Introduction
Species of the genus Cryptosporidium (phylum Apicomplexa) are protozoan parasites that infect the epithelium of the gastrointestinal tract of several vertebrate hosts, including humans and domestic and wild animals. A total of 44 Cryptosporidium species that infect fish, amphibians, reptiles, birds and mammals are currently recognized. In addition, more than 70 genotypes have been described in different hosts (Ryan et al., Reference Ryan, Fayer and Xiao2014; Chalmers et al., Reference Chalmers, Pérez-Cordón, Cacciò, Klotz and Robertson2018; Holubová et al., Reference Holubová, Tůmová, Sak, Hejzlarová, Konečný, McEvoy and Kváč2020; Ježková et al., Reference Ježková, Limpouchová, Prediger, Holubová, Sak, Konečný, Květoňová, Hlásková, Rost, McEvoy, Rajský, Feng and Kváč2021; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021). Cryptosporidium infection, which is transmitted by the fecal–oral route, can be acquired both directly, through contact with infected hosts, and indirectly, through the ingestion of food- and water-contaminated with oocysts (the infective form of the parasite) (Cacciò and Putignani, Reference Cacciò, Putignani, Cacciò and Widmer2014; Gerace et al., Reference Gerace, Lo Presti and Biondo2019). Although infection is asymptomatic in some immunocompetent hosts, watery diarrhoea, abdominal pain and vomiting are the most common symptoms of cryptosporidiosis, which can be severe or even fatal in immunocompromised patients (Shrivastava et al., Reference Shrivastava, Kumar, Smith and Sahu2017). Detection of oocysts, antigens and/or nucleic acids of the parasite in stool samples is the most frequently used method for diagnosing this infection (Gerace et al., Reference Gerace, Lo Presti and Biondo2019).
Outbreaks of cryptosporidiosis linked to drinking and recreational waters are common due to the ubiquity of the parasite, the low-infectious dose and the resistance of the oocysts to environmental pressures (temperature, desiccation and humidity conditions) and to conventional water disinfection treatments (mainly chlorination and ozonization) (King and Monis, Reference King and Monis2007; Carneiro Santos et al., Reference Carneiro Santos, Mastropasqua Rebello, Bergamo Bomfim, Bastidas Pacheco and Kamboh2020). Cryptosporidium is, thus, one of the most frequently detected pathogens in waterborne outbreaks of parasitic aetiology in developed countries, being involved in 63% of the reported outbreaks between 2011 and 2016 (Efstratiou et al., Reference Efstratiou, Ongerth and Karanis2017). Moreover, in a study on the global burden of foodborne diseases, it was estimated that Cryptosporidium was responsible for 8.6 million cases of illness and 3759 deaths in 2010 (World Health Organization, 2015). Food can be contaminated with Cryptosporidium oocysts at all points of food chain (ranging from primary production in agriculture and aquaculture to the transformation, distribution and sale of food) via direct contact with fecal material from infected hosts or indirectly via infected food handlers (Zahedi and Ryan, Reference Zahedi and Ryan2020). In developing countries, Cryptosporidium, along with Rotavirus and Shigella, are the three pathogens that most commonly cause diarrhoeal disease in children under 2 years old, with Cryptosporidium being responsible for 30–50% of childhood mortality in these countries (Sow et al., Reference Sow, Muhsen, Nasrin, Blackwelder, Wu, Farag, Panchalingam, Sur, Zaidi, Faruque, Saha, Adegbola, Alonso, Breiman, Bassat, Tamboura, Sanogo, Onwuchekwa, Manna, Ramamurthy, Kanungo, Ahmed, Qureshi, Quadri, Hossain, Das, Antonio, Hossain, Mandomando, Nhampossa, Acácio, Omore, Oundo, Ochieng, Mintz, O'Reilly, Berkeley, Livio, Tennant, Sommerfelt, Nataro, Ziv-Baran, Robins-Browne, Mishcherkin, Zhang, Liu, Houpt, Kotloff and Levine2016; Kotloff et al., Reference Kotloff, Platts-Mills, Nasrin, Roose, Blackwelder and Levine2017).
In farmed ruminants, Cryptosporidium is also recognized as one of the main enteropathogens involved in the aetiology of neonatal diarrhoea syndrome, which causes significant economic losses on farms, owing to the associated high morbidity and mortality and the delayed growth of animals (De Graaf et al., Reference De Graaf, Vanopdenbosch, Ortega-Mora, Abbassi and Peeters1999; Meganck et al., Reference Meganck, Hoflack, Piepers and Opsomer2015). Furthermore, Cryptosporidium has been reported to be widely distributed in wildlife, which may act as both an important reservoir and source of infection (Zahedi et al., Reference Zahedi, Paparini, Jian, Robertson and Ryan2016).
Brief historical review of Cryptosporidium in fish
Cryptosporidium parasites were first described by Clarke (Reference Clarke1895) as ‘swarm-spores’ in the gastric epithelium of laboratory mice (Mus musculus). Tyzzer (Reference Tyzzer1907) subsequently established the genus Cryptosporidium to refer to this protozoan parasite infecting the same host. However, it was not until seven decades later that the first study on Cryptosporidium in fish was published, in which Hoover et al. (Reference Hoover, Hoerr, Carlton, Hinsman and Ferguson1981) described Cryptosporidium nasoris (syn. Cryptosporidium nasorum), a species found in the intestine of the tropical marine fish naso tang (Naso lituratus). Nevertheless, this species is currently considered as nomen nudum for the following reasons: (1) only developmental stages of the parasite on the microvillus surface of intestinal epithelial cells have been described by light and electron microscopy; (2) no measurements of viable oocysts have been provided; (3) no other taxonomically useful diagnostic features have been presented and (4) the lack of deposited museum specimens and molecular studies prevent completion of the description. Under these circumstances, it is impossible to differentiate between C. nasorum and other Cryptosporidium spp. (Ryan et al., Reference Ryan, O'Hara and Xiao2004, Reference Ryan, Fayer and Xiao2014; Xiao et al., Reference Xiao, Fayer, Ryan and Upton2004; Fayer, Reference Fayer2010; Ryan, Reference Ryan2010).
After the first description, different histological studies detected developmental stages compatible with Cryptosporidium in the stomach and/or intestine of several fish species: carp (Cyprinus carpio; prevalence = 14.3%) (Pavlásek, Reference Pavlásek1983); cichlid fish (Oreochromis spp.; prevalence = 58.8%) (Landsberg and Paperna, Reference Landsberg and Paperna1986); barramundi (Lates calcarifer; prevalence not provided) (Glazebrook and Campbell, Reference Glazebrook and Campbell1987); black Nile catfish (Bagrus bayad; prevalence = 10.0%), North African catfish (Clarias lazera; prevalence = 20.0%), Nile tilapia (Tilapia nilotica syn. Oreochromis niloticus; prevalence = 30.0%) (Hefnawy, Reference Hefnawy1989); brown trout (Salmo trutta; prevalence = 38.9%) (Rush et al., Reference Rush, Chapman and Ineson1990); red drum (Sciaenops ocellatus; prevalence = 21.7%) (Camus and López, Reference Camus and López1996) and catfish (Plecostomus sp.; prevalence = 100%) (Muench and White, Reference Muench and White1997).
Moreover, the results of ultrastructural studies led Paperna and Vilenkin (Reference Paperna and Vilenkin1996) to propose a new genus, Piscicryptosporidium, for species of this parasite infecting fish, on the basis of the following features: (1) piscine Cryptosporidium spp. are found in the stomachs of the hosts; (2) the species differs from all other known Cryptosporidium species in that the parasitic parasitophorous vacuole has microvilli on the surface and (3) the sporulated oocysts gradually penetrate the basal part of the gut epithelium or lamina propria rather than being released into the gut lumen (Paperna and Vilenkin, Reference Paperna and Vilenkin1996). The organisms found in the stomachs of the gourami (Trichogaster leeri) and cichlid fish (Oreochromis spp.) were designated as Piscicryptosporidium reinchenbachklinkei and Piscicryptosporidium cichlidis, respectively. The validity of the genus remains to be determined and the two Piscicryptosporidium species may be considered nomina nuda because some of these features have been described in mammalian Cryptosporidium (i.e. gastric location) (Valigurová et al., Reference Valigurová, Jirků, Koudela, Gelnar, Modrý and Šlapeta2008) and there are no molecular studies to support this proposal.
In the current century, a new species, Cryptosporidium molnari, identified by Álvarez-Pellitero and Sitjá-Bobadilla (Reference Álvarez-Pellitero and Sitjà-Bobadilla2002), was found in the stomach of farmed marine fish, specifically gilthead sea bream (Sparus aurata) and European sea bass (Dicentrarchus labrax). Álvarez-Pellitero et al. (Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004) subsequently described the species Cryptosporidium scophthalmi after detecting developmental stages of this protozoon in the intestine of cultured turbot (Scophthalmus maximus syn. Psetta maxima). In the same year, Ryan et al. (Reference Ryan, O'Hara and Xiao2004) reported the results of histological, genetic and phylogenetic studies of a C. molnari-like isolate from the stomach of the ornamental fish guppy (Poecilia reticulata), which was designated as Cryptosporidium piscine genotype 1 in the absence of molecular data for C. molnari. In a later study, Palenzuela et al. (Reference Palenzuela, Álvarez-Pellitero and Sitjà-Bobadilla2010) carried out the molecular characterization of the C. molnari isolates, and Ryan et al. (Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015) subsequently elevated the piscine genotype 1 to species level and named it Cryptosporidium huwi on the basis of genetic and morphological differences relative to other gastric and intestinal species of Cryptosporidium in these hosts. Although Costa and Saraiva (Reference Costa and Saraiva2015a, Reference Costa and Saraiva2015b) described C. scophthalmi-like sequences in turbot, no genetic information on the original isolate of C. scophthalmi is yet available. Therefore, this species may be considered as not valid, owing to the observed high level of genetic heterogeneity and oocyst morphological similarity between Cryptosporidium species (Fayer, Reference Fayer2010; Ryan et al., Reference Ryan, Fayer and Xiao2014, Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015). The species names Cryptosporidium bollandi, for piscine genotype 2 found in angelfish (Pterophyllum scalare) and Oscar fish (Astronotus ocellatus), and Cryptosporidium abrahamseni, for piscine genotype 7 found in red-eye tetra (Moenkhausia sanctaefilomenae), have been recently established (see Table 1) (Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021).
Cryptosporidium species and genotypes currently recognized in piscine hosts

s.d., standard deviation; C, cultured; FW, freshwater; M, marine; O, ornamental; W, wild; P, prevalence rate; ND, no data.
a Primary host species.
* GenBank accession number identical to the original sequence (no asterisk).
The use of molecular techniques has also enabled the identification of a total of nine piscine Cryptosporidium genotypes (3–6, 8–10 and marine 1–2 genotypes), five different C. molnari-like genotypes and more than 29 unnamed novel genotypes in both freshwater and marine fish (Reid et al., Reference Reid, Lymbery, Ng, Tweedle and Ryan2010; Zanguee et al., Reference Zanguee, Lymbery, Lau, Suzuki, Yang, Ng and Ryan2010; Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013; Yang et al., Reference Yang, Palermo, Chen, Edwards, Paparini, Tong, Gibson-Kueh, Lymbery and Ryan2015, Reference Yang, Dorrestein and Ryan2016; Couso-Pérez et al., Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2018, Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019; Certad et al., Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019, Reference Certad, Zahedi, Gantois, Sawant, Creusy, Duval, Benamrouz-Vanneste, Ryan and Viscogliosi2020). The species and genotypes of Cryptosporidium currently recognized in fish are listed in Table 1.
Molecular studies have revealed the considerable genetic distance between piscine Cryptosporidium and remaining species of the genus infecting other host classes. Phylogenetic analysis of piscine-derived Cryptosporidium species/genotypes showed that the piscine clade has a basal position relative to all other Cryptosporidium species, which form two main broad branches: intestinal and gastric species. This suggests that piscine species may be the evolutionary ancestors of Cryptosporidium species infecting other host classes (Figs 1 and 2) (Palenzuela et al., Reference Palenzuela, Álvarez-Pellitero and Sitjà-Bobadilla2010; Reid et al., Reference Reid, Lymbery, Ng, Tweedle and Ryan2010; Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013; Certad et al., Reference Certad, Dupouy-Camet, Gantois, Hammouma-Ghelboun, Pottier, Guyot, Benamrouz, Osman, Delaire, Creusy, Viscogliosi, Dei-Cas, Aliouat-Denis and Follet2015, Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019, Reference Certad, Zahedi, Gantois, Sawant, Creusy, Duval, Benamrouz-Vanneste, Ryan and Viscogliosi2020; Ryan et al., Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015; Couso-Pérez et al., Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2018, Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021).
Phylogenetic relationships in the genus Cryptosporidium inferred by neighbour-joining analysis of the small subunit ribosomal RNA (18S rRNA) gene on the basis of genetic distances calculated by the Tamura 3-parameter model (gamma distributed with five rate categories) using MEGA X software (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018). The tree was generated using a total of 433 positions in the final dataset. The percentages of replicate trees in which associated taxa clustered together in the bootstrap test (10 000 replicates) are shown at the internal nodes for distance (>50%). Accession numbers are given in parentheses.
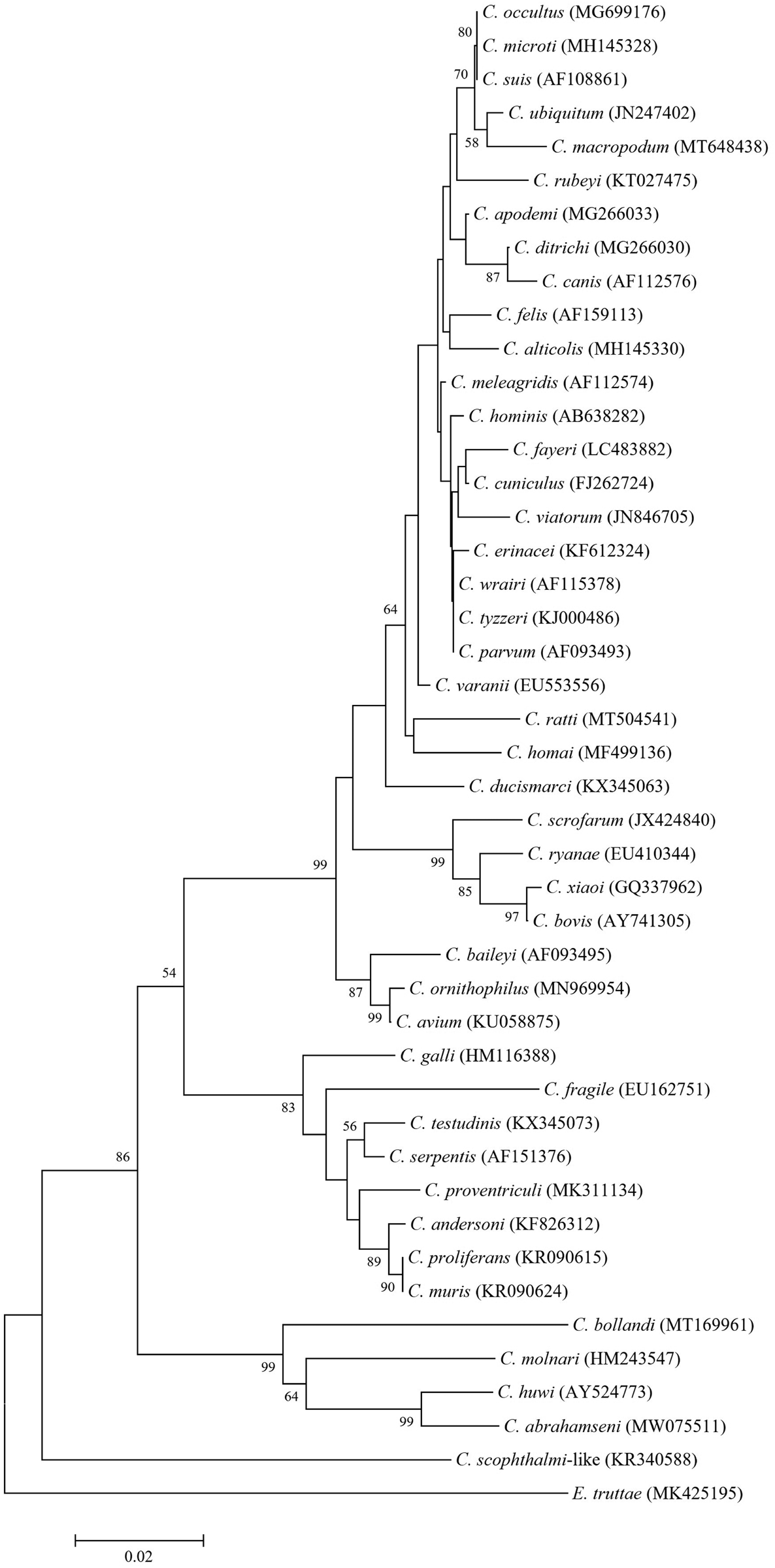
Phylogenetic relationships of piscine Cryptosporidium species and genotypes inferred by neighbour-joining analysis of the small subunit ribosomal RNA (18S rRNA) gene on the basis of genetic distances calculated by the Tamura 3-parameter model (gamma distributed with five rate categories) using MEGA X software (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018). The tree was generated using a total of 251 positions in the final dataset. The percentage numbers of replicate trees in which associated taxa clustered together in the bootstrap test (10 000 replicates) are shown at the internal nodes for distance (>50%). Accession numbers, host species and geographical regions are shown in parentheses.

Moreover, several species and genotypes of Cryptosporidium typical of other hosts such as Cryptosporidium parvum (major host: livestock; Tyzzer, Reference Tyzzer1912), Cryptosporidium hominis (major host: humans; Morgan-Ryan et al., Reference Morgan-Ryan, Fall, Ward, Hijjawi, Sulaiman, Fayer, Thompson, Olson, Lal and Xiao2002), Cryptosporidium xiaoi (major host: sheep; Fayer and Santín, Reference Fayer and Santín2009), Cryptosporidium scrofarum (major host: pig; Kváč et al., Reference Kváč, Kestřánová, Pinková, Květoňová, Kalinová, Wagnerová, Kotková, Vítovec, Ditrich, McEvoy, Stenger and Sak2013) and rat genotype 3 have been identified in fish (Table 2) (Reid et al., Reference Reid, Lymbery, Ng, Tweedle and Ryan2010; Gibson-Kueh et al., Reference Gibson-Kueh, Yang, Thuy, Jones, Nicholls and Ryan2011; Morine et al., Reference Morine, Yang, Ng, Kueh, Lymbery and Ryan2012; Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013; Certad et al., Reference Certad, Dupouy-Camet, Gantois, Hammouma-Ghelboun, Pottier, Guyot, Benamrouz, Osman, Delaire, Creusy, Viscogliosi, Dei-Cas, Aliouat-Denis and Follet2015, Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019; Palermo, Reference Palermo2016; Couso-Pérez et al., Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2018, Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019; Shahbazi et al., Reference Shahbazi, Sheikhzadeh, Nematollahi and Helan2020). Without ruling out the possibility that fish only act as mechanical carriers of these Cryptosporidium spp., the role of fish as potential hosts of zoonotic Cryptosporidium spp. will be discussed later.
Mammalian Cryptosporidium species and genotypes detected in fish

C, cultured; FW, freshwater; M, marine; O, ornamental; W, wild; P, prevalence rate; ND, no data.
Life cycle of piscine Cryptosporidium
Considering the different developmental stages of Cryptosporidium observed in ultrastructural studies (Landsberg and Paperna, Reference Landsberg and Paperna1986; Paperna and Vilenkin, Reference Paperna and Vilenkin1996; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004), the life cycle of Cryptosporidium spp. in fish can be assumed to involve the following different stages: (1) excystation and release of sporozoites; (2) schizogony or merogony; (3) gamogony; (4) zygote formation; (5) oocyst wall formation and (6) sporulation (Fig. 3). However, there are some notable differences in the life cycles of Cryptosporidium species in fish and in mammals, which we will highlight below. The cycle begins when a sporulated oocyst is ingested by a susceptible fish host. The oocyst undergoes excystation, releasing the sporozoites, which then reach the apical surface of the cells of the gastrointestinal epithelium through gliding motility. The host cell envelops the sporozoite inside a vacuolar space formed by invagination of the cytoplasmic membrane; this parasitophorous vacuole has an intracellular, but extra-cytoplasmic location, in which the subsequent developmental stages take place (Landsberg and Paperna, Reference Landsberg and Paperna1986; Paperna and Vilenkin, Reference Paperna and Vilenkin1996; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004). The first remarkable difference observed in fish relative to mammal hosts occurs at the level of the membrane of the parasitophorous vacuole. Ultrastructural studies have shown that in piscine hosts the membrane of the parasitophorous vacuole has regularly spaced microvilli containing a dense, finely granular substance. In addition, the zone of attachment between the host cell and the parasite is also very different. In fish, rather than being continuous with the interlamellar layer of the membrane of the parasitophorous vacuole, the zone of attachment appears to consist of two electron-dense bands, where the inner band is connected to two plates by osmophilic points. The feeder organelle is composed of multiple parallel folds with vesiculate widening at their endings, being longer and more numerous in piscine Cryptosporidium than in other members of the genus (Fig. 4) (Landsberg and Paperna, Reference Landsberg and Paperna1986; Paperna and Vilenkin, Reference Paperna and Vilenkin1996; Muench and White, Reference Muench and White1997; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004; Gabor et al., Reference Gabor, Srivastava, Titmarsh, Dennis, Gabor and Landos2011).
Hypothetical life cycle of piscine Cryptosporidium. The sporozoites (A) reach the apical surface of the epithelial cells (B). The sporozoite is enveloped inside a parasitophorous vacuole (PV), the membrane of which has regularly spaced microvilli. Within the PV, the sporozoite is differentiated into a trophozoite (C), which undergoes nuclear division through merogony, producing a type I meront (D). Type I merozoites infect adjacent cells and new type I or type II meronts (E) are produced. Type II merozoites invade other cells and undergo gamogony, forming microgamonts and macrogamonts, which produce microgametes (F) and macrogametes (G). After fertilization, a zygote is formed (H), generating (by sporogony) sporulated oocysts containing four naked sporozoites (I). These oocysts are found within a vacuolar space located deep within the cytoplasm in the epithelial cells, and more than one oocyst can appear in clusters in the same infected cell (J). Thick-walled oocysts are released with the feces of the host and can infect other susceptible hosts (K). Some oocysts have a thin wall that is easily broken, thus enabling the cycle to be reinitiated by endogenous autoinfection (L).

Histological sections of the stomach of Oreochromis aureus × Oreochromis niloticus naturally infected with Cryptosporidium sp. showing (A) trophozoite and (B) early differentiating meront under transmission electron microscopy (29 300×). PE, parasitophorous membrane; ER, rough endoplasmic reticulum; N, nucleus; CW, parasite plasmalemma; F, feeder organelle; H, host cell; M, differentiating merozoite (Landsberg and Paperna, Reference Landsberg and Paperna1986) (© Inter-Research).

The sporozoite differentiates into a trophozoite with a large nucleus. The nucleus undergoes division, through a process called schizogony or merogony, to produce two types of multinucleated meronts (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002). Type I meronts have eight nuclei and produce eight type I merozoites (Paperna and Vilenkin, Reference Paperna and Vilenkin1996), which infect adjacent cells where they again divide asexually, producing new type I or type II meronts. Type II meronts mature to produce four type II merozoites (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002), which probably invade other cells and undergo sexual reproduction by gamogony, to produce microgamonts and macrogamonts. As a consequence of the nuclear division of the microgamont, aflagellate microgametes are generated (up to 12 microgametes were observed by Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002). These forms leave the parasitophorous vacuole to fertilize the differentiated macrogamont or macrogamete. After fertilization, the zygote formed is internalized and undergoes sporogony to produce sporulated oocysts containing four naked sporozoites (Landsberg and Paperna, Reference Landsberg and Paperna1986; Paperna and Vilenkin, Reference Paperna and Vilenkin1996; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004). Sporulation deep within the epithelium appears to be characteristic of the piscine clade (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Ryan et al., Reference Ryan, O'Hara and Xiao2004, Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021). Thus, in Cryptosporidium species that infect fish, unlike in mammal hosts, the oocysts are found within a vacuolar space located deep within the cytoplasm in the epithelial cells (Paperna and Vilenkin, Reference Paperna and Vilenkin1996; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Ryan et al., Reference Ryan, O'Hara and Xiao2004, Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015; Palenzuela et al., Reference Palenzuela, Álvarez-Pellitero and Sitjà-Bobadilla2010; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021). In addition, more than one oocyst can appear in clusters in the same infected cell, along with aggregates of cellular residues and necrotic substances (Fig. 5). The infected cells gradually degenerate before finally disintegrating, leaving the oocysts or clusters of oocysts in the intercellular spaces or allowing these to escape to the lumen (Landsberg and Paperna, Reference Landsberg and Paperna1986; Paperna and Vilenkin, Reference Paperna and Vilenkin1996; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002).
Histological sections of the stomach of (A) guppy (Poecilia reticulata) and (B) angelfish (Pterophyllum scalare) stained with haematoxylin–eosin, showing large numbers of Cryptosporidium life cycle stages along the lining of the gastric mucosa with clusters of oocysts located deep within the epithelium (arrows). Scale bar = 10 μm (Ryan et al., Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020) (© Elsevier).

Most of the sporulated oocysts eliminated with the feces and capable of infecting other animals have a thick (trilamellar) wall. However, as occurs in mammals, some oocysts have a thin (bilamellar) envelope that is easily broken when the oocysts are released into the gastrointestinal lumen, and free sporozoites can then infect adjacent cells, reinitiating a cycle of endogenous autoinfection (Fig. 3) (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004).
Cryptosporidium species that infect fish have been detected in different sections of the gastrointestinal tract. Thus, histological observations have revealed developmental stages of C. molnari lining the stomach epithelium, whereas zygotes and fully sporulated oocysts (4.7 ± 0.5 μm × 4.5 ± 0.5 μm), sometimes in groups, have been located in the basal portion of the epithelial cell (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002). Parasitic stages of C. huwi have been found dispersed on the apical surface of the stomach mucosa, and groups of zygotes and oocysts (4.6 ± 0.3 μm × 4.4 ± 0.4 μm) have also been located deep within the gastric epithelium (Ryan et al., Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015). Similarly, trophozoites, meronts and gamonts of C. bollandi have been detected in large numbers attached to the gastric mucosa, with zygotes and oocysts (3.1 ± 0.5 μm × 2.8 ± 0.4 μm) also located deep within the epithelium (Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020). By contrast, histological analysis has revealed meronts and micro- and macrogamonts of C. scophthalmi in an extracytoplasmic position in the intestine, whereas oocysts (4.4 ± 0.3 μm × 3.9 ± 0.4 μm) have been observed deeply embedded within the epithelium (Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004). More recently, Zahedi et al. (Reference Zahedi, Bolland, Oskam and Ryan2021) observed the existence of a large number of C. abrahamseni oocysts (3.8 ± 0.2 μm × 3.2 ± 0.2 μm) and clusters of parasites also located deep within the epithelium of the small intestine.
Piscine cryptosporidiosis
Transmission
Water is an excellent vehicle for the dissemination of pathogenic organisms, which are transmitted via the fecal–oral route. Cryptosporidium is one of the infectious agents most frequently detected in water and has been reported in different types of water worldwide (river, recreational, drinking and wastewater) (Omarova et al., Reference Omarova, Tussupova, Berndtsson, Kalishev and Sharapatova2018; Vermeulen et al., Reference Vermeulen, van Hengel, Kroeze, Medema, Spanier, van Vliet and Hofstra2019). This waterborne protozoan parasite can occur in surface waters due to contamination with fecal matter of human or animal origin (both domestic and wild animals). The oocysts can spread to water bodies directly or indirectly via run-off from contaminated land surfaces with livestock manure (Lu et al., Reference Lu, Ryu, Hill, Schoen, Ashbolt, Edge and Domingo2011; Ahmed et al., Reference Ahmed, Sritharan, Palmer, Sidhu and Toze2013; Sidhu et al., Reference Sidhu, Ahmed, Gernjak, Aryal, McCarthy, Palmer, Kolotelo and Toze2013). In addition, the aquatic environment can become contaminated by effluents from wastewater treatment plants or from insufficient or deficient sewage treatment systems (Ahmed et al., Reference Ahmed, Goonetilleke and Gardner2010; Schneeberger et al., Reference Schneeberger, O'Driscoll, Humphrey, Henry, Deal, Seiber, Hill and Zarate-Bermudez2015; Vermeulen et al., Reference Vermeulen, van Hengel, Kroeze, Medema, Spanier, van Vliet and Hofstra2019).
Regarding fish farm facilities, the presence of Cryptosporidium may be associated with water quality. As previously reported by Sitjà-Bobadilla et al. (Reference Sitjà-Bobadilla, Padrós, Aguilera and Álvarez-Pellitero2005) and Álvarez-Pellitero et al. (Reference Álvarez-Pellitero, Pérez, Quiroga, Redondo, Vázquez, Riaza, Palenzuela, Sitjà-Bobadilla and Nieto2009), transmission occurs through the farm water supply (i.e. rivers and seas), and the use of filtration and ultraviolet irradiation in hatcheries and nurseries is not sufficient to prevent the entry of infective oocysts. Once the parasite is present inside the facilities, recirculation systems can contribute to dispersing and concentrating oocysts in the water. In addition, transmission of the parasite may be facilitated by the stress conditions that occur in aquaculture systems, in which fish cohabit in dense groups, and by cannibalism, a common phenomenon in piscine communities, which may also play a role in transmission (Sitjà-Bobadilla and Álvarez-Pellitero, Reference Sitjà-Bobadilla and Álvarez-Pellitero2003; Sitjà-Bobadilla et al., Reference Sitjà-Bobadilla, Padrós, Aguilera and Álvarez-Pellitero2005).
Cryptosporidium oocysts have been found in other eukaryotic organisms living in the aquatic environment, and their participation in the transmission of this protozoan parasite cannot be ruled out. Several rotifers, species of the genera Epiphanes and Euchlanis, and some free-living amoebas, such as Acanthamoeba spp., can retain C. parvum oocysts when they coexist in contaminated waters (Fayer et al., Reference Fayer, Trout, Walsh and Cole2000; Gómez-Couso et al., Reference Gómez-Couso, Paniagua-Crespo and Ares-Mazás2007). Similarly, the brine shrimp Artemia franciscana can act as a carrier in the transmission of Cryptosporidium infection in cultured fish when it is used as a live diet (Méndez-Hermida et al., Reference Méndez-Hermida, Gómez-Couso and Ares-Mazás2006, Reference Méndez-Hermida, Gómez-Couso and Ares-Mazás2007). Reboredo-Fernández et al. (Reference Reboredo-Fernández, Prado-Merini, García-Bernadal, Gómez-Couso and Ares-Mazás2014) demonstrated the presence of Cryptosporidium oocysts in benthic macroinvertebrates in Galician rivers (NW Spain) and concluded that these organisms could contribute to the transmission of the parasite through the aquatic food chain.
Prevalence
Wild fish
So far, only seven studies have reported the presence of Cryptosporidium in both marine and freshwater wild fish.
In marine fish, molecular characterization of Cryptosporidium isolates from 20 different species collected on the coasts of Western Australia and Papua New Guinea led to the identification of C. bollandi and piscine genotypes 3 and 5 in sea mullet (Mugil cephalus) (0.5–33.3%) and piscine genotype 8 in silver biddy (Gerres oblongus) (3.6%) as well as Cryptosporidium species typical of other hosts, specifically C. parvum (1.8–6.9%), C. xiaoi (1.8%) and C. scrofarum (3.6%) (see Tables 1 and 2) (Reid et al., Reference Reid, Lymbery, Ng, Tweedle and Ryan2010; Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013; Yang et al., Reference Yang, Palermo, Chen, Edwards, Paparini, Tong, Gibson-Kueh, Lymbery and Ryan2015; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020). A recent study on Cryptosporidium was carried out in commercially important edible fish across marine areas around France collected in two surveys (Certad et al., Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019). The study reported, by molecular analysis, overall prevalences of Cryptosporidium of 2.3 and 3.2% in the first and second surveys, respectively, identifying C. parvum (0.7–6.5%) (see Table 2) and also seven new piscine genotypes that exhibit genetic distances from C. molnari of between 0.5 and 12.5%. Thus, according to the terminology used by the authors, Cryptofish 1 genotype was found in saithe (Pollachius virens) (12.5%) and in blue ling (Molva dypterygia) (1.4%); Cryptofish 2 genotype was detected in ling (Molva molva) (10.9%) and in whiting (Merlangius merlangus) (0.8%); Cryptofish 3 and Cryptofish 4 genotypes were identified in ling (M. molva) (4.3%) and in blue ling (M. dypterygia) (1.4%), respectively; Cryptofish 5 genotype was found in saithe (P. virens) (6.3%) and in hake (Merluccius merluccius) (0.7%); finally, Cryptofish 6 and Cryptofish 7 genotypes were identified in cod (Gadus morhua) (0.8%) and in mackerel (Scomber scombrus) (0.9%), respectively (Certad et al., Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019). Subsequent phylogenetic analysis at the 18S rDNA and actin loci led to the designation of these genotypes as novel marine genotype 1 (Cryptofish 1, 2, 4 and 5) and marine genotype 2 (Cryptofish 7) (see Table 1) (Certad et al., Reference Certad, Zahedi, Gantois, Sawant, Creusy, Duval, Benamrouz-Vanneste, Ryan and Viscogliosi2020).
A lower diversity of Cryptosporidium species/genotypes (including C. molnari, piscine genotype 10 and C. parvum), although with higher overall prevalences (40.0%; 0.2 and 20.0–100%, respectively), has been identified by molecular methods in freshwater fish from lake Leman (France), Papua New Guinea and Galicia (NW Spain) (see Tables 1 and 2) (Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013; Certad et al., Reference Certad, Dupouy-Camet, Gantois, Hammouma-Ghelboun, Pottier, Guyot, Benamrouz, Osman, Delaire, Creusy, Viscogliosi, Dei-Cas, Aliouat-Denis and Follet2015; Couso-Pérez et al., Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019). Moreover, Cryptosporidium sp. was detected by flotation technique and enzyme-linked immunosorbent assay in alewife (Alosa pseudoharengus) (50.0%) in the New York State (USA) (Ziegler et al., Reference Ziegler, Wade, Schaaf, Stern, Nadareski and Mohammed2007).
Cultured fish
Data on Cryptosporidium in farmed fish are also very scarce, with information available for only three marine and six freshwater species (see Tables 1 and 2).
Álvarez-Pellitero and Sitjà-Bobadilla (Reference Álvarez-Pellitero and Sitjà-Bobadilla2002) carried out the first study on gilthead sea bream (S. aurata) and European sea bass (D. labrax) (both marine species), reporting prevalences of C. molnari in the range of 6.5–25.4 and 4.6–12.0%, respectively. Sitjà-Bobadilla et al. (Reference Sitjà-Bobadilla, Padrós, Aguilera and Álvarez-Pellitero2005) later conducted an epidemiological study on these same hosts, observing that the prevalence of C. molnari differed in specimens from hatcheries and in those from ongrowing tanks. Thus, prevalences of 11.6–100 and 11.1–50.0% were detected in the smallest specimens of gilthead sea bream and European sea bass, respectively, whereas those corresponding to fish from the ongrowing tanks were in the range of 3.6–55.0% for gilthead sea bream to 57.9% for European sea bass (Table 2). In another cultured species, the turbot P. maxima, the prevalence rate of C. scophthalmi was in the range of 15.0–100%, with the higher values corresponding to the juvenile specimens, observing a decrease in the prevalence as the size of the fish increased (Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004).
Regarding freshwater species, the presence of Cryptosporidium was investigated in angelfish (P. scalare) by Murphy et al. (Reference Murphy, Bradway, Walsh, Sanders and Snekvik2009), who described piscine genotype 2, currently recognized as C. bollandi (Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020). Sequences similar to C. molnari have been identified in stomach samples from Murray cod (Maccullochella peelii) (95.4%) (Barugahare et al., Reference Barugahare, Michelle, Becker and Šlapeta2011). Molecular characterization of the Cryptosporidium isolates from Asian sea bass (L. calcarifer) and Nile tilapia (O. niloticus) enabled identification of C. parvum in these species (20.0 and 2.4%, respectively) (Gibson-Kueh et al., Reference Gibson-Kueh, Yang, Thuy, Jones, Nicholls and Ryan2011; Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013) (see Tables 1 and 2). Moreover, after histological examination, Cryptosporidium-like organisms were observed in the apical border of gastric and enteric epithelial sections of intensively reared barramundi (L. calcarifer) (prevalence of 92.5%) (Gabor et al., Reference Gabor, Srivastava, Titmarsh, Dennis, Gabor and Landos2011). More recently, Couso-Pérez et al. (Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2018) conducted a study on Cryptosporidium in farmed rainbow trout (Oncorhynchus mykiss), reporting an overall prevalence of 9.2% by immunofluorescence microscopy, although a higher rate was observed in the smallest fish (14.2%), and identifying C. parvum (1.9%) and piscine genotype 9 (0.6%) (see Tables 1 and 2).
Ornamental fish
Most studies on piscine Cryptosporidium have involved ornamental fish, including about 70 species collected from various aquariums and pet shops in Perth (Western Australia) (Zanguee et al., Reference Zanguee, Lymbery, Lau, Suzuki, Yang, Ng and Ryan2010; Morine et al., Reference Morine, Yang, Ng, Kueh, Lymbery and Ryan2012; Yang et al., Reference Yang, Palermo, Chen, Edwards, Paparini, Tong, Gibson-Kueh, Lymbery and Ryan2015, Reference Yang, Dorrestein and Ryan2016; Palermo, Reference Palermo2016; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021) and in different cities in Iran (Nematollahi et al., Reference Nematollahi, Jaberi, Ashrafi Helan and Sheikhzadeh2016; Shahbazi et al., Reference Shahbazi, Sheikhzadeh, Nematollahi and Helan2020). Cryptosporidium was detected by molecular methods in a total of 12 and 16 species of marine and freshwater ornamental fish, respectively, with prevalences in the range of 9.1–100% in marine fish and 0.9–100% in freshwater species (Zanguee et al., Reference Zanguee, Lymbery, Lau, Suzuki, Yang, Ng and Ryan2010; Morine et al., Reference Morine, Yang, Ng, Kueh, Lymbery and Ryan2012; Yang et al., Reference Yang, Palermo, Chen, Edwards, Paparini, Tong, Gibson-Kueh, Lymbery and Ryan2015, Reference Yang, Dorrestein and Ryan2016; Palermo, Reference Palermo2016; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Shahbazi et al., Reference Shahbazi, Sheikhzadeh, Nematollahi and Helan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021). Molecular characterization of the isolates enabled identification of C. molnari (15.4–100%), C. molnari-like (0.9–100%), C. huwi (1.9–50.0%), C. bollandi (0.5–75.0%) and C. abrahamseni (27.3–62.5%) (see Table 1). Moreover, several authors have characterized a total of three genotypes of Cryptosporidium, specifically piscine genotypes 4 (20.0–100%), 5 (0.5–100%) and 6 (2.3–33.3%), as well as piscine genotype 3-like (0.9%), initially described in the sea mullet (M. cephalus) (see Table 1) (Zanguee et al., Reference Zanguee, Lymbery, Lau, Suzuki, Yang, Ng and Ryan2010; Morine et al., Reference Morine, Yang, Ng, Kueh, Lymbery and Ryan2012; Yang et al., Reference Yang, Palermo, Chen, Edwards, Paparini, Tong, Gibson-Kueh, Lymbery and Ryan2015, Reference Yang, Dorrestein and Ryan2016; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021). In the goldfish (Carassius auratus), C. hominis (4.6%), C. parvum (0.9%) and rat genotype 3 (5.3%) were detected in addition to C. molnari-like (2.7%) and piscine genotypes 3-like and 5 (0.9 and 2.7%, respectively) (see Tables 1 and 2) (Morine et al., Reference Morine, Yang, Ng, Kueh, Lymbery and Ryan2012; Palermo, Reference Palermo2016). Although Shahbazi et al. (Reference Shahbazi, Sheikhzadeh, Nematollahi and Helan2020) reported the presence of C. parvum in C. auratus, the sequence analysis revealed 99% similarity with the sequence corresponding to accession number AF222998, which is actually C. hominis (2.0%) (see Table 2). Furthermore, in histological studies on parasites in freshwater ornamental fish collected in aquarium fish shops in Iran, Cryptosporidium spp. were detected by histology in the 16% of examined fish, specifically in sailfin molly (Poecilia latipinna), Siamese fighting fish (Betta splendens), gourami (T. leeri), rosy barb (Puntius conchonius), platy fish (Xiphophorus maculatus), angelfish (P. scalare), electric yellow (Labidochromis caeruleus), goldfish (C. auratus), Oscar fish (A. ocellatus) and slender rainbow (Melanotaenia gracilis) (Nematollahi et al., Reference Nematollahi, Jaberi, Ashrafi Helan and Sheikhzadeh2016).
Pathology and clinical manifestations
Although several authors have reported high morbidity and mortality rates, mainly in juvenile fish specimens, the pathology of cryptosporidiosis in fish has not been extensively studied (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Ryan et al., Reference Ryan, O'Hara and Xiao2004; Yang et al., Reference Yang, Dorrestein and Ryan2016). However, infection by Cryptosporidium is known to be influenced by the following: (1) the species/genotype considered; (2) the host fish species; (3) the age/size of the fish and (4) co-infections with other pathogens (Sitjà-Bobadilla et al., Reference Sitjà-Bobadilla, Padrós, Aguilera and Álvarez-Pellitero2005, Reference Sitjà-Bobadilla, Pujalte, Macián, Pascual, Álvarez-Pellitero and Garay2006).
In most cases of Cryptosporidium infection, no clinical signs of disease are observed, although several authors have described clinical manifestations such as emaciation, atrophy of skeletal muscle, flattening of the abdomen, low growth rate, anorexia, listlessness, whitish feces, abdominal swelling and ascites (Hoover et al., Reference Hoover, Hoerr, Carlton, Hinsman and Ferguson1981; Gratzek, Reference Gratzek and Stoskopf1993; Camus and López, Reference Camus and López1996; Muench and White, Reference Muench and White1997; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004; Ryan et al., Reference Ryan, O'Hara and Xiao2004; Murphy et al., Reference Murphy, Bradway, Walsh, Sanders and Snekvik2009; Gabor et al., Reference Gabor, Srivastava, Titmarsh, Dennis, Gabor and Landos2011; Nematollahi et al., Reference Nematollahi, Jaberi, Ashrafi Helan and Sheikhzadeh2016).
Some studies have demonstrated the existence of histopathological damage induced by the accumulation of oocysts of C. molnari and C. scophthalmi (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004). The presence of large vacuoles disturbing the usual architecture of the mucosa, followed by massive necrosis of the epithelial cells and the consequent cellular detachment in specimens with high intensities of infection have been reported (Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004). Furthermore, a strong inflammatory response and intense leucocyte infiltration have also been observed (Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004; Yang et al., Reference Yang, Dorrestein and Ryan2016).
Fish as potential hosts of zoonotic Cryptosporidium species
The use of molecular subtyping tools has led to a better understanding of the transmission dynamics of Cryptosporidium infections in humans and animals. The most widely used marker is a hypervariable portion of the 60-kDa glycoprotein (GP60) gene, sequencing of which enables identification of human-specific, animal-specific and zoonotic subtypes. Regarding Cryptosporidium in fish, the GP60 gene has been amplified in very few studies. Thus, C. hominis subtype IdA15G1 was detected in wild marine fish, specifically in mackerel scad (Decapterus macarellus) from Papua New Guinea (Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013), and subtype IbA10G2 was identified in a freshwater goldfish (C. auratus) from a farm in Western Australia (Palermo, Reference Palermo2016). Other studies have identified different C. parvum subtypes in both freshwater and marine fish, including subtypes IIaA13G1R1, IIaA14G2R1, IIaA15G2R1, IIaA16G2R1, IIaA17G2R1, IIaA18G3R1 and IIaA19G4R1 (see Table 3) (Reid et al., Reference Reid, Lymbery, Ng, Tweedle and Ryan2010; Koinari et al., Reference Koinari, Karl, Ng-Hublin, Lymbery and Ryan2013; Certad et al., Reference Certad, Dupouy-Camet, Gantois, Hammouma-Ghelboun, Pottier, Guyot, Benamrouz, Osman, Delaire, Creusy, Viscogliosi, Dei-Cas, Aliouat-Denis and Follet2015, Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019; Couso-Pérez et al., Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019). However, to date there are no reports on the identification of piscine Cryptosporidium species/genotypes in human hosts (Ryan et al., Reference Ryan, Zahedi, Feng and Xiao2021).
Subtypes of C. parvum identified in piscine hosts by analysis of the GP60 gene
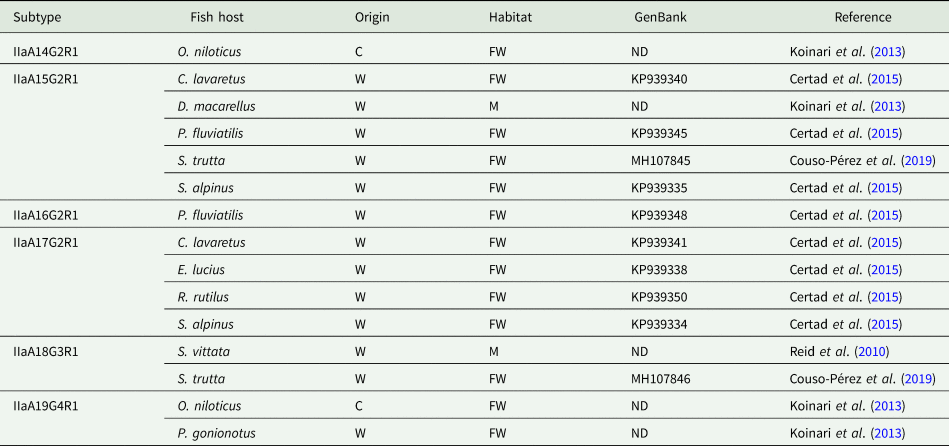
C, cultured; FW, freshwater; M, marine; O, ornamental; W, wild; ND, no data.
Detection of zoonotic Cryptosporidium subtypes in fish raises the following questions: Are fish actually infected by Cryptosporidium spp. that infect mammals or are they simply carriers of this protozoan parasite? If so, would this be important from a public health point of view? Detection of zoonotic Cryptosporidium species/genotypes in fish collected from the environment does not necessarily indicate real infection, because fish can act as mechanical carriers, as low prevalences and parasite loads are usually detected. Thus, experimental cross-transmission and/or histological studies are needed to confirm the role of fish as competent hosts for zoonotic Cryptosporidium species. Several authors have considered whether Cryptosporidium of mammalian origin can infect lower vertebrate hosts. Graczyk et al. (Reference Graczyk, Fayer and Cranfield1996) inoculated (by gastric intubation) some fish, amphibians and reptiles with C. parvum (AUCP-1 strain) oocysts obtained from the feces of experimentally infected Holstein calves, which were infectious to neonatal BALB/c mice. Nevertheless, histological sections from gastrointestinal tissues were negative for developmental stages of Cryptosporidium. These authors consider that C. parvum does not infect fish, but that under some circumstances such as after the ingestion of C. parvum-infected prey, lower vertebrates may disseminate oocysts in the environment by acting as mechanical carriers.
However, other authors consider that C. parvum can infect piscine hosts. Thus, Freire-Santos et al. (Reference Freire-Santos, Vergara-Castiblanco, Tojo-Rodríguez, Santamarina-Fernández and Ares-Mazás1998) experimentally infected rainbow trout (O. mykiss) with C. parvum oocysts collected from a naturally infected Friesian-Holstein neonatal calf, after evaluating the viability and infectivity of the oocysts in a suckling murine model. During the histological examination, no life-cycle stages of Cryptosporidium were observed in any part of the apical border of the digestive tract sections. However, large numbers of 5–7 μm spherical structures compatible with Cryptosporidium developmental stages were found located deep within the epithelial tissue of pyloric caeca sections. Moreover, an indirect fluorescent antibody test with immunoglobulin (Ig) M and IgG anti-Cryptosporidium antibodies revealed fluorescence reactivity in these structures, the number of which increased remarkably when the specimens were subjected to stress conditions.
Similarly, Couso-Pérez et al. (Reference Couso-Pérez, García-Estévez, Leiro-Vidal, Ares-Mazás and Gómez-Couso2016) experimentally infected 25 young specimens of cultured turbot (P. maxima), of weight 20–40 g, with C. parvum oocysts (subtype IIaA16G3R1) obtained from a naturally infected neonatal calf. The fish were maintained for 2 h under stress conditions in a tank contaminated with 25 × 106 purified C. parvum oocysts and then placed in another tank with clean sea water. The application of a direct immunofluorescence method with monoclonal anti-Cryptosporidium antibodies led to microscopic detection of C. parvum oocysts in the intestine and the pyloric caeca from fish collected 7 and 10 days post-exposure, respectively. Considering that C. parvum requires between 48 and 72 h to complete its life cycle in a suitable host and that the emptying time of turbot gastrointestinal tract is approximately 24 h, the authors suggested that C. parvum infection in young cultured turbot is possible, although infection must be confirmed by further histological studies (Couso-Pérez et al., Reference Couso-Pérez, García-Estévez, Leiro-Vidal, Ares-Mazás and Gómez-Couso2016). Moreover, in response to high mortality rates detected in different fish farm systems in Galicia (NW Spain), the presence of Cryptosporidium in pyloric caeca and intestinal homogenates from cultured turbot (P. maxima) was investigated, revealing the presence of Cryptosporidium sp. oocysts in 21 of 29 (72.4%) juvenile specimens examined, with a mean intensity of 14.6 oocysts/turbot. Subsequent molecular characterization identified the hypertransmissible subtype IIaA15G2R1 of C. parvum (unpublished results).
Recent epidemiological studies have suggested that C. parvum can complete its life cycle and multiply in piscine hosts. Thus, histological analysis of gastric and intestine sections from C. parvum polymerase chain reaction positive freshwater and marine fish carried out by Certad et al. (Reference Certad, Dupouy-Camet, Gantois, Hammouma-Ghelboun, Pottier, Guyot, Benamrouz, Osman, Delaire, Creusy, Viscogliosi, Dei-Cas, Aliouat-Denis and Follet2015, Reference Certad, Follet, Gantois, Hammouma-Ghelboun, Guyot, Benamrouz-Vanneste, Fréalle, Seesao, Delaire, Creusy, Even, Verrez-Bagnis, Ryan, Gay, Aliouat-Denis and Viscogliosi2019) revealed the presence of round bodies resembling developmental stages of Cryptosporidium in an apical position within the cells. As a consequence of these observations, the authors suggested that C. parvum was actually infecting fish, rather than being passively transmitted. Couso-Pérez et al. (Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2018, Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019) subjected gastrointestinal samples of brown and rainbow trout (S. trutta and O. mykiss, respectively) to a homogenization treatment using a bar homogenizer and applied a monoclonal anti-Cryptosporidium antibody test, detecting clusters of oocysts in the pyloric caeca of both fish species. Molecular analysis revealed that oocysts belonged to the species C. parvum, specifically subtypes IIaA15G2R1 and IIaA18G3R1. These findings suggest the existence of real infections, because fully sporulated oocysts, which can appear in clusters in the same infected cell, are only found within the vacuolar space located deep in the mucosal epithelium in piscine hosts (Landsberg and Paperna, Reference Landsberg and Paperna1986; Álvarez-Pellitero and Sitjà-Bobadilla, Reference Álvarez-Pellitero and Sitjà-Bobadilla2002; Álvarez-Pellitero et al., Reference Álvarez-Pellitero, Quiroga, Sitjà-Bobadilla, Redondo, Palenzuela, Padrós, Vázquez and Nieto2004; Ryan et al., Reference Ryan, O'Hara and Xiao2004, Reference Ryan, Paparini, Tong, Yang, Gibson-Kueh, O'hara, Lymbery and Xiao2015; Bolland et al., Reference Bolland, Zahedi, Oskam, Murphy and Ryan2020; Zahedi et al., Reference Zahedi, Bolland, Oskam and Ryan2021). The homogenization treatment to which the samples were subjected would have ruptured the epithelial cells, thereby releasing oocysts, individually or in clusters. Unfortunately, histological analysis of the samples was not possible as the tissue had been deteriorated by autolysis (Couso-Pérez et al., Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2018, Reference Couso-Pérez, Ares-Mazás and Gómez-Couso2019).
Identification of zoonotic C. parvum subtypes in piscine hosts suggests that fish are a potential source of infection in humans, with an associated risk to public health. Anglers and food handlers could be infected directly while eviscerating or preparing the captured specimens or indirectly via contact with contaminated surfaces or fomites during the storage of the fish. The only study that has quantified the risk of infection by Cryptosporidium was carried out by Roberts et al. (Reference Roberts, Silbergeld and Graczyk2007) and was based on fish and hand wash samples taken from urban anglers in Baltimore (USA). Using the United States Environmental Protection Agency's dose–response model, these authors determined that the mean probability of acquiring Cryptosporidium infection in anglers was 41.0% (although it could reach 100%) on the basis of positive fish samples. A significantly higher mean probability of infection (91.0%) was estimated on the basis of positive hand wash samples. When all data (positive fish and hand wash samples) were considered and, depending on host factors, such as immune status and pre-existing illnesses, it was estimated that on average 1–8 out of 10 anglers could become infected by Cryptosporidium (Roberts et al., Reference Roberts, Silbergeld and Graczyk2007). Thus, edible fish would extend the range of foodstuffs involved in the transmission of cryptosporidiosis, the aetiological agent of which is responsible for 8.6 million cases of foodborne illness annually worldwide (Ryan et al., Reference Ryan, Hijjawi and Xiao2018; Moratal et al., Reference Moratal, Dea-Ayuela, Cardells, Marco-Hirs, Puigcercós, Lizana and López-Ramon2020; Zahedi and Ryan, Reference Zahedi and Ryan2020).
Conclusions
Research on Cryptosporidium in piscine hosts has increased in recent years, reaffirming the ubiquitous nature of this protozoan parasite, which has been detected in a large number of free-living, cultured and ornamental fish species worldwide, from both marine and freshwater environments. Future studies will increase the range of piscine hosts, and novel Cryptosporidium species and genotypes will be proposed. Therefore, the taxonomy and evolutionary relationships in the genus must be clarified to enable consensus to be reached regarding the nomenclature used to designate new piscine species/genotypes. Whole genome sequencing of Cryptosporidium species/genotypes from fish is also required to assist the taxonomic clarification. Moreover, experimental cross-transmission and/or histological studies are needed to confirm the role of fish as competent hosts for zoonotic Cryptosporidium species as very few such studies have been conducted to date.
The existence of Cryptosporidium infections in cultured fish may have a significant economic impact on the aquaculture industry due to the morbidity and mortality rates reported in some fish species. The implementation of control programmes may be necessary to remove the Cryptosporidium oocysts from aquaculture facilities, where fish cohabit in dense groups and are subjected to other stress factors that can enhance transmission of this parasite.
Finally, the identification of zoonotic Cryptosporidium species in edible fish extends the range of foodstuffs potentially involved in the transmission of cryptosporidiosis, representing a risk to public health, although further risk assessment studies are required to confirm this possibility.
Author contributions
All authors contributed equally to the conceptualization, investigation (data collection), writing and editing this review article.
Financial support
This study was funded by the Autonomous Government of Galicia (grant ED431C 2021/26).
Conflict of interest
The authors declare there are no conflicts of interest.