INTRODUCTION
Trypanosomatids are unicellular flagellates and obligate parasites that infect various animals and plants. They include Trypanosoma and Leishmania, species of which cause potent vector-borne diseases in humans, livestock and wildlife; diseases that are responsible for substantial mortality and morbidity across the world. Trypanosoma cruzi causes Chagas disease in South and Central America; Trypanosoma brucei causes Human African Trypanosomiasis in sub-Saharan Africa (and, along with related species, a similar disease in livestock); while Leishmania spp. cause various forms of leishmaniasis in humans. Other species of Trypanosoma and Leishmania infect a wide range of vertebrate hosts, and all are transmitted by invertebrate vectors; predominantly these are biting insects, although some aquatic species are transmitted by leeches (Lom, Reference Lom, Lumsden and Evans1979). Phytomonas spp. are plant parasites transmitted by phloem-sucking insects and are occasionally an agricultural problem in South and Central America (Camargo, Reference Camargo1999). Besides these dixenic (i.e. two-host) parasites that cycle between insect/leech and vertebrate/plant hosts, the trypanosomatids include various other genera, such as Crithidia, Leptomonas, Herpetomonas, Angomonas and Strigomonas that are cosmopolitan, monoxenic (i.e. one host) parasites of insects (Maslov et al. Reference Maslov, Votýpka, Yurchenko and Lukeš2013). The diverse associations of trypanosomatids indicate that the origin of parasitism is singular and ancient (Simpson et al. Reference Simpson, Stevens and Lukes2006).
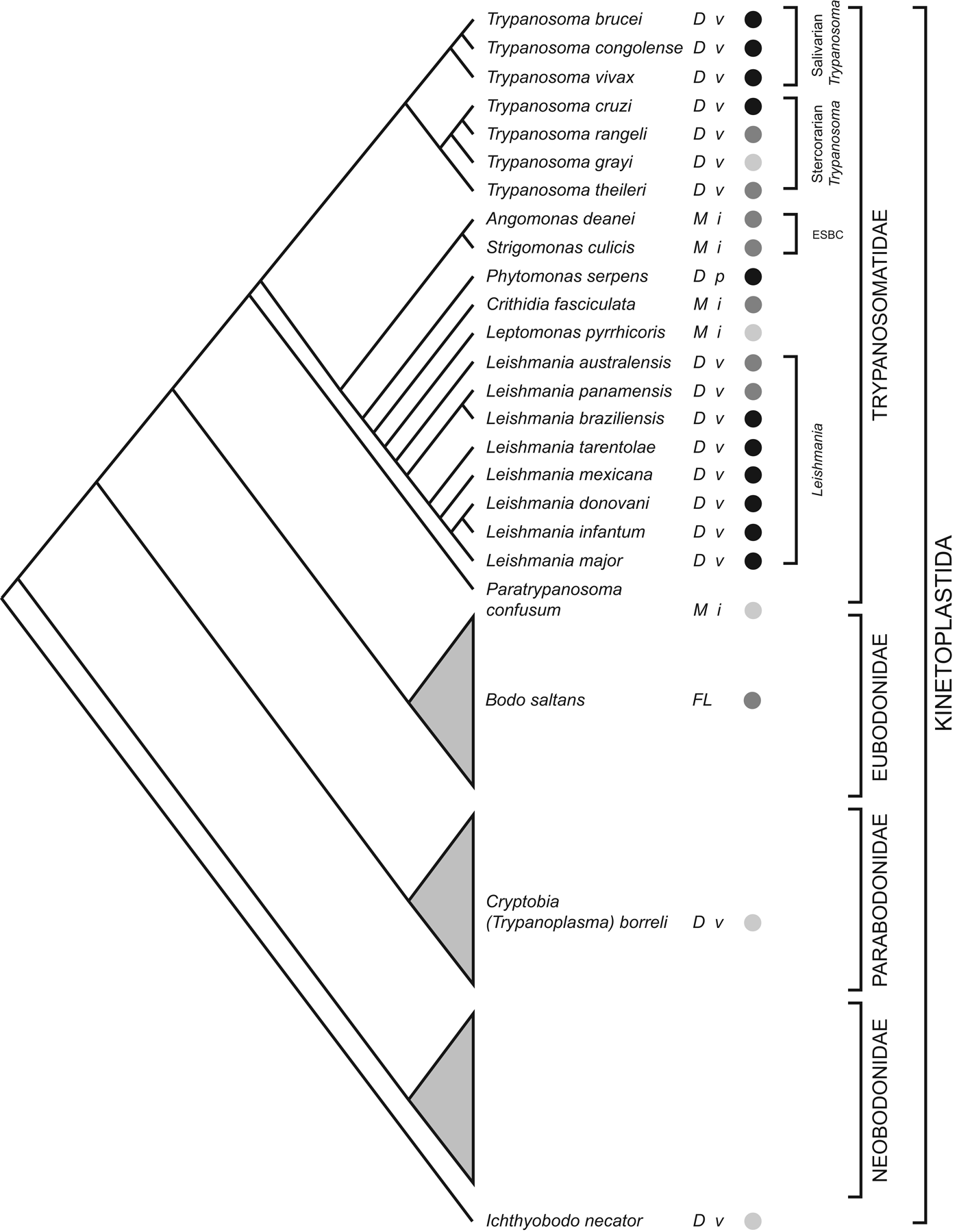
The order Trypanosomatidae is one part of the phylum Kinetoplastida; most other Kinetoplastids live freely or as commensals in marine, terrestrial and aquatic environments. The current consensus on Kinetoplastid phylogeny is summarized in Fig. 1; trypanosomatids are monophyletic and the sister clade to eubodonids (Callahan et al. Reference Callahan, Litaker and Noga2002; Simpson et al. Reference Simpson, Gill, Callahan, Litaker and Roger2004; Moreira et al. Reference Moreira, López-García and Vickerman2004; von der Heyden et al. Reference Von der Heyden, Chao, Vickerman and Cavalier-Smith2004; Deschamps et al. Reference Deschamps, Lara, Marande, López-García, Ekelund and Moreira2011). The closest known relative among eubodonids is Bodo saltans, a free-living bacteriovore of terrestrial and freshwater microbiota. Hence, the phylogeny indicates that parasitism in trypanosomatids had a single origin; although the position of the fish parasites Cryptobia spp. and Ichthyobodo spp. show that parasitism has appeared on other occasions within the Kinetoplastida (Simpson et al. Reference Simpson, Stevens and Lukes2006; von der Heyden et al. Reference Von der Heyden, Chao, Vickerman and Cavalier-Smith2004). This is the context in which I review the contribution of trypanosomatid genome sequences to our understanding of how parasitism evolved and subsequently diversified.
Kinetoplastid phylogeny. A cladogram depicting the current consensus on Kinetoplastid phylogenetic relationships (adapted from von der Heyden et al. Reference Von der Heyden, Chao, Vickerman and Cavalier-Smith2004; Simpson et al. Reference Simpson, Gill, Callahan, Litaker and Roger2004, Reference Simpson, Stevens and Lukes2006; Deschamps et al. Reference Deschamps, Lara, Marande, López-García, Ekelund and Moreira2011; Flegontov et al. Reference Flegontov, Votýpka, Skalický, Logacheva, Penin, Tanifuji, Onodera, Kondrashov, Volf, Archibald and Lukeš2013). Each bodonid order is shown as a grey triangle, representing an indeterminate, but large, number of species. The status of genome sequencing projects for each named species is indicated by filled circles (black: finished; dark grey: unfinished draft; light grey: sequencing in progress). The life cycle of each species is indicated (D: dixenic; M: monoxenic; FL: free-living), as well as the host type(s) (v: vertebrate; i: insect; p: plant). ESBC: ‘Endosymbiont-bearing clade’.
Since the publication of the ‘TriTryp’ genome sequences for T. cruzi, T. brucei and Leishmania major in 2005 (Berriman et al. Reference Berriman, Ghedin, Hertz-Fowler, Blandin, Renauld, Bartholomeu, Lennard, Caler, Hamlin, Haas, Böhme, Hannick, Aslett, Shallom, Marcello, Hou, Wickstead, Alsmark, Arrowsmith, Atkin, Barron, Bringaud, Brooks, Carrington, Cherevach, Chillingworth, Churcher, Clark, Corton, Cronin, Davies, Doggett, Djikeng, Feldblyum, Field, Fraser, Goodhead, Hance, Harper, Harris, Hauser, Hostetler, Ivens, Jagels, Johnson, Johnson, Jones, Kerhornou, Koo, Larke, Landfear, Larkin, Leech, Line, Lord, Macleod, Mooney, Moule, Martin, Morgan, Mungall, Norbertczak, Ormond, Pai, Peacock, Peterson, Quail, Rabbinowitsch, Rajandream, Reitter, Salzberg, Sanders, Schobel, Sharp, Simmonds, Simpson, Tallon, Turner, Tait, Tivey, Van Aken, Walker, Wanless, Wang, White, White, Whitehead, Woodward, Wortman, Adams, Embley, Gull, Ullu, Barry, Fairlamb, Opperdoes, Barrell, Donelson, Hall, Fraser, Melville and El-Sayed2005; El-Sayed et al. Reference El-Sayed, Myler, Bartholomeu, Nilsson, Aggarwal, Tran, Ghedin, Worthey, Delcher, Blandin, Westenberger, Caler, Cerqueira, Branche, Haas, Anupama, Arner, Aslund, Attipoe, Bontempi, Bringaud, Burton, Cadag, Campbell, Carrington, Crabtree, Darban, da Silveira, de Jong, Edwards, Englund, Fazelina, Feldblyum, Ferella, Frasch, Gull, Horn, Hou, Huang, Kindlund, Klingbeil, Kluge, Koo, Lacerda, Levin, Lorenzi, Louie, Machado, McCulloch, McKenna, Mizuno, Mottram, Nelson, Ochaya, Osoegawa, Pai, Parsons, Pentony, Pettersson, Pop, Ramirez, Rinta, Robertson, Salzberg, Sanchez, Seyler, Sharma, Shetty, Simpson, Sisk, Tammi, Tarleton, Teixeira, Van Aken, Vogt, Ward, Wickstead, Wortman, White, Fraser, Stuart and Andersson2005a ; Ivens et al. Reference Ivens, Peacock, Worthey, Murphy, Aggarwal, Berriman, Sisk, Rajandream, Adlem, Aert, Anupama, Apostolou, Attipoe, Bason, Bauser, Beck, Beverley, Bianchettin, Borzym, Bothe, Bruschi, Collins, Cadag, Ciarloni, Clayton, Coulson, Cronin, Cruz, Davies, De Gaudenzi, Dobson, Duesterhoeft, Fazelina, Fosker, Frasch, Fraser, Fuchs, Gabel, Goble, Goffeau, Harris, Hertz-Fowler, Hilbert, Horn, Huang, Klages, Knights, Kube, Larke, Litvin, Lord, Louie, Marra, Masuy, Matthews, Michaeli, Mottram, Müller-Auer, Munden, Nelson, Norbertczak, Oliver, O'neil, Pentony, Pohl, Price, Purnelle, Quail, Rabbinowitsch, Reinhardt, Rieger, Rinta, Robben, Robertson, Ruiz, Rutter, Saunders, Schäfer, Schein, Schwartz, Seeger, Seyler, Sharp, Shin, Sivam, Squares, Squares, Tosato, Vogt, Volckaert, Wambutt, Warren, Wedler, Woodward, Zhou, Zimmermann, Smith, Blackwell, Stuart, Barrell and Myler2005), there has been much comparative analysis of these seminal resources. They have been complemented by transcriptomic (Holzer et al. Reference Holzer, McMaster and Forney2006; Leifso et al. Reference Leifso, Cohen-Freue, Dogra, Murray and McMaster2007; Saxena et al. Reference Saxena, Lahav, Holland, Aggarwal, Anupama, Huang, Volpin, Myler and Zilberstein2007; Rochette et al. Reference Rochette, Raymond, Ubeda, Smith, Messier, Boisvert, Rigault, Corbeil, Ouellette and Papadopoulou2008, Reference Rochette, Raymond, Corbeil, Ouellette and Papadopoulou2009; Alcolea et al. Reference Alcolea, Alonso, Sánchez-Gorostiaga, Moreno-Paz, Gómez, Ramos, Parro and Larraga2009, Reference Alcolea, Alonso, Gómez, Moreno, Domínguez, Parro and Larraga2010; Depledge et al. Reference Depledge, Evans, Ivens, Aziz, Maroof, Kaye and Smith2009; Jensen et al. Reference Jensen, Sivam, Kifer, Myler and Parsons2009; Kabani et al. Reference Kabani, Fenn, Ross, Ivens, Smith, Ghazal and Matthews2009; Minning et al. Reference Minning, Weatherly, Atwood, Orlando and Tarleton2009; Veitch et al. Reference Veitch, Johnson, Trivedi, Terry, Wildridge and MacLeod2010; Adaui et al. Reference Adaui, Castillo, Zimic, Gutierrez, Decuypere, Vanaerschot, De Doncker, Schnorbusch, Maes, Van der Auwera, Maes, Llanos-Cuentas, Arevalo and Dujardin2011) and proteomic analyses (Atwood et al. Reference Atwood, Weatherly, Minning, Bundy, Cavola, Opperdoes, Orlando and Tarleton2005; Rosenzweig et al. Reference Rosenzweig, Smith, Opperdoes, Stern, Olafson and Zilberstein2008a , Reference Rosenzweig, Smith, Myler, Olafson and Zilberstein b ; Alcolea et al. Reference Alcolea, Alonso and Larraga2011; Eyford et al. Reference Eyford, Sakurai, Smith, Loveless, Hertz-Fowler, Donelson, Inoue and Pearson2011; Urbaniak et al. Reference Urbaniak, Guther and Ferguson2012; Butter et al. Reference Butter, Bucerius, Michel, Cicova, Mann and Janzen2013) of gene expression at various life-cycle stages. Genome sequences for additional species of Trypanosoma (Jackson et al. Reference Jackson, Sanders, Berry, McQuillan, Aslett, Quail, Chukualim, Capewell, MacLeod, Melville, Gibson, Barry, Berriman and Hertz-Fowler2009, Reference Jackson, Berry, Aslett, Allison, Burton, Vavrova-Anderson, Brown, Browne, Corton, Hauser, Gamble, Gilderthorp, Marcello, McQuillan, Otto, Quail, Sanders, van Tonder, Ginger, Field, Barry, Hertz-Fowler and Berriman2012), Leishmania (Peacock et al. Reference Peacock, Seeger, Harris, Murphy, Ruiz, Quail, Peters, Adlem, Tivey, Aslett, Kerhornou, Ivens, Fraser, Rajandream, Carver, Norbertczak, Chillingworth, Hance, Jagels, Moule, Ormond, Rutter, Squares, Whitehead, Rabbinowitsch, Arrowsmith, White, Thurston, Bringaud, Baldauf, Faulconbridge, Jeffares, Depledge, Oyola, Hilley, Brito, Tosi, Barrell, Cruz, Mottram, Smith and Berriman2007; Downing et al. Reference Downing, Imamura, Decuypere, Clark, Coombs, Cotton, Hilley, de Doncker, Maes, Mottram, Quail, Rijal, Sanders, Schönian, Stark, Sundar, Vanaerschot, Hertz-Fowler, Dujardin and Berriman2011; Rogers et al. Reference Rogers, Hilley, Dickens, Wilkes, Bates, Depledge, Harris, Her, Herzyk, Imamura, Otto, Sanders, Seeger, Dujardin, Berriman, Smith, Hertz-Fowler and Mottram2011; Raymond et al. Reference Raymond, Boisvert, Roy, Ritt, Légaré, Isnard, Stanke, Olivier, Tremblay, Papadopoulou, Ouellette and Corbeil2012; Real et al. Reference Real, Vidal, Carazzolle, Mondego, Costa, Herai, Würtele, de Carvalho, E Ferreira, Mortara, Barbiéri, Mieczkowski, da Silveira, Briones, Pereira and Bahia2013) and Phytomonas (Porcel et al. Reference Porcel, Denoeud, Opperdoes, Noel, Madoui, Hammarton, Field, Da Silva, Couloux, Poulain, Katinka, Jabbari, Aury, Campbell, Cintron, Dickens, Docampo, Sturm, Koumandou, Fabre, Flegontov, Lukeš, Michaeli, Mottram, Szöőr, Zilberstein, Bringaud, Wincker and Dollet2014) have been produced, with several more in progress (see Fig. 1).
Comparison of the Tritryp genomes showed that both gene order and gene repertoire are broadly conserved within chromosomal cores (El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ). It is generally thought that the considerable co-linearity displayed by trypanosomatid genomes, despite their apparently ancient divergences, reflects strong and fundamental selective constraints on genome structure (Ghedin et al. Reference Ghedin, Bringaud, Peterson, Myler, Berriman, Ivens, Andersson, Bontempi, Eisen, Angiuoli, Wanless, Von Arxa, Murphy, Lennard, Salzberg, Adams, White, Hall, Stuart, Fraser and El-Sayed2004). Analysis of gene order conservation across Eukaryotic genomes indicates that highly conserved gene pairs are retained for both functional and transcriptional regulation (Dávila-López et al. Reference Dávila-López, Martínez-Guerra and Samuelsson2010). While there is little to suggest that the conserved proximity of genes in trypanosomatids reflects their shared or related functions, it has been suggested that their polycistronic organization necessitates the co-directionality of replication and transcription (Ghedin et al. Reference Ghedin, Bringaud, Peterson, Myler, Berriman, Ivens, Andersson, Bontempi, Eisen, Angiuoli, Wanless, Von Arxa, Murphy, Lennard, Salzberg, Adams, White, Hall, Stuart, Fraser and El-Sayed2004), and that this structural peculiarity of trypanosomatids (the cause of which remains unsolved), is responsible for the strong purifying selection that maintains gene order.
Beyond the chromosomal cores, within sub-telomeric regions for instance, there are numerous species-specific features (El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ). From the outset it was appreciated that these genes are very often associated with disease mechanisms (El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ) and are the basis for the distinctive cell surface architectures displayed by each parasite (Acosta-Serrano et al. Reference Acosta-Serrano, Hutchinson, Nakayasu, Almeida, Carrington, Barry, McCulloch, Mottram and Acosta-Serrano2007; Handman et al. Reference Handman, Papenfuss, Speed, Goding, Myler and Fasel2008). Thus, after 10 years of comparative and experimental analysis of these genomes the principal genomic features that distinguish the stem trypanosomatid lineages, and which are most likely to have been instrumental in the evolution of parasitism, are apparent.
GENOMIC REDUCTION
Parasites were once thought to be ‘degenerate’; while this view is no longer prevalent, it remains intuitive that some characters vital to free-living organisms, but no longer necessary for parasites within a host environment, are lost when the selection pressure to retain them is removed. Hence, we expect phenotypic reduction, which is often observed of parasites, to be reflected in genomic reduction. For example, the genomes of both schistosomes and cestodes, which are phenotypically reduced relative to free-living platyhelminthes, lack elements of canonical metazoan metabolism and developmental regulation (Berriman et al. Reference Berriman, Haas, LoVerde, Wilson, Dillon, Cerqueira, Mashiyama, Al-Lazikani, Andrade, Ashton, Aslett, Bartholomeu, Blandin, Caffrey, Coghlan, Coulson, Day, Delcher, DeMarco, Djikeng, Eyre, Gamble, Ghedin, Gu, Hertz-Fowler, Hirai, Hirai, Houston, Ivens, Johnston, Lacerda, Macedo, McVeigh, Ning, Oliveira, Overington, Parkhill, Pertea, Pierce, Protasio, Quail, Rajandream, Rogers, Sajid, Salzberg, Stanke, Tivey, White, Williams, Wortman, Wu, Zamanian, Zerlotini, Fraser-Liggett, Barrell and El-Sayed2009; Tsai et al. Reference Tsai, Zarowiecki, Holroyd, Garciarrubio, Sanchez-Flores, Brooks, Tracey, Bobes, Fragoso, Sciutto, Aslett, Beasley, Bennett, Cai, Camicia, Clark, Cucher, De Silva, Day, Deplazes, Estrada, Fernández, Holland, Hou, Hu, Huckvale, Hung, Kamenetzky, Keane, Kiss, Koziol, Lambert, Liu, Luo, Luo, Macchiaroli, Nichol, Paps, Parkinson, Pouchkina-Stantcheva, Riddiford, Rosenzvit, Salinas, Wasmuth, Zamanian, Zheng, Cai, Soberón, Olson, Laclette, Brehm and Berriman2013). Genome reduction reaches its apogee in the microsporidian parasites, which in some cases have reduced their genomes to the physiological minimum required for life, and this corresponds with their extreme host dependence (Nakjang et al. Reference Nakjang, Williams, Heinz, Watson, Foster, Sendra, Heaps, Hirt and Embley2013). At such extremes, we also observe physical compaction of the genome, in addition to the loss of genes (Keeling and Slamovits, Reference Keeling and Slamovits2005).
Trypanosomatids do not appear to be reduced physically; the size of their genomes (25–35 mb in the haploid state) and the gene density (2·8–4·6 Kb/gene) is comparable with free-living unicellular eukaryotes, for instance Saccharomyces cerevisiae (12·5 mb/2·09 Kb/gene) and Dictyostelium discoideum (33·8 mb/2·72 Kb/gene). However, trypanosomatid genomes might still be functionally reduced, having lost genes essential to free-living Kinetoplastids.
Before the advent of genome sequences, it was known that trypanosomatids lacked certain common metabolic capabilities. For example, they are auxotrophic for pteridine and folate, which are essential co-factors in macromolecule biosynthesis, because they lack the ability to synthesize tetrahydrobiopterin (Beck and Ullman, Reference Beck and Ullman1990, Bello et al. Reference Bello, Nare, Freedman, Hardy and Beverley1994, Nare et al. Reference Nare, Hardy and Beverley1997; Ouellette et al. Reference Ouellette, Drummelsmith, El-Fadili, Kündig, Richard and Roy2002). Similarly, they must scavenge haem from their hosts (or obtain it from bacterial endosymbionts; Alves et al. Reference Alves, Voegtly, Matveyev, Lara, da Silva, Serrano, Buck, Teixeira and Camargo2011), because they lack a native haem biosynthesis pathway (Chang et al. Reference Chang, Chang and Sassa1975; Korený et al. Reference Korený, Lukes and Oborník2010). Trypanosomatids are also auxotrophic for purines (Marr et al. Reference Marr, Berens and Nelson1978; Gutteridge and Gaborak, Reference Gutteridge and Gaborak1979), vital in the biosynthesis of nucleic acids and energy metabolism. Other aspects of model eukaryotic physiology are also absent, for example, a system of redox homoeostasis based on catalase and glutathione reductase. Instead, trypanosomatids rely on a unique thiol-based redox metabolism based on trypanothione for the deactivation of oxidizing agents (Oza et al. Reference Oza, Shaw, Wyllie and Fairlamb2005; Krauth-Siegel and Comini, Reference Krauth-Siegel and Comini2008; Comini and Flohé, Reference Comini, Flohé, Jäger, Oliver and Flohé2013). The initial Tritryp comparison showed that trypanosomatids do not possess receptor-linked tyrosine kinases (Parsons et al. Reference Parsons, Worthey, Ward and Mottram2005), canonical mitochondrial import systems (Pusnik et al. Reference Pusnik, Charrière, Mäser, Waller, Dagley, Lithgow and Schneider2009), known telomere end binding proteins such as POT1 (Lira et al. Reference Lira, Giardini, Neto, Conte and Cano2007), certain genes that regulate autophagy (Herman et al. Reference Herman, Gillies, Michels and Rigden2006) and others controlling apoptosis (i.e. TNF-related family receptors, Bcl-2 family members and caspases; Smirlis et al. Reference Smirlis, Duszenko, Ruiz, Scoulica, Bastien, Fasel and Soteriadou2010).
The question relating to these and any other missing features is whether they represent evolutionary losses, or instead, reflect the branching position of the Kinetoplastida in the eukaryotic phylogeny. It may be that certain widely conserved genes are absent from trypanosomatids because Kinetoplastids separated from other eukaryotic lineages early in evolutionary history and before those genes evolved. Furthermore, it could be that we have systematically underestimated genomic and physiological diversity in eukaryotes, and the apparent deficiencies of trypanosomatids reflect a biased perception based on a narrow sampling of animal and plant genomes. In short, the absence of ‘typical’ features from trypanosomatids need not represent evolutionary loss. In fact, detailed comparisons in the years following publication of the Tritryp genome sequences showed that, while trypanosomatids often lack some conserved features and have numerous clade-specific derivations, they are nevertheless comparable to free-living protists in the number and diversity of protein kinases (Parsons et al. Reference Parsons, Worthey, Ward and Mottram2005; Bahia et al. Reference Bahia, Oliveira, Lima, Oliveira, Silveira, Mortara and Ruiz2009), phosphatases (Brenchley et al. Reference Brenchley, Tariq, McElhinney, Szöor, Huxley-Jones, Stevens, Matthews and Tabernero2007), GTPases and other genes involved in intracellular trafficking (Field, Reference Field2005; Field et al. Reference Field, Natesan, Gabernet-Castello and Koumandou2007) and DNA helicases (Gargantini et al. Reference Gargantini, Lujan and Pereira2012).
In summary, these genomes are not reduced in size or substantially reduced in function. While trypanosomatids employ unique solutions in redox homoeostasis, mitochondrial protein import and telomere regulation, they nonetheless have a broadly typical eukaryotic physiology. Where there are disparities, it is not clear whether these genes were lost or never existed and this will only become clear after we have sampled the genomes of free-living Kinetoplastids for comparison. Instead, there is abundant evidence that trypanosomatid genomes have expanded during their evolution both physically, through the evolution of sub-telomeres and accessory chromosomes, and functionally, with the acquisition of new genes through duplication and horizontal gene transfer.
GENOMIC INNOVATION: SPECIES-SPECIFIC GENE FAMILIES
Trypanosomatid cell surfaces include various polymorphic proteins combined with diverse glycolipid conjugates (Ferguson, Reference Ferguson1997). These structures are enigmatic and their origins are mysterious because they are not seen in other organisms; indeed, the highly abundant cell-surface glycoproteins of T. brucei, T. cruzi and L. major are mutually exclusive, making it very hard to infer what the ancestral cell surface looked like (El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ). The Tritryp genomes revealed the genes that encode these surface features and their non-random distribution in the genome, which has been reviewed in detail elsewhere (Acosta-Serrano et al. Reference Acosta-Serrano, Hutchinson, Nakayasu, Almeida, Carrington, Barry, McCulloch, Mottram and Acosta-Serrano2007; Handman et al. Reference Handman, Papenfuss, Speed, Goding, Myler and Fasel2008; De Pablos and Osuna, Reference De Pablos and Osuna2012). These cell surface proteins attract considerable interest because they are implicated in disease, virulence and mechanisms of pathogenesis (De Pablos and Osuna, Reference De Pablos and Osuna2012). Species-specific genes provide the clearest insight into genomic innovations associated with parasitism and the multi-copy gene families that encode these cell surface proteins dominate such species-specific genes in comparative analyses (El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ).
The life cycles of the Tritryp species and the points at which species-specific cell surface proteins are expressed are shown in Fig. 2. Species-specific genes in T. cruzi are dominated by gene families that encode the mucin-based surface coat during its trypomastigote stage (Cerqueira et al. Reference Cerqueira, Bartholomeu, DaRocha, Hou, Freitas-Silva, Machado, El-Sayed and Teixeira2008; Nakayasu et al. Reference Nakayasu, Yashunsky, Nohara, Torrecilhas, Nikolaev and Almeida2009; De Pablos and Osuna, Reference De Pablos and Osuna2012); primarily mucins (TcMUC; Acosta-Serrano et al. Reference Acosta-Serrano, Almeida, Freitas-Junior, Yoshida and Schenkman2001; Buscaglia et al. Reference Buscaglia, Campo, Frasch and Di Noia2006) and trans-sialidases (TS; Kim et al. Reference Kim, Chiurillo, El-Sayed, Jones, Santos, Porcile, Andersson, Myler, da Silveira and Ramírez2005; Montagna et al. Reference Montagna, Donelson and Frasch2006; Freitas et al. Reference Freitas, dos Santos, Rodrigues-Luiz, Mendes, Rodrigues, Gazzinelli, Teixeira, Fujiwara and Bartholomeu2011; Oppezzo et al. Reference Oppezzo, Obal, Baraibar, Pritsch, Alzari and Buschiazzo2011; Ammar et al. Reference Ammar, Plazolles, Baltz and Coustou2013; Oliveira et al. Reference Oliveira, Freire-de-Lima, Penha, Dias and Todeschini2014), but also a ‘dispersed gene family’ (DGF-1; El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ; Kawashita et al. Reference Kawashita, da Silva, Mortara, Burleigh and Briones2009; Lander et al. Reference Lander, Bernal, Diez, Añez, Docampo and Ramírez2010), the mucin-associated surface protein family (MASP; El-Sayed et al. Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall2005b ; Bartholomeu et al. Reference Bartholomeu, Cerqueira, Leão, daRocha, Pais, Macedo, Djikeng, Teixeira and El-Sayed2009; dos Santos et al. Reference dos Santos, Freitas, Lobo, Rodrigues-Luiz, Mendes, Oliveira, Andrade, Chiari, Gazzinelli, Teixeira, Fujiwara and Bartholomeu2012), and the T. cruzi Trypomastigote Alanine, Serine and Valine-rich proteins (TcTASV; García et al. Reference García, Ziliani, Agüero, Bernabó, Sánchez and Tekiel2010; Bernabó et al. Reference Bernabó, Levy, Ziliani, Caeiro, Sánchez and Tekiel2013). Gene families specifically expressed in the other life stages include amastin in the intracellular amastigote stage, and T. cruzi Small MUcin-like Genes (TcSMUG; Urban et al. Reference Urban, Santurio, Chidichimo, Yu, Chen, Mucci, Agüero and Buscaglia2011) in the replicative epimastigote. In addition to these developmentally regulated, surface-expressed gene families, expansions of Retrotransposon Hotspot (RHS) genes and Elongation Factor 1 gamma (EF1γ) genes are prominent innovations of the T. cruzi genome.
Trypanosomatid life cycles. Each circle represents the movement of parasites between insect (black) and vertebrate (white) hosts, showing the transition of parasite life stages, the position of each stage within the hosts and the timing of expression of cell surface-expressed protein families mentioned in the text. Note that for T. brucei and L. major, the parasites move anteriorly from the insect gut as they develop, while T. cruzi migrates posteriorly as it prepares for transmission into the vertebrate.
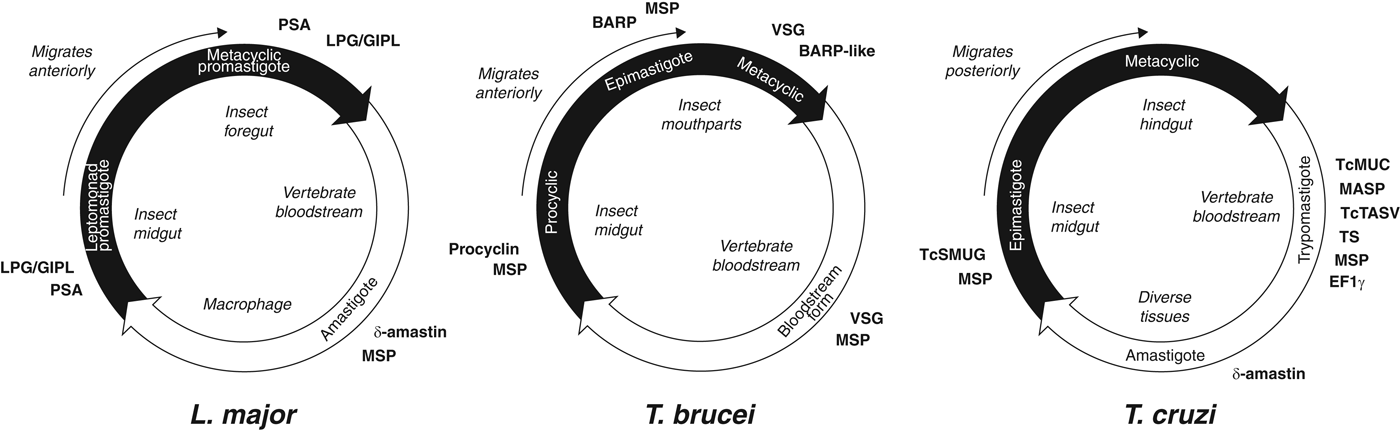
In T. brucei, species-specific genes are dominated by those encoding the Variant Surface Glycoproteins (VSG) that form the surface glycocalyx of all salivarian trypanosomes during their bloodstream stage in the mammal host (Hutchinson et al. Reference Hutchinson, Picozzi, Jones, Mott, Sharma, Welburn and Carrington2007; Marcello and Barry, Reference Marcello and Barry2007; Jackson et al. Reference Jackson, Berry, Aslett, Allison, Burton, Vavrova-Anderson, Brown, Browne, Corton, Hauser, Gamble, Gilderthorp, Marcello, McQuillan, Otto, Quail, Sanders, van Tonder, Ginger, Field, Barry, Hertz-Fowler and Berriman2012; Weirather et al. Reference Weirather, Wilson and Donelson2012; Hall et al. Reference Hall, Wang and Barry2013). Other species-specific genes like the Invariant Surface Glycoprotein (ISG) genes (Jackson et al. Reference Jackson, Windle and Voorheis1993; Ziegelbauer and Overath, Reference Ziegelbauer and Overath1993) and Expression-Site Associated Genes (ESAGs; Pays et al. Reference Pays, Lips, Nolan, Vanhamme and Pérez-Morga2001; see below) are also preferentially expressed in the bloodstream stage. In the insect host, species-specific genes are dominated by procyclin, encoding the major surface glycoprotein of the procyclic stage while in the insect midgut (Roditi et al. Reference Roditi, Furger, Ruepp, Schürch and Bütikofer1998; Berriman et al. Reference Berriman, Ghedin, Hertz-Fowler, Blandin, Renauld, Bartholomeu, Lennard, Caler, Hamlin, Haas, Böhme, Hannick, Aslett, Shallom, Marcello, Hou, Wickstead, Alsmark, Arrowsmith, Atkin, Barron, Bringaud, Brooks, Carrington, Cherevach, Chillingworth, Churcher, Clark, Corton, Cronin, Davies, Doggett, Djikeng, Feldblyum, Field, Fraser, Goodhead, Hance, Harper, Harris, Hauser, Hostetler, Ivens, Jagels, Johnson, Johnson, Jones, Kerhornou, Koo, Larke, Landfear, Larkin, Leech, Line, Lord, Macleod, Mooney, Moule, Martin, Morgan, Mungall, Norbertczak, Ormond, Pai, Peacock, Peterson, Quail, Rabbinowitsch, Rajandream, Reitter, Salzberg, Sanders, Schobel, Sharp, Simmonds, Simpson, Tallon, Turner, Tait, Tivey, Van Aken, Walker, Wanless, Wang, White, White, Whitehead, Woodward, Wortman, Adams, Embley, Gull, Ullu, Barry, Fairlamb, Opperdoes, Barrell, Donelson, Hall, Fraser, Melville and El-Sayed2005), and the Brucei Alanine-Rich Protein (BARP) that, along with related forms, is specifically expressed by the epimastigote and metacyclic stages while in the insect mouthparts (Urwyler et al. Reference Urwyler, Studer, Renggli and Roditi2007; Jackson et al. Reference Jackson, Allison, Barry, Field, Hertz-Fowler and Berriman2013).
The cell surface of Leishmania is dominated by non-protein lipophosphoglycan (LPG) and glycoinositolphospholipid (GIPL) molecules (de Assis et al. Reference de Assis, Ibraim, Nogueira, Soares and Turco2012). The LPG/GIPL coat is complemented by species-specific, multi-copy proteins such as δ-amastin, which is specifically expressed during the intracellular amastigote stage (Rochette et al. Reference Rochette, McNicoll, Girard, Breton, Leblanc, Bergeron and Papadopoulou2005). While its function is unknown, the evolution of δ-amastin is thought to be an adaptation for infection of, or survival within, macrophages since it is absent from monoxenic species (Crithidia and Leptomonas spp.) lacking a vertebrate stage (Jackson, Reference Jackson2010) and less abundant in Leishmania species that do not routinely infect macrophages (Raymond et al. Reference Raymond, Boisvert, Roy, Ritt, Légaré, Isnard, Stanke, Olivier, Tremblay, Papadopoulou, Ouellette and Corbeil2012). Furthermore, a parallel expansion of δ-amastin has occurred in T. cruzi, which also has an amastigote stage, and this is associated with virulence (Kangussu-Marcolino et al. Reference Kangussu-Marcolino, de Paiva, Araújo, de Mendonça-Neto, Lemos, Bartholomeu, Mortara, daRocha and Teixeira2013). Another Leishmania-specific family, tuzin (Ivens et al. Reference Ivens, Peacock, Worthey, Murphy, Aggarwal, Berriman, Sisk, Rajandream, Adlem, Aert, Anupama, Apostolou, Attipoe, Bason, Bauser, Beck, Beverley, Bianchettin, Borzym, Bothe, Bruschi, Collins, Cadag, Ciarloni, Clayton, Coulson, Cronin, Cruz, Davies, De Gaudenzi, Dobson, Duesterhoeft, Fazelina, Fosker, Frasch, Fraser, Fuchs, Gabel, Goble, Goffeau, Harris, Hertz-Fowler, Hilbert, Horn, Huang, Klages, Knights, Kube, Larke, Litvin, Lord, Louie, Marra, Masuy, Matthews, Michaeli, Mottram, Müller-Auer, Munden, Nelson, Norbertczak, Oliver, O'neil, Pentony, Pohl, Price, Purnelle, Quail, Rabbinowitsch, Reinhardt, Rieger, Rinta, Robben, Robertson, Ruiz, Rutter, Saunders, Schäfer, Schein, Schwartz, Seeger, Seyler, Sharp, Shin, Sivam, Squares, Squares, Tosato, Vogt, Volckaert, Wambutt, Warren, Wedler, Woodward, Zhou, Zimmermann, Smith, Blackwell, Stuart, Barrell and Myler2005), is linked to δ-amastin loci physically and phylogenetically (Jackson, Reference Jackson2010); hence, tuzin might be involved in the same adaptation. In the insect life stage, the promastigote surface antigen (PSA or gp46) is preferentially expressed in metacyclic promastigotes (Handman et al. Reference Handman, Osborn, Symons, van Driel and Cappai1995) and is encoded by a diverse gene family in human-infecting species (Devault and Bañuls, Reference Devault and Bañuls2008). Also specifically expressed in metacyclics are the HASP (Hydrophilic Acylated Surface Protein) and SHERP (Small Hydrophilic ER-associated Protein) gene families (Depledge et al. Reference Depledge, MacLean, Hodgkinson, Smith, Jackson, Ma, Uliana and Smith2010; Sádlová et al. Reference Sádlová, Price, Smith, Votýpka, Volf and Smith2010).
While the precise functions of these enigmatic gene families are unknown, several contribute to parasite fitness. This may be because they initiate infection, for instance, the TcMUC and TS proteins interact to transfer host sialic acid residues to parasite mucins, which is essential for attachment and invasion by T. cruzi trypomastigotes (Acosta-Serrano et al. Reference Acosta-Serrano, Almeida, Freitas-Junior, Yoshida and Schenkman2001; Oliveira et al. Reference Oliveira, Freire-de-Lima, Penha, Dias and Todeschini2014). Other cell surface protein families are essential for parasite development and transmission through the insect host; for example, HASP and SHERP are required for L. major to form infective metacyclics while in the insect foregut (Sádlová et al. Reference Sádlová, Price, Smith, Votýpka, Volf and Smith2010). However, given their prominent roles at the cell surface, most of these species-specific proteins are likely to have immunological roles. These may be in suppressing innate responses, for example by degrading antimicrobial peptides or other effectors of complement-mediated lysis, as has been shown for PSA (Lincoln et al. Reference Lincoln, Ozaki, Donelson and Beetham2004), or in manipulating cell-mediated immune responses. For instance, TcMUC represses T-cell expansion and cytokine production (Nunes et al. Reference Nunes, Fortes, Silva-Filho, Terra-Granado, Santos, Conde, de Araújo Oliveira, Freire-de-Lima, Martins, Pinheiro, Takyia, Freire-de-Lima, Todeschini, Dosreis and Morrot2013). Salivarian trypanosomes employ VSG in antigenic variation, and have evolved sophisticated mechanisms for regulating VSG expression (see below). The abundance and variety of TcMUC, TS and MASP genes has led some to suggest that a subtler form of antigenic variation operates in T. cruzi as well (Buscaglia et al. Reference Buscaglia, Campo, Di Noia, Torrecilhas, De Marchi, Ferguson, Frasch and Almeida2004, Reference Buscaglia, Campo, Frasch and Di Noia2006; dos Santos et al. Reference dos Santos, Freitas, Lobo, Rodrigues-Luiz, Mendes, Oliveira, Andrade, Chiari, Gazzinelli, Teixeira, Fujiwara and Bartholomeu2012).
GENOMIC INNOVATION: CONTINGENCY ZONES
Trypanosomatids have substantially modified the genome to accommodate these abundant families of cell-surface effectors, by creating genomic sub-domains segregated from the core genome by distance, but also by sequence composition and epigenetic modification (Figueiredo et al. Reference Figueiredo, Cross and Janzen2009; Rudenko, Reference Rudenko2010). We can call these sub-domains ‘contingency zones’ because they provide the environment for flexible expression of what are known as contingency genes (Deitsch et al. Reference Deitsch, Moxon and Wellems1997). In this trypanosomatids are not alone; diverse parasites possess polymorphic effector protein families that display specialized expression profiles across a wide range of physiological conditions (Deitsch et al. Reference Deitsch, Moxon and Wellems1997; Kissinger and DeBarry, Reference Kissinger and DeBarry2011). It has often been observed that contingency genes aggregate towards the telomeres, a position that promotes both the specific regulation of their expression and their diversification through recombination and gene duplication (Barry et al. Reference Barry, Ginger, Burton and McCulloch2003; Kissinger and DeBarry, Reference Kissinger and DeBarry2011). Thus, both T. brucei and T. cruzi have expanded sub-telomeric regions to contain and regulate their diverse contingency genes (Berriman et al. Reference Berriman, Ghedin, Hertz-Fowler, Blandin, Renauld, Bartholomeu, Lennard, Caler, Hamlin, Haas, Böhme, Hannick, Aslett, Shallom, Marcello, Hou, Wickstead, Alsmark, Arrowsmith, Atkin, Barron, Bringaud, Brooks, Carrington, Cherevach, Chillingworth, Churcher, Clark, Corton, Cronin, Davies, Doggett, Djikeng, Feldblyum, Field, Fraser, Goodhead, Hance, Harper, Harris, Hauser, Hostetler, Ivens, Jagels, Johnson, Johnson, Jones, Kerhornou, Koo, Larke, Landfear, Larkin, Leech, Line, Lord, Macleod, Mooney, Moule, Martin, Morgan, Mungall, Norbertczak, Ormond, Pai, Peacock, Peterson, Quail, Rabbinowitsch, Rajandream, Reitter, Salzberg, Sanders, Schobel, Sharp, Simmonds, Simpson, Tallon, Turner, Tait, Tivey, Van Aken, Walker, Wanless, Wang, White, White, Whitehead, Woodward, Wortman, Adams, Embley, Gull, Ullu, Barry, Fairlamb, Opperdoes, Barrell, Donelson, Hall, Fraser, Melville and El-Sayed2005; El-sayed et al. Reference El-Sayed, Myler, Bartholomeu, Nilsson, Aggarwal, Tran, Ghedin, Worthey, Delcher, Blandin, Westenberger, Caler, Cerqueira, Branche, Haas, Anupama, Arner, Aslund, Attipoe, Bontempi, Bringaud, Burton, Cadag, Campbell, Carrington, Crabtree, Darban, da Silveira, de Jong, Edwards, Englund, Fazelina, Feldblyum, Ferella, Frasch, Gull, Horn, Hou, Huang, Kindlund, Klingbeil, Kluge, Koo, Lacerda, Levin, Lorenzi, Louie, Machado, McCulloch, McKenna, Mizuno, Mottram, Nelson, Ochaya, Osoegawa, Pai, Parsons, Pentony, Pettersson, Pop, Ramirez, Rinta, Robertson, Salzberg, Sanchez, Seyler, Sharma, Shetty, Simpson, Sisk, Tammi, Tarleton, Teixeira, Van Aken, Vogt, Ward, Wickstead, Wortman, White, Fraser, Stuart and Andersson2005a , Reference El-Sayed, Myler, Blandin, Berriman, Crabtree, Aggarwal, Caler, Renauld, Worthey, Hertz-Fowler, Ghedin, Peacock, Bartholomeu, Haas, Tran, Wortman, Alsmark, Angiuoli, Anupama, Badger, Bringaud, Cadag, Carlton, Cerqueira, Creasy, Delcher, Djikeng, Embley, Hauser, Ivens, Kummerfeld, Pereira-Leal, Nilsson, Peterson, Salzberg, Shallom, Silva, Sundaram, Westenberger, White, Melville, Donelson, Andersson, Stuart and Hall b ; Moraes Barros et al. Reference Moraes Barros, Marini, Antônio, Cortez, Miyake, Lima, Ruiz, Bartholomeu, Chiurillo, Ramirez and da Silveira2012). It is likely that the strand-switch regions that occur between polycistrons on trypanosomatid chromosomes also serve as incubators of novelty, since they often harbour species-specific genes (Peacock et al. Reference Peacock, Seeger, Harris, Murphy, Ruiz, Quail, Peters, Adlem, Tivey, Aslett, Kerhornou, Ivens, Fraser, Rajandream, Carver, Norbertczak, Chillingworth, Hance, Jagels, Moule, Ormond, Rutter, Squares, Whitehead, Rabbinowitsch, Arrowsmith, White, Thurston, Bringaud, Baldauf, Faulconbridge, Jeffares, Depledge, Oyola, Hilley, Brito, Tosi, Barrell, Cruz, Mottram, Smith and Berriman2007; Jackson et al. Reference Jackson, Sanders, Berry, McQuillan, Aslett, Quail, Chukualim, Capewell, MacLeod, Melville, Gibson, Barry, Berriman and Hertz-Fowler2009).
Perhaps the best example of structural innovation in trypanosomatid genomes is the VSG expression site (ES) in T. brucei. African trypanosomes evade the humoral immune response by periodically switching the VSG monolayer that masks their cell surfaces. This demands that only a single VSG is expressed at a time, while all others are silenced (i.e. monoallelic expression). The function of the ES is to ensure monoallelic expression by providing a dedicated locus for VSG transcription. Thus, the active VSG is transcribed solely from one of several, alternative ESs and antigenic switching occurs when a different VSG from among the many hundreds of silent, sub-telomeric loci, replaces the ES copy through ectopic gene conversion, or by activating an alternative ES (Horn and McCulloch, Reference Horn and McCulloch2010; Rudenko, Reference Rudenko2011). Analysis of ES sequences from several T. brucei strains has identified a canonical ES structure (Graham et al. Reference Graham, Terry and Barry1999; Berriman et al. Reference Berriman, Hall, Sheader, Bringaud, Tiwari, Isobe, Bowman, Corton, Clark, Cross, Hoek, Zanders, Berberof, Borst and Rudenko2002; Hertz-Fowler et al. Reference Hertz-Fowler, Figueiredo, Quail, Becker, Jackson, Bason, Brooks, Churcher, Fahkro, Goodhead, Heath, Kartvelishvili, Mungall, Harris, Hauser, Sanders, Saunders, Seeger, Sharp, Taylor, Walker, White, Young, Cross, Rudenko, Barry, Louis and Berriman2008), which includes not only the VSG and repeat sequences required to promote recombination with sub-telomeric VSG loci, but also the ESAGs (reviewed in Pays et al. Reference Pays, Lips, Nolan, Vanhamme and Pérez-Morga2001; McCulloch and Horn, Reference McCulloch and Horn2009). The functions of most ESAGs are unclear; however, all are transcribed preferentially in the bloodstream stage (Jensen et al. Reference Jensen, Sivam, Kifer, Myler and Parsons2009; Siegel et al. Reference Siegel, Hekstra, Wang, Dewell and Cross2010; Veitch et al. Reference Veitch, Johnson, Trivedi, Terry, Wildridge and MacLeod2010) and it is known that they are T. brucei-specific innovations, often derived from conserved gene families with pre-existing cell surface roles (Barker et al. Reference Barker, Wickstead, Gluenz and Gull2008; Barnwell et al. Reference Barnwell, van Deursen, Jeacock, Smith, Maizels, Acosta-Serrano and Matthews2010; Salmon et al. Reference Salmon, Bachmaier, Krumbholz, Kador, Gossmann, Uzureau, Pays and Boshart2012; Jackson et al. Reference Jackson, Allison, Barry, Field, Hertz-Fowler and Berriman2013). Hence, it may be that they support antigenic variation or that the specific regulatory environment of the ES has been exploited secondarily to up-regulate proteins with established and diverse roles during the bloodstream stage.
GENOMIC INNOVATION: THE MAJOR SURFACE PROTEASES
Alongside the many species-specific cell surface proteins, there is one family conserved in all trypansomatid genomes that must have experienced substantial evolution since the origin of parasitism. The Major Surface Protease (MSP) gene family encode a range of metalloproteases that are implicated in various aspects of pathogenesis and virulence in Leishmania (Yao, Reference Yao2010). MSP subverts the normal host defensive mechanisms by degrading components of immune cell signalling pathways (Gomez et al. Reference Gomez, Contreras, Hallé, Tremblay, McMaster and Olivier2009; Hallé et al. Reference Hallé, Gomez, Stuible, Shimizu, McMaster, Olivier and Tremblay2009; Contreras et al. Reference Contreras, Gómez, Nguyen, Shio, McMaster and Olivier2010), and suppresses other aspects of innate immunity (Kulkarni et al. Reference Kulkarni, McMaster, Kamysz, Kamysz, Engman and McGwire2006; Lieke et al. Reference Lieke, Nylén, Eidsmo, McMaster, Mohammadi, Khamesipour, Berg and Akuffo2008). In Trypanosoma, MSP is equally abundant in gene copy number and protein abundance but its function is less well understood; it is known to remove the VSG coat from the T. brucei surface during differentiation into the procyclic form (PCF) (Grandgenett et al. Reference Grandgenett, Otsu, Wilson, Wilson and Donelson2007) and is thought to have a role in cell invasion by T. cruzi (Cuevas et al. Reference Cuevas, Cazzulo and Sánchez2003; Kulkarni et al. Reference Kulkarni, Olson, Engman and McGwire2009). As it is present in all trypanosomatids, we can infer the diversification of MSP from its phylogeny, and this too indicates that MSP has been instrumental in parasite adaptation.
The MSP phylogeny is described in Fig. 3. It shows how, beginning from a much smaller gene repertoire, MSP has differentiated into distinct clades in both Leishmania and Trypanosoma (Victoir et al. Reference Victoir, Arevalo, De Doncker, Barker, Laurent, Godfroid, Bollen, Le Ray and Dujardin2005; Ma et al. Reference Ma, Chen, Meng, Liu, Tang, Hu and Yu2011); each clade is associated with a conserved locus, and we know that some of these distinct lineages are developmentally regulated (Yao, Reference Yao2010). For instance, MSP-A and MSP-C are up-regulated in bloodstream form (BSF) T. brucei, while MSP-B is predominantly seen in the procyclic form (LaCount et al. Reference LaCount, Gruszynski, Grandgenett, Bangs and Donelson2003; Urbaniak et al. Reference Urbaniak, Guther and Ferguson2012). Hence, the trypanosomatids have elaborated their MSP repertoire by creating new loci at least in part to regulate function during the life cycle. Moreover, these different forms have been duplicated to create multiple isoforms, often in species-specific ways; for instance, MSP-C is polymorphic in Trypanosoma vivax while single copy in other salivarian species, and the single-copy MSP gene found on chromosome 28 in Leishmania has been greatly expanded in Phytomonas. However, the phylogeny also demonstrates that MSP in Leishmania and Trypanosoma cluster by genus, and therefore, there is no orthologous MSP shared by all. Thus, MSP repertoires in Leishmania and Trypanosoma have evolved independently, and their similarities in genomic structure, developmental regulation and pathogenesis represent parallel evolution, reflecting a common need for diverse surface proteases throughout trypanosomatid diversification.
Major Surface Protease (MSP) gene family phylogeny. This maximum likelihood phylogeny of MSP amino acid sequences sampled from completed genome sequences was estimated using PHYML and a LG+Γ model of amino acid substitution (Guindon et al. Reference Guindon, Dufayard, Lefort, Anisimova, Hordijk and Gascuel2010). Node robustness was assessed with non-parametric bootstraps; branches with bootstrap support >75 are shown with bold lines. Sequences are labelled with coloured symbols, according to the key. The tree is mid-point rooted, which corresponds with the Trypanosoma-specific MSP-D locus (Marcoux et al. Reference Marcoux, Wei, Tabel and Bull2010).

DEVELOPMENTAL REGULATION OF GENE EXPRESSION
Trypanosomatids display morphological plasticity that is often associated with developmental transition through a complex life cycle. This is important for the origins of parasitism but not an issue that comparative genomics can illuminate dramatically, without including a comparator lacking developmental complexity. The recent discovery of Paratrypanosoma confusum parasitizing the gut of a Culex pipiens mosquito strengthens the argument that the ancestral trypanosomatid was a monoxenic insect parasite, since P. confusum is a robust outgroup to all other trypanosomatids (Flegontov et al. Reference Flegontov, Votýpka, Skalický, Logacheva, Penin, Tanifuji, Onodera, Kondrashov, Volf, Archibald and Lukeš2013). As long as P. confusum has no second host, this shows that a dixenic life cycle has evolved on three separate occasions in Trypanosoma, Leishmania and Phytomonas. Trypanosomatids are capable of assuming multiple developmental forms and transition between forms coincides with passing between distinct environments, whether they are in different hosts or a single host, for example from the hindgut to the foregut of an insect. Experimental approaches are beginning to reveal the non-coding sequences (Bringaud et al. Reference Bringaud, Müller, Cerqueira, Smith, Rochette, El-Sayed, Papadopoulou and Ghedin2007; Holzer et al. Reference Holzer, Mishra, LeBowitz and Forney2008; Smith et al. Reference Smith, Bringaud and Papadopoulou2009; Li et al. Reference Li, De Gaudenzi, Alvarez, Mendiondo, Wang, Kissinger, Frasch and Docampo2012; Pastro et al. Reference Pastro, Smircich, Pérez-Díaz, Duhagon and Garat2013) and RNA-binding proteins (reviewed in Kolev et al. Reference Kolev, Ullu and Tschudi2014) that interact to regulate gene expression, as well as genes specifically required for differentiation from one life stage to another (Goldenberg and Avila, Reference Goldenberg and Avila2011; Kolev et al. Reference Kolev, Ramey-Butler, Cross, Ullu and Tschudi2012; Rico et al. Reference Rico, Rojas, Mony, Szoor, MacGregor and Matthews2013). Comparison of life-stage-specific transcriptomes (Holzer et al. Reference Holzer, McMaster and Forney2006; Leifso et al. Reference Leifso, Cohen-Freue, Dogra, Murray and McMaster2007; Saxena et al. Reference Saxena, Lahav, Holland, Aggarwal, Anupama, Huang, Volpin, Myler and Zilberstein2007; Rochette et al. Reference Rochette, Raymond, Ubeda, Smith, Messier, Boisvert, Rigault, Corbeil, Ouellette and Papadopoulou2008, Reference Rochette, Raymond, Corbeil, Ouellette and Papadopoulou2009; Alcolea et al. Reference Alcolea, Alonso, Sánchez-Gorostiaga, Moreno-Paz, Gómez, Ramos, Parro and Larraga2009, Reference Alcolea, Alonso, Gómez, Moreno, Domínguez, Parro and Larraga2010; Depledge et al. Reference Depledge, Evans, Ivens, Aziz, Maroof, Kaye and Smith2009; Jensen et al. Reference Jensen, Sivam, Kifer, Myler and Parsons2009; Kabani et al. Reference Kabani, Fenn, Ross, Ivens, Smith, Ghazal and Matthews2009; Minning et al. Reference Minning, Weatherly, Atwood, Orlando and Tarleton2009; Veitch et al. Reference Veitch, Johnson, Trivedi, Terry, Wildridge and MacLeod2010; Adaui et al. Reference Adaui, Castillo, Zimic, Gutierrez, Decuypere, Vanaerschot, De Doncker, Schnorbusch, Maes, Van der Auwera, Maes, Llanos-Cuentas, Arevalo and Dujardin2011;) and proteomes (Atwood et al. Reference Atwood, Weatherly, Minning, Bundy, Cavola, Opperdoes, Orlando and Tarleton2005; Rosenzweig et al. Reference Rosenzweig, Smith, Opperdoes, Stern, Olafson and Zilberstein2008a , Reference Rosenzweig, Smith, Myler, Olafson and Zilberstein b ; Alcolea et al. Reference Alcolea, Alonso and Larraga2011; Urbaniak et al. Reference Urbaniak, Guther and Ferguson2012; Gunasekera et al. Reference Gunasekera, Wüthrich, Braga-Lagache, Heller and Ochsenreiter2012; Butter et al. Reference Butter, Bucerius, Michel, Cicova, Mann and Janzen2013) in various species have estimated the proportion of genes showing preferential expression in the insect or vertebrate stages to be between 2 and 44%; the breadth of these values reflects the diverse conditions and approaches employed. However, it is clear that a significant minority of genes are developmentally regulated. We can predict that this regulation is achieved with layers of interaction between genomic loci, mRNA, non-coding RNA and DNA and RNA-binding proteins. Hence, to understand the origins of complex life cycles we will need to compare the interaction networks of free-living, monoxenic and dixenic Kinetoplastids, and in this P. confusum and the free-living Bodo saltans will be instrumental.
MECHANISMS OF GENOMIC EVOLUTION: GENE DUPLICATION
Besides the genomic innovations themselves, comparative analysis also reveals the molecular mechanisms that create them. These evolutionary events range in size from single amino acid substitutions to chromosomal duplications, and include both coding and non-coding regions, but it is gene duplication above all that creates the raw material for evolutionary novelty (Ohno, Reference Ohno1970). After duplication, paralogs may acquire new functions (neofunctionalization), segregate existing functions (subfunctionalization) or lose function under mutation pressure (pseudogenization) (Lynch and Conery, Reference Lynch and Conery2000). Since developmental regulation of gene expression is widespread, it is unsurprising that many gene duplicates are distinguished in the timing or location of their expression. For example, TcMCA5 is an epimastigote-specific metacaspase implicated in programmed cell death of T. cruzi that has evolved from a constitutively expressed metacaspase gene family (Kosec et al. Reference Kosec, Alvarez, Agüero, Sánchez, Dolinar, Turk, Turk and Cazzulo2006). In Leishmania, Zinoviev et al. (Reference Zinoviev, Akum, Yahav and Shapira2012) identified two functionally redundant RNA helicases that have evolved purely to perform the same role in insect and vertebrate stages respectively. By contrast, TcPRACA and TcPRACB are two paralogous proline racemases involved in immune-suppression by T. cruzi (Reina-San-Martín et al. Reference Reina-San-Martín, Degrave, Rougeot, Cosson, Chamond, Cordeiro-Da-Silva, Arala-Chaves, Coutinho and Minoprio2000); here, function is segregated by location, TcPRACB being expressed intracellularly and TcPRACA secreted (Chamond et al. Reference Chamond, Goytia, Coatnoan, Barale, Cosson, Degrave and Minoprio2005).
Of course, the derivation of many gene duplicates may be multifactorial; in the example of proline racemases, secretion of TcPRACA may coincide with a new role in the differentiation of infective metacyclics (Chamond et al. Reference Chamond, Goytia, Coatnoan, Barale, Cosson, Degrave and Minoprio2005). Thus, it is difficult to unambiguously distinguish neofunctionalization from the segregation of the same function by time, space or substrate. However, the transferrin receptor (TFR) in T. brucei, which is required for salvaging haem from the host and is homologous to the VSG (Salmon et al. Reference Salmon, Hanocq-Quertier, Paturiaux-Hanocq, Pays, Tebabi, Nolan, Michel and Pays1997), is one example. Recently, it was confirmed that the TFR had evolved from an a-type VSG in the ancestor of T. brucei and Trypanosoma congolense, and that, despite their homology, TFR and VSG genes do not recombine, supporting a functionally distinct role from the variant antigen repertoire (Jackson et al. Reference Jackson, Berry, Aslett, Allison, Burton, Vavrova-Anderson, Brown, Browne, Corton, Hauser, Gamble, Gilderthorp, Marcello, McQuillan, Otto, Quail, Sanders, van Tonder, Ginger, Field, Barry, Hertz-Fowler and Berriman2012, Reference Jackson, Allison, Barry, Field, Hertz-Fowler and Berriman2013). As suggested above, the conspicuous abundance and diversity of certain T. cruzi gene families, such as TS, EF1γ and MSP, could indicate that these genes have secondarily evolved a novel role in immune evasion as a consequence of being at the cell surface for their pre-existing functions, i.e. to transfer sialic acid to TcMUC in the case of TS (Oliveira et al. Reference Oliveira, Freire-de-Lima, Penha, Dias and Todeschini2014). Furthermore, many TS, EF1γ and MSP genes in T. cruzi are not predicted to encode proteins capable of their putative functions (El-Sayed et al. 2005). At first sight, this would appear to indicate frequent pseudogenization, yet a population of pseudogenes acquiring substitutions under neutral conditions would be expected to display a spectrum of mutational decay that is not seen (El-Sayed et al. 2005). This suggests that these genes may remain under purifying selection for another role, which could represent neofunctionalization.
The evolution of gene duplicates is particularly obvious in the abundant tandem gene arrays of trypanosomatid genomes. Tandem duplication is very common in trypanosomatids, perhaps as a means of increasing transcript abundance for highly expressed genes in the presence of polycistronic transcription. Comparative analysis of homologous arrays shows that tandem duplicates can evolve new functions, despite the propensity for concerted evolution of tandemly arrayed genes (Jackson, Reference Jackson2007a ), and that this follows a consistent pattern of structural segregation. Figure 4 shows two examples of functional divergence within tandem gene arrays. The expression profiles of adenylate cyclase gene paralogues from the rac array of Leishmania spp. correspond with their position in the array. The 3′-most gene (rac-A) and the gene positioned upstream of rac-A in the array (rac-B1) are expressed specifically in the promastigote (Sanchez et al. Reference Sanchez, Zeoli, Klamo, Kavanaugh and Landfear1995; Akopyants et al. Reference Akopyants, Matlib, Bukanova, Smeds, Brownstein, Stormo and Beverley2004), while transcripts for the remaining copies are more abundant in the amastigote (Akopyants et al. Reference Akopyants, Matlib, Bukanova, Smeds, Brownstein, Stormo and Beverley2004). Interestingly, rac-A and rac-B1 may have differentiated in a complementary fashion, since rac-B1 negatively regulates the activity of rac-A in the promastigote (Sanchez et al. Reference Sanchez, Zeoli, Klamo, Kavanaugh and Landfear1995). In Trypanosoma, the 5′-most copy of a cation transporter gene array is preferentially expressed in the PCF (Jensen et al. Reference Jensen, Sivam, Kifer, Myler and Parsons2009; Urbaniak et al. Reference Urbaniak, Guther and Ferguson2012) (indeed, it is essential to its growth; Alsford et al. Reference Alsford, Turner, Obado, Sanchez-Flores, Glover, Berriman, Hertz-Fowler and Horn2011), while transcripts for all downstream copies are up-regulated in the bloodstream stage (Jensen et al. Reference Jensen, Sivam, Kifer, Myler and Parsons2009; Veitch et al. Reference Veitch, Johnson, Trivedi, Terry, Wildridge and MacLeod2010).
Structural differentiation of paralogues in tandem gene arrays. a. Receptor-type adenylate cyclase (rac) genes in four Leishmania species. The rac array (i.e. LmjF.17·0200) is located at the extreme left-end of chromosome 17 in L. major; its conserved position is defined downstream by an EF1α gene array (i) and upstream by a metalloprotease (ii). The structure of the array in four species is depicted, rac genes are shown in black, and other loci are shown in white. Vertical grey shading represents homology between flanking loci, to demonstrate positional conservation. The dashed black line separates array positions that correspond with distinct clades in the phylogeny. A maximum likelihood phylogeny estimated from amino acid sequences using a LG+Γ model is shown, with non-parametric bootstraps applied to nodes where support is >75. Terminal nodes are labelled with species name initials. The tree is midpoint rooted. b. Cation transporter genes in four Trypanosoma species. The transporter array (i.e. Tb927.11.9000) is located on chromosome 11 in T. brucei; its conserved position is defined downstream by a palmitoyl acyltransferase 4 gene (i) and upstream by an EF1γ2 gene (ii). The phylogeny and genome comparison are as depicted in a. except that the tree is rooted with a single-copy orthology from L. major.

The phylogenies of these gene duplicates show that those gene copies that are functionally differentiated retain orthology across species (i.e. they cluster together despite being in different genomes), while undifferentiated copies cluster by species. This shows that gene duplicates that have diverged in their structures and expression for a novel function are preserved by selection over the course of trypanosomatid evolution, despite the pressure exerted by allelic gene conversion in these situations. In fact, when tandem gene duplicates differentiate, this often occurs at either end of the array (Jackson, Reference Jackson2007a ), even occurring in otherwise invariant arrays that are exposed to frequent gene conversion; for example, differentiation of the terminal 3′UTR in the β-tubulin array in Leishmania spp. has created a promastigote-specific β-tubulin isoform (Jackson et al. Reference Jackson, Vaughan and Gull2006).
Duplication events do not only affect individual genes. A 0·5 mb segmental duplication in T. brucei was identified that has created duplicons shared by chromosomes 4 and 8 (Jackson, Reference Jackson2007b ). Originally, this region contained approximately 158 genes but subsequent deletions from either duplicon have returned many loci to their original copy number. However, 74 loci have been retained as paralogues in both duplicons. Comparison of their coding and flanking sequences indicated that substantial divergence had occurred and this was assumed to reflect functional divergence (Jackson, Reference Jackson2007b ). They include CAP5.5, a cysteine peptidase essential for cell morphogenesis, which has been shown to have two paralogues expressed specifically in the insect and vertebrate stages respectively (Hertz-Fowler et al. Reference Hertz-Fowler, Ersfeld and Gull2001; Olego-Fernandez et al. Reference Olego-Fernandez, Vaughan, Shaw, Gull and Ginger2009). Figure 5 shows how recent proteomic evidence now confirms that several of the paralogues retained after segmental duplication have evolved stage-specific expression profiles, indicating subfunctionalization by life stage. Gene expression in trypanosomatids is largely regulated by sequences within the 3′ untranslated region (UTR) of transcripts (Vanhamme and Pays, Reference Vanhamme and Pays1995; Haile and Papadopoulou, Reference Haile and Papadopoulou2007). Accordingly, it is the paralogous pairs with no sequence identity in their 3′ UTRs that have the greatest differences (loci #13, 36, 39, 49 and 71 in Fig. 5), while those paralogues with similar 3′ UTR sequences display similar abundance in both cases (loci #23, 24, 62 and 65 in Fig. 5).
Peptide abundance in procyclic form (PCF) and bloodstream form (BSF) T. brucei for selected paralogues resulting from a segmental duplication. 74 loci are present in two forms in T. brucei due to a segmental duplication. The loci listed here are those detected in proteomic analyses. Locus number and sequence identity values refer to the segmental duplication described in Jackson (Reference Jackson2007b ). Fold change in peptide abundance between PCF and BSF cells taken from * Urbaniak et al. (Reference Urbaniak, Guther and Ferguson2012) or ** Butter et al. (Reference Butter, Bucerius, Michel, Cicova, Mann and Janzen2013). Preferential expression in PCF and BSF is indicated by blue and red shading respectively. Constitutive expression is indicated by orange shading.

MECHANISMS OF GENOMIC EVOLUTION: HORIZONTAL GENE TRANSFER
Horizontal gene transfer (HGT) is another mechanism by which many eukaryotic genomes have acquired new functionality. Berriman et al. (2005) identified 49 putative HGT from bacteria and other eukaryotes in trypanosomatid genomes. Confirming HGT rests on sound phylogenetic reconstruction; the most convincing cases are those where the donated gene is closely related to donor genes in unrelated genomes, and nested among these in a phylogeny. Some putative HGT in trypanosomatids achieve this, notably the haem-biosynthesis pathway, absent from Trypanosoma but partially restored in Leishmania and related genera through HGT of three genes (hemF, hemG and hemH encoding coproporphyrinogen oxidase, protoporphyrinogen oxidase and ferrochelatase, respectively) from gamma-proteobacteria. In phylogenies, HemF-H are nested among bacterial homologues and apart from related eukaryotic genes (Ivens et al. Reference Ivens, Peacock, Worthey, Murphy, Aggarwal, Berriman, Sisk, Rajandream, Adlem, Aert, Anupama, Apostolou, Attipoe, Bason, Bauser, Beck, Beverley, Bianchettin, Borzym, Bothe, Bruschi, Collins, Cadag, Ciarloni, Clayton, Coulson, Cronin, Cruz, Davies, De Gaudenzi, Dobson, Duesterhoeft, Fazelina, Fosker, Frasch, Fraser, Fuchs, Gabel, Goble, Goffeau, Harris, Hertz-Fowler, Hilbert, Horn, Huang, Klages, Knights, Kube, Larke, Litvin, Lord, Louie, Marra, Masuy, Matthews, Michaeli, Mottram, Müller-Auer, Munden, Nelson, Norbertczak, Oliver, O'neil, Pentony, Pohl, Price, Purnelle, Quail, Rabbinowitsch, Reinhardt, Rieger, Rinta, Robben, Robertson, Ruiz, Rutter, Saunders, Schäfer, Schein, Schwartz, Seeger, Seyler, Sharp, Shin, Sivam, Squares, Squares, Tosato, Vogt, Volckaert, Wambutt, Warren, Wedler, Woodward, Zhou, Zimmermann, Smith, Blackwell, Stuart, Barrell and Myler2005, Korený et al. Reference Korený, Lukes and Oborník2010). In salivarian trypanosomes, a phospholipase A1 (PLA1) gene is thought to have been acquired from proteobacteria (Richmond and Smith, Reference Richmond and Smith2007). In support of this, the PLA1 gene is absent from all other Kinetoplastids (indeed most other eukaryotes) and it nests among proteobacterial sequences in sequence comparisons. Moreover, the PLA1 locus (Tb927.1.4830) occurs precisely at the boundary between chromosomal core and sub-telomere in African trypanosome genomes, suggesting perhaps that it was recently transposed.
Other good examples of HGT include a cytosolic dihydroorotate dehydrogenase in the pyrimidine biosynthetic pathway, which is unique to Kinetoplastids, and replaces the mitochondrial dihydroorotate dehydrogenase that is typical of euglenids and other eukaryotes. In phylogenies, the cytosolic genes are nested among bacterial taxa, while the mitochondrial genes form a eukaryotic clade (Annoura et al. Reference Annoura, Nara, Makiuchi, Hashimoto and Aoki2005). Likewise, ornithine decarboxylase genes from salivarian trypanosomes do not cluster with homologues from other trypanosomatids, but instead they are nested among metazoan genes and are the sister taxon to ornithine decarboxylase from vertebrates (Steglich and Schaeffer, Reference Steglich and Schaeffer2006). In fact, ornithine decarboxylase is known to be absent from T. cruzi (Carrillo et al. Reference Carrillo, Cejas, González and Algranati1999), indicating that this HGT from vertebrates has restored function in African trypanosomes that was lost after the origin of Trypanosoma. However, since the African trypanosome genes are not nested within the vertebrate clade, we can rule out any recent transfer from contemporary hosts and suggest instead a more distant transfer from an ancient chordate.
In other cases of putative HGT the donated gene is not nested among would-be donors, just closest to them in phylogenies. Here, it is possible that the punctate distribution is due to lineage sorting, i.e. patchy inheritance of an ancestral lineage by daughter lineages. When, as is common, eukaryotic diversity is inadequately sampled, it is difficult to distinguish HGT and lineage sorting. For example, trypanosomatid genomes possess four superoxide dismutase genes required for antioxidant defence (soda, sodb1, sodb2 and sodc), which localize to distinct cellular compartments (Dufernez et al. Reference Dufernez, Yernaux, Gerbod, Noël, Chauvenet, Wintjens, Edgcomb, Capron, Opperdoes and Viscogliosi2006). The four sod genes do not cluster together; soda/sodc cluster most closely to Trichomonas vaginalis, while sodb1/sodb2 cluster with diverse eukaryotes (Dufernez et al. Reference Dufernez, Yernaux, Gerbod, Noël, Chauvenet, Wintjens, Edgcomb, Capron, Opperdoes and Viscogliosi2006). This suggests sorting of ancestral sod lineages but not necessarily HGT. Similarly, two metallocarboxypeptidases (TcMCP-1 and TcMCP-2) in T. cruzi are found only in Kinetoplastids and prokaryotes, but homologues from the two taxa are sister clades, rather than nested (Niemirowicz et al. Reference Niemirowicz, Parussini, Agüero and Cazzulo2007). While the original study recognized the possibility of both HGT and lineage sorting, they rejected the latter due to the number of deletions this would require. These losses may not be necessary, however, if eukaryotic diversity were exhaustively sampled. Finally, an uncharacterized protein, META1, is up-regulated in Leishmania metacyclics and is homologous to a bacterial heat-inducible protein, itself similar to a component of the type III secretion system in Shigella (Puri et al. Reference Puri, Goyal, Sankaranarayanan, Enright and Vaidya2011). META1 is hypothesized to have evolved via HGT and may be involved in secretory processes in Leishmania since mutagenesis of select hydrophobic residues in META1 affects the secretion of the secreted acid phosphatase (Puri et al. Reference Puri, Goyal, Sankaranarayanan, Enright and Vaidya2011). However, META1 is not nested among bacterial sequences and, at this stage, the HGT hypothesis rests on it remaining absent from all other eukaryotes.
Although poor sampling continues to limit our ability to distinguish HGT and lineage sorting (Opperdoes and Michels, Reference Opperdoes and Michels2007), HGT has clearly contributed to trypanosomatid genomes; for example, substantial integration of genes from a bacterial endosymbiont has recently been demonstrated in Angomonas deanei (Alves et al. Reference Alves, Voegtly, Matveyev, Lara, da Silva, Serrano, Buck, Teixeira and Camargo2011). The role of HGT in the origins of parasitism will be clarified through comparison of trypanosomatids with free-living Kinetoplastids and other neglected unicellular eukaryotes, to reject the lineage sorting hypothesis and to confirm that the HGT is uniquely associated with parasites, such as hemF-H or PLA1, and not Kinetoplastids generally.
CONCLUSION
The genetic content of trypanosomatid genomes indicates that they have been elaborated relative to their common ancestor in terms of both physical structure and physiological capacity. Species-specific gene families, instrumental in cell surface architecture, are central to this history of innovation, and implicitly linked to the origins of complex life cycles and disease. By definition, these unique innovations are mutually exclusive, yet there are themes that cut across species. These gene families are functionally differentiated to perform multiple roles in different host environments through the parasite life cycle. They are positioned in sub-telomeres, tandem gene arrays or other contingency zones that perhaps promote regulatory flexibility and sequence diversity. Their sequences are diverse and often contain low complexity repeats that may promote greater diversity through recombination. In their phylogenies, these gene families display rapid turnover – the gain and loss of lineages – that hint at the importance of host-parasite interactions in genomic evolution. These themes, which would, in fact, apply to parasites of all kinds, suggest how each trypanosomatid lineage has used similar molecular mechanisms to meet the demands of transmission and survival. There are issues in comparative analysis we have not addressed, like protein-protein interactions, the regulatory roles of non-coding regions and regulatory proteins, genomic plasticity or indeed the ~50% of trypanosomatid genes that have no known function. There are also some genes, such as the TcMUC family in T. cruzi, procyclin in T. brucei and T. congolense, and the HASP and SHERP families in L. major, that defy any explanation using a comparative approach, and which may have evolved de novo from non-coding regions. Yet, we have learned enough from the structure and content of trypanosomatid genomes to conclude that becoming parasitic was more an innovative and elaborative process, than one of loss and reduction. With the addition of free-living Kinetoplastids to our comparative analyses, the mechanisms by which these enigmatic genomic adaptations for parasitism came about will be revealed.
FINANCIAL SUPPORT
The author is a Wellcome Trust Tenure-Track Research Fellow, funded by the University of Liverpool and the Wellcome Trust [097826/Z/11/A].







