Introduction
The phylum Acanthocephala represents obligatory parasites infesting intestines of vertebrates (Kennedy, Reference Kennedy2006). So far, 22 fish-parasitizing species belonging to the families Echinorhynchidae (mostly fish-parasitizing) and Pomphorhynchidae (exclusively fish-parasitizing) are reported for Europe (Gibson et al., Reference Gibson, Bray, Hunt, Georgiev, Scholz, Harris, Bakke, Pojmanska, Niewiadomska, Kostadinova, Tkach, Bain, Durette-Desset, Gibbons, Moravec, Petter, Dimitrova, Buchmann, Valtonen and de Jong2014). Due to their ambiguous morphological characters (Kennedy, Reference Kennedy2006), individual variability (Cleave, Reference Cleave1952) and the potential occurrence of cryptic species (e.g. Steinauer et al., Reference Steinauer, Nickol and Ortí2007; Zittel et al., Reference Zittel, Grabner, Wlecklik, Sures, Leese, Taraschewski and Weigand2018), taxonomic identification of acanthocephalans remains challenging. DNA barcoding has proven as an important tool for species identification, as seen in recent studies on the genus Pomphorhynchus that revealed previous contradictory determinations with a reliable DNA barcoding library (e.g. David et al., Reference David, Staentzel, Schlumberger, Perrot-Minnot, Beisel and Hardion2017; Perrot-Minnot et al., Reference Perrot-Minnot, Špakulová, Wattier, Kotlík, Düşen, Aydoğdu and Tougard2018; Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). Such inconsistencies are often attributed to strain formations in parasitic taxa due to different ecological demands and/or adaptations to host species (e.g. O'Mahony et al., Reference O'Mahony, Bradley, Kennedy and Holland2004; Steinauer et al., Reference Steinauer, Nickol and Ortí2007; Perrot-Minnot et al., Reference Perrot-Minnot, Špakulová, Wattier, Kotlík, Düşen, Aydoğdu and Tougard2018; Grabner et al., Reference Grabner, Doliwa, Bulantová, Horák and Sures2020). However, DNA barcode sequences of acanthocephalans in Europe are so far rare and the standard mitochondrial barcoding marker gene cytochrome c oxidase subunit 1 (COI) was rarely applied in the identification of this phylum in Europe. Most COI sequences of Acanthocephala were processed to investigate phylogenetic relationships. However, morphological data belonging to these sequences are lacking.
According to Kritscher (Reference Kritscher1985), who performed the latest classification of Acanthocephala in Austria 35 years ago, three classes, nine families, 18 genera and 37 species of Acanthocephala were recorded for Austria. Thereof, five species of four genera were recognized as fish parasites: Neoechinorhynchus rutili (Müller, 1780) a member of the class Eoacanthocephala, and Acanthocephalus anguillae (Müller, 1780), Acanthocephalus lucii (Müller, 1776), Echinorhynchus truttae Schrank, 1788 and Pomphorhynchus laevis (Müller, 1776), all belonging to the class Palaeacanthocephala (Kritscher, Reference Kritscher1985). In addition, Rydlo (Reference Rydlo1990) reported Pseudoechinorhynchus clavula (Dujardin, 1845) from Lota lota (L.) obtained from different waters in Austria, but mentioned that according to a personal comment of Prof O. N. Bauer from Russia it might be assigned to Echinorhynchus borealis Linstow, 1901 (syn. Echinorhynchus cinctulus Porta, 1905).
So far, there is only scarce data regarding the distribution and occurrence of fish-parasitizing Acanthocephala in Austria. In the course of the Austrian Barcode of Life (ABOL) initiative we performed an investigation of acanthocephalans from a selection of Austrian freshwater fish and identified them morphologically, followed by molecular genetic analyses. First results of that investigation were published by Reier et al. (Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) and focused on the genus Pomphorhynchus. That study revealed the occurrence of two additional species in Austria by combining morphological and molecular genetic methods: Pomphorhynchus tereticollis (Rudolphi, 1809) and Pomphorhynchus bosniacus Kiskároly & Čanković, 1967. Here we present data on all fish-parasitizing Acanthocephala detected during the ABOL survey. The sequences processed in this study were combined with those from our previous study on Pomphorhynchus (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) and compared to other published sequences from GenBank.
The aims of this study were 2-fold: (1) we aimed to get new insights on taxa occurring in Austrian freshwater fish, and to complement the recent knowledge about the occurrence and host range of acanthocephalans in Austria (and thus Central Europe). (2) We established reliable DNA barcodes for the ABOL and Barcode of Life (BOLD) databases, which can be applied in the identification of acanthocephalans. The DNA barcodes generated should simplify future identifications in Acanthocephala.
Materials and methods
Host and parasite sampling
In this study (combined with our previous study), 203 fish specimens or fish intestines from different rivers and lakes of Styria, Burgenland, Carinthia, Vienna and Lower Austria were provided by various working groups and persons in Austria (see Acknowledgements). Identifications of fish specimens were performed by the providers. Altogether eight fish species were included in this study (Table 1). Forty-eight of the dissected fish or intestines contained 302 helminths (242 Acanthocephala, 11 Nematoda, 3 Cestoda and 46 Trematoda) which were preserved in 80% ethanol and stored at 4°C. The current study provides an overview on acanthocephalans, while all other helminths will be presented elsewhere. Sixty-four specimens of the genus Pomphorhynchus were processed in our previous study morphologically and/or genetically. In this study additional 37 specimens of Acanthocephala were investigated morphologically and genetically. The taxa recorded in the course of this study as well as the host species are summarized in Table 1. All analysed specimens are deposited in the collection Evertebrata Varia of the Natural History Museum Vienna (museum collection numbers are given in Table 1). Table S1 of Supporting information lists all analysed specimens in the former and current study.
List of all genetically analysed specimens from this study

S, Styria; B, Burgenland; LA, Lower Austria.
IDs for lab, museum, BOLD and GenBank are given.
Morphological methods
Specimens were identified morphologically according to species-specific morphological characters mentioned in the literature (Lühe, Reference Lühe1911; Meyer, Reference Meyer1933; Petrochenko, Reference Petrochenko1956; Golvan, Reference Golvan1969; Grabda-Kazubska and Ejsymont, Reference Grabda-Kazubska and Ejsymont1969; Špakulová et al., Reference Špakulová, Perrot-Minnot and Neuhaus2011; Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). In Pomphorhynchus, morphological assignment at the species level is challenging due to morphological similarity and faint differentiating traits, especially regarding the species P. tereticollis, P. laevis and P. bosniacus (Špakulová et al., Reference Špakulová, Perrot-Minnot and Neuhaus2011; Hohenadler et al., Reference Hohenadler, Nachev, Thielen, Taraschewski, Grabner and Sures2017; Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). Reier et al. (Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) did thorough morphological investigations [including confocal laser scanning microscopy (CLSM) and 3-D reconstruction] and had already established reliable DNA barcodes of those species. Therefore, the taxonomic assignment for Austrian Pomphorhynchus species of the current study relies on those reference sequences.
Since a short-term treatment with glycerol has been shown to have no negative impact on DNA quality (Reier et al., Reference Reier, Sattmann and Haring2020), specimens were cleared in a mixture of glycerol and 80% ethanol (1:1) and incubated overnight at 38°C until ethanol was evaporated. Subsequently, specimens were transferred to a microscope slide with pure glycerol as mounting medium for investigations under the light microscope. Microphotographs were taken with a Nikon Eclipse Ni-U microscope equipped with a Nikon DS-Ri2 microscope camera and measurements were performed in NIS Elements software (Nikon Instruments Europe BV, Amsterdam, Netherlands). Length and width of body trunk, proboscis, proboscis receptaculum, lemnisci and testes as well as hook length were measured for specimens of Echinorhynchus cinctulus.
Analysis of autofluorescence spectra of different excitation wavelengths was conducted for E. cinctulus with the CLSM Leica TCS SP5 II (Leica Microsystems, Wetzlar, Germany). Image stacks were merged into maximum intensity projection images and further imported and processed in FIJI (Schindelin et al., Reference Schindelin, Arganda-Carreras, Frise, Kaynig, Longair, Pietzsch, Preibisch, Rueden, Saalfeld, Schmid, Tinevez, White, Hartenstein, Eliceiri, Tomancak and Cardona2012), or by volume rendering using AMIRA 6.11 (FEI; Hillsboro, Oregon, USA). The CLSM stacks are deposited on the Phaidra repository of the University of Vienna (Table S2 of Supporting information).
Molecular genetic methods
DNA extraction was carried out in a clean room using the QIAmp DNeasy Blood and Tissue Kit (QIAGEN, Hilden, Germany). For DNA barcoding a partial sequence of the mitochondrial cytochrome c oxidase unit 1 gene (COI) was used. Primer pairs for different taxa and the corresponding annealing temperatures as well as amplicon sizes are given in Table 2. Details of DNA extraction and polymerase chain reaction (PCR) amplification are the same as described in Reier et al. (Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). The PCR products were sequenced (both directions) by Microsynth (Balgach, Switzerland) using the PCR primers if not stated otherwise in Table 2.
Primer sets used in this study
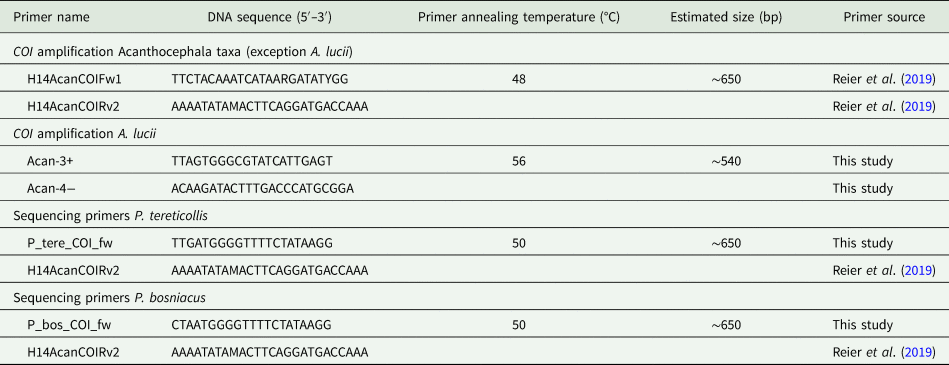

Sequences were processed in Geneious 2.10.3 (https://www.geneious.com). For understanding the position of examined genera among already published sequences we included COI sequences from GenBank of the following genera into our final alignment: Pomphorhynchus spp. [MK612497–MK612545 (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019)], Acanthocephalus spp. [KP261016 (Wayland et al., Reference Wayland, Vainio, Gibson, Herniou, Littlewood and Väinölä2015), AM039836, AM039837, AM039864–AM039866 (Benesh et al., Reference Benesh, Hasu, Suomalainen, Valtonen and Tiirola2006), DQ089718 (García-Varela and Nadler, Reference García-Varela and Nadler2006), MN416027–MN416030 (Amin et al., Reference Amin, Heckmann, Fiser, Zaksek, Herlyn and Kostanjsek2019)] and Echinorhynchus spp. [KP261013, KP261014, KP261017–KP261019 (Wayland et al., Reference Wayland, Vainio, Gibson, Herniou, Littlewood and Väinölä2015), AY218095 (Giribet et al., Reference Giribet, Sorensen, Funch, Kristensen and Sterrer2004), DQ089710 (García-Varela and Nadler, Reference García-Varela and Nadler2006)]. The rotifer Lecane bulla (Gosse, 1851) (HQ944350) was used as outgroup to root the tree. The final alignment contained 102 sequences and was trimmed to 544 bp due to different lengths of sequences. A neighbour joining (NJ) tree was calculated in MEGA 10.0.5 (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018). FigTree 1.4.3 (http://tree.bio.ed.ac.uk) was used for graphical processing of the tree and InkScape 0.92 (https://inkscape.org) for the final layout. Inter- and intraspecific P-distances (pairwise deletion) were calculated using MEGA 10.0.5 (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018).
Median joining (MJ) haplotype networks (Bandelt et al., Reference Bandelt, Forster and Rohl1999) were calculated for the species P. tereticollis, P. bosniacus, E. cinctulus, A. anguillae sensu lato (according to Amin et al., Reference Amin, Heckmann, Fiser, Zaksek, Herlyn and Kostanjsek2019) and A. lucii using PopART 1.7 software (http://www.popart.otago.ac.nz). For A. lucii, to allow inclusion of shorter GenBank sequences from Finland [AM039878, AM039880–AM039884 (Benesh et al., Reference Benesh, Hasu, Suomalainen, Valtonen and Tiirola2006)], the alignment was trimmed to 428 bp.
Results
Morphological determination
Acanthocephalans of three different fish species from Lower Austria and Burgenland were clearly assigned to the genus Acanthocephalus. Two species were differentiated morphologically as A. lucii and A. anguillae according to characters described in the literature (Meyer, Reference Meyer1933; Petrochenko, Reference Petrochenko1956; Golvan, Reference Golvan1969). Four specimens of A. anguillae [from Anguilla anguilla (L.)] were juvenile and exhibited a wrinkled body form. Nevertheless, species determination was possible based on the characteristic lateral extensions on the hooks of the proboscis and the constant number of six large hooks in the lateral rows. Only the last hook was considerably smaller without any extensions.
Sixteen specimens [obtained from A. anguilla, Gymnocephalus cernua (L.) and Perca fluviatilis (L.)] were determined as A. lucii. Morphological determination was straightforward since the analysed specimens were adults and corresponded to the descriptions and measurements known from the above-mentioned literature: we counted eight hooks in a row with wing-like shaped hook roots as shown in Meyer (Reference Meyer1933) as a typical character of A. lucii. Also, the hooks were longer than the hook roots and only the last row of hooks exhibited considerably smaller hooks.
One additional specimen (from A. anguilla) could not be identified due to its bad condition. It had a length of 5.15 mm and the proboscis was invaginated.
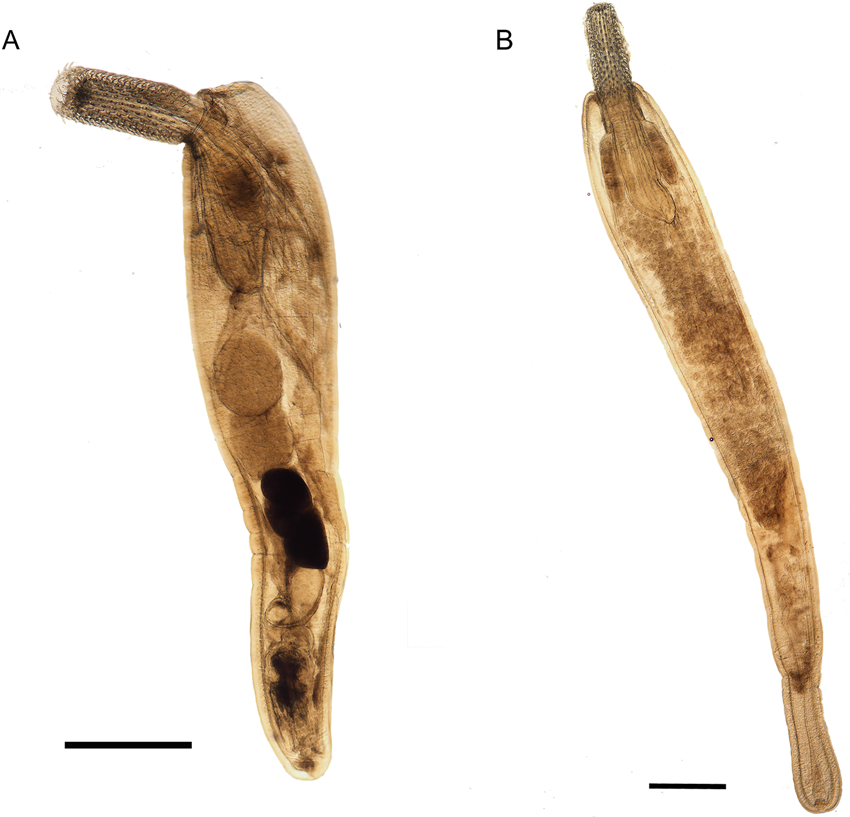
Eleven specimens were morphologically assigned to the genus Echinorhynchus. Five specimens from Styria obtained from L. lota and Zingel streber (Siebold 1863) were juvenile with an invaginated proboscis, therefore a thorough morphological investigation was impossible. The six specimens from Lower Austria obtained from Cottus gobio (L.) were adults (Fig. 1). Measurements of length and width of body trunk, proboscis, proboscis receptaculum, lemnisci and of testes of four males and seven females are given in Table 3. The appearance of the hooks was mostly uniform with simple hook roots without appendices. Furthermore, confocal laser scans were processed for two specimens (even a juvenile from L. lota) and the invaginated hooks and hook rows were counted. In these scans 12–14 hooks per row became apparent. The basal hooks were considerably smaller (26–34 μm) than the largest hooks in the centre of the proboscis (53–58 μm). Furthermore, we counted an additional rudimental hook on the ventral surface of the proboscis with a length of 14–15 μm. All measurements were in concordance with the thorough analysis of Echinorhynchus borealis Linstow, 1901 by Grabda-Kazubska and Ejsymont (Reference Grabda-Kazubska and Ejsymont1969). This species was later synonymized with Echinorhynchus cinctulus Porta, 1905, and therefore, we assigned the specimens (including the juveniles) to E. cinctulus.
Echinorhynchus cinctulus from C. gobio: (A) male and (B) female. Scale bar: 500 μm

Ranges of measurements conducted on seven female and four male specimens of E. cinctulus
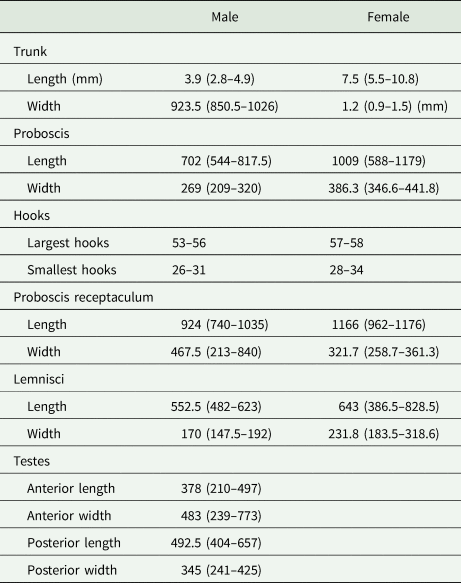
Range of measurements are given in parentheses. Measurements are in μm unless otherwise stated.
Five specimens were morphologically assigned to the genus Pomphorhynchus due to the genus-specific traits like the long neck and the bulb. Morphological assignment at the species level is rather challenging due to only faint distinguishing characteristics. Species assignment was performed according to Reier et al. (Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) combining morphological investigation and DNA barcodes. Three out of the five specimens were assigned to P. tereticollis [two from Oncorhynchus mykiss (Walbaum, 1792) and one from Barbatula barbatula (L.)], and two were determined as P. bosniacus (both from O. mykiss).
DNA analysis
The COI sequences of the 36 individuals analysed in the current study were combined with sequences from our earlier study and other sequences from GenBank. The final alignment of 544 bp comprised COI sequences of 102 specimens. The distances between lineages are illustrated in a NJ tree (Fig. 2). Nearly all major nodes had high bootstrap support. With one exception (one sequence from GenBank of Echinorhynchus salmonis appeared as a quite distinct lineage), genera were supported by high bootstrap values, too. While the NJ tree provides an overview, the sequences of the three genera detected in the current study (Fig. 2A, marked with different colours) are hereafter described in detail based on the corresponding MJ networks.
Phylogenetic relationships between genera of fish-parasitizing Acanthocephala based on a 544 bp COI dataset. (A) NJ tree comprising 102 COI sequences showing uncorrected P-distances between the genera Pomphorhynchus, Echinorhynchus and Acanthocephalus. Bootstrap values [1000 replicates, in %, only values with high support (>80%) are given] are shown next to the nodes. The dataset includes sequences generated in this study and sequences obtained from NCBI GenBank. Clades comprising sequences processed in this study are coloured. (B and C) MJ haplotype network of P. bosniacus and P. tereticollis including sequences of this study and our previous study. (D) MJ haplotype network of E. cinctulus showing two subclades (E1 and E2) of this study. (E) MJ haplotype network of A. anguillae sequences from Austria (this study) and Germany (GenBank), A. anguillae balkanicus and A. dirus. (D) MJ haplotype network (428 bp) of A. lucii (and two sequences of A. anguillae) from different localities. Subclade A1 comprises sequences generated in this study. Mutation steps are indicated with vertical lines. Black dots represent haplotypes missing in the study sampling.
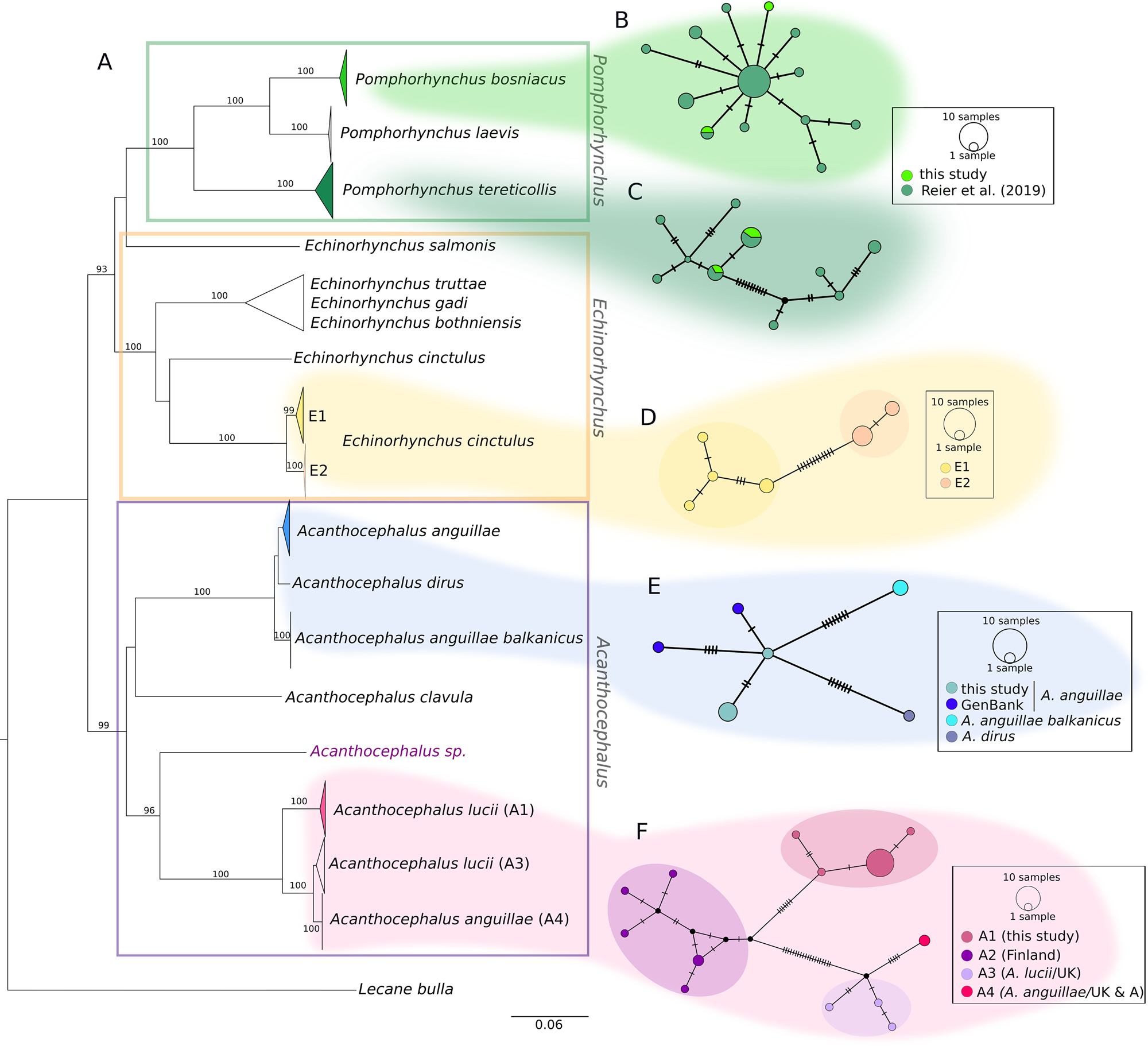
Two main clades and two lineages represented by one GenBank sequence each (one A. clavula, the other one not determined at the species level) were distinguished for the genus Acanthocephalus in the NJ tree (Fig. 2). The individuals analysed in the current study are found in the two main clades shown in detail in the networks (Fig. 2E and F). Interestingly, A. anguillae is found in both main clades.
The MJ network reveals a high variation for A. anguillae sensu lato (Fig. 2E). We identified six haplotypes. The network is star-like with the haplotype of one A. anguillae individual of this study in the centre (Fig. 2C). The other haplotype of A. anguillae detected in this study (shared by two individuals) is separated by two mutation steps. Sequences of A. anguillae from Germany are found in two haplotypes, separated from the central haplotype by three and one mutation steps, respectively (mean p- distance 0.5%, Table 4). Eight mutation steps (1.8%) separate the central haplotype from a sequence from GenBank assigned to the recently described subspecies Acanthocephalus anguillae balkanicus (Amin et al., Reference Amin, Heckmann, Fiser, Zaksek, Herlyn and Kostanjsek2019). Finally, a GenBank sequence of A. dirus is separated by seven mutation steps (mean genetic p-distance of 1.6%).
Genetic distances (p-distances in %) for the COI dataset of the genus Acanthocephalus
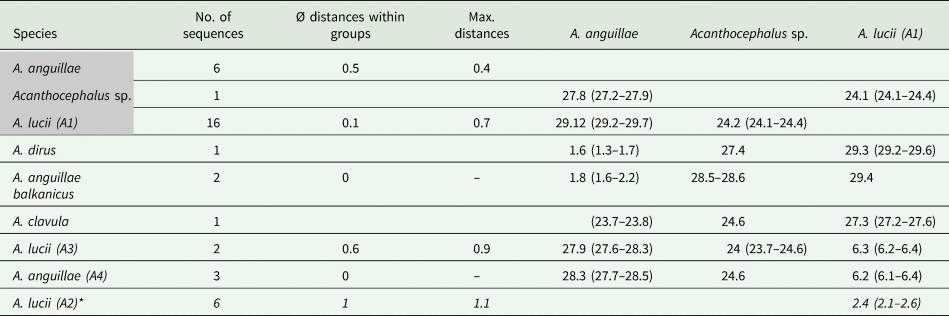
Groups containing specimens of this study are highlighted in grey. Sequences of A. lucii (A2) derived from GenBank based on a shorter alignment are indicated with an asterisk.
For the MJ network of the second main clade we included shorter GenBank sequences of A. lucii from Finland into the alignment, thus the network shows one more haplogroup (A2) compared to the tree (Fig. 2D). Altogether 13 haplotypes were found. Haplogroup A1 contains 16 sequences of this study that were morphologically assigned to A. lucii. It is separated by seven mutation steps (2.9%) from the Finnish A. lucii haplogroup (A2). The haplogroup comprising GenBank sequences assigned to A. lucii from the UK (A3) and a haplotype assigned to A. anguillae from UK (A4) is separated by 20 mutation steps from subclade A2 and by 27 steps from subclade A1. Mean p-distance between subclades A1 and A3 was 2.6% and between subclades A1 and A4 6.3%, while the mean p-distance between A3 and A4 was only 1.4% (Table 4).
One single sequence belonging to a specimen of this study (Acanthocephalus sp.) represented a distinct lineage not closely related to any clade of Acanthocephalus of the current study or GenBank, respectively (Fig. 2A).
Specimens of the current study and determined as E. cinctulus are found within one clade (E) which is divided into two subclades (E1 and E2). The sister group of this clade is a published sequence of E. cinctulus (KP261014, Table 4) which, however, is very distantly related (mean p-distance 20%) to the specimens analysed in the current study. Clade E can be divided into two subclades, one (subclade E1) containing specimens from Styria obtained from L. lota and Z. streber, and the other one (subclade E2) including specimens from Lower Austria from C. gobio. The haplotype network shows the relationships between the six haplotypes of clade E. The two haplogroups (E1 and E2) are separated by 12 mutation steps (mean distance 2.9%, Table 5). Mean genetic distances within the two haplogroups are given in Table 5.
Genetic distances (P-distances in %) for the COI dataset of the genus Echinorhynchus

Groups containing specimens of this study are highlighted in grey.
The other lineages of Echinorhynchus are formed by sequences from GenBank: besides, the highly distant E. cinctulus sequence, there is one clade comprising sequences of three different species of Echinorhynchus (E. bothniensis sensu lato, E. truttae and E. gadi). Furthermore, a single sequence from GenBank of Echinorhynchus salmonis Müller 1784 falls outside of the main Echinorhynchus clade (Fig. 2A).
Concerning Pomphorhynchus, results of most samples have been presented earlier (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). For the current study, five additional individuals were included, which were assigned to the species P. bosniacus and P. tereticollis. In the NJ tree (Fig. 2A–C) we present all Pomphorhynchus sequences from Austria available so far (this study as well as Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019), representing the three species P. bosniacus, P. tereticollis and P. laevis. The sequences of newly analysed individuals cluster in the tree and the MJ network with conspecific representatives (46 individuals) detected previously by Reier et al. (Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). Both species, P. bosniacus and P. tereticollis, were found at the same locality in Styria in consecutive years (Table 1). An additional specimen of P. tereticollis was obtained from B. barbatula from Burgenland. In the haplotype network of P. tereticollis the two haplotypes detected in this study are shared with specimens from our previous study from Styria (Fig. 2C). The two specimens belonging to P. bosniacus had two different haplotypes, one of which was shared with a specimen from our previous study originating from the river Danube (Vienna) (Fig. 2B). Mean intraspecific genetic p-distances are shown in Table 6, compared to Reier et al. (Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) they are lower as only sequences of specimens from Austria are included here.
Genetic distances (p-distances in %) for the COI dataset of the genus Pomphorhynchus

Groups containing specimens of this study are highlighted in grey.
Discussion
In the current study, we confirm the occurrence of six species of fish-parasitizing Acanthocephala in Austria including the first record of E. cinctulus in Austria. One species of the genus Acanthocephalus does not correspond – according to the DNA barcode – to any hitherto recorded species from Austria and public databases did not deliver any closely related sequence. Therefore, we could not determine it at the species level. Furthermore, we present the first record of P. bosniacus for Styria, a species which was so far only known from the Danube in Austria (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019).
Occurrences of Acanthocephala in Austria
Three species belonging to the genus Acanthocephalus were sampled and identified during this survey. Two of these species, A. anguillae and A. lucii, were also listed by Kritscher (Reference Kritscher1985). The third species, of which only one specimen was found, had its proboscis invaginated which made a morphological identification impossible. Since DNA sequence comparisons did not provided any hints, the specimen must remain undetermined and may represent a new species.
Specimens of A. anguillae clustered in the phylogenetic tree with A. dirus and A. a. balkanicus sequences retrieved from GenBank (Fig. 2A). Furthermore, we found a high genetic variation in the haplotypes of A. anguillae sensu lato (Fig. 2B). The clustering of A. dirus within this clade was unexpected, since it is not yet reported for Europe and known as the most common Acanthocephalus species in North America (Amin, Reference Amin1984, Reference Amin1985). Also, it has simple hook roots without extensions (e.g. Amin, Reference Amin1975, Reference Amin1984; Golvan, Reference Golvan1969), while A. anguillae possess two lateral extensions as observed in the specimens of this study. Therefore, the assignment of the specimens determined in this study as A. anguillae was straightforward. The clustering of A. dirus within the A. anguillae clade was already attributed to a probable misidentification by Amin et al. (Reference Amin, Heckmann, Fiser, Zaksek, Herlyn and Kostanjsek2019). The isopod Asellus aquaticus (L.) was reported as the host of the latter specimen (García-Varela and Nadler, Reference García-Varela and Nadler2006) and, since the life cycle of these helminths is not completed yet in the intermediate host, the specimen probably was at larval stage which might have impeded correct morphological identification. Another reason for misidentification might be due to morphological variation within A. anguillae sensu lato: the two lateral extensions were previously described as one of the most characteristic identification traits of the species (e.g. Meyer, Reference Meyer1933; Petrochenko, Reference Petrochenko1956), but they were not observed (or only rudimentary) in A. a. balkanicus, a recently described subspecies of A. anguillae parasitizing in Proteus anguinus Laurenti, 1768 (Amin et al., Reference Amin, Heckmann, Fiser, Zaksek, Herlyn and Kostanjsek2019). Therefore, the specimen from the USA could represent an additional lineage of A. anguillae sensu lato which lacks the lateral extensions, too. Also, the occurrence of A. anguillae in North America might be possible (Meyer, Reference Meyer1933; Petrochenko, Reference Petrochenko1956).
The common species A. lucii was collected in Lower Austria and in Burgenland. The shape of the hook roots as well as the anatomy corresponds to previous descriptions of A. lucii in the literature (Meyer, Reference Meyer1933; Petrochenko, Reference Petrochenko1956; Golvan, Reference Golvan1969). Sequences of the COI marker of A. lucii from Finland (approximately 450 bp; Benesh et al., Reference Benesh, Hasu, Suomalainen, Valtonen and Tiirola2006) were included in the MJ network analysis. They form a separate subclade (A2) closely related to subclade A1 (P-distance 2.5%), which supports their taxonomic assignment. However, NJ analysis revealed a considerably high genetic p-distance of 6.2% between our specimens and sequences derived from GenBank determined as A. lucii from the UK (subclade A3). Benesh et al. (Reference Benesh, Hasu, Suomalainen, Valtonen and Tiirola2006) mentioned the option of a misidentification of the English specimens or the existence of an English strain of A. lucii. The latter assumption is supported by an English sequence of another study determined as A. lucii (Wayland et al., Reference Wayland, Vainio, Gibson, Herniou, Littlewood and Väinölä2015), found in the same subclade (A3).
Interestingly, two GenBank sequences of A. anguillae from the UK and Austria (subclade A4) occurred within the A. lucii clade what had been explained by mitochondrial (mt) introgression due to the co-existence of the two species in one host (Benesh et al., Reference Benesh, Hasu, Suomalainen, Valtonen and Tiirola2006). Competitive behaviour between the two species has been reported, whereby A. lucii seemed to be the stronger competitor (Kennedy, Reference Kennedy1992). In Austria, a co-existence of A. anguillae and A. lucii in European Eels was observed in Lake Neusiedl in Illmitz (Burgenland) during a long-term survey (Schabuss et al., Reference Schabuss, Kennedy, Konecny, Grillitsch, Reckendorfer, Schiemer and Herzig2005). Results of the latter study did not indicate an overrepresentation of A. lucii in relation to A. anguillae, but an alternating presence of the two species in Illmitz with a higher abundance of A. anguillae in spring and a switch in autumn, when A. lucii becomes the dominant species (Schabuss et al., Reference Schabuss, Kennedy, Konecny, Grillitsch, Reckendorfer, Schiemer and Herzig2005). Anyhow, the high genetic distance between A. lucii and A. anguillae (mean p-distance of 29.3%) argues against an mt introgression between the two species. Therefore, it is more likely that the sequences of clade A4 were misidentified as A. anguillae. Also, the origin of the sequence available in GenBank, determined as A. anguillae from Austria would be interesting. As there is no information available on morphology of the published sequences in clade A3 and A4, their taxonomic assignment remains open.
Benesh et al. (Reference Benesh, Hasu, Suomalainen, Valtonen and Tiirola2006) mentioned the problem of nuclear pseudogenes (numts) of the mitochondrial COI gene in A. lucii. However, in the current study indications for numts were not encountered. The use of nested primers specific for A. lucii might have overcome such problems.
We found one species of Echinorhynchus in the current study. Of the two subclades of E. cinctulus, one occurred in the river Mur in Styria (E1) and the other one in Lower Austria in brooks around Lunz entering the river Ybbs (E2). The genetic p-distance of 2.9% between the two subclades E1 and E2 implies the existence of a phylogeographic structure. Considerable intraspecific variation was observed in some Acanthocephala and were especially investigated in the genus Pomphorhynchus (Dudiňák and Šnábel, Reference Dudiňák and Šnábel2001; O'Mahony et al., Reference O'Mahony, Bradley, Kennedy and Holland2004; Vardić Smrzlić et al., Reference Vardić Smrzlić, Valić, Kapetanović, Filipović Marijić, Gjurčević and Teskeredžić2015; Perrot-Minnot et al., Reference Perrot-Minnot, Špakulová, Wattier, Kotlík, Düşen, Aydoğdu and Tougard2018) as well as in Polymorphus minutus sensu lato (Zittel et al., Reference Zittel, Grabner, Wlecklik, Sures, Leese, Taraschewski and Weigand2018; Grabner et al., Reference Grabner, Doliwa, Bulantová, Horák and Sures2020). Also, intraspecific differences were revealed for species of Echinorhynchus, explained by post-glacial population bottlenecks (Väinölä et al., Reference Väinölä, Valtonen and Gibson1994). Unfortunately, due to the juvenile life stage of individuals of subclade E1, a more detailed morphological comparison of the two lineages was impossible. The examined specimens of lineage E2 were adult and their thorough morphological investigation indicated that the measurements were in concordance with the detailed description of E. cinctulus (formerly E. borealis; Grabda-Kazubska and Ejsymont, Reference Grabda-Kazubska and Ejsymont1969). Echinorhynchus borealis was synonymized with E. cinctulus since the name E. borealis was preoccupied by Echinorhynchus borealis Gmelin 1791 (Golvan, Reference Golvan1994; Amin, Reference Amin2013). The high genetic distance to a COI sequence from GenBank assigned to E. cinctulus (20%) raised doubts on whether these two distinct mitochondrial lineages belong to the same species. Cryptic species are known for various species of Acanthocephala (e.g. Steinauer et al., Reference Steinauer, Nickol and Ortí2007; Zittel et al., Reference Zittel, Grabner, Wlecklik, Sures, Leese, Taraschewski and Weigand2018). For example, the wide distances found between mitochondrial lineages of P. laevis (up to 20%) lead to the assumption of cryptic species (Perrot-Minnot et al., Reference Perrot-Minnot, Špakulová, Wattier, Kotlík, Düşen, Aydoğdu and Tougard2018; Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). Remarkable morphological variation within E. cinctulus was revealed in the work of Grabda-Kazubska and Ejsymont (Reference Grabda-Kazubska and Ejsymont1969) and it presently remains unknown whether the authors examined cryptic species or if the species is featured by a high intraspecific variation. The specimen of E. cinctulus processed by Wayland et al. (Reference Wayland, Vainio, Gibson, Herniou, Littlewood and Väinölä2015) was obtained from L. lota in Finland. Wayland et al. (Reference Wayland, Vainio, Gibson, Herniou, Littlewood and Väinölä2015) considered the systematics of the whole genus Echinorhynchus as controversial and recommended a total revision of Echinorhynchus sensu lato (Petrochenko, Reference Petrochenko1956; Golvan, Reference Golvan1969) by using an integrative taxonomic approach and to split the genus into smaller units, if monophyletic groups with distinct morphological traits can be distinguished. Our results confirm the need for a comprehensive survey of Echinorhynchus spp. throughout Europe gathering enough material to assess geographic variation regarding morphology, genetics as well as host range. Until now, E. truttae has been described as only representative of Echinorhynchus s. l. in Austria (Kritscher, Reference Kritscher1985). The taxonomic status of the specimens determined by Prof O. N. Bauer as E. borealis, as reported by Rydlo (Reference Rydlo1990) remains unconfirmed. The synonymy of E. borealis and E. cinctulus (Golvan, Reference Golvan1994; Amin, Reference Amin2013) indicates that the specimens determined by Bauer also belong to E. cinctulus. According to all available data, cryptic diversity within and taxonomic status of E. cinctulus remains open.
The identified specimens of Pomphorhynchus clustered in the known clades from previous studies (e.g. David et al., Reference David, Staentzel, Schlumberger, Perrot-Minnot, Beisel and Hardion2017; Perrot-Minnot et al., Reference Perrot-Minnot, Špakulová, Wattier, Kotlík, Düşen, Aydoğdu and Tougard2018; Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). The two species P. tereticollis and P. bosniacus were distinguished. Both species were found in a fish farm in consecutive years in rainbow trouts: P. bosniacus was obtained during 2017, whereas P. tereticollis was found during the year 2018. The question of whether both species co-exist in the fish farm or whether one species replaced the other in the following year could not be clarified. An explanation for the alternating occurrence of the two parasite species could be the purchase and stocking of fish from different suppliers during shortages (fish farmer, personal communication). Another scenario might be the introduction of cystacanths through intermediate hosts, since the fish farm is connected to white water. Interestingly, the specimens belonging to P. tereticollis shared haplotypes with specimens obtained from fish caught in the nearby Styrian rivers Mur and Sulm in former years (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019; Fig. 2C). This could be an indication that the specimens in the fish farm derived from white water. A replacement of dominant species was also reported in other studies (Schabuss et al., Reference Schabuss, Kennedy, Konecny, Grillitsch, Reckendorfer, Schiemer and Herzig2005). The specimens belonging to P. bosniacus were closely related to specimens from the Danube obtained in former years (Fig. 2B), which could be an indication that they were introduced into the fish farm. Nevertheless, it is important to note that one very common haplotype is shared by many specimens of P. bosniacus sampled along the Danube and, moreover, many closely related haplotypes (one to two mutation steps to this common haplotype) were detected in various European countries (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). Therefore, it is difficult to trace the origins of haplotypes of this species. A long-term study in correlation with data of fish stocking and a monitoring of the potential intermediate host fauna should be conducted to elucidate the alternating occurrences in fish farming.
Pomphorhynchus tereticollis was the most frequently detected species of Pomphorhynchus in Styria in a previous study (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019). Pomphorhynchus laevis which was previously detected in Styria (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) was not recorded in the current study. For P. bosniacus we report here the first record for Styria. Furthermore, we report the occurrence of P. tereticollis in Burgenland with B. barbatula as host species.
Reliability of DNA barcodes
Identification of helminths based on morphology is challenging, and the taxonomic expertise is mostly restricted to a few persons. Not without reason Acanthocephala are denoted as ‘a taxonomists nightmare’ (Kennedy, Reference Kennedy2006), showing high individual variability in proboscis size, and number and arrangement of the hooks on the proboscis (Cleave, Reference Cleave1952), which are respectively the only hard structures on their body (e.g. Brown, Reference Brown1987; Kennedy, Reference Kennedy2006). Reliable DNA barcodes can simplify species assignment of e.g. animals with invaginated proboscis or damaged specimens or of larval stages (Alcántar-Escalera et al., Reference Alcántar-Escalera, García-Varela, Vázquez-Domínguez and Pérez-Ponce de León2013). Meyer and Paulay (Reference Meyer and Paulay2005) criticized the solely usefulness of DNA barcoding in well-examined and efficient sampled taxa, which was supported in this study for the phylum Acanthocephala. It is a known issue that there exists a high number of sequences with wrong taxonomic determination in GenBank since quality checks are not standard and it is not mandatory to provide details on the origin of the specimen from which a sequence was obtained (Harris, Reference Harris2003). The BOLD data systems established more strict requirements (Ratnasingham and Hebert, Reference Ratnasingham and Hebert2007), but since data are also mined from GenBank, sequences of misidentified specimens accumulate there, too. This, in turn, could lead scientists to draw false conclusions from published sequences.
Indeed, our study indicated that DNA barcoding might be a promising tool in the identification of Acanthocephala, as shown for the genus Pomphoryhnchus. Yet, only a thorough morphological investigation followed by the determination of DNA barcodes (Reier et al., Reference Reier, Sattmann, Schwaha, Harl, Konecny and Haring2019) provided reliable basis for subsequent species identification. Inconsistencies and uncertain species assignment as shown for the genera Acanthocephalus and Echinorhynchus underline the need for more DNA sequence data of morphologically investigated and documented specimens. Therefore, DNA sequence data should be used with caution when no morphological data are provided.
Conclusion
Our study showed that DNA barcoding might be a promising tool in the identification of parasitic taxa, but for the taxon Acanthocephala we are still far from being able to use DNA barcoding for species identification. There still exist species with an uncertain species validity (Amin, Reference Amin2013) for which it is recommended to use a combined approach by investigating freshly collected material as well as museum collection material as references. Furthermore, our study further indicates cryptic species within taxa of Acanthocephala. Therefore, we suggest a revision of the phylum in Europe, based on an integrative taxonomic approach, and a broad sampling throughout Europe should be conducted to establish reliable DNA barcodes for Acanthocephala.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182020001316
Acknowledgements
We thank Julia Schindelar and Barbara Tautscher for technical assistance in the lab. We also express our gratitude to Stefan Koblmüller and Lukas Zangl of the University of Graz, Robert Konecny of the Environment Agency Austria, Michael Schabuss of Profisch OG, Vienna, Nadine Ebm of WasserCluster Lunz and Clemens Gumpinger of blattfisch e.U., Wels, for providing material and valuable discussions.
Financial support
Financial support was provided by the Austrian Federal Ministry of Education, Science and Research via the ABOL initiative (Austrian Barcode of Life; www.abol.ac.at). SR was supported by the FEMtech initiative of the Austrian Federal Ministry for Transport, Innovation and Technology (BMVIT). Open access funding provided by the University of Vienna.
Conflict of interest
None.
Ethical standards
None.










