Introduction
Cryptosporidiosis is a diarrhoeal disease caused by Cryptosporidium species, ubiquitous enteric parasites that can infect a variety of hosts, including humans (Chalmers and Giles, Reference Chalmers and Giles2010; Xiao, Reference Xiao2010). The global pooled prevalence of Cryptosporidium infection in people is estimated at 7.6% (Dong et al., Reference Dong, Yang, Wang, Yang, Yang, Shi, Li, Li, Chen, Jiang and Zhou2020). In high-income countries, the reported prevalence ranges from 0.1 to 14.1% with an average estimate of 4.3%, whilst in low-income countries the reported prevalence ranges from 1.3 to 31.5% and an average estimate of 10.4% (Fayer, Reference Fayer2004; Dong et al., Reference Dong, Yang, Wang, Yang, Yang, Shi, Li, Li, Chen, Jiang and Zhou2020; Liu et al., Reference Liu, Gong, Liu, Shen, Wu, Zhang and Cao2020). Asymptomatic carriage is usually low, particularly in high-income countries, and has been reported to range from 0 to 4% (Enserink et al., Reference Enserink, van den Wijngaard, Bruijning-Verhagen, van Asten, Mughini-Gras, Duizer, Kortbeek, Scholts, Nagelkerke, Smit, Kooistra-Smid and van Pelt2015; Reh et al., Reference Reh, Muadica, Köster, Balasegaram, Verlander, Chércoles and Carmena2019). Further, while it has been reported that Cryptosporidium is present in 1% of stools of immunocompetent individuals in high-income countries, the pathogen is detected in 5–10% of stools of people in low-income countries (Checkley et al., Reference Checkley, White, Jaganath, Arrowood, Chalmers, Chen, Fayer, Griffiths, Guerrant, Hedstrom, Huston, Kotloff, Kang, Mead, Miller, Petri, Priest, Roos, Striepen, Thompson, Ward, Van Voorhis, Xiao, Zhu and Houpt2015). However, the diagnosis of pathogens causing gastroenteritis may be as low as 1% of true prevalence in high-income countries and even lower in low-income countries (Scallan et al., Reference Scallan, Hoekstra, Angulo, Tauxe, Widdowson, Roy, Jones and Griffin2011).
Cryptosporidium is responsible for several well-documented food- and waterborne as well as occupational outbreaks (Karanis et al., Reference Karanis, Kourenti and Smith2006; Baldursson and Karanis, Reference Baldursson and Karanis2011; Widerström et al., Reference Widerström, Schönning, Lilja, Lebbad, Ljung, Allestam, Ferm, Björkholm, Hansen, Hiltula, Långmark, Löfdahl, Omberg, Reuterwall, Samuelsson, Widgren, Wallensten and Lindh2014; Efstratiou et al., Reference Efstratiou, Ongerth and Karanis2017; Hancock-Allen et al., Reference Hancock-Allen, Alden and Cronquist2017). A cryptosporidiosis outbreak can be defined as 2 or more cases epidemiologically linked to a common source by location and time of exposure (Gharpure et al., Reference Gharpure, Perez, Miller, Wikswo, Silver and Hlavsa2019). Transmission can occur by ingestion of contaminated water or food and through contact with the feces of infected people or animals (Fayer et al., Reference Fayer, Morgan and Upton2000). There were 120 Cryptosporidium waterborne outbreaks reported worldwide from 2004 to 2010 (Baldursson and Karanis, Reference Baldursson and Karanis2011). Outbreaks are more commonly notified in high-income countries due to better notification and surveillance systems (Efstratiou et al., Reference Efstratiou, Ongerth and Karanis2017). Waterborne outbreaks can be very large with some of the largest reported from Europe and the USA. For example, a waterborne cryptosporidiosis outbreak in Milwaukee, USA, affected more than 400 000 people during 2 months in 1993, with an associated cost estimated over US$96 million (Corso et al., Reference Corso, Kramer, Blair, Addiss, Davis and Haddix2003). In Sweden, 2 waterborne outbreaks were reported in 2010 and 2011 affecting about 47 000 people (Widerström et al., Reference Widerström, Schönning, Lilja, Lebbad, Ljung, Allestam, Ferm, Björkholm, Hansen, Hiltula, Långmark, Löfdahl, Omberg, Reuterwall, Samuelsson, Widgren, Wallensten and Lindh2014). Between 2004 and 2014, 28.5% of the worldwide waterborne cryptosporidiosis outbreaks were reported in New Zealand (Efstratiou et al., Reference Efstratiou, Ongerth and Karanis2017).
Cryptosporidiosis has been a notifiable disease in New Zealand since 1996 (Learmonth et al., Reference Learmonth, Ionas, Ebbett and Kwan2004). The reported incidence of the disease in humans from New Zealand is very high, even though it is estimated that only 2% of acute gastrointestinal illnesses, including cryptosporidiosis, are notified (Snel et al., Reference Snel, Baker and Venugopal2009a, Reference Snel, Baker, Kamalesh, French and Learmonth2009b). Compared to other high-income countries, New Zealand reports between 26.1 and 32.3 new cases per 100 000 population per year, whilst Australia reports 12.8 per 100 000 population, England and Wales 8.0 per 100 000, USA 2.9 per 100 000, and Canada 2.7 per 100 000 (Learmonth et al., Reference Learmonth, Ionas, Ebbett and Kwan2004; Lal et al., Reference Lal, Cornish, Fearnley, Glass and Kirk2015; Garcia-R et al., Reference Garcia-R, Pita, Velathanthiri, French and Hayman2020). Cryptosporidiosis incidence in New Zealand is suggested to be increasing partly because of outbreaks from untreated water and farming. However, the number of cases involved in most outbreaks is small compared to outbreaks in other high-income countries (Stefanogiannis et al., Reference Stefanogiannis, McLean and Van Mil2001; Snel et al., Reference Snel, Baker, Kamalesh, French and Learmonth2009b; Grinberg et al., Reference Grinberg, Pomroy, Squires, Scuffham, Pita and Kwan2011). This might not only reflect high disease burden, but also effective surveillance and primary health care reporting.
Here, we use reports from outbreaks occurring since 2010 submitted by the Institute of Environmental Science and Research Ltd (ESR) to the Public Health Surveillance website (https://surv.esr.cri.nz/index.php) to highlight emerging aspects of cryptosporidiosis in New Zealand. We also employed molecular data records collected by our laboratory to describe epidemiologic features and gain a better understanding of species and subtypes involved in cryptosporidiosis outbreaks in New Zealand.
Methods
We used the cryptosporidiosis outbreak data provided by the Public Health Surveillance website (https://surv.esr.cri.nz/index.php) over the years 2010–2021. We also extracted data on the modes of transmission and hospitalized cases when available from the Public Health Surveillance website. Before 2017, the data about hospitalized cases and modes of transmissions were individually documented for each pathogen, including Cryptosporidium, but since 2018 this information was supplied for all notifiable diseases, not allowing us to collect the data for outbreaks caused by Cryptosporidium only. Sources and modes of transmission are reported following the ESR Guidelines for the Investigation and Control of Disease Outbreaks (Institute of Environmental Science & Research Limited, 2012).
Our laboratory, the Protozoa Research Unit at Hopkirk Research Institute (Massey University), has used molecular characterization of Cryptosporidium-positive stool samples from symptomatic humans visiting private and public general practitioners (GP) throughout New Zealand under a Ministry of Health contract for genetic surveillance. Some of the samples sent to our lab are from cases associated with outbreaks. We collected the genetic variant(s) involved in 87 human samples from historic cases of cryptosporidiosis outbreaks occurring between 2010 and 2021 (Table 1) previously analysed by extracting DNA and polymerase chain reaction (PCR) molecular typing of the gp60 gene. The sequences have been previously submitted to GenBank (Garcia-R et al., Reference Garcia-R, French, Pita, Velathanthiri, Shrestha and Hayman2017, Reference Garcia-R, Pita, Velathanthiri, French and Hayman2020). High quality and accessible archive of gp60 forward chromatograms generated in our laboratory using Sanger sequencing for 23 samples involved in the outbreaks were used in the Tracking of Indels by DEcomposition (TIDE) algorithm (Dettwiler et al., Reference Dettwiler, Troell, Robinson, Chalmers, Basso, Rentería-Solís, Daugschies, Mühlethaler, Dale, Basapathi Raghavendra, Ruf, Poppert, Meylan and Olias2021) to identify mixed subtype infections and stutter artefacts. We re-investigate the chromatograms generated by Sanger sequencing in 6 outbreaks (Taranaki 2013 and 2021, Auckland 2015 and 2017, Wellington 2013 and Blenheim 2017) to identify shifts in the target sequences and the likelihood of multiplicity of infections using the TIDE algorithm. Forward strand chromatograms from other outbreaks were not located or have low quality, making them unsuitable for the TIDE analysis. We follow Dettwiler et al. (Reference Dettwiler, Troell, Robinson, Chalmers, Basso, Rentería-Solís, Daugschies, Mühlethaler, Dale, Basapathi Raghavendra, Ruf, Poppert, Meylan and Olias2021) in applying a conservative threshold for differentiating mixed infections from stutter (a reverse stutter filter of 15% of sequences for N–3 and 8% of sequences for N–6).
Cryptosporidiosis outbreaks between 2010 and 2021 reported in this study

Results
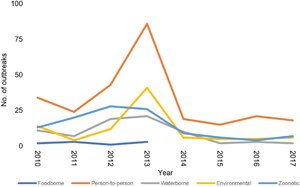
Cryptosporidiosis outbreak-associated cases from 2010 to 2021 show distinct peaks in various years with the highest number of notifications in 2013 (n = 547), followed by 2010 (n = 294) and 2018 (n = 209) (Fig. 1). For the period 2010–2017, 20 hospitalizations and multiple modes of transmission associated with different risk factors were reported. The most important mode of transmission was person-to-person (primary infections and secondary or close contacts infections), related to 260 outbreaks and 1320 cases, followed by 113 outbreaks associated with animals, resulting in 436 human cases (Table 2). From 2018 to 2021, there were 37 cryptosporidiosis outbreaks associated with 324 cases (Fig. 1).
Reported cryptosporidiosis outbreaks (grey) and the number of associated cases with outbreaks (black) by year. The decrease in cryptosporidiosis cases since 2020 is associated with Covid-19 restrictions (Knox et al., Reference Knox, Garcia-R, Ogbuigwe, Pita, Velathanthiri and Hayman2021).

Cryptosporidiosis outbreaks by mode of transmission and exposure from 2010 to 2017

Note that outbreaks can be associated with different modes of transmission when evidence to implicate 1 specific contributing factor is complicated or insufficient.
Our laboratory has been involved in the molecular characterization of samples associated with 11 cryptosporidiosis outbreaks. Two outbreaks during March 2010 that occurred in Auckland and Christchurch showed C. hominis 1bA10G2 as the most predominant variant. The outbreak in Auckland was associated with a contaminated swimming pool (Table 1). We also describe the molecular epidemiology of 4 outbreaks during the summer/autumn period of 2013 in Hawkes Bay, Waikato, Wellington and Taranaki regions. The outbreak in Hawkes Bay was associated with C. hominis IgA17, the outbreak in Taranaki with C. hominis 1bA10G2, whilst the outbreaks in Wellington and Waikato showed both subtypes. In 2015, we were involved in an outbreak in Auckland related to the consumption of raw milk. Samples were positive for C. parvum IIaA18G3R1. In May 2017, we identified C. hominis subtypes IgA20 and IbA10G2 in an outbreak in Auckland. In June 2017, a swimming pool complex in Blenheim was linked to a cryptosporidiosis outbreak with C. hominis subtype IbA10G2 and C. parvum subtype IIdA24G1. In May and September 2021, 2 outbreaks associated with the consumption of raw milk in Taranaki were found to be caused by both C. parvum IIaA18G3R1 and IIaA19G4R1 subtypes. All gp60 chromatograms analysed using the TIDE algorithm (Fig. 2) were found with stutter artefacts without potential coinfections with different subtypes within individual people. We mainly found reverse N–3 (main allele minus 3 nucleotides) and N–6 (allele minus 6 nucleotides) stutter artefact peaks in TIDE. Curiously, the outbreak in Wellington 2013 showed peaks at N–2 and N–3 (Supplementary Fig. S1), and multiple peaks at varying positions on samples in the outbreak at Taranaki 2021 (Table S1).
Indel spectrums from TIDE analyses of sequences from different outbreaks in New Zealand. Outbreak location, year of collection and sample numbers are shown. Y-axis is the percentage of sequences, X-axis the relative position and bar colours show P value <0.001 (red) or not (blue). Underlying codon-size variations (e.g. −6 nt and −3 nt) suggest stutter artefacts, not coinfection with multiple strains in the sample.

Discussion
Cryptosporidium is one of the most common pathogens causing infectious disease outbreaks in New Zealand, along with Giardia and norovirus. From 2010 to 2017, Cryptosporidium was implicated in 318 (6%) of the 5230 infectious disease outbreaks reported in New Zealand. In 2013, Cryptosporidium was the second most reported agent causing gastrointestinal diseases, accounting for 15% of outbreaks (98/652) and 7.7% of associated cases (547/7137) (https://surv.esr.cri.nz/surveillance/annual_surveillance.php). During 2013 there were a high number of cryptosporidiosis cases (1348 cases in total), many (547 cases) associated with outbreaks (Garcia-R et al., Reference Garcia-R, Pita, Velathanthiri, French and Hayman2020).
Cryptosporidiosis outbreaks in New Zealand have been reported from farm environments, food sources, contaminated water and following anthroponotic transmission. Our molecular work identified 8 of the 11 outbreaks to be caused by C. hominis subtypes. This and the high incidence of person-to-person transmission as a mode of transmission indicates that C. hominis is the most common species in outbreaks and dominated by anthropogenic transmission, which differs to surveillance cases where C. parvum is more dominant (Garcia-R and Hayman, Reference Garcia-R and Hayman2017; Garcia-R et al., Reference Garcia-R, French, Pita, Velathanthiri, Shrestha and Hayman2017, Reference Garcia-R, Pita, Velathanthiri, French and Hayman2020). The importance of C. hominis in person-to-person transmission can also be inferred by the absence of C. hominis cases reported and the lack of C. hominis outbreaks in 2020 during the Covid-19 restrictions in New Zealand (Knox et al., Reference Knox, Garcia-R, Ogbuigwe, Pita, Velathanthiri and Hayman2021). Limitations on international travel and nationwide lockdown limited person-to-person transmission, but not human–animal contacts, evidenced by continued C. parvum cases. Transmission rates in the UK have also been found to be higher for C. hominis compared with C. parvum and other species, mainly in households, child-care facilities, nursing homes and schools (McKerr et al., Reference McKerr, Chalmers, Elwin, Ayres, Vivancos, O'Brien and Christley2022).
Waterborne and foodborne outbreaks of cryptosporidiosis have occurred less frequently than person-to-person transmission. However, this may be related to an absence of notification and a lack of investigation rather than to truly different epidemiological situations. Sporadic cases associated with water and food are likely unrecognized. Cryptosporidium has been responsible for numerous waterborne outbreaks worldwide (Baldursson and Karanis, Reference Baldursson and Karanis2011; Efstratiou et al., Reference Efstratiou, Ongerth and Karanis2017). Outbreaks linked to recreational waters, especially swimming pools, have increased in the UK and the USA (Cacciò and Chalmers, Reference Cacciò and Chalmers2016). In the USA, a review of 444 cryptosporidiosis outbreaks resulting in 7465 cases between 2009 and 2017 revealed that contaminated recreational water (treated and untreated) was responsible for 60% (4495/7465) of cases, 38% of outbreaks and 64% (n = 186) of hospitalizations (Gharpure et al., Reference Gharpure, Perez, Miller, Wikswo, Silver and Hlavsa2019). In addition, treated recreational water-associated outbreaks increased by ~14% per year between 2009 and 2016 (Gharpure et al., Reference Gharpure, Perez, Miller, Wikswo, Silver and Hlavsa2019). In Australia, a review between 2001 and 2007 of waterborne outbreaks identified 78% attributed to recreational water, 98% of them caused by Cryptosporidium (Dale et al., Reference Dale, Kirk, Sinclair, Hall and Leder2010). Recreational water-associated outbreaks worldwide have predominantly been caused by C. hominis (Causer et al., Reference Causer, Handzel, Welch, Carr, Culp, Lucht, Mudahar, Robinson, Neavear, Fenton, Rose, Craig, Arrowood, Wahlquist, Xiao, Lee, Mirel, Levy, Beach, Poquette and Dworkin2006; Cope et al., Reference Cope, Prosser, Nowicki, Roberts, Roberts, Scheer, Anderson, Longsworth, Parsons, Goldschmidt, Johnston, Bishop, Xiao, Hill, Beach and Hlavsa2015; Chalmers et al., Reference Chalmers, Robinson, Elwin and Elson2019; Zahedi and Ryan, Reference Zahedi and Ryan2020; Ryan et al., Reference Ryan, Hill and Deere2022). For instance, C. hominis is responsible for all recreational water-associated outbreaks in Australia and, similarly to the UK and NZ, it has been found to have a higher infectivity for humans than C. parvum (Ryan et al., Reference Ryan, Hill and Deere2022).
Cryptosporidium hominis subtype IbA10G2 was found in outbreaks in Auckland, Christchurch and Taranaki and it is highly prevalent in outbreak cases in Europe, Australia and the USA (Chalmers et al., Reference Chalmers, Hadfield, Jackson, Elwin, Xiao and Hunter2008; Ng et al., Reference Ng, MacKenzie and Ryan2010). Subtype IgA17 of C. hominis was involved in a large outbreak in Hawkes Bay (reported here) and other subsequent outbreaks in the North Island of New Zealand during the summer and autumn of 2013. Before 2013, the subtype Ig was not previously registered in New Zealand (Garcia-R et al., Reference Garcia-R, French, Pita, Velathanthiri, Shrestha and Hayman2017, Reference Garcia-R, Pita, Velathanthiri, French and Hayman2020). This subtype was relatively rare overseas, but it has been isolated from human cases in Australia (Ng et al., Reference Ng, MacKenzie and Ryan2010). It might indicate that IgA17 was introduced in New Zealand from Australia with subsequent widespread disseminations from the Hawkes Bay to other regions, potentially following several swimming pool-related outbreaks. Cryptosporidium parvum was less commonly found in waterborne outbreaks and mainly associated with consumption of raw milk. Nonetheless, subtype IIaA18G3R1, identified in some outbreaks in this study, has been reported as a cause of waterborne outbreaks of human cryptosporidiosis in Europe and Australia (Glaberman et al., Reference Glaberman, Moore, Lowery, Chalmers, Sulaiman, Elwin, Rooney, Millar, Dooley, Lal and Xiao2002; Chalmers et al., Reference Chalmers, Ferguson, Cacciò, Gasser, Abs EL-Osta, Heijnen, Xiao, Elwin, Hadfield, Sinclair and Stevens2005). Zoonotic and environmental transmissions account for almost half of Cryptosporidium outbreaks, possibly linked to farming and contamination of drinking and recreational water (Phiri et al., Reference Phiri, Pita, Hayman, Biggs, Davis, Fayaz, Canning, French and Death2020; Grout et al., Reference Grout, Marshall, Hales, Baker and French2022).
Variability in the sequence of the gp60 gene has been useful for inferring transmission routes. However, different gp60 subtypes can be linked to the same outbreak, which may indicate a coinfection of multiple subtypes, or an artefact caused by DNA polymerase slippage during the PCR elongation step. TIDE allowed us to differentiate between stutter effects and coinfection by multiple subtypes in the samples. The TIDE analyses in our study identified stutters within outbreak sample sequences but did not support evidence of mixed subtype infections. Most effects were N-3 or, less frequently, N−6. For example, the outbreak in Taranaki 2013 was caused by C. hominis subtype IbA10G2 and there was evidence of N−3 stutter. However, the outbreak in Wellington 2013 showed peaks at N−2 (unexpected result because they should be a multiple of 3) and N−3 and the outbreak in Taranaki 2021 presented multiple peaks. None reached the level of support suggested by Dettwiler et al. (Reference Dettwiler, Troell, Robinson, Chalmers, Basso, Rentería-Solís, Daugschies, Mühlethaler, Dale, Basapathi Raghavendra, Ruf, Poppert, Meylan and Olias2021); however, further analyses such as metabarcoding might identify mixed infections, including mixed family subtypes and species (Garcia-R et al., Reference Garcia-R, Ogbuigwe, Pita, Velathanthiri, Knox, Biggs, French and Hayman2021; Ogbuigwe et al., Reference Ogbuigwe, Biggs, Garcia-Ramirez, Knox, Pita, Velathanthiri, French and Hayman2022).
Although cryptosporidiosis in New Zealand is a notifiable disease, there are some variations in diagnoses that may affect the identification, classification and characterization of outbreaks and implicated cases (Shirley et al., Reference Shirley, Moonah and Kotloff2012). For instance, the infection may be asymptomatic but meeting the clinical description in New Zealand requires the patient to show compatible symptoms accompanied by laboratory definitive evidence. Localized outbreaks can be missed by routine surveillance and/or sporadic cases (including infected individuals with no access to a local GP) being part of unrecognized outbreaks and likely unreported (Tam et al., Reference Tam, Rodrigues, Viviani, Dodds, Evans, Hunter, Gray, Letley, Rait, Tompkins and Brien2012; Briggs et al., Reference Briggs, Boxall, Van Santen, Chalmers and McCarthy2014). Additionally, most pathology laboratories in New Zealand used antigen and microscopy-based techniques for the diagnosis of cryptosporidiosis at the time of the study period, which are 83.7% less sensitive and 98.9% less specificity compared to PCR (Morgan et al., Reference Morgan, Pallant, Dwyer, Forbes, Rich and Thompson1998; Mergen et al., Reference Mergen, Espina, Teal and Madison-Antenucci2020), and can overlook positive outbreak samples. Nevertheless, diagnostic laboratories are starting to switch to multiplexed qPCR (Hayman et al., Reference Hayman, Garcia-Ramirez, Pita, Velathanthiri, Knox, Ogbuigwe, Baker, Rostami, Deroles-Main and Gilpin2020). The change among laboratories to molecular techniques will improve control programmes. However, the availability and value of subtyping provide a more in-depth and robust understanding of cryptosporidiosis epidemiology (e.g. the epidemiological interpretation of subtype occurrence and distribution trends), sources and transmission of outbreaks, and potentially, outbreaks detection to public health agencies. In the future, the application of whole-genome sequencing will likely allow population-based screening of Cryptosporidium species/subtypes and advance our understanding of cryptosporidiosis even further.
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/S0031182023000288.
Data availability
The nucleotide sequences of the partial gp60 gene were deposited in the GenBank database under accession numbers that can be found in Garcia-R et al., 2017; Garcia-R et al., 2020.
Author's contribution
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by J. C. G.-R. The first draft of the manuscript was written by J. C. G.-R. and D. T. S. H. commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Financial support
This work was supported by funds from the New Zealand Ministry of Health contract number 355766-02, Lewis Fitch, Palmerston North Medical Research Foundation, Wairarapa Vet Assn Research Fund, Royal Society Te Apārangi Rutherford Discovery Fellowship (RDF-MAU1701) and the Percival Carmine Chair in Epidemiology and Public Health.
Conflict of interest
None.
Ethical standards
The authors confirm that the ethical policies of the journal, as noted on the journal's author guidelines page, have been adhered to. No ethical approval was required as this is a review article with no original research data.