µl of blood per card). Once filled, cards were left to dry for 2–4 h at room temperature and placed in an airtight aluminium bag with a drying sachet and returned by post to their corresponding recruiting centre. The samples were then sent via courier service to Vitas, where the following metabolic markers were measured: total cholesterol, carotenoids (lutein, zeaxanthin, β-cryptoxanthin, α-carotene, β-carotene, lycopene) and twenty fatty acids, as shown in Table 1. The metabolic markers were measured using the following methods: cholesterol (LC-UV), carotenoids (HPLC-diode array detector-MS/MS) and fatty acids (GC-flame ionisation detector).
Table 1 Clustering variables and other metabolitesFootnote * (Mean values and standard deviations)

1,2,3 Superscripts indicate which cluster is significantly different using Bonferroni post hoc tests for pairwise comparison between clusters.
* Subjects were clustered based on the above variables. One-way ANOVA was used to examine the differences across the clusters.
† Highest values across the clusters.
‡ Lowest values across the clusters.
§ Total saturated fat was calculated as the sum of myristic acid (C14 : 0), palmitic acid (C16 : 0) and stearic acid (C18 : 0).
|| Omega-3 Index was calculated by the following formula: Omega-3 Index=1·4473+0·8303×(EPA+DPA+DHA).
¶ Total carotenoids was calculated by the following formula: total carotenoids=α-carotene+β-carotene+lutein+zeaxanthin+β-cryptoxanthin+lycopene.
Participants randomised to levels 1, 2 and 3 received personalised reports based on decision trees to allow for the delivery of systematic tailored advice. The personalised reports were sent via email at months 0, 3 and 6. Standard operating procedures were developed for use of the decision trees and these were standardised across the seven recruitment centres to ensure consistency in the personalised advice given across all centres. Those individuals in level 1 received feedback based on their current dietary intake and physical activity levels. Level 2 participants received feedback based on their current diet, physical activity levels and phenotypic measures such as anthropometry and metabolic markers (total cholesterol, carotenoids including lutein, zeaxanthin, β-cryptoxanthin, α-carotene, β-carotene, lycopene and twenty fatty acids, as shown in Table 1). Level 3 participants received the same feedback as level 2 with the addition of genotypic information (information on FTO/FADS1/TCF7L2/ApoE(e4)/MTHFR). The final section of the report contained a personalised goals section where participants were given three nutrient-related goals. The personalised goals were selected by a predefined ranking system, where those nutrients and metabolic markers that most warranted change were prioritised. Participants were asked to focus on making changes to these three nutrients in the personalised reports in line with the patient-centred counselling models for facilitating behaviour change( Reference Rosal, Ebbeling and Lofgren 28 ).
Development and testing of targeted dietary advice
Targeted dietary advice was developed for each cluster based on the characteristics of the cluster and using a decision tree process. Two decision trees were developed per cluster based on the following: (1) metabolic markers and anthropometric information and (2) dietary information. This resulted in forty-nine messages for cluster 1, twenty messages for cluster 2 and twenty-four messages for cluster 3. As there are no defined cut-offs for total saturated fat (%) from DBS data, cluster 1 was described as high saturated fat, cluster 2 as low saturated fat and cluster 3 as medium saturated fat based on the mean values across the clusters, as shown in Table 1.
The appropriateness of the targeted dietary advice developed per cluster was then tested by comparison with the three nutrient-related goals, which were delivered to all level 2 participants (n 180) by trained nutritionists, as part of their personalised feedback reports. The development of these personalised feedback reports has been published elsewhere( Reference Forster, Walsh and O’Donovan 29 ). The agreement between the two methods was assessed based on the following questions:
-
(1) How many of the nutrient-related goals given as part of the personalised advice reports within the Food4Me study were given as part of the targeted dietary advice derived from this study?
-
(2) How many dietary messages were given as part of the targeted dietary advice in comparison with the personalised advice within Food4Me? (That is number of messages given as per the targeted dietary advice.)
Statistics
Baseline data were analysed using SPSS software package version 20.0 for Windows (SPSS, Inc.). In all, twenty-seven metabolic markers including cholesterol and individual carotenoids and fatty acids from the DBS analysis were chosen for clustering, as presented in Table 1. A full data set was available for 1354 participants (see online Supplementary Table S3). Following standardisation using z-scores, two-step cluster analysis revealed the presence of three clusters, and k-means cluster analysis was then used to characterise the clusters. The differences between the clusters were assessed using one-way ANOVA with Bonferroni post hoc tests. χ 2 distributions were used to assess categorical variables across the clusters including sex and country. As age, sex, BMI and country were significantly different across the clusters, these variables were controlled for in the general linear models with Bonferroni post hoc tests. P values were also adjusted for multiple comparisons using the Bonferroni approach.
Results
Characterisation of the clusters
Three clusters were identified in the Food4Me population (Table 1). Cluster 1 (n 326) was the group with the highest cholesterol, highest circulating trans-fatty acids (0·85 (sd 0·25) %) and lowest Omega-3 Index (5·16 (sd 0·93) %). Cluster 2 (n 433) was the most metabolically healthy group as they had the highest average Omega-3 Index (6·56 (sd 1·29) %), highest total carotenoid concentrations (2·15 (sd 0·71) µm) and lowest total saturated fat. Cluster 3 subjects (n 595) had the lowest average cholesterol concentrations (4·25 (sd 0·78) mm) and highest stearic acid (Table 1). Age was significantly different across the groups, with clusters 1 and 2 being older on average (Table 2). BMI and waist circumference were also significantly different across the clusters. Cluster 1 had the highest BMI of 27·7 (sd 5·3) kg/m2 and waist circumference of 0·93 (sd 0·14 m, whereas participants in cluster 2 had the lowest BMI and waist circumference (Table 2). With the exception of the Netherlands and the UK, the distribution of nationality differed significantly across the clusters.
Table 2 Demographical information across the clustersFootnote * (Mean values and standard deviations; percentages and numbers)
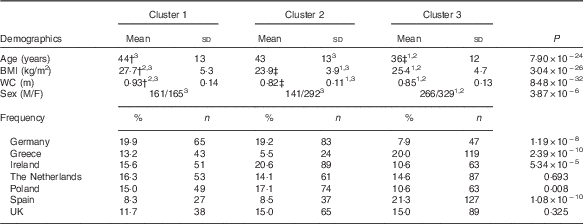
WC, waist circumference; M, male; F, female.
1,2,3 Superscripts indicate which cluster is significantly different using Bonferroni post hoc tests for pairwise comparison between clusters.
* One-way ANOVA was used to examine the differences across the clusters with exception of sex and country where χ 2 was used instead.
† Highest values across the clusters.
‡ Lowest values across the clusters.
Reported dietary intakes across the clusters are presented in Table 3. There were no differences in total energy intake and macronutrients across the clusters. However, percentage energy contribution from alcohol and PUFA was found to be significantly different (P=0·048). Furthermore, intakes of many micronutrients differed significantly across the clusters, including fat-soluble vitamins A, D and E, as well as some water-soluble vitamins such as folate, and vitamin C. Participants in cluster 1 had the higher percentage contribution of energy from alcohol (4·2 (sd 4·5) %) compared with individuals in clusters 2 and 3. The diets of cluster 2 participants were considered to be healthier as these individuals had the highest intakes of dietary fibre (32 (sd 15) g), fat-soluble vitamins D and E, folate and vitamin C.
Table 3 Dietary intakes across the clustersFootnote * (Mean values and standard deviations)

1,2,3 Superscripts indicate which cluster is significantly different using Bonferroni post hoc tests for pairwise comparison between clusters.
* P values provided by general linear models controlling for age, sex, BMI and country where appropriate. Bonferroni post hoc tests were used to examine pairwise comparisons between groups with the exception of vitamin D, where Fisher’s least significant difference post hoc tests were used instead.
† General linear models were calculated on log-transformed values where necessary and adjusted for multiple comparisons.
‡ Highest values across the clusters.
§ Lowest values across the clusters.
|| Adjusted for energy (kJ). Dietary intakes were examined across the three clusters.
Intakes of the food groups savouries (P=1·27×10−4), fruit (P=1·39×10−8), fish, fish dishes and products (P=8·16×10−4) differed significantly between the clusters, as illustrated in Table 4. Similar to their nutrient intakes, participants in cluster 2 had the healthiest food intakes with the lowest intakes of savouries (11 (sd 13) g) and white bread/rolls/scones/croissants (34 (sd 73) g) and highest intakes of yogurt (91 (sd 107) g), fruit (355 (sd 306) g), fish, fish dishes and products (71 (sd 53) g). Clusters also differed in terms of supplement users (P=9·31×10−8), with the highest percentage found in cluster 2 (54·3 %), in which the individuals also had the highest Omega-3 Index.
Table 4 Food group intakes across the clustersFootnote * (Mean values and standard deviations)

1,2,3 Superscripts indicate which cluster is significantly different using Bonferroni post hoc tests for pairwise comparison between clusters.
* P values provided by general linear models controlling for age, sex, BMI and country where appropriate. Frequency of supplement users was assessed using χ 2 analysis.
† General linear models were calculated on logged values where necessary and adjusted for multiple comparisons.
‡ Lowest values across the clusters.
§ Highest values across the clusters.
Development of the targeted dietary advice
Targeted dietary advice was developed on the basis of the characteristics (anthropometric, metabolic and nutrient intake data) of each cluster using a decision tree method (see Fig. 1). Two decision trees were constructed per cluster: a combined metabolic and anthropometric decision tree and a dietary decision tree. Ranges of the metabolic markers and nutrients were calculated for each of the clusters and these values were then used to determine whether individuals in each cluster were within the desirable or high/low range for that particular variable, as shown in Table 5. The cut-offs used in the current study were based on those used within the Food4Me study (online Supplementary Table S2), but were simplified for the purposes of the development of the targeted dietary advice. For the targeted dietary advice, the cut-offs were set as either ‘desirable’ or ‘high/low’(Table 5), whereas in Food4Me the cut-offs were developed using a more complex gradation scale (online Supplementary Table S2). This information was then used to construct the branches of each of the decision trees per cluster. Using this method, dietary advice was developed based on four metabolic markers (total cholesterol, total saturated fat, Omega-3 Index and carotenoids) and five key nutrients (salt, dietary fibre, Fe, Ca and folate). The online Supplementary Fig. S1(a) and (b) demonstrate the metabolic and anthropometric decision tree and dietary decision trees for cluster 2, respectively. Furthermore, examples of targeted messages from each of the decision trees for cluster 2 are given in the online Supplementary Fig. 1(a) and (b). Combining the information from the characteristics of each cluster and the messages from the decision trees resulted in a comprehensive message. An example of advice for a participant in cluster 1 is given in the online Supplementary Table S4.

Fig. 1 Development of targeted dietary advice and comparison with individualised advice. Overview of the process for the delivery of targeted advice and comparison with individualised dietary advice using data from the Food4Me Study. Individuals are placed in metabotypes on the basis of their metabolic profiles. In this example, three distinctly different clusters are identified (cluster 1 had high cholesterol, high trans-fat and low n-3; cluster 2 had high n-3 and high total carotenoids; cluster 3 had low cholesterol and high stearic acid). Dietary advice encompasses characteristics of the metabotype and the decision trees, which include dietary factors not captured by the metabolites and anthropometric characteristics. The appropriateness of the targeted dietary advice was then compared with the individualised dietary advice of randomly selected Food4Me participants (n 180).
Table 5 Range of values across the clusters and cut-offs used for the development of the targeted dietary advice

WC, waist circumference.
Comparison of the targeted dietary advice and personalised feedback reports
Level 2 participants from Food4Me (n 180) were selected to test the appropriateness of the targeted dietary advice developed within this study. Excellent agreement was found between the personalised advice delivered by trained nutritionists in Food4Me and the targeted method developed in this study, with an average match of 82 % in relation to the dietary messages given (Table 6). Examining the clusters individually, good agreement was also found with an average match of 83 % for cluster 1, 74 % for cluster 2 and 88 % for cluster 3 for the dietary messages given. The number of messages given as part of the targeted dietary advice is depicted in Table 7. In general, more messages were given using the targeted approach compared with the individualised method used in Food4Me, where a restriction to three nutrient-related goals was imposed.
Table 6 Agreement between the proposed targeted dietary advice and the individualised dietary advice method adopted within the Food4Me studyFootnote *

* The agreement between the targeted and individualised method is at the level of the delivery of the same dietary message.
Table 7 Number of messages given as per the targeted dietary adviceFootnote * (Numbers and percentages)

* Number of dietary messages given using the proposed targeted dietary advice method.
Discussion
The present study demonstrates a successful method for the delivery of targeted nutrition advice using a combination of metabotyping and decision trees. Excellent agreement between this method and that of a personalised method delivered by a team of trained nutritionists and dietitians in the Food4Me study was found, with an average match of 82 %, at the level of agreement of the same dietary message given. To the best of our knowledge, this is the first study to identify metabotypes in the European population and to develop tailored dietary solutions appropriate for participants from diverse cultures and dietary intakes. This work paves the way for further development of this approach and potential delivery of personalised nutrition advice to large population groups.
Using cluster analysis, three distinctly different metabotypes were identified based on a range of blood-based metabolic markers. Individuals in cluster 1 were found to have an unhealthy metabolic profile as these individuals had the highest cholesterol levels, highest saturated fat levels and lowest Omega-3 Index. On the other hand, individuals in cluster 2 were identified as the healthiest group and had the lowest saturated fat levels, highest carotenoid concentrations and highest Omega-3 Index. Subjects in cluster 3 were found to be the youngest with the lowest cholesterol levels and poor dietary quality in terms of fruit intake fibre levels and carotenoid levels. These findings are similar to previously published studies on metabotypes( Reference Morris, O’Grada and Ryan 6 , Reference van Bochove, van Schalkwijk and Parnell 17 ). Morris et al.( Reference Morris, O’Grada and Ryan 6 ) identified four metabotypes consisting of four different responses to an oral glucose tolerance test (OGTT). Classification of individuals based on their response curves to an OGTT revealed an ‘at-risk’ metabolic phenotype, which had the highest BMI, TAG levels, C-reactive protein, C-peptide, insulin and homeostatic model assessment of insulin resistance (HOMA-IR) score( Reference Morris, O’Grada and Ryan 6 ). In a similar manner, van Bochove et al.( Reference van Bochove, van Schalkwijk and Parnell 17 ) identified three clusters based on their lipoprotein profiles and reported one cluster in which individuals did not respond favourably to fenofibrate treatment. In our previous study, one cluster with a metabolically unfavourable profile and another cluster in which the individuals were relatively healthy with respect to a range of metabolic markers were also identified( Reference O’Donovan, Walsh and Nugent 22 ). The consistency of identification of clusters across a range of studies adds validation to the approach and supports the clusters found in the present study.
An important finding from the current study is the evidence that there was a relationship between the metabolic profiles of each cluster and the corresponding nutrient and food group intakes of those clusters. For example, in line with their high carotenoid concentrations, participants in cluster 2 were also found to have the highest intakes of vitamin C, folate and dietary fibre. Similarly, individuals in cluster 2 had the highest intake of the fish, fish dishes and products, which was also reflected in their metabolic profile as this group had the highest average Omega-3 Index. However, individuals in cluster 2 had the highest intakes of supplements, which were likely to contribute to their high n-3 levels. The agreement between the metabolic profiles and dietary intake supports the concept of using blood-based metabotypes as a basis for targeted nutrition advice.
Good agreement between the proposed framework and the individualised advice delivered in the Food4Me study was observed. In Food4Me, personalised dietary advice was delivered by trained dietitians and nutritionists across seven research centres in Europe and was based on a decision tree method, which resulted in 295 possible dietary messages( Reference Celis-Morales, Livingstone and Marsaux 23 ). In the final section of the personalised reports, participants were given three key pieces of dietary advice that they were encouraged to focus on, which were selected based a priority system, developed specially for the purposes of the Food4Me study( Reference Celis-Morales, Livingstone and Marsaux 23 ). In contrast to this, a more simplified method is proposed here, in which blood-based metabolic data in conjunction with minimal dietary information could be used to deliver tailored dietary advice. This more simplified approach showed an average match of 82 % at the level of the dietary advice given, with the actual advice delivered within the Food4Me study. On the basis of this, it is proposed that tailored dietary advice could be given based primarily on the metabolic markers and information on the intakes of five key nutrients.
A framework for the delivery of targeted dietary advice in the Irish population, by the identification of three diverse metabotypes, and development of tailored dietary advice based on decision trees was previously presented( Reference O’Donovan, Walsh and Nugent 22 ). In the current paper, a similar method to identify metabotypes was used, but we have advanced this concept by the inclusion of a broader range of metabolic data. Furthermore, the decision trees for the delivery of the advice were expanded to include specific key nutrients. Furthermore, the decision trees were designed to accommodate sex-specific recommendations. This approach has potential to improve public health through the provision of tailored dietary advice to patients, in a quick and efficient manner, with minimal effort required by healthcare providers.
In this study, the metabolic markers were collected using DBS cards by the participants in their own homes. Collection of samples by DBS has a number of advantages including reduced costs, possibility of collection of large sample sizes, no blood processing and minimal storage facilities required( Reference McDade, Williams and Snodgrass 30 , Reference McDade 31 ). This presents another opportunity for the proposed framework to be adopted in the community setting where community health nurses could deliver the targeted dietary advice. Community nurses are suitable candidates to deliver tailored advice as they routinely see patients who may benefit from dietary/lifestyle change, have regular contact with patients over long periods, visit patients in their own homes and can involve their family in the intervention, and visit those who may not be physically capable of attending their doctor( Reference Chan, Laws and Williams 32 ). Chan et al. conducted a study to investigate the scope for risk management practices by nurses based in the community( Reference Chan, Laws and Williams 32 ). They reported that levels of obesity and prevalence of risk factors including smoking status and low physical activity levels were higher in the individuals (n 804) who took part in the study, compared with the general population, and that the majority of individuals with at least one risk factor had not received advice or been referred in the past 3 months( Reference Chan, Laws and Williams 32 ). This suggests that there is considerable scope to deliver dietary and lifestyle interventions in the community. In addition, when provided with appropriate training, community nurses were shown to be confident in assessing lifestyle factors such as smoking, anthropometric measures and dietary intake( Reference Chan, Jayasinghe and Christl 33 ). It is envisaged that the proposed framework, in our study, could easily be adopted by nurses in the community setting, to deliver tailored dietary advice with minimal training required, and have the potential to reach many more individuals who could benefit from tailored dietary advice.
A major strength of this study is its applicability to the European population. Furthermore, good agreement was reported between the proposed targeted method and an individualised method delivered by a team of nutritionists across seven research centres in the Food4Me study. A limitation of this study is that the dietary intake data were collected using an online FFQ, which assessed dietary intake of the previous month. Furthermore, the dietary advice developed did not take into account cooking abilities, likes/dislikes or cost of meals.
The present study developed a framework for the identification of metabotypes in the European population and the development of tailored dietary advice. Good agreement was found in comparison with an individualised personalised nutrition approach, which has been used to deliver advice across seven countries. The demonstration of this approach in a pan European study offers significant credibility to the framework. In our previous study, we envisaged translation of this approach for use by healthcare professionals and the present study further supports such a concept. With this in mind, future work should test this framework in such a setting to ascertain whether the advice is effective in motivating changes in diet and lifestyle factors.
Acknowledgements
The Food4Me Study is supported by the European Commission under the Food, Agriculture, Fisheries and Biotechnology Theme of the 7th Framework Programme for Research and Technological Development, grant no. 265494.
C. B. O. D., E. R. G. and L. B. developed and tested the targeted dietary approach, carried out the statistical analyses and drafted the manuscript. C. C.-M., R. F., A. L. M., C. F. M. M., S. N.-C., R. S.-C., C. B. O. D., H. F., C. W., S. K., L. T., C. M., C. P. L., G. M., M. G., A. S., M. C. W. and J. C. M. conducted the intervention. I. T., C. A. D., H. D., Y. M., J. A. M., W. H. M. S., J. A. L., J. C. M., M. J. G., E. R. G. and L. B. contributed to the research design of the Food4Me study. All authors contributed to a critical review of the manuscript during the writing process and approved the final version to be published.
The authors declare that there are no conflicts of interest.
Supplementary Material
For supplementary material/s referred to in this article, please visit https://doi.org/10.1017/S0007114517002069









