Introduction
Marteilia is a genus of protistan parasites in the order Paramyxida (Rhizaria, Endomyxa, Ascetosporea; Bass et al. (Reference Bass, Ward and Burki2019)), including species associated with mortalities of aquatic mollusc species around the globe. Characterized by their cell-in-cell morphology, 8 species of Marteilia have so far been described and have available molecular sequence data: M. refringens (Grizel et al. Reference Grizel, Comps, Bonami, Cousserans, Duthoit and Le Pennec1974), M. pararefringens (Kerr et al. Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018), Marteilia sydneyi (Perkins and Wolf, Reference Perkins and Wolf1976), M. cochillia (Carrasco et al. Reference Carrasco, Hine, Durfort, Andree, Malchus, Lacuesta, González, Roque, Rodgers and Furones2013), M. (Eomarteilia) granula (Itoh et al. Reference Itoh, Yamamoto, Kang, Choi, Green, Carrasco, Awaji and Chow2014), M. octospora (Ruiz et al. Reference Ruiz, López, Lee, Rodríguez and Darriba2016), M. tapetis (Kang et al. Reference Kang, Itoh, Limpanont, Lee, Whang and Choi2019) and M. cocosarum (Skujina et al. Reference Skujina, Hooper, Bass, Feist, Bateman, Villalba, Carballal, Igleias, Cao, Ward, Ryder, Bignell, Kerr, Ross, Hazelgrove, Macarie, Prentice, King, Thorpe, Malham, Mckeown and Ironside2022). Two other species of Marteilia have been described morphologically, but do not have any available sequence data: M. christenseni (Comps, Reference Comps1983) and M. lengehi (Comps, Reference Comps1976). Marteilia have a broad geographical range and have been detected in invertebrate hosts in Europe, Oceania, Asia, Africa and the USA (Grizel et al. Reference Grizel, Comps, Bonami, Cousserans, Duthoit and Le Pennec1974; Perkins and Wolf, Reference Perkins and Wolf1976; Moyer et al. Reference Moyer, Blake and Arnold1993; Audemard et al. Reference Audemard, Le, Barnaud, Collins, Sautour, Sauria, De, Coustau, Combes and Berthe2002; Elgharsalli et al. Reference Elgharsalli, Aloui-Bejaoui, Salah, Chollet, Joly, Robert, Couraleau and Arzul2013; Adlard and Nolan, Reference Adlard and Nolan2015).
Historical taxonomy
Described in the context of mortalities of the flat oyster (Ostrea edulis) in 1969, M. refringens was the first species of Marteilia to be identified (Comps, Reference Comps1970; Grizel et al. Reference Grizel, Comps, Bonami, Cousserans, Duthoit and Le Pennec1974). Marteilia refringens is a listed pathogen by both the World Organization for Animal Health (WOAH; previously OIE) (Woah, 2023) and the European Union (under EU regulation 2016/429) as a significant pathogen of bivalve molluscs. Subsequent to the identification of M. refringens in flat oysters, a Marteilia species was detected in the mussels Mytilus edulis (Comps et al. Reference Comps, Grizel, Tige and Duthoit1975) and My. galloprovincialis (Comps et al. Reference Comps, Grizel and Papayanni1982). Based on host specificity and ultrastructural characteristics, the Marteilia infecting mussels was described as a separate species: M. maurini (Comps et al. Reference Comps, Grizel and Papayanni1982). Further evidence that the Marteilia lineages infecting oysters and mussels were distinct species was provided by Le Roux et al. (Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001), who sequenced partial internal transcribed spacer 1 (ITS1) and found consistent nucleotide differences between them. However, Longshaw et al. (Reference Longshaw, Feist, Matthews and Figueras2001) challenged the characterization of 2 different species, finding no significant differences in ultrastructure between M. refringens and M. maurini, albeit based on small sample numbers and limited lifecycle stages. Berthe et al. (Reference Berthe, Le Roux, Peyretaillade, Peyret, Rodriguez, Gouy and Vivarés2000) also disputed there being 2 species, based on no clear differences in small ribosomal subunit gene (18S) sequence data between Marteilia isolated from oysters and mussels. It also became apparent that M. maurini could be amplified from O. edulis, and M. refringens could be amplified from My. galloprovincialis (López-Flores et al. Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004; Novoa et al. Reference Novoa, Posada and Figueras2005; Balseiro et al. Reference Balseiro, Montes, Ceschia, Gestal, Novoa and Figueras2007), so host specificity was disregarded as a species-defining characteristic. It was determined by the European Food Safety Association (European Food Safety, Reference European Food Safety2008), based on Balseiro et al. (Reference Balseiro, Montes, Ceschia, Gestal, Novoa and Figueras2007), that there was insufficient evidence to maintain separate species status. Marteilia refringens and M. maurini were therefore synonymized as M. refringens in 2007, referred to as ‘O-type’ (previously the ‘oyster-infecting’ M. refringens) and ‘M-type’ (previously ‘mussel-infecting’ M. maurini), defined by the ITS1 sequence differences identified by Le Roux et al. (Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001).
Synonymization of the 2 Marteilia species had important consequences – many studies after the synonymization have not distinguished between M and O type (only 28 out of 57 publications since the synonymization include typing), thereby conflating differences between the 2 lineages. Seventeen years after the synonymization, differences between the pathology and host range of the 2 lineages are insufficiently understood. The change also resulted in discovery of either M- type or O-type ‘M. refringens’ requiring reporting to WOAH and to the European Commission under Directive EC/2006/88, even in the absence of mortalities (European Food Safety, Reference European Food Safety2008). The result of the synonymization was an enlargement of the apparent host range of M. refringens, which not only included O. edulis but also My. edulis and My. galloprovincialis. As a consequence, several sites in Northern Europe, where M. refringens type M (i.e. M. pararefringens) was detected were considered positive for M. refringens, despite no Marteilia detection by histology in oysters (Laing et al. Reference Laing, Dunn, Peeler, Feist and Longshaw2014). Disputing the synonymization, a phylogenetic study carried out by Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) sequenced almost full length rRNA gene assemblies from O- and M-type M. refringens, and determined that the 2 lineages within M. refringens were robustly distinguishable at species level, with the full ITS1 sequence being a reliable marker to discriminate between the 2 species. As such, they were formally redefined as separate species: M. refringens (previously O-type) and M. pararefringens (previously M-type). As there was no material available for M. maurini to test the hypothesis that it was synonymous with M-type (and thus synonymous with M. pararefringens). To prevent confusion, for the present study we herein adopt the same terminology as in Bøgwald et al. (Reference Bøgwald, Skår, Karlsbakk, Alfjorden, Feist, Bass and Mortensen2022), referring to M. refringens as described in Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) as M. refringens sensu stricto, and M. refringens in publications where it is not possible to determine the genotype as M. refringens sensu lato. Marteilia refringens sensu stricto and M. pararefringens have overlapping but distinct host and geographical distributions; however, data on confirmed infections in potential host species (supported by both histopathology and molecular data) remain sparse (Le Roux et al. Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001; Audemard et al. Reference Audemard, Le, Barnaud, Collins, Sautour, Sauria, De, Coustau, Combes and Berthe2002; López-Flores et al. Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004, Reference López-Flores, Garrido-Ramos, De La Herran, Ruiz-Rejón, Ruiz-Rejón and Navas2008a, Reference López-Flores, Robles, Valencia, Grau, Villalba, De La Herran, Garrido-Ramos, Ruiz-Rejon, Ruiz-Rejon and Navas2008b; Kerr et al. Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018; Bøgwald et al. Reference Bøgwald, Larsson and Mortensen2024).
Molecular markers for species demarcation
As there is no generally accepted basis for delimiting species in parasitic protists (or for microeukaryotes in general), the definition of species boundaries is a highly debated topic (Boenigk et al. Reference Boenigk, Ereshefsky, Hoef-Emden, Mallet and Bass2012). However, as molecular techniques for the detection of parasites are progressing (i.e. metagenomes, 18S amplicon-based studies, etc.), discovery and characterization of organisms closely related to known parasites is likely to become more common. An understanding of how these closely related parasites have evolved is important to be able to correctly describe them taxonomically. Many molecular markers, particularly within Ascetosporea, target the 18S and ITS1 regions of the rRNA array. However, few sequences exist for the remainder of the array, leaving sequence variability in these organisms poorly understood. The present study applies a method developed by Hooper et al. (Reference Hooper, Ward, Foster, Skujina, Ironside, Berney and Bass2023) to sequence the rRNA array in its entirety in 2 overlapping amplicons, enabling rapid generation of sequence data.
A criticism of the phylogenetic study carried out by Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018), which formally classified M. refringens sensu stricto and M. pararefringens as separate species, was the small number of samples that the analysis was performed on: almost complete arrays were produced for only 2 independent M. pararefringens samples (9,756–9,777 bp) and 1 M. refringens sensu stricto sample (8,559 bp); incomplete arrays were produced for 3 independent M. refringens samples (4,031–5,910 bp) and 1 M. pararefringens sample (6,811 bp). The incompleteness of the arrays was the result of the primer walking method used to generate them – gaps were present where primer sets failed to amplify regions of the array. Due to the small sample size used and the incompleteness of the arrays, it has not previously been possible to determine consistently different nucleotide differences between M. refringens sensu stricto and M. pararefringens outside of the ITS1 region. The inferred phylogenetic distinctiveness of the 2 species was based on a very small sample set, and the inference may have been undermined by sequence variation in additional samples.
In this study, we sequenced full rRNA arrays, 18S (small ribosomal subunit) – ITS1 (first internal transcribed spacer) – 5.8S (component of the large ribosome subunit) – ITS2 (second internal transcribed spacer) – 28S (component of the large ribosome subunit) – IGS (intergenic spacer containing the NTS (non-transcribed spacer) and ETS (external transcribed spacer)), for a larger number of M. refringens sensu stricto and M. pararefringens samples from France, Greece, Italy, Norway, Spain and the United Kingdom using the method outlined in Hooper et al. (Reference Hooper, Ward, Foster, Skujina, Ironside, Berney and Bass2023). Sequence and phylogenetic analyses were carried out on full rRNA arrays, and the most discriminating regions of the rRNA array, of M. refringens sensu stricto and M. pararefringens, from all available samples covering the known hosts and covering the full documented European geographical range of these parasites. Results of these analyses were used to determine new – and confirm the appropriateness of current – markers to discriminate each species and inform the development of improved diagnostic assays.
Materials and methods
Samples
Samples that had previously been shown to be PCR positive for M. refringens sensu stricto or M. pararefringens were used in this study and are outlined in Supplementary Table 1. This sample set was as comprehensive as possible, covering as much of the host range and geographical distribution of the 2 Marteilia species held by the WOAH reference laboratory for marteiliosis and contributed by EU partners.
PCR to generate long rRNA amplicons
PCR reactions to generate long amplicons were carried out in triplicate in 25 µl volume reactions comprising 1 × PrimerSTAR GXL Buffer (Takara Bio, Shiga, Japan), 0.2 µM dNTP mixture (Takara Bio), 0.4 µM of each primer and 1.25 U PrimeSTAR GXL DNA polymerase (Takara Bio). Cycling conditions for all long amplicon PCRs were carried out by initial denaturing at 95°C for 5 min, followed by 40 cycles of 95°C for 45 s, a variable annealing temperature (described below) for 45 s and 68°C for 150 s; and final extension of 10 min at 68°C.
Amplicons covering 18S-ITS1-5.8S-ITS2-28S or ITS1-5.8S-ITS2-28S were amplified with EuroMarteilia30F/LSU8799degen with an annealing temperature of 64°C or MartDBITSf1/LSU8799degen with an annealing temperature of 65°C, respectively. Amplicons covering 28S-IGS-18S or 28S-IGS-18S-ITS1 were amplified using 7939 F/ParaGenRGW with an annealing temperature of 64°C or 7939 F/MartDBITSr1 with an annealing temperature of 63°C. The orientation of primers and the regions of the rRNA array they amplify are detailed in Figure 1, and all primer sequences are given in Table 1.
Schematic illustrating the position of primers used for long-range PCR on the of the ribosomal RNA array and the approximate length of amplicons generated by them. 18S – gene for the small subunit of the ribosome, 28S and 5.8S – genes of the components of the large subunit of the ribosome, ITS1 - first internal transcribed spacer, ITS2 - second internal transcribed spacer, IGS - intergenic spacer, NTS - non-transcribed spacer of IGS, ETS - external transcribed spacer of the IGS.

Primer sequences, their position of the primer on the rRNA and the direction in which they amplify

Sequencing using Illumina and nanopore technologies
Amplicons were cleaned using AMPure XP beads following the manufacturer’s protocol using a 1 × bead volume. All long amplicons were prepared for sequencing on an Illumina MiSeq (Illumina, California, USA) using a Nextera XT DNA Library Preparation Kit (Illumina), following the manufacturer’s protocol, but using half volume reactions. Libraries were sequenced on v2 Nano flow cells (500-cycle) (Illumina) with 2 × 250 bp read length.
A subset of amplicons covering 28S-IGS-18S and 28S-IGS-18S-ITS1 were prepared for Nanopore sequencing using the PCR Barcoding Kit (Oxford Nanopore Technologies, Oxford, UK) and sequenced on Flongle Flow Cells (12 samples per flow cell) (Oxford Nanopore Technologies) using a MinIon Mk1C device (Oxford Nanopore Technologies). Base calling was completed in real-time on the Mk1C device using Guppy (Oxford Nanopore Technologies).
Sequence analysis of Illumina and nanopore reads
For amplicons sequenced using Illumina MiSeq, raw paired-end sequence reads were trimmed to remove adaptor and low-quality sequences using Trimmomatic v0.39 (using a sliding window of 4, minimum quality of 15, leading and trailing values of 3 and a minimum length of 50 bases; Bolger et al. (Reference Bolger, Lohse and Usadel2014)). The quality of trimmed and filtered reads was assessed using FastQC v0.11.8 (default parameters; https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) prior to assembly using SPAdes v3.13.1 (in – meta mode, using kmer sizes of 21, 33, 55, 77, 88 and 127; Prjibelski et al. (Reference Prjibelski, Antipov, Meleshko, Lapidus and Korobeynikov2020)).
Nanopore-generated raw reads were trimmed, corrected and a consensus generated using Canu v2.1 (default parameters with a genome size of 0.0065 m; Koren et al. (Reference Koren, Walenz, Berlin, Miller, Bergman and Phillippy2017)). Trimmed and filtered Illumina paired reads were concatenated using a python script (https://github.com/isovic/racon/blob/master/scripts/racon_preprocess.py) and mapped to the consensus Nanopore contig using minimap2 v2.17-r941 (using -x sr; Li (Reference Li2018)). Using the mapped reads, the Nanopore-generated contig was polished once using racon v1.4.13 (using default parameters; Vaser et al. (Reference Vaser, Sović, Nagarajan and Šikić2017)). Nanopore reads were only required to determine the sequence of the non-transcribed spacer (NTS) of the rRNA array, which contains tandem repeats (Hooper et al. Reference Hooper, Ward, Foster, Skujina, Ironside, Berney and Bass2023). The NTS region was determined not to be useful for this study due to the variation in tandem repeat number within a given sample, so Illumina sequencing alone was used to obtain ETS-28S sequence data due to its higher accuracy rate compared to Nanopore-based sequencing.
Paired reads from each sample were mapped to their respective consensus sequence using BWA-MEM v0.7.17 (Li and Durbin, Reference Li and Durbin2009) and SAMtools v1.9 (Li et al. Reference Li, Handsaker, Wysoker, Fennell, Ruan, Homer, Marth, Abecasis and Durbin2009) with default parameters. The output from BWA-MEM was visualized with Integrative Genomics Viewer (IGV) v2.5.2 (Robinson et al. Reference Robinson, Thorvaldsdóttir, Winckler, Guttman, Lander, Getz and Mesirov2011) and the presence of any single-nucleotide polymorphisms (SNPs) within an individual sample was assessed with, any SNPs present at ≥5% considered. Coverage, assembly quality and accuracy, and using QualiMap v2.2.2 (García-Alcalde et al. Reference García-Alcalde, Okonechnikov, Carbonell, Cruz, Götz, Tarazona, Dopazo, Meyer and Conesa2012).
PCR to generate short amplicons covering diagnostic regions
PCR reactions to generate shorter amplicons covering the most variable regions of the rRNA array (ITS1 and ETS) were carried out in 20 µl reactions containing 1 μl DNA template, 1 × Green GoTaq® Flexi Buffer (Promega, Wisconsin, USA), 2.5 mM MgCl2 (Promega), 0.4 mM dNTP mix (Bioline, London, UK), 0.5 µM of each primer, 10 µg Bovine Serum Albumin (BSA) (New England Biolabs, Massachusetts, USA) and 0.5 u GoTaq® G2 DNA Polymerase (Promega). For ITS1, the nested primer set MartDBITSf1/MartDBITSr1 and MartDBITSf2/MartDBITSr2 was used to amplify a 1,034-bp fragment (Kerr et al. Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018). For ETS, the nested primer set INT2_RC/18S and ETS_FN/EuroMartRC was used to amplify a 1,132-bp fragment. Primer ‘18S’ was published by López-Flores et al. (Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004), and ‘INT2RC’ and ‘EuroMartRC’ are the reverse complement of ‘INT2’ and ‘EuroMarteilia30F’ primers published by López-Flores et al. (Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004) and Skujina et al. (Reference Skujina, Hooper, Bass, Feist, Bateman, Villalba, Carballal, Igleias, Cao, Ward, Ryder, Bignell, Kerr, Ross, Hazelgrove, Macarie, Prentice, King, Thorpe, Malham, Mckeown and Ironside2022), respectively. Primers were checked for specificity to Marteilia using Primer-BLAST (Ye et al. Reference Ye, Coulouris, Zaretskaya, Cutcutache, Rozen and Madden2012).
Amplicons covering full-length ITS1 generated using MartDBITSf1/MartDBITSr1 and MartDBITSf2/ MartDBITSr2 primers were amplified by an initial denaturing at 94°C for 5 min, followed by 35 cycles of 94°C for 1 min, 65°C for 1 min and 72°C for 1 min; and a final extension of 7 min at 72°C. Amplicons covering a variable region of the ETS were generated using INT2RC/18S primers by an initial denaturing at 95°C for 5 minutes, followed by 35 cycles of 95°C for 1 minute, 63°C for 1 minute and 72°C for 1 minute; and a final extension of 10 minutes at 72°C. The nested amplicon produced by ETSfN/EuroMartRC was amplified as above but with an annealing temperature of 60°C. All primer sequences are detailed in Table 1.
Sequencing and analysis of ITS1 and ETS amplicons
Amplicons produced by ITS1 and ETS nested PCRs were cleaned using AMPure XP beads following the manufacturer’s protocol using a 1 × bead volume. Cleaned amplicons were sequenced bidirectionally using the nested amplification primers using Eurofins PlateSeq or Eurofins TubeSeq Service (Eurofins Genomics, Wolverhampton, UK).
Cloning of ITS1 amplicons
When the presence of multiple sequence types was indicated by mixed chromatograms, the first round and nested PCRs were repeated with a proof-reading polymerase, followed by cloning to ensure high sequence accuracy. PCR products that were cloned are indicated with (*) in Table S1. Each 25 µl PCR contained 2 × NEBNext® Ultra™ II Q5® Master Mix (New England Biolabs), 0.5 µM of each primer and 1.25 µl of template DNA. Full ITS1 was amplified by an initial denaturing at 98°C for 30 s, followed by 35 cycles of 98°C for 10 s, 65°C for 30 s and 68°C for 45 s; and a final extension of 2 min at 68°C. PCR products were cleaned using AMPure XP beads following the manufacturer’s protocol using a 1 × bead volume. Cleaned amplicons were heat shocked into plasmids using CloneJet™ PCR cloning kit (Thermo Fisher Scientific, Massachusetts, USA), and DH10B competent cells (Thermo Fisher Scientific) were transformed following the manufacturer’s protocol for blunt ended PCR products. Twenty-two colonies from each sample were taken forward for colony PCR using the MartDBITSf2 and MartDBITSr2 primer set, as above, cleaned using 0.8 × AMPure XP beads, and sequenced bidirectionally using Eurofins PlateSeq service (Eurofins Genomics).
Phylogenetics
Given the inherent difficulties achieving convergence using a Maximum Likelihood (ML) approach to tree construction from multiple sequence alignments with low-information data (Beerli, Reference Beerli2006) (e.g. regions of the rRNA array that appear to have low mutation rates), a Bayesian approach was taken to construct consensus trees for full transcribed rRNA arrays and the most informative regions of the rRNA array. A Bayesian consensus tree was constructed from ETS-28S rRNA array alignments, produced using MAFFT v7.0 (L-INS-I algorithm; Katoh and Standley (Reference Katoh and Standley2013)), with MrBayes v3.2.7 (Ronquist et al. Reference Ronquist, Teslenko, Van Der Mark, Ayres, Darling, Höhna, Larget, Liu, Suchard and Huelsenbeck2012) on the CIPRES Science Gateway (Miller et al. Reference Miller, Pfeiffer and Schwartz2010). The tree was constructed using 2 separate MC3 runs, carried out for 2 million generations using 1 cold and 3 hot chains. The first 500,000 generations were discarded as burn-in, and trees were sampled every 1000 generations. Robust clade distinction was based on maximal support of branching events (posterior probabilities of 1).
For the regions of the rRNA that appeared to be most consistently different between M. pararefringens and M. refringens sensu stricto (ITS1 and ETS), all publicly available sequences were downloaded from NCBI and aligned against the sequences generated in this study with MAFFT, as above. Ambiguous sequences were removed and Bayesian consensus trees for these regions were constructed as above.
Coalescent-based species delimitation
Species boundaries within the Marteilia clade were inferred using 2 methods. First, a Bayesian Poisson Tree Process (bPTP) model was used to delimit species on the rooted Bayesian consensus tree constructed from a Marteilia ETS-28S rRNA array multiple sequence alignment. bPTP analysis was performed on the bPTP web server (Zhang et al. Reference Zhang, Kapli, Pavlidis and Stamatakis2013), modelled using 100,000 MCMC generations, a thinning factor of 100, a burn-in of 0.1, and the seed set to 123. Convergence was checked by visually examining the likelihood trace plot. The values corresponding to partitions found by a simple heuristic search were added to the branches of the Bayesian consensus tree.
Secondly, BPP (Flouri et al. Reference Flouri, Jiao, Rannala and Yang2018), a Bayesian Markov chain Monte Carlo (MCMC) program, was used to determine the posterior probability of inferred species by applying the multispecies coalescent model. The topology of the Bayesian consensus tree generated from Marteilia ETS-28S rRNA array multiple sequence alignment was used as a guide tree for species delimitation, and the inferred 5 Marteilia species from bPTP analysis were used to assign species to each full array for the Imap input file. To evaluate the influence of prior distributions for ancestral population sizes (θ) and the root age (τ0), 3 different combinations of priors were used for the analysis. Priors are assigned a gamma G(α, β) distribution and were applied to the full array to account for:
i. large ancestral population sizes and deep divergence between species, θ ∼ G(1, 10) and τ0 ∼ G(1, 10);
ii. large ancestral population sizes and shallow divergence between species, θ ∼ G(1, 10) and τ0 ∼ G(2, 2000);
iii. small ancestral population sizes and shallow divergence between species, θ ∼ G(2, 2000) and τ0 ∼ G(2, 2000).
The analyses were performed with the following parameters: rjMCMC algorithm = 0, fine-tuning parameter ɛ = 15, usedata = 1, cleandata = 0 and speciesmodelprior = 1. The rjMCMC analyses were run for 50,000 generations (sampling interval of 5) with a burn-in period of 5,000. Each analysis was run in duplicate to ensure consistent results across analyses initiated with different starting seeds.
Results
Assembly of arrays and multiple sequence analysis
Full transcribed arrays (ETS-28S; approximately 8,380 bp) were assembled from 9 independent samples for M. refringens sensu stricto (M. refringens as described in Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018)) and 28 independent samples for M. pararefringens by merging the overlapping fragments for each sample to produce 1 contig. The non-transcribed spacer (NTS) region of the intergenic spacer (IGS) was excluded from analysis due to its tandem repeat structure, making it an unsuitable region for determining differences between the 2 Marteilia species. The full transcribed regions of the rRNA arrays generated in this study were deposited to GenBank under accession numbers PP549144-80, with details of the host, location from which they were amplified and average contig coverage given in Supplementary Table 1.
A total of 20 consistent nucleotide differences (differences seen in all samples of each Marteilia species) were observed across the 8,410 nucleotide positions of transcribed rRNA (ETS-28S) alignments of M. refringens sensu stricto and M. pararefringens, including the 7 complete or near complete rRNA arrays generated by Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018). The majority of these differences were within the ETS (11 nucleotide differences over 1,508 bp) and ITS1 (6 nucleotide differences over 690 bp). The remaining nucleotide differences were located in ITS2 (1 nucleotide difference) and the 5ʹ end of the 28S (2 nucleotide differences). Marteilia refringens sensu stricto and M. pararefringens had 100% sequence similarity in 18S (except samples from Norway, described below) and 5.8S regions of the rRNA array.
The ITS2 region of the rRNA had only 1 consistent nucleotide difference between species across its 800-bp length. Mapping Illumina reads to the consensus contigs obtained from SPAdes assembly revealed that ITS2 in M. pararefringens had regions containing indels (shown as dark green blocks in Figure 2A), which occurred in short tandem repeat regions. For example, within sample 2006_031_11, 1, 2 or 3 copies of ACA (ungapped position 4,376 bp on the rRNA) were present. Within 28S, 2 short regions (∼50 bp) at the 3ʹ end of the gene appeared to have bases that were variable within a sample. This variability was consistent across both M. pararefringens and M. refringens sensu stricto samples, with the majority of ambiguity occurring as T to G base changes. A shorter region of variability (∼20 bp) was also observed at the 5ʹ end of the 18S gene. Within this region, there were no consistent nucleotide differences between M. pararefringens and M. refringens sensu stricto; however, M. pararefringens from Norway showed 3 consistently different nucleotide differences with respect to all other M. pararefringens sequences, these differences are shown in Figure 2C. Across the remainder of the array, there were 3 other nucleotide positions, 2 in ETS and 1 in ITS2, which were consistently different in samples from Norway than other M. pararefringens sequences.
Consistently different nucleotide differences between Marteilia pararefringens and M. refringens sensu stricto. (A) Schematic representation of full transcribed regions of the ribosomal RNA (rRNA) arrays for M. pararefringens and M. refringens sensu stricto. Vertical black lines represent consistent differences between species. Darker blocks in ITS1 and ITS2 represent tandem repeat regions. Grey and white horizontal blocks in under the ETS represent regions amplified by López-Flores et al. (Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004) nested primer set and dark and light blue blocks under the 18S and ITS1 represent regions amplified by Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) nested primer set. Dark and light blue horizontal blocks under the ETS and 18S represent the region amplified by nested primers designed in this study, and the white horizontal block under ITS1 represents the region amplified by the Le Roux et al. (Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001) pr4/pr5 primer set. (B) Details of the consistently different nucleotide positions depicted in A. Red bases are invariant within each species, with the position of these bases relative to sample 18/35/12 (accession number PP549162). Numbers above and below the highlighted bases depict the number of samples for each species that were available for these regions to determine the differences. (C) Consistent nucleotide differences in the 18S gene between M. pararefringens amplified from samples from Norway compared to all other M. pararefringens, and M. refringens sensu stricto samples.

Amplification of the most discriminating regions of the rRNA array
To confirm the differences in ETS and ITS1 over a larger sample set, amplification and Sanger sequencing using nested PCR assays was carried out. ITS1 and ETS sequences generated in this study were deposited to GenBank under accessions PP544135-53 and PP549181-98, respectively, with details of the host and location from which they were amplified given in Supplementary Table 1.
Internal transcribed spacer 1
As the MartDBITSf1/MartDBITSr1 and MartDBITSf2/MartDBITSr2 ITS1 nested primer set (Kerr et al. Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) amplified all consistently different nucleotides in the ITS1 region between the 2 Marteilia species identified by long-range PCR and sequencing, this primer set was applied to all available samples. The resulting sequences were aligned with the ITS1 region of the full-length arrays and all available ITS1 sequences on GenBank, totalling 120 M. pararefringens amplified from 6 different species, and 112 M. refringens sensu stricto sequences amplified from 3 different species (Table 2). Identical sequences were collapsed into sequence types; the sequences that constitute each type are outlined in Supplementary Table 2. Four ITS1 sequences were removed from the analysis: DQ426583, DQ426587 and DQ426596 due to sequencing reads appearing to be chimeric clones (a single DNA sequence originating from the joining of multiple DNA sequences caused by dissociation of polymerases from the template during sequencing). A proportion of the sequence matched an M. pararefringens clone, and the remaining proportion matched the M. refringens sensu stricto clone generated from the same sample. MH304865 was removed as the chromatogram showed mixed sequence types.
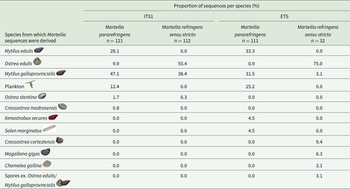
Species from which M. pararefringens and M. refringens sensu stricto were amplified from for ITS1 and ETS phylogenetic analysis. Data include sequences from NCBI and those generated in this study. Spores ex Ostrea edulis/Mytilus galloprovincialis refer to a sequence derived from Marteilia spores purified from a pool of tissues from both species

External transcribed spacer
Primers were designed to amplify 1,577 bp (single PCR) or 1,132 bp (nested PCR) of the ETS, producing an amplicon that covered most of the potential nucleotide difference between ETS regions of M. refringens sensu stricto and M. pararefringens suggested as species-specific markers by scrutiny of full-length rRNA arrays. The regions amplified by the single round and nested PCRs are shown as dark and light blue horizontal blocks on Figure 2A. Two differences at the 5ʹ end of the ETS could not be amplified due to low GC content and homopolymers upstream of this site, making it unsuitable for primer design. The primer set was applied to all available samples and the resulting sequences were aligned with the ETS amplicon region of the full-length arrays and all available ETS sequences on GenBank, totalling 111 M. pararefringens amplified from 6 different species, and 32 M. refringens sensu stricto sequences amplified from 6 different species (Table 2). Identical sequences were collapsed into sequence types; the sequences that constitute each type are outlined in Supplementary Table 3. Sequencing additional samples and comparison to ETS sequences in GenBank revealed that 5-nucleotide differences initially thought to be unique to each species in the full-length arrays were not unique with a greater sample number.
Consistently different nucleotide differences in the larger sample set are shown in Figure 2A–B. Three publicly available ETS sequences were removed from further analysis: accessions AJ629374-5 were removed due to there being degenerative bases present in clonal sequences, and MH304860, as revisiting the chromogram revealed mixed sequence types. Confirmation and/or disproof of the consistently different nucleotides observed in the limited number of full-length arrays by Sanger sequencing of shorter regions resulted in a total of 15 consistently different nucleotide positions across the full array between the 2 species of Marteilia.
Phylogenetic analysis of full rRNA arrays
A Bayesian consensus demonstrated maximal support (posterior probabilities of 1) for M. refringens sensu stricto and M. pararefringens clades. The tree, generated from 13 M. refringens sensu stricto and 30 M. pararefringens ETS-28S rRNA arrays (8,860 nucleotide positions), demonstrated mutually exclusive, monophyletic clades for both species (Figure 3). All other species of Marteilia, where full rRNA arrays were available, branched with maximal support for each species, with similar phylogenetic distances between M. refringens sensu stricto and M. pararefringens as between other closely related Marteilia species (M. cochillia and M. cocosarum). One long M. pararefringens sequence (accession numbers MH304630, MH329401, MH304638 and MH304857) branched separately from other M. pararefringens sequences, although still with maximal support with all other M. pararefringens sequences. However, this rRNA assembly is partial – lacking the informative region of ITS1 – which is likely to explain this branching pattern. Within the M. pararefringens clade, samples from Norway branched separately from other M. pararefringens samples, again with maximal support.
Bayesian consensus tree constructed from the full transcribed region (ETS-28S) of the rRNA array (8,860 nucleotide positions) for Marteilia refringens sensu stricto (n = 13) and M. pararefringens (n = 30), M. cochillia (n = 5), M. cocosarum (n = 5) and M. octospora (n = 1). Location of sampling and host species are denoted adjacent to each branch. Posterior probabilities to two significant figures are displayed on the branches in black text, with maximal support values (1.0) shown as black circles. Branch lengths are shown on key branches in red text. The tree is rooted to Paramarteilia canceri. Posterior probability values from bPTP analysis for all branching events are shown in blue text on branches. The posterior probabilities of likely speciation events suggested by the bPTP analysis are underlined.

Phylogenetic analyses of ITS1 and ETS regions
A Bayesian tree, constructed from all full-length ITS1 sequences generated in this study and ITS1 sequences available on GenBank, produced a highly supported M. refringens sensu stricto clade (0.97 posterior probability), and a moderately supported (0.86 posterior probability) M. pararefringens clade (Figure 4). Unlike the tree generated from full-length transcribed rRNA arrays, M. pararefringens amplified from Norway did not branch separately from other M. pararefringens sequences.
Bayesian consensus tree constructed from full ITS1 sequences (945 nucleotide positions) for Marteilia refringens sensu stricto (n = 112) and M. pararefringens (n = 121). Location of sampling, host species and number of individual samples the sequence type was detected in is denoted adjacent to each branch. Posterior probabilities to two significant figures are displayed on the branches in black text, with maximal support values (1.0) shown as black circles. Branch lengths are shown on key branches in red text. Tree is rooted to M. cochillia/M. cocosarum.

A Bayesian consensus tree constructed from long ETS sequences (1,043 nucleotide positions) generated in this study, and ETS sequences available on GenBank, for M. refringens sensu stricto (n = 32) and M. pararefringens (n = 111) produced a tree with similarly moderate support values for each clade as the ITS1 tree, with both species forming mutually exclusive, monophyletic clades (Figure 5).
Bayesian consensus tree constructed from long ETS sequences (1,043 nucleotide positions) for M. refringens sensu stricto (n = 32) and M. pararefringens (n = 111). Location of sampling and host species is denoted adjacent to each branch. Posterior probabilities to two significant figures are displayed on the branches in black text, with maximal support values (1.0) shown as black circles. Branch lengths are shown on key branches in red text. Tree is rooted to M. cochillia/M. cocosarum.

Trees based on the shorter regions amplified by primers recommended in the Aquatic Animal Health Manual (Woah, 2023) (originally published by Le Roux et al. (Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001), Figure 2A) were also constructed. A tree based on short ITS1 sequences produced mutually exclusive monophyletic clades for both M. pararefringens and M. refringens sensu stricto but had no support (0.51 posterior probability) for the M. refringens sensu stricto clade (Supplementary Figure 1A). Trees constructed from short ETS amplicon regions produced trees in which M. pararefringens was paraphyletic with respect to M. refringens sensu stricto (Supplementary Figure 1B-C), with high support for the M. refringens sensu stricto clade. In both ITS1 and ETS regions, longer amplicon regions produced trees that had higher support for each species’ clade than their shorter counterparts.
Coalescent-based species delimitation
bPTP inferred 5 Marteilia species (M. refringens sensu stricto, M. pararefringens, M. cochillia, M. cocosarum and M. octospora) from analysis of the Paramarteilia canceri rooted ETS-28S Bayesian consensus tree (Figure 3). Moderate to high posterior probability values were seen for M. refringens sensu stricto and M. pararefringens (0.88 and 0.90, respectively). Despite samples from Norway branching separately from all other M. pararefringens with maximal support on the Bayesian consensus tree, bPTP analysis did not infer this branching pattern as a speciation event (posterior probability = 0.00). Posterior probability values from bPTP analysis for all branching events are shown in blue text on branches in Figure 3, with the posterior probabilities of likely speciation events suggested by the bPTP analysis indicated by an underlined posterior probability.
To investigate the validity of the 5 hypothesized Marteilia species inferred from bPTP, BPP analyses were performed. Bayesian species delimitation supported the guide tree with speciation probabilities of ≥0.95 on all nodes (Figure 6), a probability that other studies have considered as strong support for a speciation event (Leaché and Fujita, Reference Leaché and Fujita2010). M. refringens sensu stricto and M. pararefringens are supported by a speciation probability of 1.00, with different prior distributions for θ and τ0 not affecting this outcome (node E on Figure 6). Speciation probabilities of 1.00 were consistent for all tested combinations of priors for all nodes except for the M. cochillia and M. cocosarum speciation event. Speciation probabilities for M. cochillia and M. cocosarum increased from 0.96 to 0.99/1.00 using priors that assumed that these 2 species had recently diverged (node D on Figure 6).
Bayesian species delimitation results for Marteilia based on the full transcribed rRNA multiple sequence alignment and assuming a 5-species guide tree rooted to Paramarteilia canceri. The speciation probabilities are provided for each node under each combination of priors for θ and τ0: (i) θ ∼ G(1, 10) and τ0 ∼ G(1, 10); (ii) θ ∼ G(1, 10) and τ0 ∼ G(2, 2000); and (iii) θ ∼ G(2, 2000) and τ0 ∼ G(2, 2000).

Inferred host preference of M. pararefringens and M. refringens sensu stricto based on available sequence data
Phylogenetic trees in this study were generated using newly generated sequence data and all publicly available sequences. Once generated, the host from which the sequence was amplified was mapped onto the branches of the tree to determine if there were any differences between the 2 Marteilia species; the results of this are summarized in Table 2. One of the clearest differences between the 2 Marteilia species was that no publicly available M. refringens sensu stricto sequences from any part of the rRNA array exist from My. edulis or non-bivalve species, although a singular detection of M. refringens sensu stricto was reported from My. edulis by Le Roux et al. (Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001) using RFLP analysis. M. pararefringens has been frequently amplified from My. edulis, with 28.1% and 33.3% of all M. pararefringens ITS1 and ETS sequences amplified from this species.
Due to the low sample numbers for M. refringens sensu stricto, the ETS region did not provide enough information to compare potential hosts, so ITS1 was used as a proxy, with 121 sequences classified as M. pararefringens and 112 sequences classified as M. refringens sensu stricto. Although some M. pararefringens sequences were amplified from O. edulis (9.9%), a larger percentage of sequences amplified from this species branched with M. refringens sensu stricto (55.4%). There did not appear to be a clear Marteilia species preference in My. galloprovincialis with similar numbers of sequences amplified for each Marteilia species: 47.1% M. pararefringens and 38.4% M. refringens sensu stricto. As several of the sequences deposited on GenBank were generated by sequencing of clones, the exact number of animals that these sequences were derived from is unclear as multiple clones may have originated from the same Marteilia amplicon.
Discussion
In this study, we used high-throughput sequencing to generate full rRNA arrays from 9 independent Marteilia refringens sensu stricto (M. refringens as described in Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018)) and 28 independent M. pararefringens samples. Bayesian phylogenies generated from full rRNA arrays, and the most variable regions of the rRNA (ITS1 and ETS) produced mutually exclusive monophyletic clades for each species, with high to maximal support for each clade. These phylogenies continue to support the separate species status of M. refringens sensu stricto and M. pararefringens, as defined in Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018). The robust definition of the consistent nucleotide difference between M. refringens sensu stricto and M. pararefringens will allow the study of these parasites in isolation to determine their individual and combined impacts on bivalve culture in affected regions.
Genomic differences between M. refringens sensu stricto and M. pararefringens
Two publications in the early 2000s described consistent nucleotide differences observed in small sections of the rRNA array (ITS1 and ETS) between the 2 Marteilia species. In ITS1, Le Roux et al. (Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001) described 5 nucleotide differences over a 413 bp region, which were confirmed by López-Flores et al. (Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004). From the samples analysed in this study, we confirm that these differences are still robust for discriminating between the 2 species in PCR positive samples from 2002 to 2019 (Figure 2B). López-Flores et al. (Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004) described 2 differences between the 2 species over a 525 bp region of ETS. Here, we describe 6 consistent nucleotide differences across the whole region. Four of these nucleotide differences were amplified from a larger M. refringens sensu stricto and M. pararefringens sample set using a novel nested PCR assay. The similarity between M. refringens sensu stricto and M. pararefringens over the full ETS (99.60%) is higher than that of other closely related but distinct paramyxid species: Marteilia cochillia and M. cocosarum have 99.03% similarly and Paramarteilia canceri and P. orchestiae have 97.81% similarity (Hooper et al. Reference Hooper, Ward, Foster, Skujina, Ironside, Berney and Bass2023). However, when branch lengths are compared (Figure 5), M. refringens sensu stricto and M. pararefringens appear more distantly related to each other than M. cocosarum is to M. cochillia, closely related Marteilia species that are clearly delimited as separate species due their mode of infection (Skujina et al. Reference Skujina, Hooper, Bass, Feist, Bateman, Villalba, Carballal, Igleias, Cao, Ward, Ryder, Bignell, Kerr, Ross, Hazelgrove, Macarie, Prentice, King, Thorpe, Malham, Mckeown and Ironside2022).
Due to intragenomic variation, or coamplification of M. refringens sensu stricto and M. pararefringens, particularly in My. galloprovincialis, it has been necessary for some previous studies to clone PCR products to acquire a clean sequence. Many cloned sequences on NCBI have nucleotide differences that were not reproduced, even at low frequencies, in high-throughput or Sanger sequencing in this study. It is possible that these differences are artefacts from PCRs that have used standard Taq polymerases, which are prone to errors. These errors are unlikely to be seen when sequencing a PCR product directly, but errors may become more prevalent after further amplification of single ligated PCR products during the cloning process. We excluded 3 ITS1 sequences that were suspected to have sequencing artefacts from cloning from our analysis. To remove any ambiguity in sequence data, we recommend that high-fidelity, proof-reading polymerases should be used for sensitive downstream applications, such as looking at intraspecies variation and coinfections by cloning and Nanopore sequencing.
Preliminary data from Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) suggested that 28S and ITS2 may also be useful regions for discriminating between M. refringens sensu stricto and M. pararefringens; however, analysis of a larger number of samples in this study showed that both of these regions are not suitable as taxonomic markers. The 28S was shown to have only 2 consistently different nucleotides, located within a 100 bp region of the 5ʹ end, with the remainder of the gene being identical in sequence between species. Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) suggested that ITS2 may have up to 13 markers to distinguish between the 2 species; however, this was based on only 3 M. pararefringens and 2 M. refringens sensu stricto sequences. Here, we find that only one of those nucleotide differences can be reliably used based on a larger sample set. We also find that within samples, there are regions of the ITS2 that have intragenomically variable short tandem repeat regions, making it unsuitable as a marker region due to the requirement to clone PCR amplicons to this region to acquire sequence data.
Intraspecies variation
Short intragenomic variable regions are also seen in the 28S and 18S, as well as the spacer regions of the 2 Marteilia species. Within 28S, these regions are located at the 3ʹ end of the gene, and the same variability is observed in both M. refringens sensu stricto and M. pararefringens, irrespective of the location or species they were amplified from. A short ∼20 bp region of intragenomic variability is also present at the 5ʹ end of the 18S. Unlike in the 28S, this region only appears to be variable in M. pararefringens sequences. Interestingly, within this region of variability, M. pararefringens samples from Norway contain 3 nucleotide positions that are unique to samples from this geographical region and differentiate them from all other M. pararefringens and M. refringens sensu stricto sequences. These nucleotide differences were consistent across all samples from Norway, irrespective of the species they were amplified from or the year they were sampled. The differences are reflected in the full transcribed rRNA array (ETS-28S) phylogenetic tree (Figure 3), where samples from Norway branch robustly within the M. pararefringens clade, but separately from all other samples, with maximal support for the clade. Despite these differences appearing in a large number of samples from Norway, all samples used to generate full rRNA arrays originated from 1 estuarine pond (‘poll’) on the West coast of Norway. To investigate the geographic extent of this M. pararefringens population-level sequence variant in Norway we sequenced M. pararefringens positive mussels from 6 other regions in Norway recently found to have M. pararefringens-infected mussels (Bøgwald and Mortensen, Reference Bøgwald and Mortensen2024) and found that these differences were present in all samples, including 2 M. pararefringens sequences originating from My. trossulus (Bøgwald et al. Reference Bøgwald, Larsson and Mortensen2024). Hypotheses about how this sequence variant arose include (but are not limited to) M. pararefringens in Norway having undergone a population bottleneck event, or this sequence variant representing a northern European variant of M. pararefringens that may be present in other northern European countries but remains undetected due to the absence of substantial Marteilia-associated mortality events in mussels and thus limited surveillance.
Host preference
Previous publications disputed the species status of M. refringens sensu stricto and M. pararefringens due to the ability of each species to be detected in a wider host range than originally described (Balseiro et al. Reference Balseiro, Montes, Ceschia, Gestal, Novoa and Figueras2007). Marteilia refringens sensu stricto has been amplified from O. edulis (Le Roux et al. Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001) and My. galloprovincialis (López-Flores et al. Reference López-Flores, De La Herran, Garrido-Ramos, Navas, Ruiz-Rejon and Ruiz-Rejon2004). Sporadic detection of M. refringens sensu stricto has been seen in My. edulis (Le Roux et al. Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001), the clam Chamelea gallina (López-Flores et al. Reference López-Flores, Robles, Valencia, Grau, Villalba, De La Herran, Garrido-Ramos, Ruiz-Rejon, Ruiz-Rejon and Navas2008b), and the oysters Ostrea stentina (Elgharsalli et al. Reference Elgharsalli, Aloui-Bejaoui, Salah, Chollet, Joly, Robert, Couraleau and Arzul2013), Magallana gigas and Crassostrea corteziensis (Grijalva-Chon et al. Reference Grijalva-Chon, Castro-Longoria, Enríquez-Espinoza, Maeda-Martínez and Mendoza-Cano2015; Martínez-García et al. Reference Martínez-García, Grijalva-Chon, Castro-Longoria, Chávez-Villalba, Enríquez-Espinoza, Maeda-Martínez and Peña-Messina2017). M. pararefringens is commonly found in My. edulis and My. galloprovincialis (Le Roux et al. Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001). Sporadic detection of M. pararefringens has been seen in O. edulis (Le Roux et al. Reference Le Roux, Lorenzo, Peyret, Audemard, Figueras, Vivarès, Gouy and Berthe2001), the Pacific blue mussel My. trossulus (Bøgwald et al. Reference Bøgwald, Larsson and Mortensen2024), the clams Solen marginatus (López-Flores et al. Reference López-Flores, Garrido-Ramos, De La Herran, Ruiz-Rejón, Ruiz-Rejón and Navas2008a) and Ruditapes decussatus (Boyer et al. Reference Boyer, Chollet, Bonnet and Arzul2013) and the black-pygmy mussel Xenostrobus securis (Pascual et al. Reference Pascual, Villalba, Abollo, Garci, González, Nombela, Posada and Guerra2009). M. refringens sensu lato (based on detection without or with limited sequence data) has been found in the oysters Ostrea chilensis (Grizel et al. Reference Grizel, Comps, Raguenes, Leborgne, Tige and Martin1982), O. angasi (Bougrier et al. Reference Bougrier, Tige, Bachere and Grizel1986), O. puelchana (Pascual et al. Reference Pascual, Martin, Zampatti, Coatanea, Defossez and Robert1991), O. denselamellosa (Martin, Reference Martin1993), and Crassostrea virginica (Renault et al. Reference Renault, Cochennec and Chollet1995). Marteilia refringens sensu stricto has not been detected in Northern Europe, but preliminary data suggests that it may be present in the USA (Grijalva-Chon et al. Reference Grijalva-Chon, Castro-Longoria, Enríquez-Espinoza, Maeda-Martínez and Mendoza-Cano2015; Kerr et al. Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018). Marteilia pararefringens has been detected in both Northern and Southern Europe (Kerr et al. Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018). Marteilia refringens sensu lato and M. pararefringens have also been detected in a number of non-bivalve species including copepods, a cnidarian and a nematode (Audemard et al. Reference Audemard, Le, Barnaud, Collins, Sautour, Sauria, De, Coustau, Combes and Berthe2002; Carrasco et al. Reference Carrasco, López-Flores, Alcaraz, Furones, Berthe and Arzul2007; Boyer et al. Reference Boyer, Chollet, Bonnet and Arzul2013; Arzul et al. Reference Arzul, Chollet, Boyer, Bonnet, Gaillard, Baldi, Robert, Joly, Garcia and Bouchoucha2014; Bøgwald et al. Reference Bøgwald, Skår, Karlsbakk, Alfjorden, Feist, Bass and Mortensen2022).
In Table 2, we present the largest dataset to date, collated with respect to the species from which each Marteilia species was amplified. We emphasize that this table reports amplification of parasite DNA from host samples, but this does not confirm infection. Positive PCR results may be generated by extracellular parasite DNA, particularly in the case of filter-feeding hosts. Furthermore, interpretation of the results in Table 2 must take in account possible misidentification of Mytilus species in some studies, or as hybridization between Mytilus species is common (Nascimento-Schulze et al. Reference Nascimento-Schulze, Bean, Peñaloza, Paris, Whiting, Simon, Fraser, Houston, Bierne and Ellis2023), undetected hybrids may have been present in Marteilia detection studies. 38.4% of publicly available M. refringens sensu stricto sequences originated from My. galloprovincialis; however, only M. pararefringens was amplified from mussels in this study. The available sequences are derived from a limited number of studies, mostly from Spain, and using the cloning approach, so may overrepresent the detection of this species in My. galloprovincialis. There are also still no confirmed infections (histopathological identification) of a Mytilus species with M. refringens sensu stricto.
M. refringens sensu stricto from O. edulis comprised over 50% of all ITS1 sequences, whereas M. pararefringens from O. edulis only represents 9.9% of ITS1 sequences. However, although M. pararefringens has been detected by PCR from O. edulis, there are no confirmed reports of infection by histopathology, even – as reported in Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018) and Bøgwald and Mortensen (Reference Bøgwald and Mortensen2024) – at sites where M. pararefringens has been shown to be an abundant parasite of mussels (e.g. Bømlo, Norway). A recent study introduced naïve O. edulis to a site with a historically high prevalence of M. pararefringens, and despite high confirmed infections of M. pararefringens in My. edulis, there was no evidence of infection in introduced or native O. edulis, despite weak PCR detection in 10 of the 220 screened oysters (Bøgwald et al. Reference Bøgwald, Skår and Mortensen2025).
Defining M. refringens sensu stricto and M. pararefringens as distinct species
For M. refringens, the infraspecific ‘type’ terminology used for the past couple of decades is not taxonomically recognized, creating uncertainty and ambiguity, and should be avoided. The only infraspecific rank recognized by the ICZN that could be considered for M. refringens sensu stricto and M. pararefringens is subspecies, but this term is most appropriately used to distinguish morphologically distinct and geographically separate, but not reproductively isolated, populations within a species (Mayr, Reference Mayr1982; Monroe, Reference Monroe1982). These criteria are not applicable to M. refringens sensu stricto and M. pararefringens.
Following the recommendation that the bases for species definition be made on a case-by-case basis, particularly for microeukaryotes for which many existing species concepts are not applicable for a variety of reasons (Boenigk et al. Reference Boenigk, Ereshefsky, Hoef-Emden, Mallet and Bass2012), we assert that the robust phylogenetic mutual exclusivity of M. refringens sensu stricto and M. pararefringens and their different host preferences demonstrated by the new data and analyses presented in this paper, in combination with their differing (although overlapping) geographical ranges, provide more than sufficient evidence of their valid status as separate species. In addition to this, we performed coalescent-based species delimitation analysis on the complete transcribed region of the rRNA locus to confirm the validity of the 5 Marteilia species for which full rRNA arrays exist (M. refringens sensu stricto, M. pararefringens, M. cochillia, M. cocosarum, and M. octospora), suggested by phylogenetic and bPTP analysis. These analyses provided maximal support for the M. refringens sensu stricto and M. refringens speciation event across 3 prior distributions that assumed differing ancestral population sizes and divergence depth.
It is worth restating here: there is no consistent relationship between genetic differences between marker genes used for taxonomic discrimination and phenotypic divergence across eukaryotic taxa. The processes underlying sequence evolution in marker genes and those under adaptive selection are not the same, and the relative rates differ depending on the biology and ecology of the taxa involved at any point in time. Different genes/gene regions evolve at different rates in different taxa, and with different correspondences to phenotypic change. Recently, a genome for M. pararefringens has been generated (Hiltunen Thorén et al. Reference Hiltunen Thorén, Onuț-Brännström, Alfjorden, Pecková, Swords, Hooper, Holzer, Bass and Burki2024). When a comparable genome for M. refringens sensu stricto is available, further marker regions outside of the rRNA gene may be identified; however, investigation of other marker genes based on the M. pararefringens genome was outside of the scope of this study.
M. refringens sensu stricto and M. pararefringens are certainly closely related, but their degree of genetic (marker gene) divergence is commensurate with that of other species in the same genus, other microeukaryotes (e.g. the ciliate species Oxytricha granulifera and Ox. atypica, which differ by only 3 nucleotide sites over a 4127 bp region of the 18S to 28S rRNA gene array (Fan et al. Reference Fan, Yao, Luo, Dong, Xu, Chen, Bourland, Zhao and Huang2021)), the dinoflagellates Gymnodinium baicalense and G. corollarium (Annenkova et al. Reference Annenkova, Hansen and Rengefors2020), cercomonads (Bass et al. Reference Bass, Richards, Matthai, Marsh and Cavalier-Smith2007), fungi (for which multi-locus species differentiation is recommended as individual markers (e.g. ITS rDNA) may underestimate species diversity; (Matute and Sepúlveda, Reference Matute and Sepúlveda2019)), and even some macroorganisms, such as seaweed species in the genus Fucus (Leclerc et al. Reference Leclerc, Barriel, Lecointre and De Reviers1998).
Summary
We provide further robust evidence to support the separate species status of M. refringens sensu stricto and M. pararefringens: (1) Phylogenies generated from a larger number of complete rRNA arrays than in Kerr et al. (Reference Kerr, Ward, Stentiford, Alfjorden, Mortensen, Bignell, Feist, Villalba, Carballal, Cao, Arzul, Ryder and Bass2018), and all available sequences from the regions of the rRNA array with the most consistently different nucleotides between the 2 Marteilia species (ITS1), produce mutually exclusive, monophyletic clades for M. refringens sensu stricto and M. pararefringens; (2) the 2 Marteilia species branched with similar phylogenetic distances to other closely related Marteilia species i.e. M. cochillia, M. cocosarum and M. octospora; (3) coalescent-based species delimitation analyses on the complete transcribed the rRNA array confirmed the validity of 5 Marteilia species for which full rRNA arrays exist (M. refringens sensu stricto, M. pararefringens, M. cochillia, M. cocosarum, and M. octospora) and (4) analysis of sequences collated with respect to the species from which each Marteilia species was amplified showed a clearer host preference than studies had previously reported – no current sequence data for M. refringens sensu stricto has been amplified from Mytilus edulis, and M. pararefringens is amplified less frequently in Ostrea edulis than M. refringens sensu stricto. Use of the robust marker region in ITS1 allows clear distinction between the 2 species and should still be used for monitoring and research studies on both parasite species in order to clarify their respective host preferences, pathogenicity and life cycles.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182025100796.
Data availability
The full transcribed regions of the rRNA arrays generated in this study are deposited to GenBank under accession numbers PP549144-80, and ITS1 and ETS sequences are deposited under accessions PP544135-53 and PP549181-98, respectively.
Acknowledgements
Samples in this study were kindly provided by network of collaborators spanning the European range of the two species of Marteilia including from France (NRL for marine mollusc diseases, Ifremer La Tremblade), from Italy (NRL for mollusc diseases, Istituto Zooprofilattico Sperimentale delle Venezie, Adria), from Greece (NRL for mollusc diseases, Directorate of Veterinary Centre of Thessaloniki, Thessaloniki), and from Norway (NRL for mollusc diseases, Institute of Marine Research); and from Spain (Institute of Marine Research (IIM).
Author contributions
C.H., D.B. and I.A. conceived and designed the study; study materials were provided by D.S., I.A., L.C., M.B., S.M., R.A., A.F., G.M.W. and D.B.; investigation was carried out by C.H. and D.S.; data analysis and the initial manuscript was drafted was completed by C.H.; review and editing of the manuscript was completed by C.H., D.S., I.A., L.C., M.B., S.M., R.A., A.F., G.M.W. and D.B. Funding was acquisitioned by D.B. and I.A.
Financial support
This project was funded by Defra project AHPFFB2: Diagnostics and Emerging Disease, by the Genomics for Animal and Plant Disease Consortium (GAP-DC) provided by Defra and UK Research and Innovation (UKRI), by Ifremer and by EU DG SANTE through the European Reference Laboratory for mollusc diseases.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
Not applicable.