Article contents
G-protein-coupled receptors function as logic gates for nanoparticle binding using systems and synthetic biology approach
Published online by Cambridge University Press: 20 February 2019
Abstract
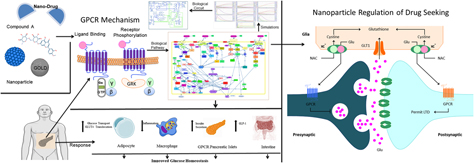
G-protein-coupled receptor 142 (GPR142) belongs to rhodopsin family. GPR142 and GPR119, both Gq-coupled receptors, are expressed in pancreatic β cells of pancreas; their activation eventually leads to triggering of insulin secretion. In this paper, through a systems and synthetic biology approach, the effect of a common hit compound has been investigated in GPR142 and GPR119 pathways. This hit that has the potential to be developed as a lead for nanodrug was obtained through high-throughput virtual screening. The hit compound was further docked with nanoparticles (GOLD, SPION, and CeO2). The probable effect of this potential hit on insulin secretion in type 2 diabetes and its dynamic behavior was explored. Kinetic simulation was performed for cross-validation of its role in both the pathways. This study opens up a probable avenue in therapy of type 2 diabetes through regulation of GPR142 and GPR119 receptors. The biological circuit constructed may further have an application as a modulator to control the up- and downregulation of the biochemical pathway and can be implemented as sensors or nanochips for therapy.
Information
- Type
- Article
- Information
- Journal of Materials Research , Volume 34 , Issue 11: Focus Issue: (Nano)materials for Biomedical Applications , 14 June 2019 , pp. 1854 - 1867
- Copyright
- Copyright © Materials Research Society 2019
References
- 3
- Cited by

