The human body contains diverse groups of commensal microbiota including both aerobes and anaerobes. The largest population of these resides in the gastrointestinal tract which is colonised by more than 400 bacterial species in the adult(Reference Sghir, Gramet and Suau1, Reference Falk, Hooper and Midtvedt2). The commensal bacteria regulate intestinal epithelial development and function and any interruption of these interactions may result in disease conditions(Reference Guarner and Malagelada3, Reference Isolauri, Sutas and Kankaanpaa4). The beneficial effects of the gut microbiota are attributed to probiotics which are defined as ‘Live micro-organisms that when administered in adequate amounts confer a health benefit on the host’(5).
Lactobacillus species are Gram-positive, non-pathogenic and desirable members of the intestinal tract. The health-promoting effects of lactobacilli have been widely explored and include stabilisation of the indigenous microbial population, protection against intestinal infection, alleviation of lactose intolerance, increased nutritional value of foods, reduction of serum cholesterol levels and non-specific enhancement of the immune systems(Reference Kim, Kim and Whang6–Reference Suvarna and Boby9). Several lactobacilli which act as probiotic bacteria are currently being explored as novel biological therapeutic agents(Reference Sullivan and Nord10, Reference Reid, Kim and Kohler11). Since not all lactobacilli possess the ability to confer health benefits to the host, it becomes necessary to screen and characterise numerous strains in order to obtain ideal probiotics. The colonisation of the gastrointestinal tract is desirable for any probiotic which depends on several factors including the ability of the bacteria to tolerate acidic pH of the stomach and bile and on the adhesion of bacteria to intestinal cells and mucus(Reference van Belkum and Nieuwenhuis12). Isolation from human hosts, the capability to tolerate acidic pH and bile, antimicrobial activity and a good adhesion ability are principle desirable properties in potential probiotics(Reference Dunne, O'Mahony and Murphy13). Several probiotic effects are mediated through immune regulation, particularly through establishing and maintaining a balance between pro- and anti-inflammatory cytokines(Reference Isolauri, Sutas and Kankaanpaa4, Reference Gackowska, Michalkiewicz and Krotkiewski14, Reference Winkler, Ghadimi and Schrezenmeir15). This is why the study of immunomodulatory properties of probiotics also has a high priority. The human intestinal epithelial cell line Caco-2 has been extensively used to study the adhesion ability of lactobacilli(Reference Coconnier, Klaenhammer and Kerneis16, Reference Tuomola and Salminen17). This is primarily because Caco-2 cells express morphological and functional differentiation of mature enterocytes including polarisation and functional brush border in vitro (Reference Sambuy, De Angelis and Ranaldi18).
In the present study, the isolation of lactobacilli was carried out from food and human faecal samples with an aim to screen potential probiotic candidates for human use. The ability of the isolates to adhere to Caco-2 monolayers, their capacity to tolerate bile salts and low pH, and their antagonism against various organisms was tested. The antagonistic property was tested against five different pathogenic test cultures. The isolates were studied for their stimulation of Caco-2 cells, THP-1 cells and human peripheral blood mononuclear cells (PBMC) through differences in the level of transcripts of selected cytokines and chemokine genes.
Materials and methods
Isolation and identification of Lactobacillus
Isolation was carried out from seven sources including one curd of buffalo milk, faeces from five breast-feeding human children aged 4–6 months and one human adult aged 27 years. About 1 g of faecal or curd sample was suspended in 10 ml sterile normal saline, vigorously mixed and allowed to settle. A 100 μl sample of the suspension was then spread on three Rogosa SL agar (a selective medium for Lactobacillus isolation; Himedia, Mumbai, India)(Reference Rogosa, Mitchell and Wiseman19) plates containing cycloheximide (100 μg/ml; Sisco Research Laboratories, Mumbai, India) for each sample. The plates were incubated at 37°C until sufficient growth was observed. Between six and eight isolated colonies were then picked from each Rogosa SL agar plate and transferred to de Man, Rogosa and Sharpe (MRS; Himedia) agar plates and subjected to further microscopic and biochemical tests. Molecular identification of isolates was carried out by amplification of 16S–23S rRNA gene intergenic region as reported by Tannock et al. (Reference Tannock, Tilsala-Timisjarvi and Rodtong20). Briefly, 16S–23S rRNA gene intergenic regions of the isolates were amplified using primer 16-1A (5′-GAATCGCTAGTAATCG-3′) and 23-1B (5′-GGGTTCCCCCATTCGGA-3′) by colony PCR(Reference Tannock, Tilsala-Timisjarvi and Rodtong20). Amplification conditions were as follows: initial denaturation at 94°C for 5 min, followed by thirty cycles of denaturation at 94°C for 45 s, annealing for 30 s at 55°C, followed by extension at 72°C for 1 min and a final extension at 72°C for 6 min. The smaller band was excised, eluted and re-amplified using the same set of primers and the resultant amplicon was sequenced. The sequences were compared with the 16S–23S rRNA gene intergenic small spacer region sequences held in GenBank. The sequences of the 16S–23S rRNA gene intergenic region of the isolates are held in GenBank (for accession numbers, see Table 1). Standard strains Lactobacillus rhamnosus GG (LGG) and Lactobacillus plantarum American Type Culture Collection (ATCC) 8014 were obtained as kind gifts from Dr Shira Doron (MD, Department of Medicine, Tufts Medical Center, Boston, MA, USA)(Reference Doron, Snydman and Gorbach21) and Food and Drugs Laboratory (FDL; Vadodara, India), respectively.
Identification of Lactobacillus isolates based on percentage similarity to 16S–23S rRNA small intergenic spacer region sequences with database in GenBank

Cell culture
The human colonic adenocarcinoma cell line Caco-2 and monocyte-like cell line THP-1 were obtained from the National Centre for Cell Science, Pune, India, which were routinely cultured in Dulbecco's modified Eagle's medium and Roswell Park Memorial Institute (RPMI)-1640 (Sigma-Aldrich, St Louis, MO, USA) medium, respectively, at 37°C in a humidified atmosphere containing 5 % CO2–95 % air atmosphere. The media were supplemented with 10 % (v/v) fetal bovine serum (Sigma-Aldrich), 10 mm-non-essential amino acids, 1 mm-sodium pyruvate and gentamicin (50 μg/ml). The media lacked gentamycin whenever antibiotic-free medium was used.
Bile and pH tolerance
The minimal inhibitory concentration of bile was determined for individual lactobacilli strains by inoculating 1 × 106 bacterial cells in MRS broth containing 1–5 % (w/v) bile salts (sodium cholate and sodium deoxycholate; Himedia) for 24 h. Based on this subsequently, bacterial cells grown overnight in normal MRS broth were washed with PBS and 20 μl each of selected lactobacilli (1 × 108 colony-forming units (cfu)/ml) were transferred to 980 μl MRS broth containing 3 % bile salts and incubated at 37°C for 2 h. To examine the survival rate of different lactobacilli under acidic condition at 37°C, 20 μl each of selected lactobacilli (1 × 108 cfu/ml) were transferred to 980 μl of acidic buffer (consisting of 3·5 g d-glucose, 2·05 g NaCl, 0·6 g KH2PO4, 0·11 g CaCl2 and 0·37 g KCl per litre; the pH was adjusted to 2·0 and 2·5 using HCl)(Reference Casey, Casey and Gardiner22). For both bile and acid tolerance, the samples were taken at 0 min and 2 h and cultures were plated after appropriate dilution on MRS agar plate; enumeration was done following 48 h incubation at 37°C.
Adhesion assay
For the adhesion assay, Caco-2 cells were seeded at a density of 104 cells per well in twenty-four-well standard tissue culture plates (Corning Inc., Corning, NY, USA) and maintained for 2 weeks following confluence. Before the adhesion assay, Caco-2 monolayers were pre-incubated with antibiotic-free medium for 4 h. The pH of the media used for the adhesion assay was adjusted to 6·5 with 1 m-HCl before use. Lactobacilli cells were harvested by centrifugation (10 000 g for 2 min at 4°C), washed twice with Dulbecco's PBS, pH 7·0 (Sigma-Aldrich) and cell density was adjusted to the desired level by measuring absorbance at 600 nm. The exact number of viable lactobacilli used in the assays was determined for each experiment by plate counting on MRS agar. Wells with Caco-2 monolayers were inoculated with 1 × 108 viable cells of each bacterial cell suspension and incubated at 37°C for 90 min in 5 % CO2–95 % air atmosphere. Un-adhered bacterial cells were then withdrawn from the wells and the Caco-2 monolayers were washed twice with 1 ml PBS each. The Caco-2 cells were lysed by treatment with 0·5 ml 0·05 % (v/v) Triton X-100 in PBS for 20 min at 37°C. The Caco-2 lysate including bound lactobacilli was plated after appropriate dilution on MRS agar plate; enumeration was done following 48 h incubation at 37°C. It was also determined that a 30 min treatment of 0·05 % (v/v) Triton X-100 in PBS at 37°C did not affect the viability of lactobacilli (data not shown). At the end of each experiment, three randomly preselected unused wells were trypsinised and numbers of Caco-2 cells were counted on a haemocytometer. The average value of the Caco-2 cell count was used for expressing the adhered bacteria per Caco-2 cell.
Antimicrobial activity
The antimicrobial activity of lactobacilli was determined by the agar spot test as described by Schillinger & Lücke(Reference Schillinger and Lücke23) with minor modifications. Briefly, 5 μl each of selected Lactobacillus strains (1 × 108 cfu/ml) was spotted on the surface of an MRS agar plate and incubated for 24 h at 37°C for the spots to develop. A 150 μl sample of each of the indicator bacteria (listed in Table 2) grown overnight in Luria broth under shaking condition at 37°C was vigorously mixed with 15 ml of Luria soft agar (0·6 % agar, w/v) and poured over the MRS agar plates containing developed colonies of lactobacilli. The plates were then incubated for 24 h at 37°C and the zones of inhibition were measured and expressed as described by Baccigalupi et al. (Reference Baccigalupi, Di Donato and Parlato24). The test strains used were Shigella dysentery, Staphylococcus aureus (ATCC 6538), Pseudomonas aeruginosa (ATCC 25 668), Salmonella typhi and Escherichia coli S5, obtained from the culture collection facility at our department.
(Mean values and standard deviations and percentage survival rates)

cfu, Colony-forming units; LGG, Lactobacillus rhamnosus GG; ATCC, American Type Culture Collection; ATCC 8014, L. plantarum ATCC 8014; M, L. delbrueckii M; ASt1, L. fermentum ASt1; CS23, L. plantarum CS23; CS25, L. rhamnosus CS25.
* Mean value of isolates was significantly higher than that of L. plantarum ATCC 8014 (P < 0·05).
† Mean value of isolates was significantly higher than that of L. rhamnosus GG (P < 0·05).
‡ Results were obtained from three independent experiments. The strains were compared with two different controls (LGG and ATCC 8014) by means of two independent ANOVA tests. Significant ANOVA were followed by Dunnett's test for multiple comparisons v. the control group.
Stimulation of Caco-2 monolayers with lactobacilli
To obtain monolayers, 5 × 104 Caco-2 cells were seeded in a T25 tissue culture flask (Corning Incorporated) and maintained for 2 weeks post-confluence under the same conditions as described above. The Caco-2 monolayers were stimulated with 1 × 108 cfu/ml(Reference Morita, He and Fuse25) of different lactobacilli and co-incubated for 90 min in the absence of gentamycin at 37°C in a CO2 incubator. Thereafter, gentamycin (50 μg/ml) was added to prevent bacterial growth and further incubated under the same conditions for another 4 h and 30 min. At the end of incubation, the culture supernatant fraction was discarded and Caco-2 monolayers were lysed in the presence of guanidine thiocyanate which is included in the total RNA extraction kit (Bangalore Genei, Bangalore, India) that was employed. Further steps were performed as per the manufacturer's instructions.
Isolation of human peripheral blood mononuclear cells
PBMC were separated as described previously(Reference Bam and Bagchi26). Briefly, 20 ml of peripheral blood was collected in EDTA-coated vials (Himedia) from two healthy donors. The present study was conducted according to the guidelines laid down in the Declaration of Helsinki and all procedures involving human subjects were approved by the Institutional Ethics Committee for Human Research of the Faculty of Science, M. S. University of Baroda. Written informed consent was obtained from all subjects. The samples were layered over an equal volume of Histopaque-1077 (Sigma-Aldrich) and centrifuged for 20 min at 400 g at room temperature. The mononuclear cell layer was carefully pipetted out from each donor, mixed and washed twice with 10 ml PBS each. Pellets were then re-suspended in 7 ml RPMI-1640 cell culture medium.
Stimulation of peripheral blood mononuclear cells and THP-1 cells with lactobacilli
The cell number of PBMC and THP-1 cells for stimulation experiments was set to 1 × 106 cells/ml(Reference Miettinen, Vuopio-Varkila and Varkila27) in RPMI-1640 and 1 ml each of cell suspension was transferred to a twenty-four-well plate. To this, 50 μl of different lactobacilli suspensions containing 1 × 106 bacteria were added and co-incubated for 90 min in the absence of gentamycin at 37°C in a CO2 incubator. Thereafter, gentamycin (50 μg/ml) was added to prevent bacterial growth and further incubated under the same conditions for another 4 h and 30 min. Cells from the suspension culture and adhered cells were harvested, pooled and subjected to total RNA extraction using the kit as mentioned above.
RNA isolation and RT-PCR
Total RNA was isolated from control (to which no bacteria were added) Caco-2 cells, PBMC and THP-1 cells and those co-incubated with various lactobacilli and using a total RNA extraction kit (Bangalore Genei). The quality of the RNA samples was assessed by inspecting the 28S and 18S bands following agarose gel electrophoresis. A quantity of 2 μg of each RNA sample was used with Oligo (dT18) for cDNA synthesis in a 20 μl system using an M-MuLV RT-PCR kit (Bangalore Genei) following the manufacturer's instructions. Briefly, the RNA and Oligo (dT18) mixture was incubated at 65°C for 10 min, centrifuged briefly and kept at room temperature for 2 min. Into this RNAsin, dithiothreitol, RT buffer, deoxynucleotide triphosphate (dNTP) and M-MuLV RT were added, as instructed by the manufacturer and incubated further at 37°C for 1 h, followed by 5 min incubation at 95°C. Each of the cDNA preparations was then amplified for thirty cycles in a thermal cycler (Eppendorf, Hamburg, Germany) with β-actin-specific primers (forward, 5′-AGCGGGAAATCGTGCGTGACA-3′; reverse, 5′-CGCAACTAAGTCATAGTCCG-3′, generating an amplicon of 536 bp) by taking 1 μl of the cDNA in a 12·5 μl system. This was used as a control for the synthesis of cDNA. Controls for checking genomic DNA contamination included amplification of the total RNA without reverse transcription which did not give any amplicon (results not shown). PCR amplifications were then performed with specific primers for cytokines in a 12·5 μl system with 1 μl of the first-strand cDNA, 0·2 mm-deoxynucleotide triphosphates (dNTPs), 1·5 mm-MgCl2 and 1·25 pmol each of the forward and reverse primers. This was run for thirty cycles after the addition 0·5 U of Taq polymerase. Amplification conditions were as follows: initial denaturation at 94°C for 5 min, followed by thirty cycles of denaturation at 94°C for 45 s, annealing for 30 s at 66°C for β-actin, 60°C for IL-6, IL-8, IL-12p35 and transforming growth factor-β, 62°C for IL-12p40 and TNF-α and 56°C for IL-15 followed by extension at 72°C for 1 min and a final extension at 72°C for 6 min. PCR products separated on a 2 % agarose gel were stained with ethidium bromide (0·5 μg/ml), following which densitometric analysis was carried out using AlphaEaseFC 4.0 software (Alpha Innotech, San Leandro, CA, USA) and the results are expressed as integrated density value divided by the selected area of band on the gel.
Statistics
Values are given as mean values and standard deviations of triplicate independent experiments. Significant ANOVA were followed by the Dunnett test in the case of the adhesion assay and immunomodulation by different lactobacilli for Caco-2 cells, human PBMC and THP-1 cells with respect to the respective controls (P < 0·05). All analysis was conducted using SigmaStat 3.5 software (Cranes Software International, Bangalore, India).
Results and discussion
Isolation and identification of Lactobacillus
Out of 143 isolates screened, twenty-seven isolates which were Gram-positive rods, found negative for the presence of endospore and for the production of catalase, and which could grow on Rogosa SL and MRS agar plates were selected for subsequent molecular identification on the basis of 16S–23S rRNA gene intergenic region amplification. The agarose gel mobility of the amplification products generated from the 16S–23S rRNA gene intergenic region of eighteen isolates matched with that of standard strains LGG and ATCC 8014. The smaller intergenic region of ten of these eighteen isolates was sequenced. In addition, the smaller intergenic region of three of nine others whose amplified product size was distinct from that of standard strains was sequenced and following sequence alignment with the National Center for Biotechnology Information (NCBI) database only the former ten were found to belong to the Lactobacillus genus. Of these, L. plantarum CS23, L. rhamnosus CS25, L. fermentum ASt1 and L. delbrueckii M were used in the present study. The 16S–23S rRNA gene intergenic region sequences of these strains are held in GenBank and their corresponding accession numbers are shown in Table 1. A potential probiotic organism is preferred to be of human origin(Reference Tannock28, Reference Teitelbaum and Walker29) so that it can lead us to a candidate probiotic eventually targeted for human consumption. Commercially available probiotic lactobacilli such as LGG, L. plantarum 299v and L. gasseri LA39 have all been of human origin, as they have been isolated from human faeces(Reference Doron, Snydman and Gorbach21, Reference Goossens, Jonkers and Russel30, Reference Kawai, Ishii and Uemura31). The isolates have been assayed in vitro for significant probiotic properties such as the ability to adhere to Caco-2 monolayers, their tolerance to bile and an acidic environment, the degree of antagonism they exhibit against selected pathogenic organisms and the ability to modulate the immune response of human PBMC, Caco-2 and THP-1 cells.
Bile and acid tolerance
From different concentration of bile salts taken, 3 % bile salts in MRS broth was the minimal inhibitory concentration of bile for most Lactobacillus strains (data not shown); hence the survival of lactobacilli post 2 h incubation in MRS containing 3 % bile salts was analysed and the results were subjected to statistical analysis (P < 0·05). As given in Table 2, isolate CS23 showed an excellent survival of 14·04 % in contrast to standard strains LGG and ATCC 8014, which showed survival of 5·34 and 3·88 %, respectively. Survival rates of lactobacilli in acidic buffer (pH 2·5 and 2·0) were examined by the difference in viable cell counts following 0 min and 2 h incubation, as shown in Table 3. The isolate CS25 (6·90 % at pH 2·5, 5·12 % at pH 2·0) showed the highest survival in acidic pH, immediately followed by CS23, M and ASt1 (5·70, 5·40 and 4·03 % at pH 2·5, 4·90, 4·31 and 2·33 % at pH 2·0, respectively). Before the lactobacilli reach the hindgut region where they are generally known to colonise in human hosts, the organism needs to survive through acidic pH and bile. So the tolerance of an organism to these factors is equally important in selecting a potential probiotic. In studies by other authors, the survival of Lactobacillus isolates in pH 2·5 yielded similar results; however, the survival has not been more than 0·2 %(Reference Kim, Jung and Chang32, Reference Perelmuter, Fraga and Zunino33).
Spectrum of antimicrobial activity exhibited by various lactobacilli*
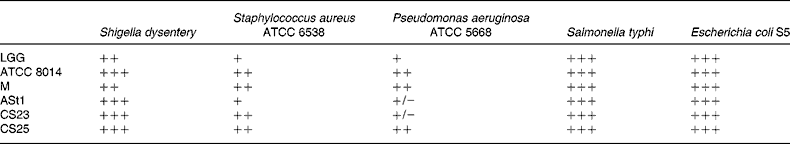
ATCC, American Type Culture Collection; LGG, Lactobacillus rhamnosus GG; ATCC 8014, L. plantarum ATCC 8014; M, L. delbrueckii M; ASt1, L. fermentum ASt1; CS23, L. plantarum CS23; CS25, L. rhamnosus CS25.
* The inhibition zones 1 mm, 2 mm, 2–5 mm and more than 5 mm were classified as strains of no (+/ − ), mild (+), strong (++) and very strong (+++) inhibition, respectively.
Adhesion assay
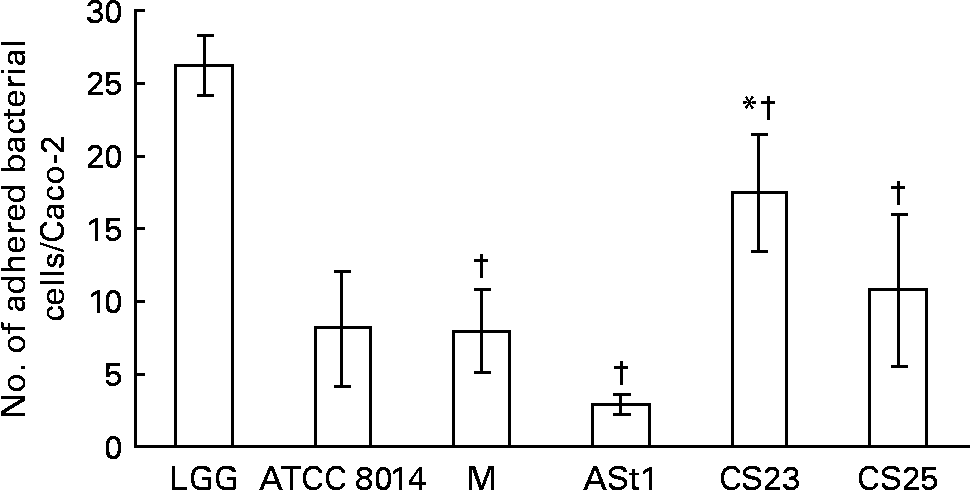
In the present study none of the isolates showed better binding to Caco-2 than LGG (Fig. 1). One of our isolates, CS23, showed statistically significant better and another two, L. rhamnosus CS25 and L. delbrueckii M, showed binding ability as good as L. plantarum ATCC 8014. The adult stool isolate ASt1 showed the poorest binding capacity. The same results were obtained when considering the percentage of the number of bacteria bound to the number of bacteria added per well (data not shown). Caco-2 cells have been widely accepted as an in vitro model for assessing the adhesion of potential probiotics, correlating with adherence to the intestinal epithelium in vivo. This is principally because Caco-2 cells express morphological and functional differentiation in vitro and express several markers that are distinctive of normal small-intestinal villi(Reference Coconnier, Klaenhammer and Kerneis16). The ability of a probiotic to adhere to intestinal epithelium enables the organism to colonise the gut as it can resist against being washed out due to peristaltic and other movements inside the gut. Buck et al. (Reference Buck, Altermann and Svingerud34) reported a significant decrease in the adhesion ability of different isogenic surface protein mutants of L. acidophilus NCFM, thereby suggesting multiple cell-surface proteins imparting the organisms with the ability of binding to intestinal epithelial cells in vitro. Since the percentage of adhesion of a probiotic under a given set of conditions does not represent an absolute value, the adhesion should be represented as a value relative to a type strain(Reference Buck, Altermann and Svingerud34, Reference Blum, Reniero and Schiffrin35).
Adhesion of Lactobacillus isolates to Caco-2 epithelial cell line compared with standard strains Lactobacillus rhamnosus GG (LGG) and L. plantarum American Type Culture Collection (ATCC) 8014 (ATCC 8014). M, L. delbrueckii M; ASt1, L. fermentum ASt1; CS23, L. plantarum CS23; CS25, L. rhamnosus CS25. Values are means of three independent experiments, with standard deviations represented by vertical bars. The strains were compared with two different controls (LGG and ATCC 8014) by means of two independent ANOVA tests. Significant ANOVA were followed by Dunnett's test for multiple comparisons v. the control group. * Mean value of isolates was significantly different than that of ATCC 8014 (P < 0·05). † Mean value of isolates was significantly different than that of LGG (P < 0·05).
Antimicrobial test
As shown in Table 3, the antagonistic activity of the lactobacilli was examined against Gram-negative Shigella dysentery, P. aeruginosa, E. coli S5 and Salmonella typhi, as well as Gram-positive Staphylococcus aureus.
The isolate CS25 and standard strain ATCC 8014 had the highest inhibitory activity against both Gram-positive and Gram-negative bacteria, followed by M and CS23. All the strains tested showed a strong inhibition towards E. coli S5. Baccigalupi et al. had observed a range of antimicrobial responses by lactobacilli to various indicator bacteria and also indicated a strong possibility that these activities were not caused by an acidic environment(Reference Baccigalupi, Di Donato and Parlato24). In the present study, though CS23 and ATCC 8014 and similarly LGG and CS25 are from the same species, respectively, the difference in their antimicrobial activity suggests that antimicrobial attributes must be strain specific. Once colonised, the probiotics get an opportunity to exhibit their antimicrobial and immunomodulatory properties in favour of the host organism. The antimicrobial activity of Lactobacillus is generally due to production of lactic acid, H2O2 and/or other antibacterial molecules such as bacteriocin(Reference Kucerova, Chumchalova and Mikova36, Reference Eschenbach, Davick and Williams37). It should be interesting to study the mode of antimicrobial activity in different strains.
Stimulation of Caco-2, human peripheral blood mononuclear cells and THP-1 co-cultured with lactobacilli
The interaction of the enterocytes with micro-organisms is known to modulate the cytokine profile of the former. Certain cytokines in turn serve as chemo-attractants and activators of other immunocompetent cells. The stimuli eventually have a potential to bring the immunocompetent cells from the gut-associated lymphoid population in direct contact with the intestinal microbiota, at least at the sites of injury. The bacteria, on the other hand, do have the potential to cross the epithelial barrier and thereafter come across gut-associated immune cells(Reference Perdigon, Medina and Vintini38). So, it becomes important to study the interaction of probiotics with the enterocytes as well as the other immunocompetent cells of the intestinal epithelium(Reference Tannock28). In the present study Caco-2 cells, THP-1 cells and the PBMC isolated from healthy human subjects have been used as in vitro models for studying the immunomodulatory capacity of different lactobacilli.
The presence of transcripts of β-actin (control) and a selected number of cytokine genes in Caco-2 cells, human PBMC and THP-1 cells co-cultured with different lactobacilli was analysed employing semi-quantitative RT-PCR. They were compared with the control where none of the bacterial culture was added (Fig. 2). TNF-α and IL-6 are pro-inflammatory cytokines, which are produced by the host in response to bacterial colonisation or invasion and hence are central to the host defence mechanism against pathogens(Reference Morita, He and Fuse25, Reference Solis-Pereyra, Aattouri and Lemonnier39). Though lipopolysaccharide of Gram-negative bacteria is known to stimulate their production, Miettinen et al. have reported an increase in IL-6 and TNF-α production in human PBMC exposed to lactobacilli and thereby suggested the use of probiotics as vaccine vectors and for the purpose of stimulating non-specific immunity(Reference Miettinen, Vuopio-Varkila and Varkila27). In the present study, all the isolates have exhibited their ability to induce in Caco-2, and PBMC, a higher level of TNF-α transcript. In the case of THP-1, however, only ATCC 8014 and isolate CS25 had the capability to up-regulate TNF-α expression. Similarly, in the case of IL-6 and in Caco-2 cells, isolates CS23 and CS25 had a strong up-regulatory potential compared with the down-regulation that is induced by ATCC 8014 and ASt1. Interestingly, there was no effect of these organisms on PBMC and THP-1 cells. It should be noted here that IL-6 is known to have both pro- as well as anti-inflammatory properties. It has a very important role in resolving the initial inflammatory reaction which leads to the activation of the adaptive immune response. Therefore, it appears that while CS23 and CS25 have the capacity to induce a strong pro-inflammatory environment, the presence of IL-6 may help them to convert an innate response to a more specific and sustained adaptive response against pathogens.
Cytokines and chemokine transcript levels in (a) Caco-2 cells co-incubated with different lactobacilli (1 × 108 colony-forming units (cfu)/ml), (b) peripheral blood mononuclear cells (PBMC) and (c) THP-1 cells co-incubated with different lactobacilli (1 × 106 cfu/ml). Different lactobacilli used in the study were: Lactobacillus plantarum American Type Culture Collection (ATCC) 8014 (![]() ); L. delbrueckii M (
); L. delbrueckii M (![]() ); L. fermentum ASt1 (
); L. fermentum ASt1 (![]() ); L. plantarum CS23 (
); L. plantarum CS23 (![]() ); L. rhamnosus CS25 (
); L. rhamnosus CS25 (![]() ); control, to which no bacteria were added (□). TGF, transforming growth factor. The integrated density values (IDV) of each cytokine or chemokine were used to express its transcript levels. Values are means, with standard deviations represented by vertical bars. Significant ANOVA were followed by Dunnett's test for multiple comparisons v. the control group. † Within each cytokine or chemokine, mean value was significantly lower than that of the control (P < 0·05). ‡ Within each cytokine or chemokine, mean value was significantly higher than that of the control (P < 0·05). No amplification products were observed for IL-12p40 in Caco-2 cells, IL-6 and IL-12p35 in PBMC and IL-6 and IL-12p40 in THP-1 cells.
); control, to which no bacteria were added (□). TGF, transforming growth factor. The integrated density values (IDV) of each cytokine or chemokine were used to express its transcript levels. Values are means, with standard deviations represented by vertical bars. Significant ANOVA were followed by Dunnett's test for multiple comparisons v. the control group. † Within each cytokine or chemokine, mean value was significantly lower than that of the control (P < 0·05). ‡ Within each cytokine or chemokine, mean value was significantly higher than that of the control (P < 0·05). No amplification products were observed for IL-12p40 in Caco-2 cells, IL-6 and IL-12p35 in PBMC and IL-6 and IL-12p40 in THP-1 cells.

As is evident from the data, this possibility is strengthened all the more by the induction ability of both CS23 and CS25 towards the production by Caco-2 of IL-8, a chemokine that is responsible for recruiting elements of the innate immune system. Of all the organisms tested for effect on IL-8 transcript levels, isolate CS25 has a stronger up-regulatory potential in Caco-2 cells and PBMC, while ASt1 and M showed up-regulation in Caco-2 and THP-1, respectively. The other combinations of organism and cell lines tested are either neutral, or down-regulatory as in ATCC 8014 and isolate M for Caco-2, and isolates CS23 and CS25 for THP-1. IL-8 has an important role in the induction of neutrophil accumulation and activation(Reference Godaly, Proudfoot and Offord40). Zhang et al. have suggested that a higher dose of LGG (107 cfu/ml) induces an increased production of IL-8 in Caco-2(Reference Zhang, Li and Caicedo41), which is in agreement with the present results wherein an increased transcription of IL-8 has been seen in Caco-2 stimulated by ASt1, CS25 and CS23, and in PBMC stimulated by CS25, and THP-1 stimulated by M. The number of bacterial cells taken in the present study is 108 cfu/ml with Caco-2 and 106 cfu/ml in studies with PBMC and THP-1. In spite of the high bacterial dose, a significant suppression of IL-8 transcription has been observed in Caco-2 co-incubated with M and ATCC 8014 and in THP-1 co-incubated with CS25 and CS23. A high expression level of IL-8 is associated with the pathogenesis of several diseases including neonatal necrotising enterocolitis and so the immunosuppressive effects of the above isolates in context of IL-8 transcription seem promising. However, further studies need to be conducted at the level of protein expression and in vivo.
IL-12 is a heterodimeric protein formed by a complex of p40 with p35, having an important role in interferon-γ production by T helper cells(Reference Trinchieri42). The cytokine is thus non-functional in the absence of either subunit. All isolates and standard strains demonstrated suppression of IL-12p35 in Caco-2 and IL-12p40 was suppressed in PBMC by M, ASt1, CS25 and ATCC 8014. Isolate CS23 in PBMC and ATCC 8014 in THP-1 cells induced higher transcription of IL-12p40 and IL-12p35, respectively. IL-12 is a pro-inflammatory cytokine and plays an important role in the pathogenesis of inflammatory diseases such as Crohn's disease. In recent clinical trials the use of antibodies against IL-12 suggests a role for this cytokine in inflammatory bowel diseases(Reference Mannon, Fuss and Mayer43) and hence the immunosuppressive property of the isolates in context of IL-12 can be appropriately utilised under similar circumstances.
IL-15 is a pleiotropic cytokine and regulates T cell and natural killer cell activation and proliferation(Reference Ma, Boone and Lodolce44). It also has a role in maintaining immune homoeostasis in the mucosal environment by activating intraepithelial cells. It therefore appears from our data that ASt1 and CS25 are capable of functioning better at the mucosal interface. Interestingly, both of them exhibit either a neutral or down-regulatory potential when exposed to monocytes. Isolates ASt1 and CS25 in Caco-2, all strains (except CS25) in PBMC and ATCC 8014 in THP-1 have been observed to induce IL-15 at the transcripts level. IL-15 plays a critical role in the maintenance of memory lymphocytes by supporting proliferation. Another cytokine that has also been looked at, transforming growth factor-β, is an anti-inflammatory immunoregulatory cytokine which regulates the production of IgA antibodies and is important in early defence against intestinal infection(Reference Sonoda, Matsumoto and Hitoshi45). In the present study, none of our isolates modulated significant mRNA expression of transforming growth factor-β, which is in support of earlier observations(Reference Borruel, Casellas and Antolin46).
Gabriella et al. have hypothesised that the intestinal epithelial cells which sense the bacterial surface with pro-inflammatory components such as EF-Tu (elongation factor thermo unstable) and GroEL and anti-inflammatory components, such as lipoteichoic acid, process the information and respond with a concurrent pro- or anti-inflammatory response(Reference Bergonzelli, Granato and Pridmore47). In the present study ASt1, CS23 and CS25 have shown strong and varied immunomodulatory characteristics that are reflected from the differences in the transcripts of different cytokines in Caco-2 cells, PBMC and THP-1 cells co-incubated with these strains as compared with the respective controls. Another point that comes across strongly from the present elaborate study is that each of these isolates is bestowed with its own combination of pro- and anti-inflammatory potential. This gives rise to the possibility that each one of them may be useful under a specific set of immunological environments in the gut.
Though LGG has shown a very good binding capability in the present study as well as in several other studies(Reference Matijasic, Narat and Zoric48, Reference Roselli, Finamore and Britti49), our isolates have been able to score more in relation to other properties such as antimicrobial properties (CS25 and M), and bile and acid tolerance (CS23 and CS25). Jacobsen et al. (Reference Jacobsen, Rosenfeldt Nielsen and Hayford50) had correlated the results of their in vitro assays for binding ability and acid tolerance with the in vivo performance of the strains and concluded that the survival of strains was strongly linked with adhesion ability to Caco-2 and tolerance to acidic pH. The human child faecal isolate CS23 showed very good binding ability, high acid and bile tolerance and strong and varied immunomodulation, and is a promising contender for a potential probiotic strain.
Acknowledgements
The authors thank the Department of Biotechnology, New Delhi, India for financial support (grant no. BT/PR-7496/PID/20/292/2006).
T. B. conceived of, conceptualised and supervised the present study. S. B. G. and A. S. D. performed the isolation, identification and analysis of probiotic properties. T. B. performed the cell culture. S. B. G. wrote the manuscript with A. S. D. and T. B. providing substantial contributions. All authors read and approved the findings of the study.
There is no conflict of interest to declare by any of the authors.






