Introduction
Cestodes of the family Taeniidae Ludwig, 1886, are parasites of terrestrial mammals that typically occur as adult tapeworms in predatory definitive hosts, with their larval stages developing in the prey. These parasitic organisms go through a life cycle that includes 2 hosts: carnivorous or omnivorous definitive hosts, which harbour the adult tapeworms in their small intestine, and herbivorous or omnivorous intermediate hosts, in which the larval stages develop in muscles, visceral organs, body cavities or the central nervous system, depending on the cestode species. The family Taeniidae, belonging to the order Cyclophyllidae, comprises 4 genera: Taenia Linnaeus, 1758; Echinococcus Rudolphi, 1801; Hydatigera Lamarck, 1816 and Versteria (Nakao et al., Reference Nakao, Lavikainen, Iwaki, Haukisalmi, Konyaev, Oku, Okamoto and Ito2013). Hydatigera, a genus recently revived by Nakao et al. (Reference Nakao, Lavikainen, Iwaki, Haukisalmi, Konyaev, Oku, Okamoto and Ito2013), comprises 4 recognized species: Hydatigera taeniaeformis s.s. (Batsch, 1786), Hydatigera kamiyai (Lavikainen et al., Reference Lavikainen, Iwaki, Haukisalmi, Konyaev, Casiraghi, Dokuchaev, Galimberti, Halajian, Henttonen, Ichikawa-Seki, Itagaki, Krivopalov, Meri, Morand, Näreaho, Olsson, Ribas, Terefe and Nakao2016), Hydatigera parva (Baer, 1924) and Hydatigera krepkogorski Schulz and Landa, 1934. Hydatigera tapeworms mature in the small intestine of felids and viverrids, while their larval stages, known as metacestodes or strobilocerci, develop in the tissues and body cavities of rodents (Nakao et al., Reference Nakao, Lavikainen, Iwaki, Haukisalmi, Konyaev, Oku, Okamoto and Ito2013; Lavikainen et al., Reference Lavikainen, Iwaki, Haukisalmi, Konyaev, Casiraghi, Dokuchaev, Galimberti, Halajian, Henttonen, Ichikawa-Seki, Itagaki, Krivopalov, Meri, Morand, Näreaho, Olsson, Ribas, Terefe and Nakao2016). Hydatigera taeniaeformis s.s. and H. kamiyai have been the focus of extensive research in recent years, leading to increased molecular studies and the discovery of cryptic species (Lavikainen et al., Reference Lavikainen, Iwaki, Haukisalmi, Konyaev, Casiraghi, Dokuchaev, Galimberti, Halajian, Henttonen, Ichikawa-Seki, Itagaki, Krivopalov, Meri, Morand, Näreaho, Olsson, Ribas, Terefe and Nakao2016). In contrast, much less attention has been paid to H. parva and H. krepkogorski, and very little is known about their genetic data, especially in the case of H. parva.
H. parva is thought to have originated in Africa and to have been introduced to the Iberian Peninsula (South-Western Europe) together with its final host, the common genet (Genetta genetta L. 1758; Carnivora: Viverridae). In this new environment, it continued using the same intermediate host, the wood mouse (Apodemus sylvaticus L. 1758), previously used in North Africa (Alvarez et al., Reference ‘Álvarez, Iglesias, Bos, Tojo and Sanmart’ín1990). The introduction of the common genet from North Africa to Europe probably occurred through several events facilitated by different civilizations, although the exact timing and pathways are still under debate (Gaubert et al., Reference Gaubert, Del Cerro, Centeno-Cuadros, Palomares, Fournier, Fonseca, Paillat and Godoy2015; Delibes et al., Reference Delibes, Centeno-Cuadros, Muxart, Delibes, Ramos-Fernández and Morales2017). Currently, stable populations of common genet are established in the Iberian Peninsula, Southwestern France and the Balearic Islands (Calzada, Reference Calzada, Palomo, Gisbert and Blanco2007; Jennings and Veron, Reference Jennings, Veron, Wilson and Mittermeier2009; Delibes et al., Reference Delibes, Centeno-Cuadros, Muxart, Delibes, Ramos-Fernández and Morales2017). The life cycle of H. parva in Europe appears to revolve primarily around the genet and the wood mouse, as this parasite is detected mostly in these hosts (Alvarez et al., Reference ‘Álvarez, Iglesias, Bos, Tojo and Sanmart’ín1990; Fuentes et al., Reference Fuentes, Cerezuela and Galan-Puchades2000, Reference Fuentes, Sáez, Trellis, Cruz, Sarmento, Casanova, Torres, Feliu and Esteban2003, Reference Fuentes, Fuentes, Sáez, Trelis, Galán-Puchades and Esteban2004; Torres et al., Reference Torres, Trelis, Espert, Ribas, Toledo, Casanova, Roman, Arrizabalga, Esteban and Feliu2003; Eira et al., Reference Eira, Torres, Vingada and Miquel2006; Millán and Casanova, Reference Millán and Casanova2007; Ribas et al., Reference Ribas, Feliu and Casanova2009), with occasional findings in the house mouse (Mus musculus) Schwarz and Schwarz, 1943 (Alvarez et al., Reference ‘Álvarez, Quinteiro-Alonso, Outeda-Macias and Sanmart’ín-Dur’án1987). Most of the papers referred to above have focused primarily on a wide range of parasite species and have only tangentially mentioned the presence of H. parva. Additionally, much of this research is over 15 years old, and genetic studies of H. parva in the Iberian Peninsula are limited to a single isolate (TpaSp) available in GenBank (Lavikainen et al., Reference Lavikainen, Haukisalmi, Lehtinen, Henttonen, Oksanen and Meri2008; Nakao et al., Reference Nakao, Lavikainen, Iwaki, Haukisalmi, Konyaev, Oku, Okamoto and Ito2013). One of the few recent studies focusing on the epidemiological and molecular findings of H. parva was conducted in Senegal representing the first application of genetic tools to characterize H. parva in autochthonous rodents on the African continent (Catalano et al., Reference Catalano, Bâ, Diouf, Léger, Verocai and Webster2019).
In general, information on the occurrence of H. parva in the Iberian Peninsula and elsewhere in Europe is limited and sporadic, with a notable lack of molecular-genetic studies. In this context, the aims of the present study were: (i) to determine the presence of H. parva in populations of autochthonous rodents in Spain; (ii) to investigate distribution patterns, prevalence rates and the influence of biotic factors on infection prevalence, in order to determine whether there is any correlation between transmission patterns of the parasite and specific habitats and host characteristics; and (iii) to perform a comprehensive analysis based on mtDNA genes (cox1, 12S DNA), which will contribute to a better understanding of its phylogenetic relationships and provide molecular data for further studies.
Material and methods
Field methods
Fieldwork was carried out in 2022 in riparian habitats along the Ebro River in La Cartuja Baja, Zaragoza province, Autonomous Region of Aragón (Northwest Spain; 41°36′16″N 0°49′21″W; Figure S1). Two types of habitats were selected: non-protected natural riparian habitats and agricultural fields close to the natural areas. These forests are characterized by ash trees (Fraxinus sp.), black poplars (Populus sp.), willow trees (Salix sp.), tamarisk shrubs (Tamarix gallica) and common reed grasses (Phragmites australis). Crops are mostly devoted to alfalfa, wheat and barley. Rodents were trapped using Sherman traps (H.B. Sherman Traps, Inc., Tallahassee, Florida). A mixture of wheat flour and vegetable oil was used as bait and a piece of hydrophobic cotton as nesting material. Traps were set up in the evening and inspected in the morning for 4 consecutive days. Animals were transferred without handling to a plastic bag and weighed using a Pesola scale to the nearest 0.5 g and anesthetized with a combination of ketamine (Domtor©, Esteve, Barcelona, Spain) and medetomidine (Imalgene©, Merial, Barcelona, Spain) (Chirife and Millán, Reference Chirife and Millán2014). Animals were then euthanized by bleeding and necropsied in detail, sexed, aged and measured. A total of 341 individuals of 2 different species were included in the study: 257 Algerian mice (Mus spretus Lataste, 1883) and 84 wood mice. Parasitological specimens were preserved in 90% ethanol and transported to the laboratory of the Department of Genetic Research, Institute for Biological Research ‘Siniša Stanković’-National Institute of the Republic of Serbia, University of Belgrade, Serbia. The import of samples to Serbia was authorized by the Veterinary Directorate of the Ministry of Agriculture, Forestry and Water Management of the Republic of Serbia (permit number: 000491352 2023 14841 004 000 000 001-02).
DNA processing: extraction, amplification and sequencing
Genomic DNA was extracted from each parasite specimen using the AccuPrep® Genomic DNA Extraction Kit (Bioneer Corporation, Daejeon, South Korea) following the manufacturer’s instructions, preceded by an overnight digestion with proteinase K and RNase. The mitochondrial cytochrome c oxidase subunit 1 (cox1) gene fragment (approx. 400 bp) was amplified using the primers JB3 (5′-TTT TTT GGG CAT CCT GAG GTT TAT-3′) and JB45 (5′-TAA AGAAAG AAC ATA ATG AAA ATG-3′) (Bowles et al., Reference Bowles, Blair and Mcmanus1992). Additionally, a fragment of approximately 350 bp of mitochondrial (mt) 12S rDNA was amplified using the primers P60for (5′-TTA AGA TAT ATG TGG TAC AGG ATT AGA TAC CC-3′) and P375rev (5′-AAC CGA GGG TGACGG GCG GTG TGT ACC-3′) (von Nickisch-rosenegk et al., Reference von Nickisch-rosenegk, Silva-Gonzalez and Lucius1999). All PCRs were performed in a final reaction volume of 25 μL, which included 2.5 μL (10× PCR Dream Taq buffer), 1.25 μL dNTPs (10 mM), 1.25 μL of each primer (20 μM), 0.2 μL (1 U) Dream Taq polymerase (Thermo Fisher Scientific), genomic DNA extract (30–100 ng) and RNAse free water up to the final volume. The PCR conditions for both molecular markers were identical: an initial denaturation at 94 °C for 3 min; followed by 40 cycles of 30 sec at 94 °C, 1 min at 56 °C and 45 sec at 72 °C; and a final extension at 72 °C for 2 min. The PCR amplification products were separated by agarose gel electrophoresis, stained with Midori Green Direct (Nippon Genetics Europe) and visualized using a Bio-Rad Gel Doc 1000 (Bio-Rad Laboratories, Hercules, California, USA). Subsequently, the remaining PCR products were purified using the ExS-Pure™ Enzymatic PCR Cleanup Kit (NimaGen) and further used as templates in the sequencing reaction with the BrilliantDye® v3.1 Dye-Terminator Cycle Sequencing Kit (NimaGen), both following the manufacturer’s instructions. The sequencing reaction was followed by purification through ethanol precipitation, as suggested in the Applied Biosystems Chemistry Guide|Third Edition – DNA Sequencing by Capillary Electrophoresis (p. 76). Finally, capillary electrophoresis was performed using the SeqStudio Genetic Analyzer (Applied Biosystems – Thermo Fisher Scientific).
Molecular, phylogenetic and genetic analysis
The DNA sequences obtained were compared with existing sequences in GenBank© using the NCBI BLAST© search tool (www.blast.ncbi.nlm.nih.gov) to confirm species identity. Subsequently, the sequences of H. parva (cox1) (344 bp) were subjected to population genetic and phylogenetic analysis with previously published sequences. Alignment and visual inspection were performed using Clustal W in MEGA software (v.11), and sequences were trimmed to a uniform length of 328 bp and 282 bp depending on the analysis type. A maximum likelihood tree was constructed with MEGA (v.11) using the HKY + G model. DnaSP 6.12.03 was used to analyse genetic diversity (including haplotype number, haplotype diversity and nucleotide diversity) and neutrality indices (Fu’s Fs and Tajima’s D) (Rozas et al., Reference Rozas, Ferrer-Mata, Sánchez-delbarrio, Guirao-Rico, Librado, Ramos-Onsins and Sánchez-Gracia2017). PopART 1.7 was used to create a median join network containing 25 cox1 nucleotide sequences from our study and 7 sequences from the GenBank© database (Bandelt et al., Reference Bandelt, Forster and Rohl1999). Pairwise nucleotide sequence divergences were calculated using the Kimura 2 parameter (K2P) (Kimura, Reference Kimura1980) model with a gamma value of 0.5 in MEGA (v.11) software.
Statistical analysis
A binary logistic regression model was used to evaluate the association between the probability of a wood mouse being parasitized and the season, type of habitat (agricultural vs natural), and the mouse’s sex and age. A backward stepwise elimination method was used in which the least significant variable was removed after each step. Statistical analysis was performed using IBM SPSS Statistics 26 for Windows® (IBM Corporation, Route 100, Somers, New York, USA). A significance level of P < 0.05 was considered statistically significant.
Results
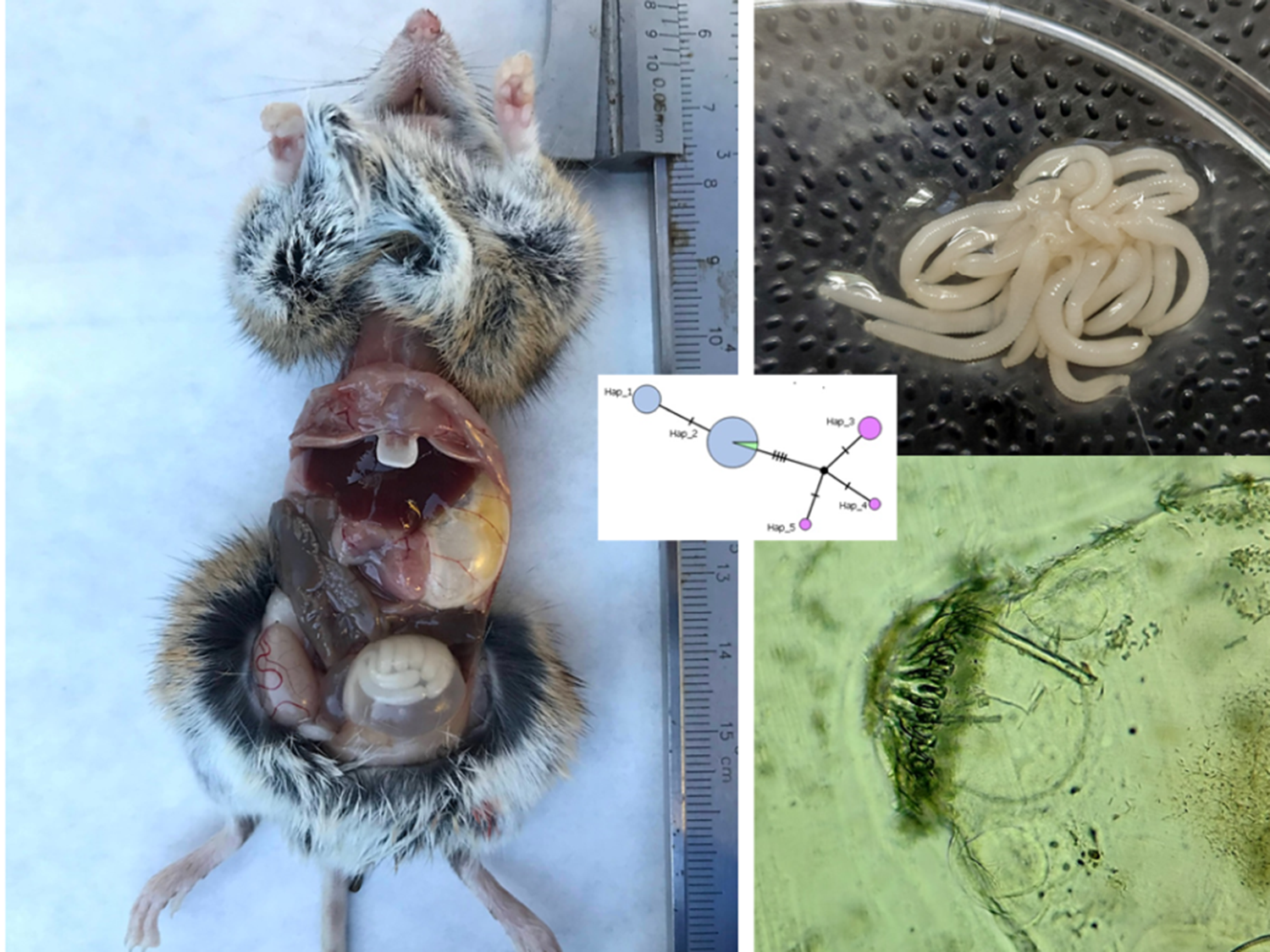
Encysted polycephalic larvae of H. parva were found in the body cavity of 27 wood mice (32.14%) (Figure 1) and a single Algerian mouse (0.39%). Prevalence was significantly higher in wood mouse (χ 2 = 84.69, df = 1, p < 0.001). All observed cysts showed the characteristic morphology of classical metacestodes, and the opening revealed the typical shape of polycephalic strobilocerci. Nine wood mice had 2 cysts, one had 3, and the remaining had 1 cyst. The cyst from the Algerian mouse had no viable larvae. The statistical analysis did not reveal any association between the presence of cysts in wood mouse and the type of habitat, the season or the animal’s sex and age.
A wood mouse with an abdominal cyst (left); larvae extracted from the cyst (top right); a microscope image of the rostellum (bottom right).
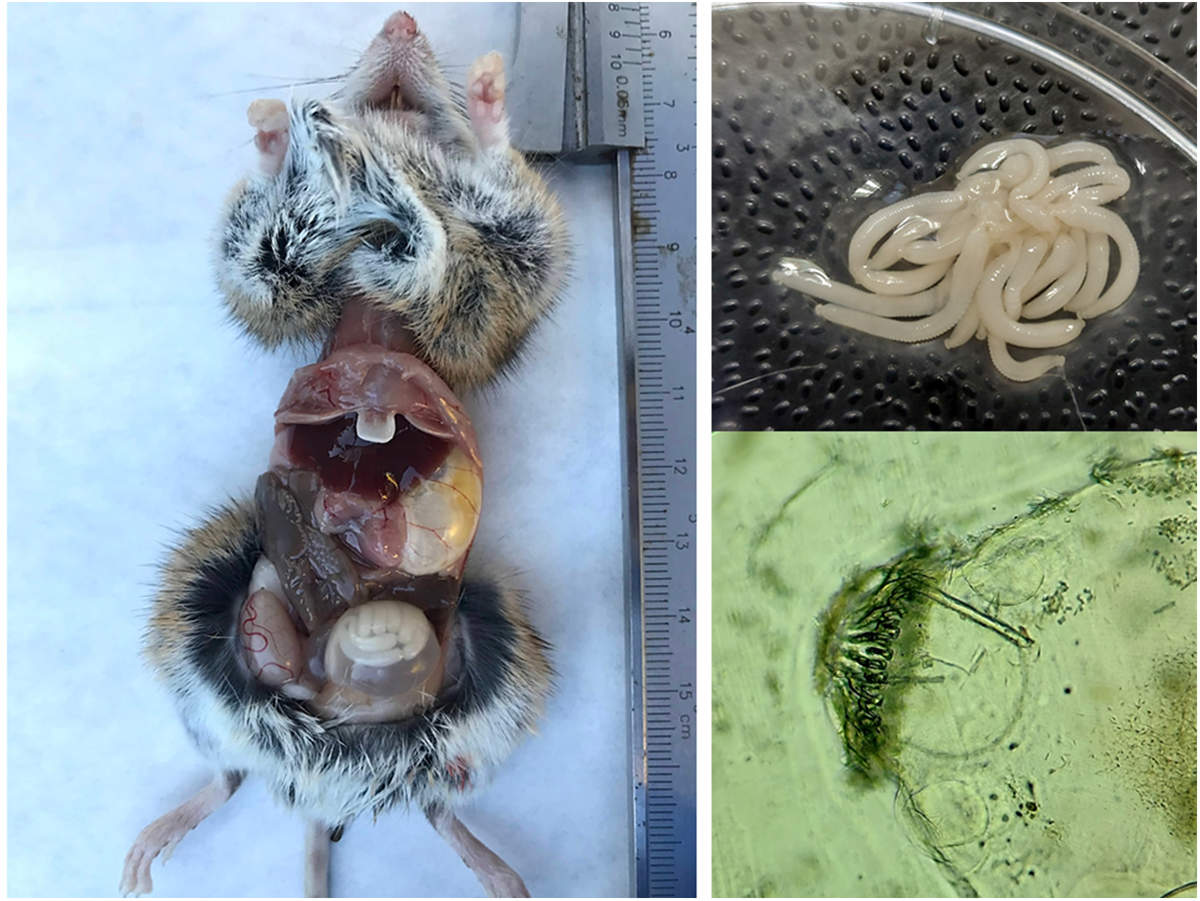

Of the 28 genetically analysed cysts, cox1 sequences (344 bp) were successfully amplified in 25 samples, while 3 samples were confirmed using the 12S rDNA gene, which was amplified only when cox1 amplification was unsuccessful. All 25 cox1 sequences and three 12S rDNA sequences of H. parva showed 99.31–100% identity with the sequence NC_021141 from GenBank© originating from a wood mouse from Galicia, Northwestern Spain. Genetic analysis of 25 cox1 sequences revealed a G + C content of 31.1%. A single variable site and 1 mutation were identified among the total positions analysed. Nucleotide diversity (π) was calculated to be 0.00110, with an average of 0.38000 nucleotide differences (k) observed. Haplotype analysis revealed 2 distinct haplotypes, resulting in a haplotype diversity (Hd) of 0.380. Statistical tests, including Tajima’s D (0.72124) and Fu’s Fs (1.032), showed no significant deviation from neutrality (Table 1).
Genetic diversity metrics of cox1 (344 bp) sequences from this study
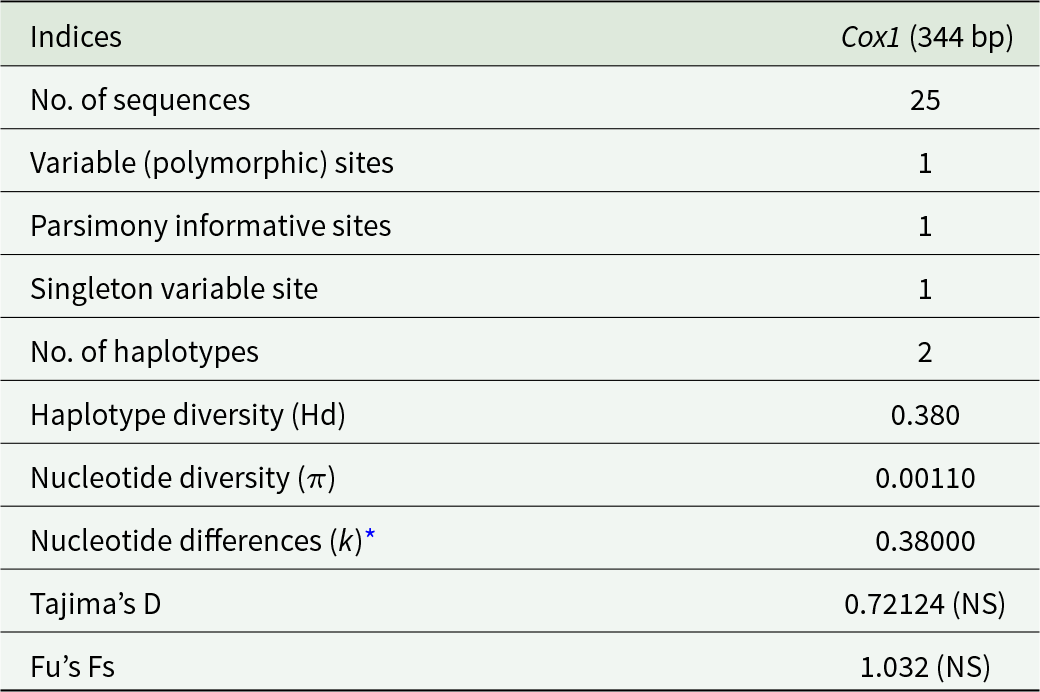
NS, not significant;
* Average number of pairwise nucleotide differences (k).
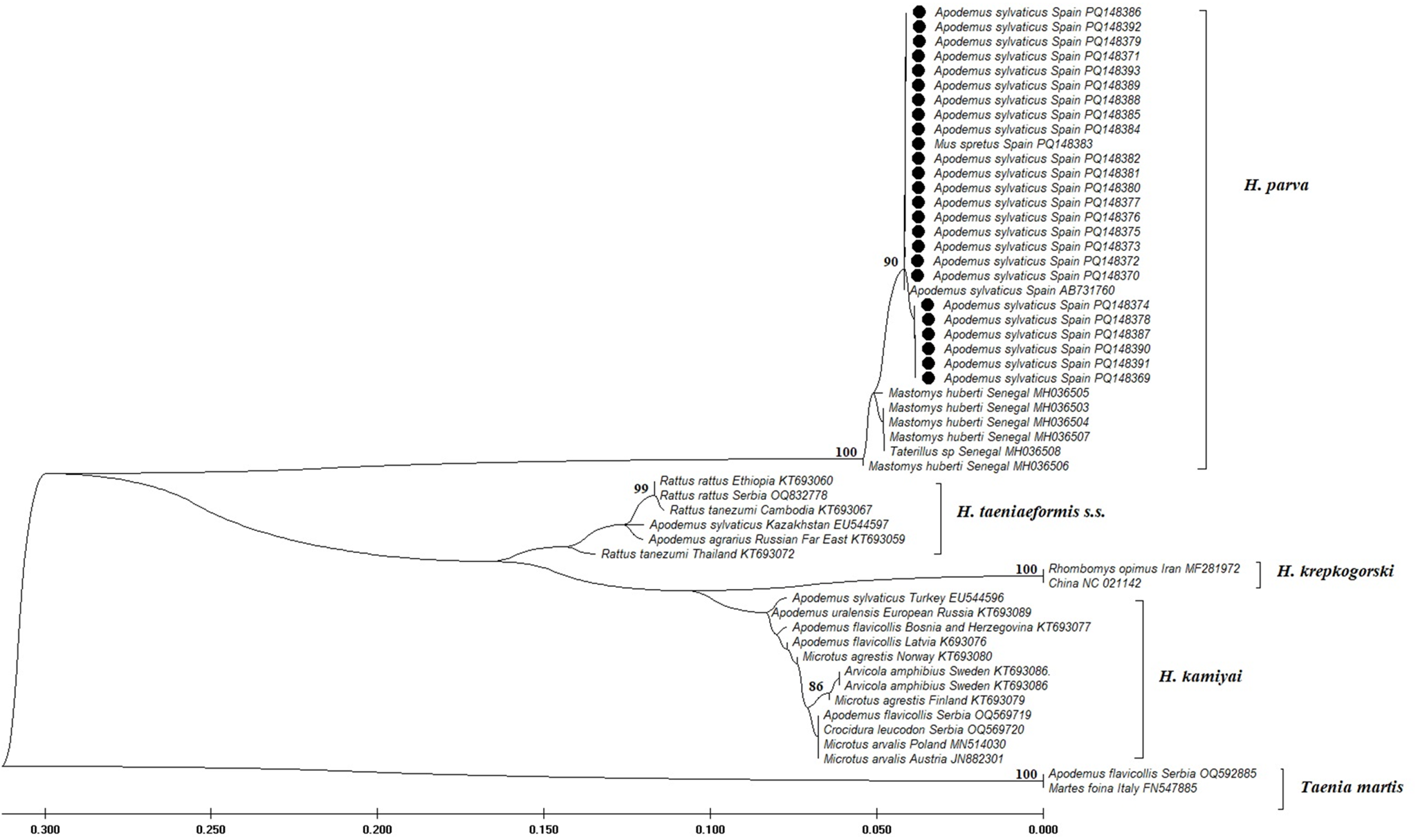
Twenty-five cox1 sequences from our study and seven sequences available in GenBank© (six from Senegal and one from Spain) were all truncated to 328 bp. According to the median-joining network analysis, these sequences revealed a total of 5 haplotypes. The single isolate from Spain in GenBank clustered with our samples, while the 6 sequences from Senegal resulted in 3 distinct haplotypes. There were 5–7 mutational steps between these 2 regional groups of haplotypes (Figure 2). Inter-population pairwise divergence analysis of H. parva samples from Spain revealed distance variations up to 0.30%, while regional analysis with samples from Africa (Senegal) indicated distances between 1.6% and 1.9%. At the inter-species level, our H. parva samples showed significant differences from congeneric samples: 17.50–18.64% from H. kamiyai, 16.46–20.41% from H. taeniaeformis s.s. and 19.19–19.74% from H. krepkogorski (Table 2). Phylogenetic analyses supported these results by clustering our samples with H. parva and identified the distances from other species within the genus Hydatigera (Figure 3).
Median-joining network of H. parva isolates from our study compared to isolates from Africa (Senegal) and an isolate from Spain from GenBank (MH036503-MH036508; AB731760), based on cox1 gene sequences (328 bp).

Phylogenetic tree of H. parva based on 282 bp cox1 gene sequences. A maximum likelihood tree was constructed using MEGA (v.11) with the HKY + G model. Values >85% are indicated. Sequences of other Hydatigera species from GenBank© studies are included in the tree, with Taenia martis used as an outgroup.
Pairwise genetic distances between H. parva and other Hydatigera species (sequences 282 bp)
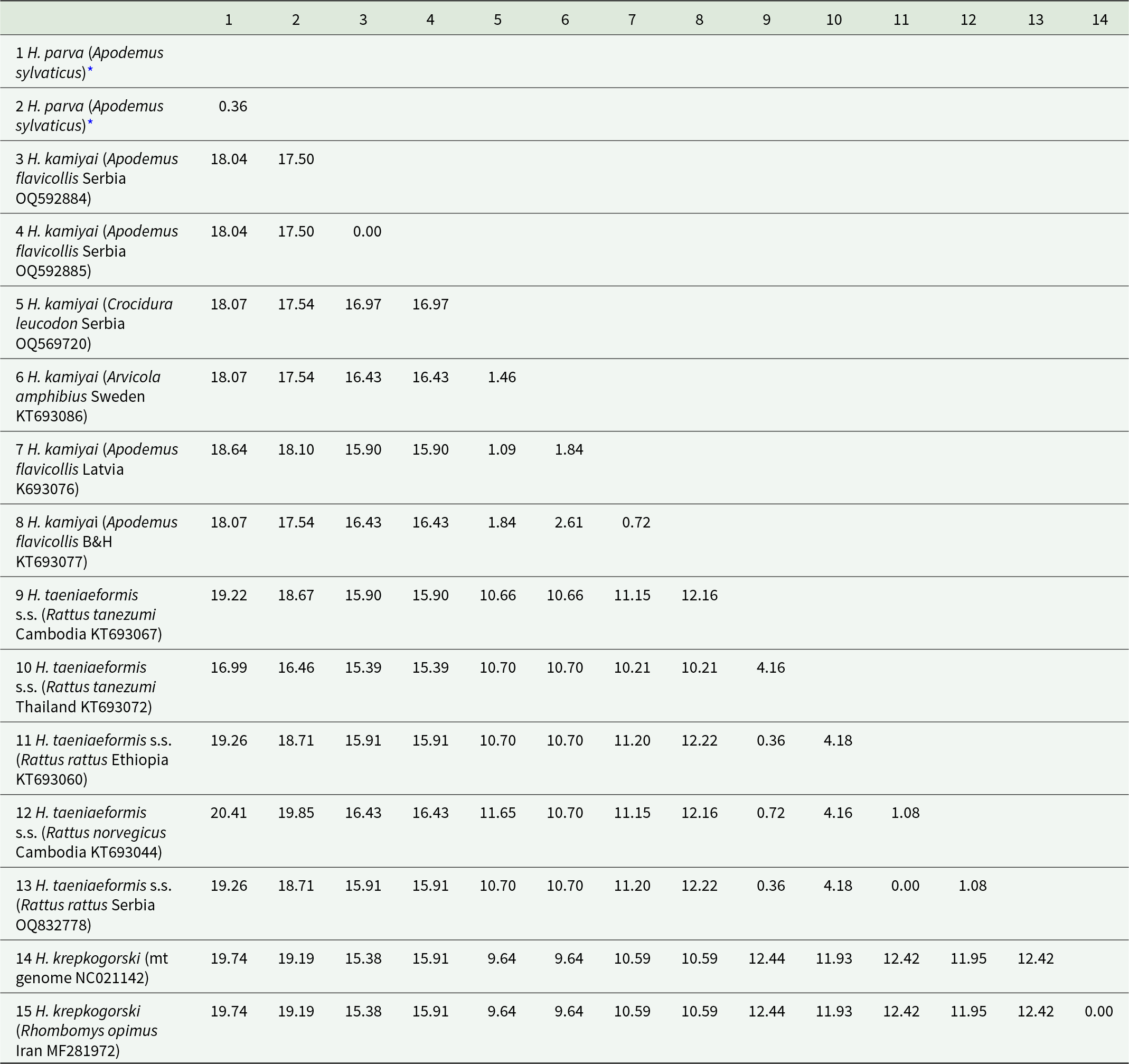
Values represent the proportional genetic distances (substitutions per site) calculated using the Kimura 2-parameter model (K2P) with gamma correction (gamma = 0.5) (Kimura, Reference Kimura1980).
* The isolates from this study represent 2 different haplotypes.
Discussion
Our results indicated a relatively high prevalence of H. parva in wood mice, compared to previous studies conducted in Spain and Portugal 2 decades ago, which showed lower rates ranging from 3% to 13.3% (Fuentes et al., Reference Fuentes, Cerezuela and Galan-Puchades2000, Reference Fuentes, Sáez, Trellis, Cruz, Sarmento, Casanova, Torres, Feliu and Esteban2003, Reference Fuentes, Fuentes, Sáez, Trelis, Galán-Puchades and Esteban2004; Eira et al., Reference Eira, Torres, Vingada and Miquel2006). This high infection rate can be attributed to the selective feeding preferences of the common genet. Despite its generalist feeding behaviour, this viverrid seems to prey predominantly on wood mice in the region (Torre Corominas et al., Reference Torre Corominas, Arrizabalaga and Ribas2015; Belbel et al., Reference Belbel, Boukheroufa, Benotmane, Sakraoui, Henada and Sakraoui2022), suggesting that it may play a crucial role in maintaining the life cycle of H. parva in the Iberian Peninsula. In our study, H. parva was detected in Algerian mice in only 1 case, contrasting with the high prevalence of the parasite in wood mice, which is consistent with its recognized role as a suitable host. The hypothesis that H. parva has continued to utilize the wood mouse as an intermediate host since its introduction alongside the definitive host (the common genet) to the Iberian Peninsula (Alvarez et al., Reference ‘Álvarez, Iglesias, Bos, Tojo and Sanmart’ín1990) is supported by our findings. This specificity suggests that the parasite may not have initiated adaptation or utilization of other micromammals from the region as intermediate hosts. It is important to remark that the isolate of H. parva detected in the Algerian mouse in our study had no viable larvae, confirming unsuccessful cyst development in this host and suggesting incidental exposure to infective eggs.
Our examination of wood mice in Spain indicated no significant differences in H. parva infection prevalence between sexes, habitats (natural and agricultural) and seasons (winter, spring, summer, autumn). In contrast, a study conducted in Portugal reported significant differences in H. parva prevalence based on some of these factors (Eira et al., Reference Eira, Torres, Vingada and Miquel2006). It was found that wood mouse males tend to show higher prevalence than females, and that prevalence can vary significantly among different seasons, with summer values being lower compared to other seasons. A study from Senegal (Africa) found that season significantly influenced the probability of infection, with the majority of infected Mastomys huberti (Wroughton, 1909) captured during spring, while prevalence of H. parva did not vary significantly with host gender, habitat or locality (Catalano et al., Reference Catalano, Bâ, Diouf, Léger, Verocai and Webster2019). All these contrasting findings suggest that the factors influencing H. parva prevalence may vary significantly between different regions and environmental conditions. Additionally, the population size of the definitive host (common genet), which serves as the main reservoir for this species, likely plays an important role. It should be noted that high prevalences of H. parva, ranging from 50% to 100%, have been reported in the common genet in Spain (Alvarez et al., Reference ‘Álvarez, Iglesias, Bos, Tojo and Sanmart’ín1990; Millán and Casanova, Reference Millán and Casanova2007; Ribas et al., Reference Ribas, Feliu and Casanova2009).
In our study, all polycephalic larvae of H. parva were confirmed by amplification and sequencing using 2 mitochondrial markers (cox1 and 12S rDNA), representing the first molecular study of this parasite from the Iberian Peninsula at the population level and one of the few genetic studies worldwide (Catalano et al., Reference Catalano, Bâ, Diouf, Léger, Verocai and Webster2019). Following the comprehensive taxonomic revision of the genus Hydatigera, including a reclassification and the identification of cryptic species (Nakao et al., Reference Nakao, Lavikainen, Iwaki, Haukisalmi, Konyaev, Oku, Okamoto and Ito2013; Lavikainen et al., Reference Lavikainen, Iwaki, Haukisalmi, Konyaev, Casiraghi, Dokuchaev, Galimberti, Halajian, Henttonen, Ichikawa-Seki, Itagaki, Krivopalov, Meri, Morand, Näreaho, Olsson, Ribas, Terefe and Nakao2016), several molecular genetic studies have subsequently focused on H. taeniaeformis s.s. and H. kamiyai in Asia and Europe (Bajer et al., Reference Bajer, Alsarraf, Dwużnik, Mierzejewska, Kołodziej-Sobocińska, Behnke-Borowczyk, Banasiak, Grzybek, Tołkacz, Kartawik, Stańczak, Opalińska, Krokowska-Paluszak, Górecki, Alsarraf and Behnke2020; Zhao et al., Reference Zhao, Zhou, Wu, Zhou, Liu, Yang and Zhang2020; Alvi et al., Reference Alvi, Li, Ohiolei, Qamar, Saqib, Tayyab, Altaf, Ashfaq, Hassan, Jamal, Wahab, Alvi, Usman, Bajwa, Fu, Yan and Jia2021; Martini et al., Reference Martini, Dumendiak, Gagliardo, Ragazzini, La Rosa, Giunchi, Thielen, Romig, Massolo and Wassermann2022; Miljević et al., Reference Miljević, Rajičić, Umhang, Bajić, Bjelić Čabrilo, Budinski and Blagojević2023). However, H. parva has remained unexplored at the genetic level. Based on cox1 sequence analysis of the 25 H. parva isolates, we identified only 2 haplotypes, resulting in low haplotype diversity (Hd: 0.380). Furthermore, the only 6 sequences from Senegal (Africa), deposited in GenBank©, showed potentially greater diversity (3 haplotypes out of 6 sequences). This contrast indicates a potentially reduced genetic diversity after the introduction of the parasite to Europe. Therefore, our results support the hypothesis that the parasite was introduced to Europe from Africa, probably following the introduction of the genet, as it is widely known that species generally have a higher genetic diversity in their place of origin (Austerlitz et al., Reference Austerlitz, Jung-Muller, Godelle and Gouyon1997). The pairwise divergence within our samples was up to 0.30%, while the genetic divergence with African isolates was slightly higher at 1.6–1.9%, illustrating geographic variation within the same species. This level of genetic divergence suggests a degree of genetic isolation, reflecting the effects of geographical distance between populations, likely driven by limited gene flow and temporal separation. Our H. parva isolates showed genetic distances to other species within the genus Hydatigera ranging from about 16% to 20%. This distinction confirms the precise taxonomic classification and suggests that the studied specimens are genetically distinct from other species of the genus Hydatigera. In addition, other authors have shown similar genetic distances within species of the genus Hydatigera (Mello et al., Reference Mello, Furtado, Rabelo and Pinto2018; Catalano et al., Reference Catalano, Bâ, Diouf, Léger, Verocai and Webster2019), further supporting our results.
In summary, the wood mouse proved to be the most suitable intermediate host species for H. parva in the study area, playing an important role in maintaining the parasite’s life cycle in the Iberian Peninsula. These preliminary results on the prevalence and genetic variation of H. parva in Spain provide valuable information for future studies on the distribution and population structure of H. parva in the Iberian Peninsula. In addition, this is the first population study of H. parva, based on mitochondrial genes in Europe and the second worldwide, representing important contribution to the understanding of genetic diversity and host suitability of this tapeworm species. Further studies are needed to better understand the genetic relationships between the populations in Africa and on the Iberian Peninsula and the evolutionary dynamics of the parasite. Our study serves as an example of the potential identification of introduction routes of a parasite with its definitive host.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182025000058.
Data availability statement
Nucleotide sequences of cox1 and 12S rDNA genes from the present study have been deposited in the GenBank database under the accession number PQ148369-PQ148393.
Author contributions
Conceptualization: MM, JM; Animal sampling: JM, RRP, JM; Laboratory work: JM, MM, MR, JB, BB. Data analysis: MM. Writing (original draft): MM, JM. Writing (editing and reviewing): RRP, MM, JB, BB.
Financial support
This study was supported by the Ministry of Science, Technological Development and Innovation of the Republic of Serbia, No. 451-03-66/2024-03/ 200007, and by Gobierno de Aragón (grant LMP90_21) and the Instituto Agroalimentario de Aragón-IA2 through one of its Research Strategic Lines.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
Capture was authorized by Gobierno de Aragón under permit 500201/24/2022/07284 and by the bioethics committee of Universidad de Zaragoza (PI16/22).








