Introduction
Big cats are rare and elusive. Although they attract a significant amount of conservation funding and have been studied across their range, much remains unknown about them. Accurate species identification of big cats is important in research on matters such as poaching and trafficking, livestock depredation, ex situ breeding and dispersal events. Genetic tools are often used to identify species of large felids, but classical tools may be inadequate for this task. A recent study by Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024) that reported the presence of a Javan tiger Panthera tigris sondaica in West Java is an example of the inadequacy of these classical genetic tools. The Javan tiger was categorized as Extinct in 2008 (Jackson & Nowell, Reference Jackson and Nowell2008), and has not been detected since the 1990s. However, in 2019 a local resident reported seeing a tiger near a village in West Java, and one of the authors of Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024) collected a hair sample from the sighting location.
To determine whether this sighting could be the extinct Javan tiger, museum samples of Javan and Sumatran tigers Panthera tigris sumatrae were also collected and DNA was extracted from the hair and the museum samples (Wirdateti et al., Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024). The authors sequenced the cytochrome B (cytB) region of the samples and performed comparative phylogenetic analysis with previously published cytB sequences of tigers and leopards, concluding that the hair belonged to a Javan tiger. However, concerns were raised regarding the study (Emont, Reference Emont2024; Sui et al., Reference Sui, Yamaguchi, Liu, Xue, Sun, Nyhus and Luo2024). Here we reanalyse the sequences and repeat some of the experiments, to highlight the difficulties of studying big cat genetics and some potential solutions to these issues.
Phylogenetic tree reconstruction
We reanalysed the Javan tiger sequences in Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024) by making a phylogenetic tree that included additional cytB and nuclear copies of mitochondrial pseudogene (Numt) sequences downloaded from the National Center for Biotechnology Information database (Table 1). We aligned the sequences using MAFFT (Katoh et al., Reference Katoh, Misawa, Kuma and Miyata2002). We conducted three batches of analysis based on the length of sequences analysed and the type of sequences used. For Dataset 1 we removed sequences MH290773, AB211408–AB211411 and FJ403465 from the analysis because of the excess of missing data. We trimmed all regions with any missing data using Jalview (Waterhouse et al., Reference Waterhouse, Procter, Martin, Clamp and Barton2009). This retained 36 sequences with 265 bp of data for analysis. For Dataset 2 we removed sequences AB211408–AB211411, FJ895266, FJ403466, FJ403467, MH290773 and FJ403465 from the analysis because of the excess of missing data. We trimmed all regions with any missing data using Jalview, retaining 33 sequences with 971 bp of data for analysis. For Dataset 3 we aligned the cytB Numt sequences using MAFFT. We trimmed sites with missing data using Jalview. This retained 453 sites with eight sequences.
List of sequences (of domestic dog Canis familiaris, leopard Panthera pardus, lion Pathera leo and six subspecies of tiger Panthera tigris) from the National Center for Biotechnology Information database used in this analysis.

We built neighbour joining trees using MAFFT. We chose the default option for multiple sequence alignment. For building the tree we chose the conserved sites option, which retained 252 sites for Dataset 1, 937 sites for Dataset 2 and 240 sites for Dataset 3. We used raw differences for the substitution model and performed bootstrap resampling 1,000 times.
DNA extract re-sequencing
The Numt sequences were amplified because of stochastic binding of the primers to the Numt region instead of the mitochondrial cytB region (Sui et al., Reference Sui, Yamaguchi, Liu, Xue, Sun, Nyhus and Luo2024) in Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024); we attempted to rectify this through repeated PCR and sequencing (Fig. 1). In Round 1 we amplified the DNA extract using 3, 4, 5 and 7 μl templates in a 30 μl PCR, using the conditions described in Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024). The amplicons from these reactions were not visible in agarose gel electrophoresis. Thus, we further amplified the 3 and 5 μl products from all four PCRs (henceforth referred to as ‘amplicon templates’) in the previous step using the same PCR conditions in a 30 μl volume (leading to a total of eight reactions in this Round 2). Five of the eight reactions in Round 2 yielded bands on the agarose gel, four of which were from the 5 μl amplicon template and in the PCR in which the 7 μl original template was used. The 3 μl amplicon template in the second PCR also yielded bands. We amplified all templates on different days to avoid cross-contamination.
DNA amplification strategy used in this study. We performed two rounds of PCR. In Round 1 we used DNA extracts from samples as templates, and in Round 2 we used the amplicons from Round 1 as templates. The numbers on the arrows indicate the volumes of the templates used for each step. (Readers of the printed journal are referred to the online article for a colour version of this figure.)

The reaction with the 5 μl original template in Round 1 and the 5 μl amplicon template in Round 2 of PCR yielded the c. 900 bp-long DNA fragment from the test hair strand sequence that is presented here. We aligned these new sequences to the sequences listed in Table 1 using MAFFT and trimmed them using Jalview. We conducted two batches of analysis on the basis of this trimming: one that retained 38 sequences and 264 bases and another that retained 35 sequences and 907 bases.
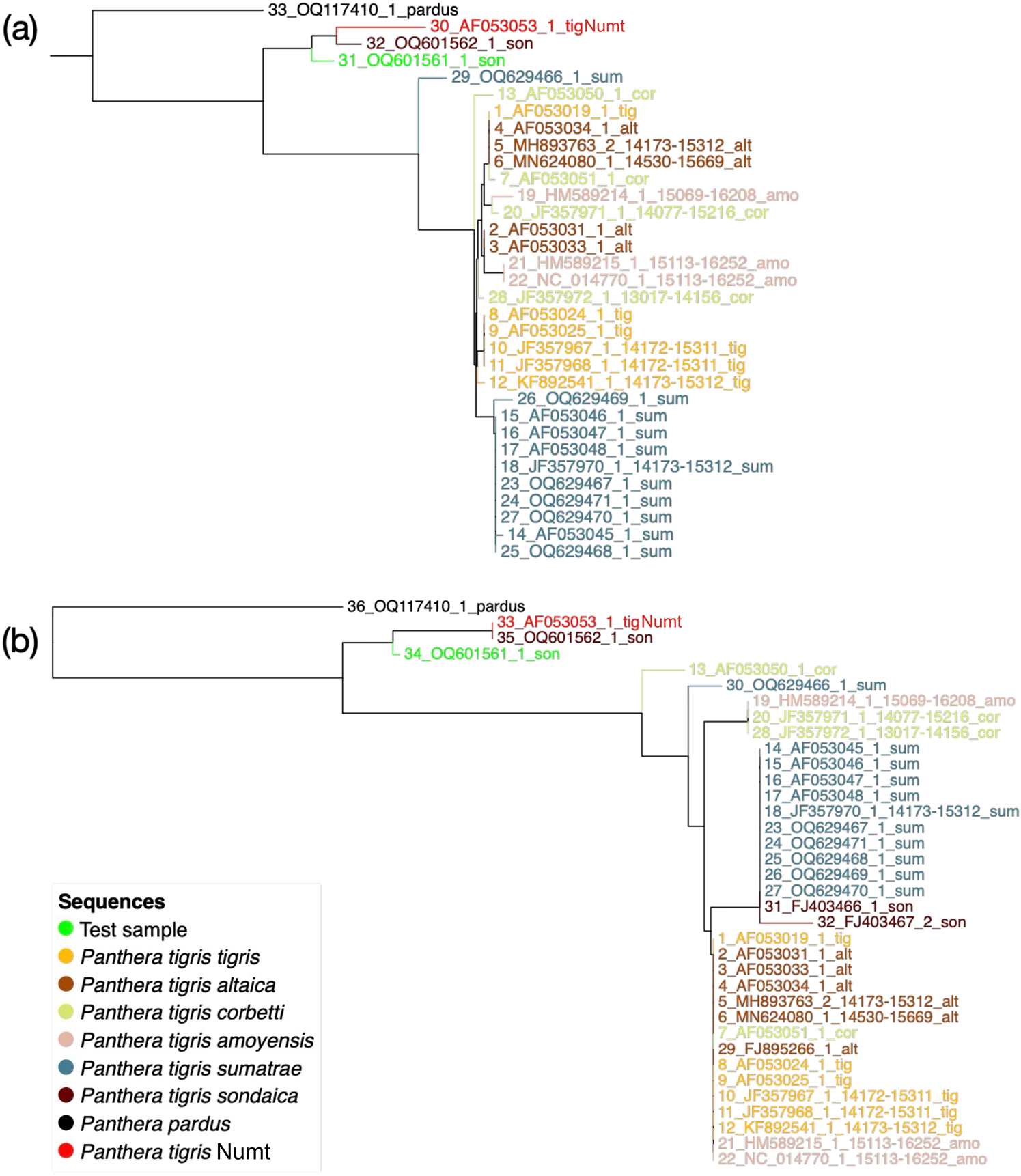
We reanalysed the sequences generated by Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024) alongside a nuclear copy of the cytB pseudogene (Numt) sequence of the Bengal tiger Panthera tigris tigris (AF053053.1) and the cytB sequences of several other tiger subspecies. The clustering of the putative Javan tiger sequence (test sample in Fig. 2) and the museum sample of the Javan tiger (OQ601562 in Fig. 2) with the Numt sequence revealed that the sequences generated for the samples were Numts and not the cytB regions that they were being compared to (Fig. 2).
Neighbour joining tree of the test sample (hair strand of putative Javan tiger Panthera tigris sondaica), the Javan tiger museum sample (OQ601562_1_son) and other tiger sequences using (a) 36 sequences and 252 sites and (b) 33 sequences and 937 sites. We included a sequence of the nuclear copy of the cytochrome B pseudogene (Numt) in the analysis. (Readers of the printed journal are referred to the online article for a colour version of this figure.)
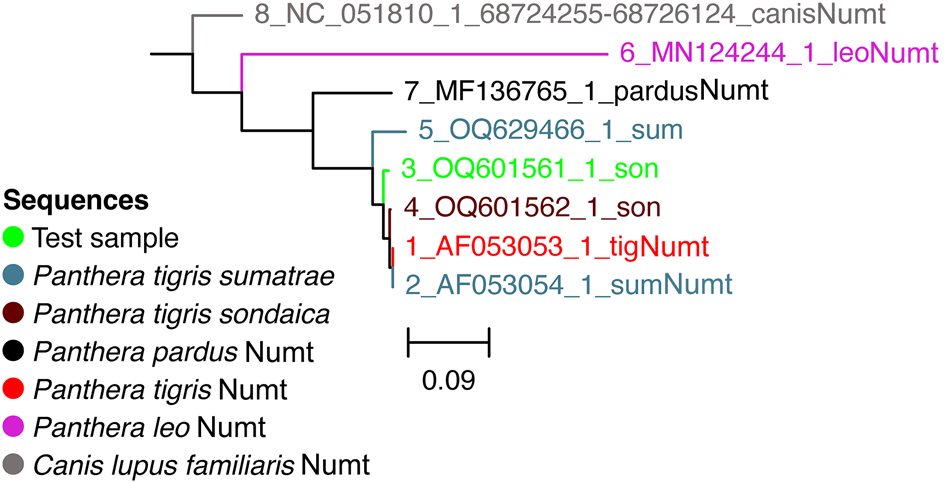
We further added Numt sequences of other Panthera species and the domestic dog Canis familiaris (as an outgroup) to test the potential of Numts for species identification (Fig. 3). Comparisons of the available annotated cytB Numt sequences of Panthera species in the National Center for Biotechnology Information database demonstrate that Numt sequences can be highly divergent within the Panthera genus (Fig. 3). We also observed that the lion Panthera leo cytB Numt sequence is more diverged from other cats and from a canid cytB Numt.
Neighbour joining tree of the test sample (hair strand of putative Javan tiger) and cytochrome B Numt sequences from other big cats and a domestic dog Canis familiaris. (Readers of the printed journal are referred to the online article for a colour version of this figure.)
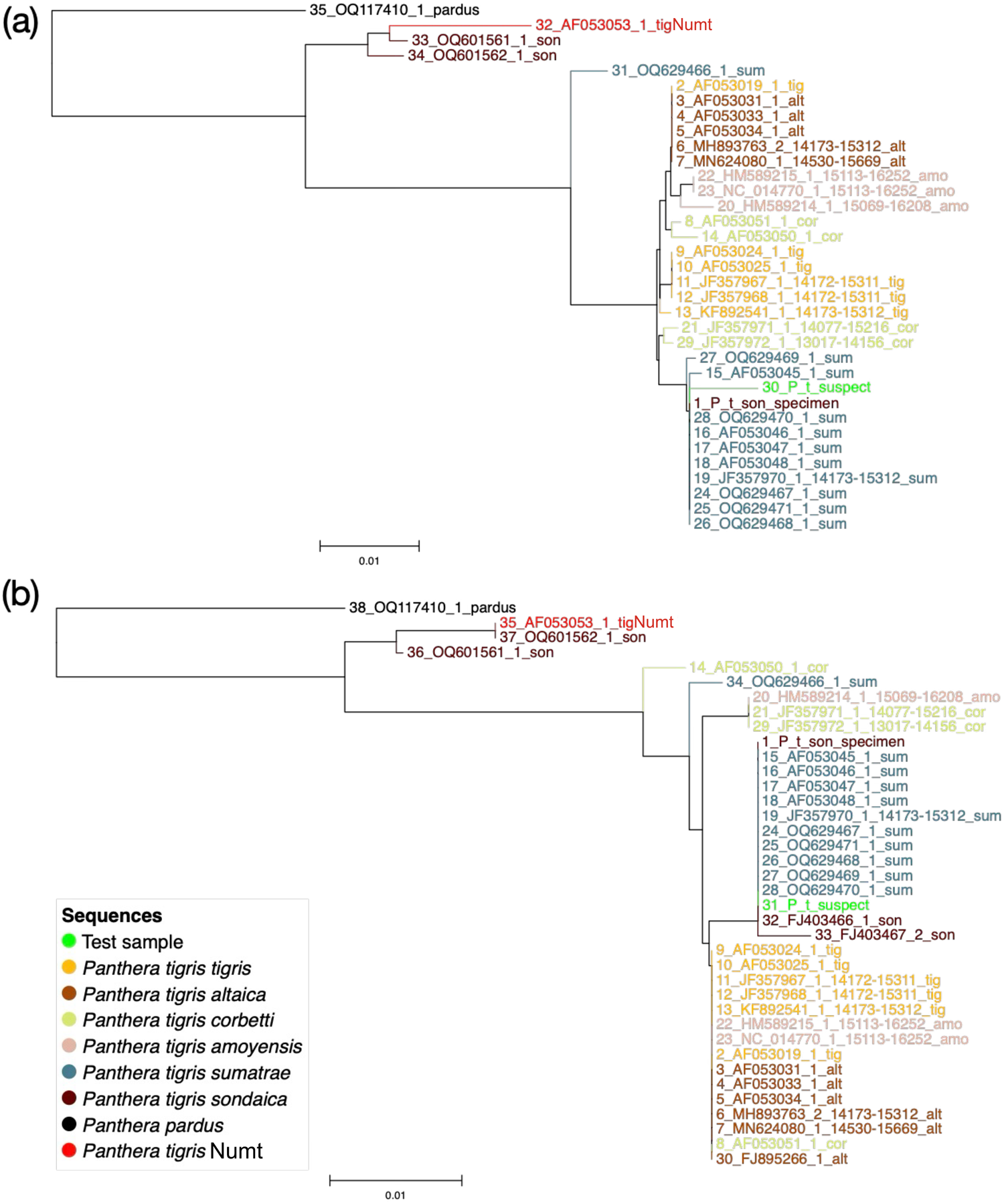
We re-sequenced DNA amplicons from the putative Javan tiger hair strand and the museum specimen of Javan tiger, and we retained sequences closely related to the cytB sequences of the other tigers (Fig. 4). The 3 μl template in Round 1 and the 5 μl amplicon template in Round 2 yielded the Numt sequence (Fig. 1). However, the mitochondrial cytB DNA does not have the power to distinguish between Sumatran and Javan tigers (Fig. 4).
Neighbour joining tree of the re-sequenced test sample (hair strand of putative Javan tiger, P_t_suspect), a museum sample of a Javan tiger (OQ601562_1_son) and other tiger sequences using (a) 38 sequences and 252 sites and (b) 35 sequences and 869 sites. (Readers of the printed journal are referred to the online article for a colour version of this figure.)
Discussion
Mitochondrial genomes are often used to delimit species and subspecies. However, they have several limitations, especially for big cats. In big cats nuclear copies of mitochondrial pseudogenes are a common problem in genetic investigations (Kim et al., Reference Kim, Antunes, Luo, Menninger, Nash, O'Brien and Johnson2006; Morgan et al., Reference Morgan, Ewart, Nguyen, Sitam, Ouitavon and Lightson2021). These regions evolve independently of the mitochondrial genome, have different rates of evolution and should not be compared directly. However, as they have sequence similarities, primers intended to amplify the mitochondrial copy of a gene might accidently amplify the nuclear copy and thus provide misleading results (Sui et al., Reference Sui, Yamaguchi, Liu, Xue, Sun, Nyhus and Luo2024).
Additionally, as mitochondrial DNA is inherited matrilineally, analysis of only mitochondrial sequences limits the ability to detect admixture events. For example, mitochondrial DNA could reveal the subspecies of the mother of a tiger sample but would indicate nothing about the father. This is especially relevant if samples of an admixed tiger are obtained for forensic analysis or captive breeding or for tracing the origins of a sample.
Extracting mitochondrial DNA is low cost, and this DNA is well preserved in non-invasive samples because of its abundance in cells compared to nuclear DNA. As mitochondrial DNA has been analysed experimentally for a long time, the protocols are familiar to most researchers. However, ancestry-informative single nucleotide polymorphism (SNP) panels (Khan et al., Reference Khan, Krishna, Ramakrishnan and Das2022), low-depth sequencing (Fuentes-Pardo & Ruzzante, Reference Fuentes-Pardo and Ruzzante2017), multiplex PCR panels (Natesh et al., Reference Natesh, Taylor, Truelove, Hadly, Palumbi, Petrov and Ramakrishnan2019) and pooled sequencing (Fuentes-Pardo & Ruzzante, Reference Fuentes-Pardo and Ruzzante2017) are low-cost alternatives that overcome the limitations of mitochondrial DNA from non-invasive samples. At present, the need for computational infrastructure and expertise, high start-up costs, high import costs of reagents and lack of access to high-throughput sequencers are major barriers to using these alternatives in many tropical countries (Khan & Tyagi, Reference Khan and Tyagi2021).
In this study, we confirm that the hair sample collected in West Java nests within the clade of Sundaland tigers, but we are unable to assign it to a subspecies. This is partially because there is no database of extinct tiger lineages. Although Sanger sequencing techniques are commonly used in several lower- and middle-income countries, the lack of databases remains a challenge. There are several specimens of the extinct Javan, Bali and Caspian Panthera tigris virgata tigers in museums (Yamaguchi et al., Reference Yamaguchi, Driscoll, Werdelin, Abramov, Csorba and Cuisin2013), but genetic resources from these lineages are lacking. This has limited our ability to explore the possibility of shared haplotypes in extant tigers and to determine the similarities and differences between extinct and extant lineages. Wilting et al. (Reference Wilting, Courtiol, Christiansen, Niedballa, Scharf and Orlando2015), for example, demonstrated the inability of short DNA sequences to resolve tiger subspecies, whereas whole-genome sequencing studies (Liu et al., Reference Liu, Sun, Driscoll, Miquelle, Xu and Martelli2018; Armstrong et al., Reference Armstrong, Khan, Taylor, Gouy, Greenbaum and Thiéry2021) have been successful in doing so, at least for extant lineages.
We highlight some of the challenges in studies of big cat genetics (Wirdateti et al., Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024): (1) There are few good-quality samples of big cats and use of non-invasive samples is the norm. Because of the abundance of mitochondrial DNA compared to nuclear DNA in cells, it has been common practice to analyse mitochondrial sequences. However, Numts are common in big cats and can lead to misrepresentation in analyses. (2) Despite the availability of methods for analysing whole genomes from non-invasive samples such as shed hair and faeces (Khan et al., Reference Khan, Patel, Bhattacharjee, Sharma, Chugani and Sivaraman2020; Tyagi et al., Reference Tyagi, Khan, Thatte and Ramakrishnan2022; Khan Reference Khan2023), the lack of sequencing facilities, computational infrastructure, DNA enrichment reagents or SNP panels render these tools inaccessible in some countries (Khan & Tyagi, Reference Khan and Tyagi2021). (3) There is a need to develop expertise in next-generation sequencing in biodiversity-rich tropical countries to support the discovery, conservation and management of the species in these regions. (4) Big cats and their parts are often subjects of forensic investigation in cases of trafficking or livestock depredation or in cases such as that of Wirdateti et al. (Reference Wirdateti, Yulianto, Raksasewu and Adriyanto2024). All such investigations could be affected by Numts if their analyses are only restricted to the mitogenome. There is a need for cost-effective SNP panels to be developed for ancestry determination and population assignment (Khan et al., Reference Khan, Krishna, Ramakrishnan and Das2022). (5) Databases such as GenBank, and research journals, need to insist and verify that the sequences they report are archived properly and made available, as the lack of genetic data hampers further studies in this field. For example, despite there being numerous genetic studies of tigers, these sequences are difficult to retrieve or use because of improper annotations or a lack of archiving; e.g. the sequences from Wilting et al. (Reference Wilting, Courtiol, Christiansen, Niedballa, Scharf and Orlando2015) need better annotation, and the genome assemblies from Armstrong et al. (Reference Armstrong, Khan, Taylor, Gouy, Greenbaum and Thiéry2021) and Zhang et al. (Reference Zhang, Lan, Lin, Fu, Yuan and Lin2023) need to be released, along with those of many other studies. Making these data available would facilitate progress in the science of large felid genetics.
Author contributions
Study conceptualization, analysis, writing: AK; coordination of author communication: SGA; laboratory experiments: YY; supervision: WW; revision: all authors.
Acknowledgements
This research received no specific grant from any funding agency or commercial or not-for-profit sectors.
Conflicts of interest
None.
Ethical standards
This research abided by the Oryx guidelines on ethical standards.
Data availability
The sequences generated will be submitted to the National Center for Biotechnology Information.






