Introduction
Microsporidians are intracellular microparasites related to the kingdom Fungi (Hirt et al., Reference Hirt, Logsdon, Healy, Dorey, Doolittle and Embley1999; Capella-Gutiérrez et al., Reference Capella-Gutiérrez, Marcet-Houben and Gabaldón2012). They infect a wide range of hosts from single-celled eukaryotes to vertebrates (Smith, Reference Smith2009). Depending on the species, they can develop in various host tissues where they form spores that are infective for the next host. Additionally, some microsporidians can be transmitted vertically from the mother to the offspring (Smith, Reference Smith2009). Microsporidia are frequently found in aquatic systems where the majority of known species infects aquatic invertebrates (Stentiford et al., Reference Stentiford, Feist, Stone, Bateman and Dunn2013; Stentiford and Dunn, Reference Stentiford, Dunn, Weiss and Becnel2014). While there are numerous studies on microsporidians in aquatic crustaceans such as amphipods (Krebes et al., Reference Krebes, Blank, Frankowski and Bastrop2010; Grabner et al., Reference Grabner, Weigand, Leese, Winking, Hering, Tollrian and Sures2015, Reference Grabner, Weber and Weigand2020; Madyarova et al., Reference Madyarova, Adelshin, Dimova, Axenov-Gribanov, Lubyaga and Timofeyev2015; Weigand et al., Reference Weigand, Kremers and Grabner2016; Bacela-Spychalska et al., Reference Bacela-Spychalska, Wróblewski, Mamos, Grabowski, Rigaud, Wattier, Rewicz, Konopacka and Ovcharenko2018; Quiles et al., Reference Quiles, Bacela-Spychalska, Teixeira, Nicolas, Grabowski, Rigaud and Wattier2019, Reference Quiles, Wattier, Bacela-Spychalska, Grabowski and Rigaud2020, Reference Quiles, Rigaud, Wattier, Grabowski and Bacela-Spychalska2021), to our knowledge the only microsporidium reported from Asellus aquaticus is Mrazekia argoisi that was detected in fat body cells of the host (Kudo, Reference Kudo1924).
The freshwater isopod A. aquaticus and amphipods of the genus Gammarus are both shredders (feeding on larger organic matter, e.g. leaves, and break it into smaller pieces), but the former inhabits predominantly slow-flowing or stagnant waters, while the latter is usually found in faster-flowing waters (Graça et al., Reference Graça, Maltby and Calow1994). Asellus aquaticus is the most widespread, and abundant freshwater isopod in Europe and Asia Minor and a recent study shows that it consists of several genetically distinct lineages or operational taxonomic units (OTUs), most of them being potentially distinct (sub-)species that have rather restricted ranges in southern Europe, i.e. on the Balkan Peninsula and the Apennine Peninsula (Sworobowicz et al., Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015). The majority of Europe, particularly its central and northern area, is inhabited by the nominative species, Asellus aquaticus aquaticus, showing no clear spatio-genetic structure (Sworobowicz et al., Reference Sworobowicz, Mamos, Grabowski and Wysocka2020). Given the fact that microsporidians are common in most groups of crustaceans (Stentiford et al., Reference Stentiford, Feist, Stone, Bateman and Dunn2013), we would expect a similar microsporidian diversity in isopods as in amphipod hosts. This assumption is supported by the occurrence of other parasite taxa in both groups of crustaceans, e.g. being intermediate hosts for closely related acanthocephalan (Sures, Reference Sures and Schmidt-Rhaesa2014) and trematode parasites (Bock, Reference Bock1984; Bojko et al., Reference Bojko, Bącela-Spychalska, Stebbing, Dunn, Grabowski, Rachalewski and Stentiford2017). Nevertheless, due to the lack of studies investigating microsporidians in isopods, it is not clear to date whether A. aquaticus is an equally suitable host for microsporidians compared to amphipods.
Therefore, the aim of the present study was to obtain a first overview of the presence and diversity of microsporidian parasites in the genetically heterogenous aquatic isopod A. aquaticus from a wide geographical range, which also allowed us to investigate potential differences of microsporidian diversity in the genetically distinct host lineages. The presence of microsporidian infections was tested on samples collected previously in the studies of Sworobowicz et al. (Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015, Reference Sworobowicz, Mamos, Grabowski and Wysocka2020), including several host OTUs originating from a large geographic area within Europe.
Materials and methods
Collection of isopods
Ethanol (99%) fixed individuals of A. aquaticus that were collected previously at various localities throughout Europe within the studies of Sworobowicz et al. (Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015, Reference Sworobowicz, Mamos, Grabowski and Wysocka2020) were used for the present study (247 individuals from 30 sites in 17 countries, for sampling details see Sworobowicz et al., Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015, Reference Sworobowicz, Mamos, Grabowski and Wysocka2020) (Table 1). In the present study, individuals clustered into OTUs as determined previously by Sworobowicz et al. (Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015): A (Asellus aquaticus aquaticus), D, F and J were used to cover most of the sampling area and the major host OTUs. Depending on the availability of specimens remaining from the study of Sworobowicz et al. (Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015), 5–10 host individuals were analysed for each sampling site (see Supplemental file 1). Specimens originating from sites where more than 1 MOTU was present were barcoded as described in Sworobowicz et al. (Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015).
Details on sampling location and microsporidian prevalence per site in Asellus aquaticus hosts.

Sample processing and molecular detection of microsporidians
To analyse the relationship between host size and parasite infection, the length of each A. aquaticus individual was measured according to images taken with a stereo microscope equipped with a camera (moticam 2300, Motic®) that was calibrated with a scaled slide. The animals were cut approximately in the sagittal plane and the intestine was removed to avoid contamination with gut content. The DNA was extracted from the remaining tissue following the procedure described in Grabner et al. (Reference Grabner, Weigand, Leese, Winking, Hering, Tollrian and Sures2015). Detection of microsporidians was conducted using the universal microsporidian primers V1 (5′-CACCAGGTTGATTCTGCCTGAC-3′) (Zhu et al., Reference Zhu, Wittner, Tanowitz, Kotler, Cali and Weiss1993) and mic-uni3R (5′-ATTACCGCGGMTGCTGGCAC-3′) (Weigand et al., Reference Weigand, Kremers and Grabner2016). PCR thermal profiles and reaction volumes were conducted as described in Weigand et al. (Reference Weigand, Kremers and Grabner2016). PCR products were purified using an E.Z.N.A. Cycle Pure kit (Omega Bio-Tek) and sent for Sanger sequencing (Microsynth Seqlab) using primer V1. Raw sequences were quality-checked using Geneious v2022.0.1 (Biomatters).
Data analysis
The dependency of habitat type or host MOTU to the frequency of microsporidian-infected A. aquaticus was tested by Pearson's χ 2 test (function “chisq.test”) in R v4.1.0 (R Core Team, 2021). Size differences of infected and uninfected host individuals were tested by a 2-sample t-test (function “t.test” in R). Graphs were generated in R using the ggplot2-package v3.3.5 (Wickham, Reference Wickham2016).
Microsporidian molecular taxonomic units (MICMOTUs) were identified from the sequences if the genetic similarity between the isolates was less than 96%. Microsporidian prevalence was calculated for each sampling site. The results were depicted in a map using QGIS v3.16.7 (QGIS.org, 2021) and Natural Earth (naturalearthdata.com). Similarity to known sequences was analysed by BLAST-search (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
MEGA X v10.2.6 (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018) was used to align the consensus sequences of the respective MICMOTUs with the ClustalW algorithm and default parameters. Calculation of p-distances (including transitions and transversions, using gamma-distribution with invariant sites for the substitution rate, partial deletion with 95% site cut-off, and 1000 bootstrap replicates) was also conducted with MEGA X.
The alignment for the phylogenetic analysis included sequences from GenBank that were most similar to the detected microsporidian isolates and representatives of the major microsporidian groups according to the phylogeny by Bojko et al. (Reference Bojko, Reinke, Stentiford, Williams, Rogers and Bass2022). Sequences were aligned using the MAFFT algorithm v7.48 (Katoh and Standley, Reference Katoh and Standley2013) from the EMBL-EBI sequence tools (Madeira et al., Reference Madeira, Pearce, Tivey, Basutkar, Lee, Edbali, Madhusoodanan, Kolesnikov and Lopez2022). Gaps and unaligned regions were removed manually, and the final alignment had a length of 315 bp and contained 153 sequences. Model selection and phylogenetic analysis were conducted with IQ-TREE v1.6.12 (Nguyen et al., Reference Nguyen, Schmidt, Von Haeseler and Minh2015; Kalyaanamoorthy et al., Reference Kalyaanamoorthy, Minh, Wong, Von Haeseler and Jermiin2017) using ultrafast bootstrap (UFBoot) to test branch support (Hoang et al., Reference Hoang, Chernomor, Von Haeseler, Minh and Vinh2018). Amphiamblys sp. (KX214674) and Chytridiopsis typographi (MH728789) were used as outgroups. The tree was visualized with the program FigTree v1.4.4 (Rambaut, Reference Rambaut2010).
Results
Prevalence of microsporidians
In total, 247 individuals of A. aquaticus from 30 sites were PCR-tested for microsporidian infections of which 66 were found positive (total prevalence 26.7%, Table 1). At 9 sites, no infected A. aquaticus-individual was detected (5–10 specimens investigated). The 2 sites with the highest prevalence were POL29 (90.0%, n = 10) and SVK7 (85.7%, n = 7) and the sites with the lowest prevalence were UKR13 and POL44 (10.0%, n = 10; see Table 1).
The analysis of the frequency of infected individuals and habitat type showed a significant relationship (Pearson's χ 2 test, (χ 2 = 36.044, df = 14, P < 0.005). The lowest proportion of infected individuals (8.7%) was found for springs, while the highest proportion was recorded for ponds (37.7%; see Table 2). Furthermore, as significant dependency of the 4 distinct host OTUs (according to Sworobowicz et al., Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015) to the frequency of microsporidian-infected individuals was detected (Pearson's χ 2 test, (χ 2 = 15.645, df = 3, P < 0.005) (see Supplementary file 1 for raw data). No infections were recorded for OTU F, even though it occurred at 5 sites (BUL2, GRE1, MNE11, 23, 32). The highest proportion of microsporidian infections was found in OTUs J (32.1%) and A (31.7%) followed by OTU D (21.7%) (Table 3).
Number of Asellus aquaticus hosts and proportion of individuals infected by microsporidians for each habitat type

Number of Asellus aquaticus hosts and proportion of individuals infected by microsporidians in each host OTU

Relation of host size and infection
A significant difference was found for the size of infected and uninfected A. aquaticus individuals (t = −2.26, P < 0.05). Infected individuals were larger than uninfected specimens (mean 7.64 mm vs 7.08 mm; Fig. 1; see Supplementary file 1 for individual measurements).
Size dependence of positive (n = 66) and negative (n = 181) results of the microsporidian PCR. Asterisks indicate significant difference (P < 0.05).

MICMOTUs and their geographic location
In total, 57 microsporidian sequences were obtained and assigned to 17 different MICMOTUs. No usable sequence could be obtained from 9 individuals due to low-quality reads and high background. The P-distances (proportion of nucleotide sites at which 2 sequences being compared are different) between the MICMOTUs were between 0.046 (MICMOTUs 5 and 7) and 0.376 (MICMOTUs 2 and 4). For details on the P-distances see Supplementary file 2.
MICMOTU 3 was the most common and was revealed from 17 host individuals, followed by MICMOTUs 1 and 4 with 8 host individuals infected by these parasites. MICMOTUs 2 and 5 were represented by 5 and 4 host individuals, respectively. MICMOTUs 6, 7, 8 and 11 were detected in 2–3 isopods, while all other MICMOTUs were single findings. MICMOTUs 1 and 5 occurred in all host OTUs, while MICMOTU 9 was found only in OTU D as a single finding. The most abundant host OTU A harboured all MICMOTUs (except 9). The map in Fig. 2 shows the geographic distribution of the MICMOTUs.
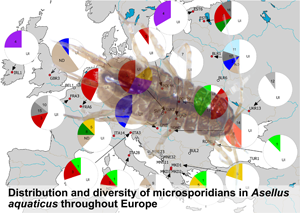
Map showing the sampling locations and pie-charts showing the prevalence of the respective microsporidian MOTUs (MICMOTU) at each site. At sites indicated with white dots, no infections were found. Red dots indicate sites where infected Asellus aquaticus were detected. Numbers in pies indicate the respective MICMOTU. UI, uninfected, ND, not determined (PCR positive but sequence was too short or of poor quality). Greyscale fill indicates MICMOTUs that were detected only in a single host individual. Please note the uncertainty of prevalence values given due to low sample size.

Based on the available dataset, there was no obvious geographical distribution of the MICMOTUs isolated from A. aquaticus. The most abundant MICMOTU 3 occurred at 8 sites throughout Europe. MICMOTU 1 was located mostly in Southern Europe (Italy, North Macedonia, Romania), but was also found at 1 site in Central Europe (Poland). MICMOTU 4 was found at 2 northern sites (Ireland, Estonia), but also in Slovakia (Fig. 2).
Phylogenetic reconstruction of MICMOTUs
The nucleotide sequence data of the MICMOTUs are available in the GenBank database under the accession numbers OM509764–OM509780 (also shown in Fig. 3). The phylogenetic inference illustrated the phylogenetic position of the microsporidian isolates from A. aquaticus found in the present study (Fig. 3). Most MICMOTUs clustered in the groups Nosematida and Enterocytozoonida while only MICMOTU 17 was found in the group Amblyosporida (taxonomy according to Bojko et al., Reference Bojko, Reinke, Stentiford, Williams, Rogers and Bass2022). MICMOTU4 was located among the Nosema spp. (in Nosematida), while MICMOTU6 was located in a position basal to both the Vairimorpha and Nosema spp. MICMOTU12 also clustered in the Nosematida close to previous isolates from amphipods.
Maximum likelihood phylogenetic tree reconstruction with 307 ultrafast bootstrap iterations of the microsporidian MOTUs (MICMOTUs) detected in Asellus aquaticus (in bold) including microsporidian sequences representing the recent microsporidian taxonomy sensu Bojko et al. (Reference Bojko, Reinke, Stentiford, Williams, Rogers and Bass2022). Substitution model was GTR + F + R5. Amphiamblys sp. (KX214674) and Chytridiopsis typographi (MH728789) were used as outgroups. GenBank accession numbers are shown in brackets and the host group/sample type for each sequence isolate is indicated. Branches that did not contain microsporidians from A. aquaticus were collapsed to make the tree clearer. The same tree with branches not collapsed can be seen in Supplementary file 3.

MICMOTU5, 7, 8, 10 and 14 were found in the Enterocytozoonida in a branch dominated by microsporidians detected in amphipods, with MICMOTU 7 being almost identical to Microsporidium sp. 1199 (FN610845) (Fig. 3). The remaining MICMOTUs were located all over the Enterocytozoonida and it is noteworthy that MICMOTUs 1, 2, 9 and 16 were most closely related to different environmental samples of microsporidians (without host record), all found in the study of Dubuffet et al. (Reference Dubuffet, Chauvet, Moné, Debroas and Lepère2021).
Discussion
The present study provides the first survey data on microsporidian diversity in A. aquaticus from a total of 30 sampling sites throughout Europe. A total of 17 MICMOTUs were identified, of which only 5 were detected in 4 or more host individuals. Eight MICMOTUs were only detected in single individuals. For the latter, it is doubtful as to whether these isolates were true infections of A. aquaticus specimens or rather contaminations by spores or DNA of microsporidians actually infecting other species. Even though the intestines of the hosts were removed prior to DNA extractions, contaminations with remaining of the intestinal content cannot be ruled out completely. Such contaminations with environmental spores, but also co-infections with 2 or more species of microsporidians might explain the failure to obtain sequencing results (high background or short reads) in some cases, indicating low amounts of microsporidian DNA in the sample or mixtures of the DNA of different parasite species. On the other hand, such rare microsporidians were also detected in several species of amphipods and it is assumed that these are true infections (Grabner et al., Reference Grabner, Weigand, Leese, Winking, Hering, Tollrian and Sures2015; Grabner, Reference Grabner2016). Particularly the problem of co-infections in the same host individual should be eliminated in follow-up studies by using metabarcoding techniques like those recently applied for microsporidians (Trzebny et al., Reference Trzebny, Slodkowicz-Kowalska, Becnel, Sanscrainte and Dabert2020). On the other hand, by removing the intestine, infections in this organ could also have been overlooked. Therefore, the microsporidian diversity in A. aquaticus might even be higher.
Compared to studies on other aquatic organisms such as amphipods or insect larvae, where the prevalence rates are often 50% or higher (Grabner et al., Reference Grabner, Weigand, Leese, Winking, Hering, Tollrian and Sures2015; Grabner, Reference Grabner2016), the prevalence found in A. aquaticus in the present study was lower, and highly variable (0–90%) between the different sampling sites. However, it must be noted that the small sample size (at few sites only 5 individuals were available) increases the uncertainty of the estimate, as a low prevalence would remain undetected. The significantly higher prevalence found in larger hosts might indicate an accumulation of infective spores during the life span of the host or an increased susceptibility of larger hosts due to a switch in feeding habits or microhabitat preference. This pattern does not seem to be universally valid for microsporidia, as in a previous study no weight difference was found between amphipods that were infected or uninfected with the microsporidium Dictyocoela duebenum (Chen et al., Reference Chen, Grabner, Nachev, Shih and Sures2015). In contrast, infections with Nosema sp. in Gammarus duebeni caused a size reduction of infected females (Terry et al., Reference Terry, Smith and Dunn1998). Therefore, the effect of microsporidian infection on host weight/size seems to depend on the respective host–parasite system. In the present study, MICMOTU4 was found to be closely related to Nosema spp., indicating that this and maybe other microsporidians in A. aquaticus might cause a size reduction of the host. Due to the overall low sample size, we did not differentiate between the different MICMOTUs for the analysis. Therefore, it has to be noted that an analysis of single MICMOTUs using a larger dataset would provide more differentiated results, as the effects of the different microsporidians for their host might be quite different.
There was a significant relationship between the host OTUs, as determined in the study of Sworobowicz et al. (Reference Sworobowicz, Grabowski, Mamos, Burzyński, Kilikowska, Sell and Wysocka2015), and the frequency of infected A. aquaticus, even though the proportions of infected individuals was similar for OTUs A, D and J. An exception refers to the host OTU F with zero infections, even though this OTU was represented by 32 A. aquaticus individuals, however with most of them originating from 3 sites in Montenegro. This might be due to local factors affecting the host populations (e.g. drought), or seasonal changes in parasite prevalence. In a study on the prevalence of microsporidians in the amphipod Paracalliope fluviatilis, prevalence varied over time and no microsporidians were detected at some time points (Park and Poulin, Reference Park and Poulin2021). A similar time course may account for the absence of Microsporidia at the 3 sites in Montenegro, which coincidentally corresponds to the presence of OTU F at this location. Another possible reason for the lack of microsporidians in OTU F might be the relative isolation of the sampling sites in Montenegro (Lake Skadar system) which is characterized by a high rate of endemic species (Grabowski et al., Reference Grabowski, Jabłońska, Wysocka, Pešić, Pešić, Karaman and Kostianoy2018). Therefore, OTU F might have lost their microsporidian parasites during the colonization of the system and so far, no microsporidians of A. aquaticus were co-introduced to the area.
The associations of host OTUs and frequency of microsporidians in the present study are in contrast to the study by Wilkinson et al. (Reference Wilkinson, Rock, Whiteley, Ovcharenko and Ironside2011), who found little support for coevolution of Microsporidia of the genus Dictyocoela with their gammarid hosts. Nevertheless, there is strong evidence for co-diversification of microsporidians and their amphipod hosts (Park et al., Reference Park, Jorge and Poulin2020; Quiles et al., Reference Quiles, Wattier, Bacela-Spychalska, Grabowski and Rigaud2020). This indicates that the distribution pattern of microsporidians in amphipods is shaped both by ancient host–parasite associations and more recent horizontal transfer between host species or lineages (Quiles et al., Reference Quiles, Rigaud, Wattier, Grabowski and Bacela-Spychalska2021). The same might be true for A. aquaticus and their microsporidians, but it would require a larger sample size and a parasite species-specific analysis to substantiate the link between host OTU and the frequency of infection observed in the present study.
The frequency of infected A. aquaticus was significantly related to habitat type, which might be explained by specific habitat characteristics (e.g. temperature, flow velocity, nutrient availability) that can affect infection rate and thereby parasite prevalence (Marcogliese, Reference Marcogliese2001; Kelly et al., Reference Kelly, Dunn and Hatcher2002; Narr et al., Reference Narr, Ebert, Bastille-Rousseau and Frost2019). Asellus aquaticus collected from spring habitats in the present study showed the lowest proportion of infected individuals, which is in contrast to findings from amphipods, where species-rich microsporidian communities were found in niphargid amphipods from such ground water-dependent habitats (Grabner et al., Reference Grabner, Weber and Weigand2020).
The genus Asellus is widely distributed throughout Europe (Sket, Reference Sket1994), therefore a spatially homogenous distribution of associated microsporidian parasites would be expected with unique parasites in remote locations. Nevertheless, the distribution of the different MICMOTUs throughout Europe did not show a conclusive pattern. This is similar for microsporidians of amphipods that show a pan-European distribution without a clear geographic pattern (Krebes et al., Reference Krebes, Blank, Frankowski and Bastrop2010; Grabner et al., Reference Grabner, Weigand, Leese, Winking, Hering, Tollrian and Sures2015; Bacela-Spychalska et al., Reference Bacela-Spychalska, Wróblewski, Mamos, Grabowski, Rigaud, Wattier, Rewicz, Konopacka and Ovcharenko2018; Prati et al., Reference Prati, Grabner, Pfeifer, Lorenz and Sures2022). Furthermore, the rather small sample size has to be taken into account, and it is likely that a higher number of sampling sites and more tested individuals would probably show a more even distribution of most MICMOTUs.
The phylogenetic analysis shows the proximity of MICMOTUs detected in A. aquaticus to various branches including microsporidians of amphipods. Some isolates were similar to those detected previously in environmental samples from aquatic habitats and they might originally be parasites of A. aquaticus (Dubuffet et al., Reference Dubuffet, Chauvet, Moné, Debroas and Lepère2021). Interestingly, MICMOTU4 from the present study was closely related to Nosema spp., a group of microsporidians that was mostly found to parasitize insects (with the exception of N. granulosis from amphipods) (Tokarev et al., Reference Tokarev, Huang, Solter, Malysh, Becnel and Vossbrinck2020). This might indicate that the host diversity within the genus Nosema and possibly also Vairimorpha will extend to other groups of arthropods, in the course of future studies.
Most of the more common MICMOTUs detected in A. aquaticus (MICMOTU1, 2, 3, 5) were highly similar to microsporidian isolates from amphipods from the group Enterocytozoonida. It raises the question, if these isopod and amphipod microsporidians are closely related but distinct species, or if the same microsporidian species is a host generalist that is able to infect different groups of aquatic crustaceans. As we know from microsporidians with well-described life cycles, both strategies (host generalists and host specialists) can be found in different microsporidian species (Wadi and Reinke, Reference Wadi and Reinke2020), but generally low host specificity was found for microsporidians infecting amphipods (Prati et al., Reference Prati, Grabner, Pfeifer, Lorenz and Sures2022). In this context, it is interesting to note that no MICMOTUs from A. aquaticus were related to microsporidians from the group Glugeida that includes common parasites of amphipods like Dictyocoela or Cucumispora spp.
Conclusion
The present study provides a first overview on the microsporidian diversity in different genetic lineages of A. aquaticus. Several issues arise from this study that should be addressed in the future: First of all, more host individuals should be analysed to detect MICMOTUs that might indicate coevolution of host and parasite lineages, and to clarify the status of “rare” microsporidians as true infections or contaminations. Furthermore, the geographic distribution of the microsporidians should be studied in closer detail to substantiate the presence (or inferred absence) of common microsporidian species throughout the study area. And finally, the ratio of co-infections of 2 or more microsporidians in the same host should be addressed.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S003118202200124X
Author's contributions
All authors contributed to the study conception and design. Sample collection was performed by Lidia Sworobowicz, Michał Grabowski and Tomasz Mamos. Sample processing and molecular analyses were performed by Annemie Doliwa and Daniel Grabner. The first draft of the paper was written by Daniel Grabner and all authors commented on previous versions of the paper. All authors read and approved the final paper.
Financial support
The fieldwork and molecular part of the study were partially supported by the National Science Center, Poland [grant number 2014/15/B/NZ8/ 00266].
Conflict of interests
The authors declare there are no conflicts of interest.
Ethical Standards
Not applicable.