Introduction
Haemosporidians (Apicomplexa, Haemosporida) comprise a diverse and widely distributed parasite group transmitted by hematophagous dipterans (e.g. mosquitoes, sandflies, black flies, tabanids, and others) that use reptiles, birds and mammals as vertebrate hosts (Garnham, Reference Garnham1966; Valkiūnas, Reference Valkiūnas2005; Telford, Reference Telford2009). Some of these parasites have medical importance, as they involve the etiological agents of human malaria – species of the genus Plasmodium Marchiafava and Celli, 1885 – responsible for significant negative impacts on public health and the global economy (World Health Organization – WHO, 2023). Therefore, these pathogens are among the best-known and studied in the world (Pacheco and Escalante, Reference Pacheco and Escalante2023). However, many aspects of the ecological-evolutionary history, taxonomy, and systematics of haemosporidians are poorly understood, especially for species that infect wildlife (Pacheco et al., Reference Pacheco, Matta, Valkiūnas, Parker, Mello, Stanley, Lentino, Garcia-Amado, Cranfield, Kosakovsky Pond and Escalante2018a, Pacheco et al., Reference Pacheco, Ceríaco, Matta, Vargas-Ramírez, Bauer and Escalante2020).
There are around 500 haemosporidian species described, classified into 11 genera and four families: Garniidae (Fallisia Lainson et al., 1974, Garnia Lainson et al., 1971, and Progarnia Lainson, 1995), Haemoproteiidae (Haemocystidium Castellani and Willey, 1904, Haemoproteus Kruse, 1890, Hepatocystis Levaditi and Schoen, 1932, Nycteria Garnham and Heisch, 1953, and Polychromophilus Dionisi, 1899), Leucocytozoidae (Leucocytozoon Berestneff, 1904 and Saurocytozoon Lainson and Shaw, Reference Lainson and Shaw1969), and Plasmodiidae (Plasmodium) (Garnham, Reference Garnham1966; Pacheco and Escalante, Reference Pacheco and Escalante2023). Classical methods of taxonomy were used to erect these taxa by combining characters visible under a light microscope, such as the presence or absence of malarial pigment (hemozoin) and the occurrence of erythrocytic merogony (Garnham, Reference Garnham1966; Lainson et al., Reference Lainson, Landau and Shaw1971). Likewise, most of the genera and species were delimited based on the morphology of the blood stages (trophozoites, gametocytes, and meronts) and by association with the vertebrate host species, vectors, and geographic region (Pacheco and Escalante, Reference Pacheco and Escalante2023).
Current phylogenetic reconstructions based on multilocus sequencing and/or nuclear, mitochondria, and apicoplast genomes show support for the validity of the Haemoproteidae, Leucocytozoidae and Plasmodiidae, despite some of their genera forming polyphyletic groups (Escalante et al., Reference Escalante, Freeland, Collins and Lal1998; Borner et al., Reference Borner, Pick, Thiede, Kolawole, Kingsley, Schulze, Cottontail, Wellinghausen, Schmidt-Chanasit, Bruchhaus and Burmester2016; Galen et al., Reference Galen, Borner, Martinsen, Schaer, Austin, West and Perkins2018; Pacheco et al., Reference Pacheco, Matta, Valkiūnas, Parker, Mello, Stanley, Lentino, Garcia-Amado, Cranfield, Kosakovsky Pond and Escalante2018a, Reference Pacheco, Ceríaco, Matta, Vargas-Ramírez, Bauer and Escalante2020). Nevertheless, these reconstructions are primarily derived from haemosporidian DNA sequences from infections in birds and mammals (Galen et al., Reference Galen, Borner, Martinsen, Schaer, Austin, West and Perkins2018; Pacheco et al., Reference Pacheco, Matta, Valkiūnas, Parker, Mello, Stanley, Lentino, Garcia-Amado, Cranfield, Kosakovsky Pond and Escalante2018a; Pacheco and Escalante, Reference Pacheco and Escalante2023). For the Garniidae, a family of unpigmented haemosporidians that mainly parasitize reptiles, the limited molecular data available indicate that some species may be part of the Plasmodiidae, suggesting the need for a comprehensive investigation into the taxonomy of these parasites (Perkins, Reference Perkins2000; Córdoba et al., Reference Córdoba, Ferreira, Pacheco, Escalante and Braga2021; Matta et al., Reference Matta, González, Vargas-Ramírez, Valkiūnas, Escalante and Pacheco2023).
Despite comprising a third of known haemosporidian species, reptile haemosporidians (n = 160/500) remain relatively understudied (Telford, Reference Telford2009; Pacheco and Escalante, Reference Pacheco and Escalante2023). Among them, only 29 species, primarily from the genera Plasmodium and Haemocystidium have associated Cytochrome b (cytb), short nuclear, and apicoplast gene sequences, and only nine have published complete mitochondrial genomes (mtDNA) (Table 1). These sequences originate from lizards and chelonians sampled in the Americas, Africa, Europe, and Oceania (Table 1), reflecting the broad distribution of these haemosporidians. Lizards, particularly those inhabiting Neotropical humid forests, account for nearly 90% (n = 144/160) of reptile haemosporidian species richness (Telford, Reference Telford2009; Lainson, Reference Lainson2012; Picelli et al., Reference Picelli, Ramires, Masseli, Pessoa, Viana and Kaefer2020). However, molecular characterization remains limited for many Neotropical lizard haemosporidians (Matta et al., Reference Matta, González, Pacheco, Escalante, Moreno, González and Calderón-Espinosa2018, Reference Matta, González, Vargas-Ramírez, Valkiūnas, Escalante and Pacheco2023; Harris et al., Reference Harris, Santos, Borges-Nojosa and Castro2019; Ferreira et al., Reference Ferreira, Alves, Jager, Franzini, Mesquita, Díaz-Delgado, Catão-Dias and Braga2020; Córdoba et al., Reference Córdoba, Ferreira, Pacheco, Escalante and Braga2021).
Table 1. Haemosporidian species described in reptiles with partial cytb gene or nearly complete mitochondrial genome sequences available. GenBank accession numbers, associated hosts, and references are provided

* Species with nearly complete mitochondrial genome.
** Sequence with ‘n’ and/or IUPAC nucleotide code.
This lack of genetic information creates difficulties in our understanding of the validity of some controversial haemosporidian taxa, like the genera Fallisia, Garnia and Saurocytozoon, that infect these Neotropical hosts (Matta et al., Reference Matta, González, Vargas-Ramírez, Valkiūnas, Escalante and Pacheco2023; Pacheco and Escalante, Reference Pacheco and Escalante2023). The genus Saurocytozoon was established by Lainson and Shaw (Reference Lainson and Shaw1969) for a leucocytozoid-like parasite infecting the white blood cells of tegu lizards. It was placed within the family Leucocytozoidae due to similarities of its blood stages (gametocytes) with leucocytozoids infecting birds (Lainson and Shaw, Reference Lainson and Shaw1969). Since then, only a few Saurocytozoon species have been described. Many authors have debated the validity of this genus, but the taxonomic discussions were based on morphological traits and their value to define and delimit this genus (Hsu et al., Reference Hsu, Campbell and Levine1973; Telford, Reference Telford1973, Reference Telford1983, Reference Telford2013; Lainson et al., Reference Lainson, Landau and Shaw1974a; Ayala, Reference Ayala and Kreier1977).
Tegus of the genus Tupinambis Daudin, 1802 are the largest Neotropical lizards, widely distributed and occurring in diverse habitats across South America, including primary and secondary forests, savannas, and anthropic areas (Ribeiro-Junior and Amaral, Reference Ribeiro-Junior and Amaral2016). These diurnal terrestrial predators are known for their resilience and play critical ecological roles in their ecosystems (Murphy et al., Reference Murphy, Jowers, Lehtinen, Charles, Colli, Peres, Hendry and Pyron2016). They have been heavily exploited, mainly due to the commercialization of their skin (Fitzgerald, Reference Fitzgerald1994; Fitzgerald et al., Reference Fitzgerald, Cook and Aquino1999), meat consumption (Alves et al., Reference Alves, Vieira, Santana, Vieira, Almeida, Souto, Montenegro and Pezzuti2012), and as a medicinal and healing resource (Valencia-Parra and de la Ossa, Reference Valencia-Parra and de la Ossa2016). The most iconic species of this group, the golden tegu Tupinambis teguixin (Linnaeus, 1758), is exceptional among the Neotropical lizards for its diverse haemoparasites (Telford, Reference Telford2009; Picelli et al., Reference Picelli, Ramires, Masseli, Pessoa, Viana and Kaefer2020), including two haemosporidian species, Plasmodium minasense tegui Carini and Rudolph, 1912 and Saurocytozoon tupinambi Lainson and Shaw, 1969 (Lainson and Shaw, Reference Lainson and Shaw1969; Telford, Reference Telford1979).
After extensive fieldwork in the Amazonia region of Brazil and the Orinoquia region of Colombia, we rediscovered a haemosporidian species parasitizing the white blood cells of golden tegus. Using an integrative approach, we reevaluated the taxonomy of this non-erythrocytic parasite and placed it as Plasmodium (Saurocytozoon) tupinambi comb. nov.
Materials and methods
Study area and specimen sampling
A total of 39 T. teguixin were captured, 26 in Brazil between 2016 and 2018 and 13 in Colombia in 2023 (Figure 1; Table 2, and Supplementary Table S1). In Brazil, lizards were sampled using pitfalls with drift fences and live traps baited with boiled eggs in primary and secondary upland (‘terra-firme’) forest sites located in the State of Amazonas (see Picelli et al., Reference Picelli, Ramires, Masseli, Pessoa, Viana and Kaefer2020 for details). Most specimens (n = 21) collected were returned to the sampling sites, while others (n = 5) were euthanized (via 2% lidocaine injection), preserved in 10% formalin, and deposited as vouchers in the Zoological Collections of Universidade Federal do Amazonas and Instituto Nacional de Pesquisas da Amazonia (INPA). Lizards from Colombia were captured from different localities in the Casanare department, within the Colombian Eastern plains, by live traps baited with fruit and boiled eggs, and all were released after blood collection.

Figure 1. Map of distribution records of Plasmodium (Saurocytozoon) tupinambi comb. nov. in Northern South America. Details of the sampling sites for this study are available in Table 2 and Supplementary Table S1. Map was generated using World Physical Map (US National Park). Image credit: Paulo Vedovello.
Table 2. Sampling locations and haemosporidian infections detected by light microscopy in golden tegus Tupinanambis teguixin from Brazil and Colombia (2016-2023). Mean parasitemia (%) ± SD is provided followed by (minimum and maximum values) for non-erythrocytic and erythrocytic forms
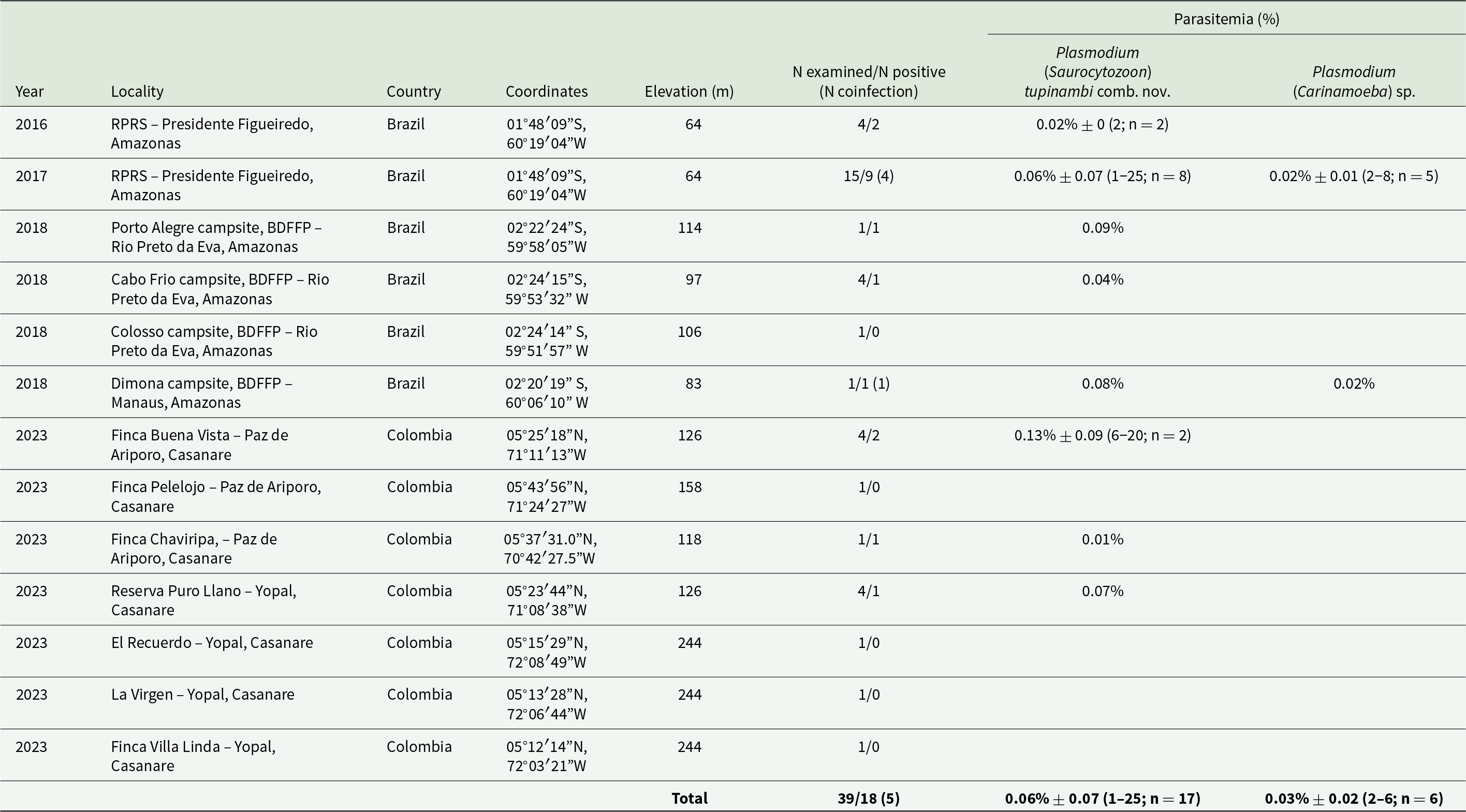
RPRS – Rio Pardo Rural Settlement; BDFFP – Biological Dynamics of Forest Fragments Project.
In both study areas, approximately 0.5 mL of blood was collected through tail venipuncture using a sterile insulin syringe (Samour et al., Reference Samour, Risley, March, Savage, Nieva and Jones1984). A portion of the blood was used to prepare thin smears, fixed in absolute methanol for 5 min, then stained with 10% Giemsa for 45 min (Rodríguez and Matta, Reference Rodríguez and Matta2001). The remaining blood was either applied onto filter paper or preserved in 96% ethanol for molecular analysis.
Microscopic analyses
Blood smears from T. teguixin from Brazil were examined using Olympus CX31 at magnifications of 400× and 1000× to detect blood parasites. Digital images were captured using an Olympus Qcolor 5 camera and processed with the QCapture (Olympus Corporation, Tokyo, Japan). The diagnosis was made for Colombian samples using an Olympus BX43 microscope with integrated camera DP27 and the software CellSens (Olympus Corporation). Measurements were taken with ImageJ software (Schneider et al., Reference Schneider, Rasband and Eliceiri2012) and analysed based on criteria outlined by Telford (Reference Telford2009) and using the description for Leucocytozoon spp. by Valkiūnas (Reference Valkiūnas2005). For each measurement, a range of observations, including sample means and standard deviations, were recorded (Table 3). Parasitemia was assessed by counting the number of parasitized cells per 10,000 total erythrocytes (Staats and Schall, Reference Staats and Schall1996) (Table 2 and Supplementary Table S1). Blood smears were deposited as vouchers in the Collections of the Institute of Biological Sciences at Universidade Federal de Minas Gerais (UFMG), Belo Horizonte, Brazil, and in the Biological Collection Grupo de Estudio Relación Parásito Hospedero (GERPH) at Universidad Nacional de Colombia, Bogotá, Colombia.
Table 3. Morphometric characteristics of the haemosporidian parasites found in the golden tegus Tupinambis teguixin sampled in this study and compared with original descriptions. Measurements are in micrometers (µm). Mean ± SD is provided followed by (minimum and maximum values)

L/W – Length and width ratio.
* This author indicated the presence of possible meronts for this species, thus, morphometric data for this parasitic stage is presented here.
Molecular detection of haemosporidian parasites
DNA from whole blood preserved in filter paper was extracted using QIAamp DNA Micro Kit (QIAGEN GmbH, Hilden, Germany) from 26 T. teguixin samples from Brazil and 13 from Colombia (Table 2, and Supplementary Table S1). Then, the extracted DNA was screened to assess the presence of haemosporidians using a nested polymerase chain reaction (PCR) protocol that targets the parasite mitochondrial cytochrome b gene (cytb, 1131 bp) with primers described by Pacheco et al. (Reference Pacheco, Cepeda, Bernotiene, Lotta, Matta, Valkiunas and Escalante2018b). Briefly, primary PCR amplifications were carried out in a 50 μl volume with 5 μl of total genomic DNA, 2.5 mM MgCl2, 1 × PCR buffer, 0.25 mM of each deoxynucleoside triphosphate, 0.4 μM of each primer, and 0.03 U/μl AmpliTaq polymerase (Applied Biosystems, Thermo Fisher Scientific, USA). Outer PCR oligos used were forward AE298 5′-TGT AAT GCC TAG ACG TAT TCC 3′ and reverse AE299 5′-GT CAA WCA AAC ATG AAT ATA GAC 3′, and inner PCR oligos forward AE064 5′-T CTA TTA ATT TAG YWA AAG CAC 3′ and reverse AE066 5′-G CTT GGG AGC TGT AAT CAT AAT 3′. Primary PCR conditions were: An initial denaturation at 94°C for 4 min and 36 cycles with 1 min at 94°C, 1 min at 53°C, and 2 min extension step at 72°C. In the last cycle, a final extension step of 10 min at 72°C was added. Nested PCR mix and conditions were the same as the primary PCR but using only 1 μl of the primary PCRs and an annealing temperature of 56°C. PCR amplified products (50 μl) were excised from agarose gels and purified by the QIAquick Gel Extraction Kit (QIAGEN GmbH, Hilden, Germany). Both strands for the cytb gene fragments were directly sequenced at Genewiz from Azenta Life Sciences (New Jersey, USA). Electropherograms were carefully inspected for all positive samples, and samples with double peaks were considered mixed infections. Sequences obtained here for single infections were compared against the GenBank database using BLAST (Altschul et al., Reference Altschul, Gish, Miller, Myers and Lipman1990) and deposited in GenBank under the accession numbers PQ680045-PQ680069.
mtDNA amplification, cloning, and sequencing
Nearly complete parasite mtDNA were amplified from 3 T. teguixin samples from Brazil and 1 from Colombia using a nested PCR protocol with Takara LA Taq™ polymerase (TaKaRa Takara Mirus Bio, San Jose, USA) following Pacheco et al. (Reference Pacheco, Matta, Valkiūnas, Parker, Mello, Stanley, Lentino, Garcia-Amado, Cranfield, Kosakovsky Pond and Escalante2018a). Outer oligos used were forward AE170-5′ GAGGATTCTCTCCACACTTCAATTCGTACTTC 3′ and reverse AE171-5′ CAGGAAAATWATAGACCGAACCTTGG ACTC 3′, and the inner oligos forward AE176-5′ TTTCATCCTTAAATCTCGTAAC 3′ and reverse AE136-5′ GACCGAACCTTGGACTCTT 3′. PCRs were carried out in 50 μL using 5 μL of the total DNA for each PCR. Negative (distilled water) and positive controls (samples from an infected primate, Plasmodium inui Halberstaedter and von Prowazek, 1907) were also included. Amplification conditions for both PCRs were an initial denaturation at 94°C for 1 min and 30 cycles with 30 s at 94°C and 7 min at 67°C, followed by a final extension of 10 min at 72°C. Two independent PCR products (50 μL) were excised from the gel (bands of ∼6 kb), purified using the QIAquick Gel extraction kit (Qiagen, GmbH, Hilden, Germany), and cloned into the pGEM-T Easy Vector systems (Promega, Madison, Wisconsin, USA) following the manufacturer’s instructions. Both strands of 4 clones for each sample were sequenced at Genewiz from Azenta Life Sciences (New Jersey, USA). All clones within samples were identical without inconsistencies, suggesting that only one parasite species per sample was found using this protocol. The mtDNA genome sequences obtained were identified as two different haplotypes of P. (S) tupinambi comb. nov. and submitted to GenBank under the accession number PQ680070–PQ680073. It is worth noting that the Plasmodium (Carinamoeba) sp. found by microscopy did not amplify using all the protocols performed in this study.
Phylogenetic analyses
Using both the parasite partial cytb gene and the nearly complete parasite mtDNA genome, the phylogenetic relationships between P. (S) tupinambi comb. nov. sequences obtained in this study and previously reported haemosporidian sequences were estimated on three alignments. All alignments were performed using ClustalX v2.0.12 and Muscle as implemented in SeaView v4.3.5 (Gouy et al., Reference Gouy, Guindon and Gascuel2010) with manual editing. The two first alignments included 112 and 88 partial cytb gene sequences from 4 Haemosporida genera (Plasmodium, Haemocystidium, Haemoproteus and Leucocytozoon) available from GenBank (Benson et al., Reference Benson, Karsch-Mizrachi, Clark, Lipman, Ostell and Sayers2013) and the cytb gene sequences obtained here. All cytb sequences obtained here were included in the first alignment, and only one for each haplotype (H1 and H2) found in this study was included in the second alignment. Haemoproteus (Parahaemoproteus) spp. and Leucocytozoon (Akiba) caulleryi (Mathis and Leger, 1909) sequences were used as an outgroup. Unfortunately, these two alignments only have 383 bp (excluding gaps), given that several lizard parasite sequences contained various sites with IUPAC code and Ns, reducing the number of informative sites for phylogenetic analyses.
A third alignment was done using the 132 nearly complete parasite mtDNA genome sequences (5081 bp excluding gaps) available in the GenBank for parasites belonging to four genera (Haemocystidium, Leucocytozoon, Haemoproteus and Plasmodium), including the genome sequences reported here for each haplotype found in this study (PQ680070–PQ680073). In this case, the phylogenetic analyses were performed with sequences from Leucocytozoon (Leucocytozoon) and Haemoproteus (Haemoproteus) parasites as an outgroup (Pacheco and Escalante, Reference Pacheco and Escalante2023).
Phylogenetic hypotheses were inferred based on these three alignments using Bayesian Inference implemented in MrBayes v3.2.7 with default priors (Ronquist and Huelsenbeck, Reference Ronquist and Huelsenbeck2003) and a general time-reversible model with gamma-distributed substitution rates with invariant sites (GTR + Γ + I) as it was the best model that fit the data with the lowest Bayesian information criterion scores estimated by MEGA v7.0.26 (Kumar et al., Reference Kumar, Stecher and Tamura2016). Bayesian supports were inferred for the nodes in MrBayes by sampling every 1000 generations from 2 independent chains of 4 × 106 Markov Chain Monte Carlo steps. The chains were assumed to have converged once the potential scale reduction factor value was between 1.00 and 1.02, and the average standard deviation of the posterior probability was <0.01. Once convergence was reached, the first 25% of the samples were discarded as a ‘burn-in’. Lineages names and GenBank accession numbers of all sequences (partial cytb gene and mtDNA genomes) used here are shown in all phylogenetic trees.
In addition, the average evolutionary divergences between P. (S) tupinambi comb. nov. haplotype sequence pairs were estimated using the partial cytb gene (745 sites in the final dataset) and the nearly complete parasite mtDNA genome sequences (5494 sites in the final dataset) with a Kimura 2-parameter model (Kimura, Reference Kimura1980) as implemented in MEGA v7.0.26 (Kumar et al., Reference Kumar, Stecher and Tamura2016). The rate variation among sites was modelled with a gamma distribution (shape parameter = 1).
Results
Microscopic detection
Haemosporidians were detected by microscopic examination of blood films in 18 T. teguixin (46%; n = 39; Table 2); four T. teguixin (30%; n = 13) from the eastern plains of Colombia and 14 (54%; n = 26) were from Central Amazonia of Brazil, showing that they are common blood parasites at both study sites. All lizards from Colombia were infected only with non-erythrocytic parasites (Figure 2). In samples collected in Brazil (Figures 3 and 4), infections by erythrocytic parasites in one lizard, non-erythrocytic parasites in eight lizards, and mixed infections in five lizards were observed. The non-erythrocytic (Figures 2 and 3) and erythrocytic parasites exhibited distinct morphologies (Figure 4). Non-erythrocytic parasites were markedly larger than erythrocytic parasites; they did not exhibit merogonic stages, lacked hemozoin pigment, and contained numerous prominent volutin granules. Morphological and morphometric data (Figures 2 and 3; Table 3) showed that the morphology of these non-erythrocytic parasites is compatible with the description of S. tupinambi (Figure 5). For the erythrocytic parasites (Figure 4), due to low parasitemia (ranging from 0.02% to 0.06%; Table 2 and Supplementary Table S1), the number and diversity of observed blood stages were insufficient for species identification, thus they were identified as Plasmodium (Carinamoeba) sp. The species morphological characterization section below shows details for erythrocytic and non-erythrocytic parasites.

Figure 2. Plasmodium (Saurocytozoon) tupinambi comb. nov. in golden tegus (Tupinambis teguixin) from Casanare, Colombia. (a–i) Macrogametocytes. (j–l) Microgametocytes. (h–i) Coinfection with Hepatozoon parasites. Black arrow – host cell nucleus; black arrowheads –parasite nucleus; red arrow – parasite nucleolus; asterisk – portion of the host cell cytoplasm. Thin blood smears stained with Giemsa. Scale bar = 10 μm.
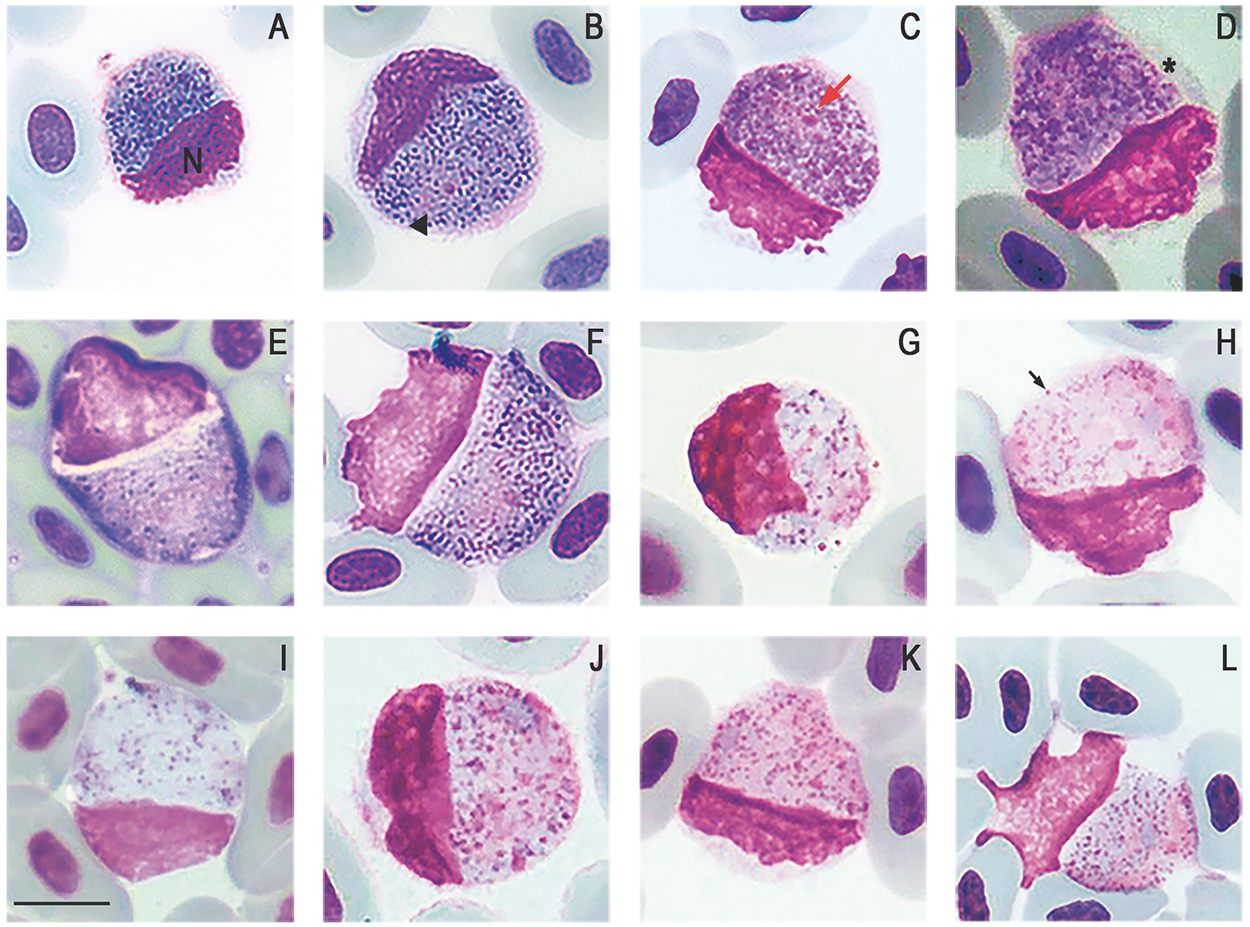
Figure 3. Plasmodium (Saurocytozoon) tupinambi comb. nov. in golden tegus (Tupinambis teguixin) from Amazonas, Brazil. (a–f) Macrogametocytes. (g–l) Microgametocytes. Black arrowhead – parasite nucleus; (n) host cell nucleus; red arrow – parasite nucleolus; black arrow – volutin granule; asterisk – portion of the host cell cytoplasm. Thin blood smears stained with Giemsa. Scale bar = 10 μm.
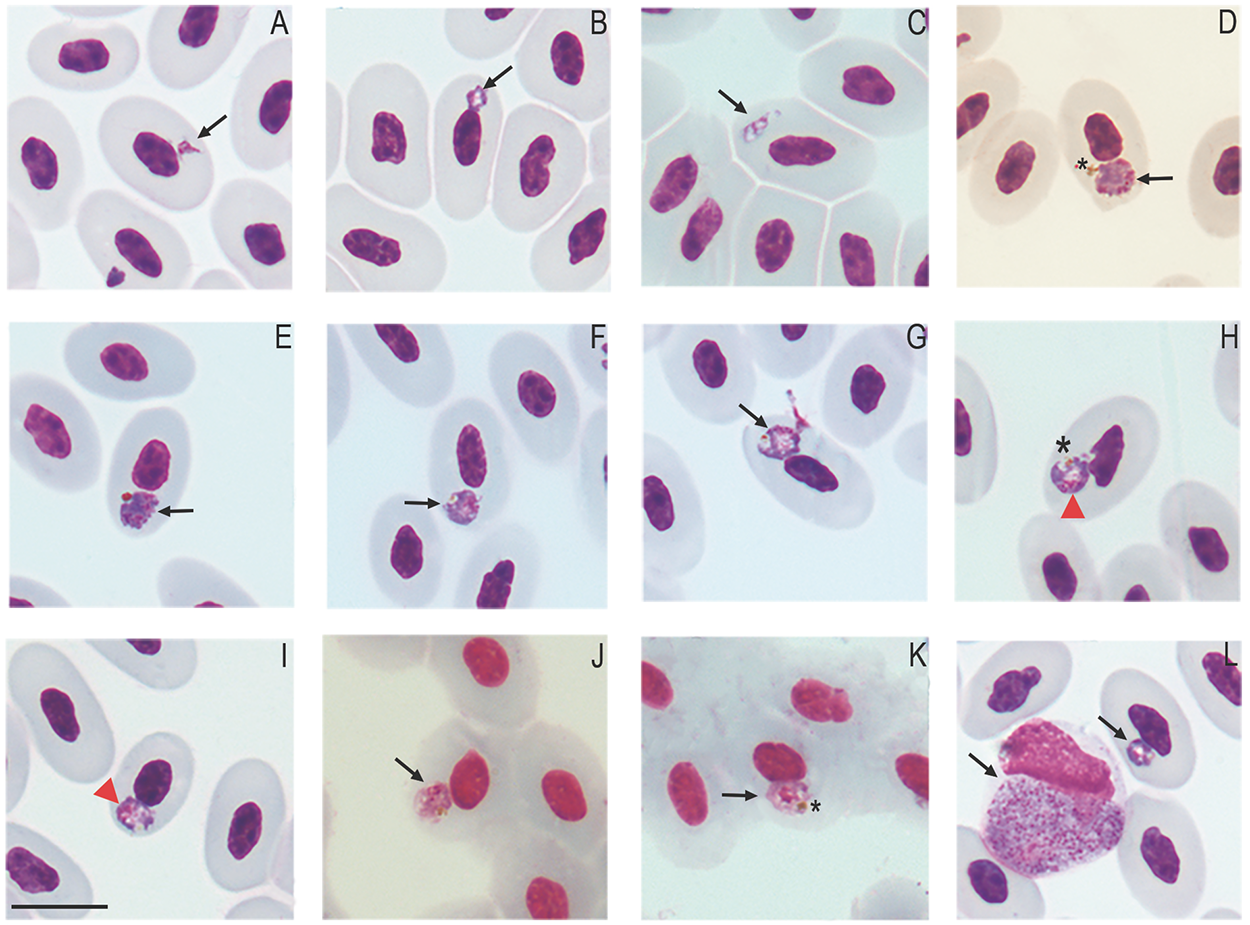
Figure 4. Erythrocytic Plasmodium (Carinamoeba) sp. infection in golden tegus (Tupinambis teguixin) from Amazonas, Brazil. (a–b) Trophozoites. (c–e) Meronts. (f–i) Macrogametocytes. (j–k) Microgametocytes. (l) Coinfection with Plasmodium (Saurocytozoon) tupinambi comb. nov. Black arrows – parasites; red arrowheads – parasite nucleus; asterisk – hemozoin pigment granules. Thin blood smears stained with Giemsa. Scale bar = 10 μm.

Figure 5. Plasmodium (Saurocytozoon) tupinambi comb. nov. in golden tegus (Tupinambis teguixin) from Para, Brazil, hapantotype (no. 949) from Garnham’s collection, NHM. (a–c) Macrogametocytes. (d–e) Microgametocytes. Black arrowhead – parasite nucleus; (N) host cell nucleus; red arrow – parasite nucleolus; black arrow – volutin granule; asterisk – portion of the host cell cytoplasm. Thin blood smears stained with Giemsa. Scale bar = 10 μm.
Molecular and phylogenetic analyses
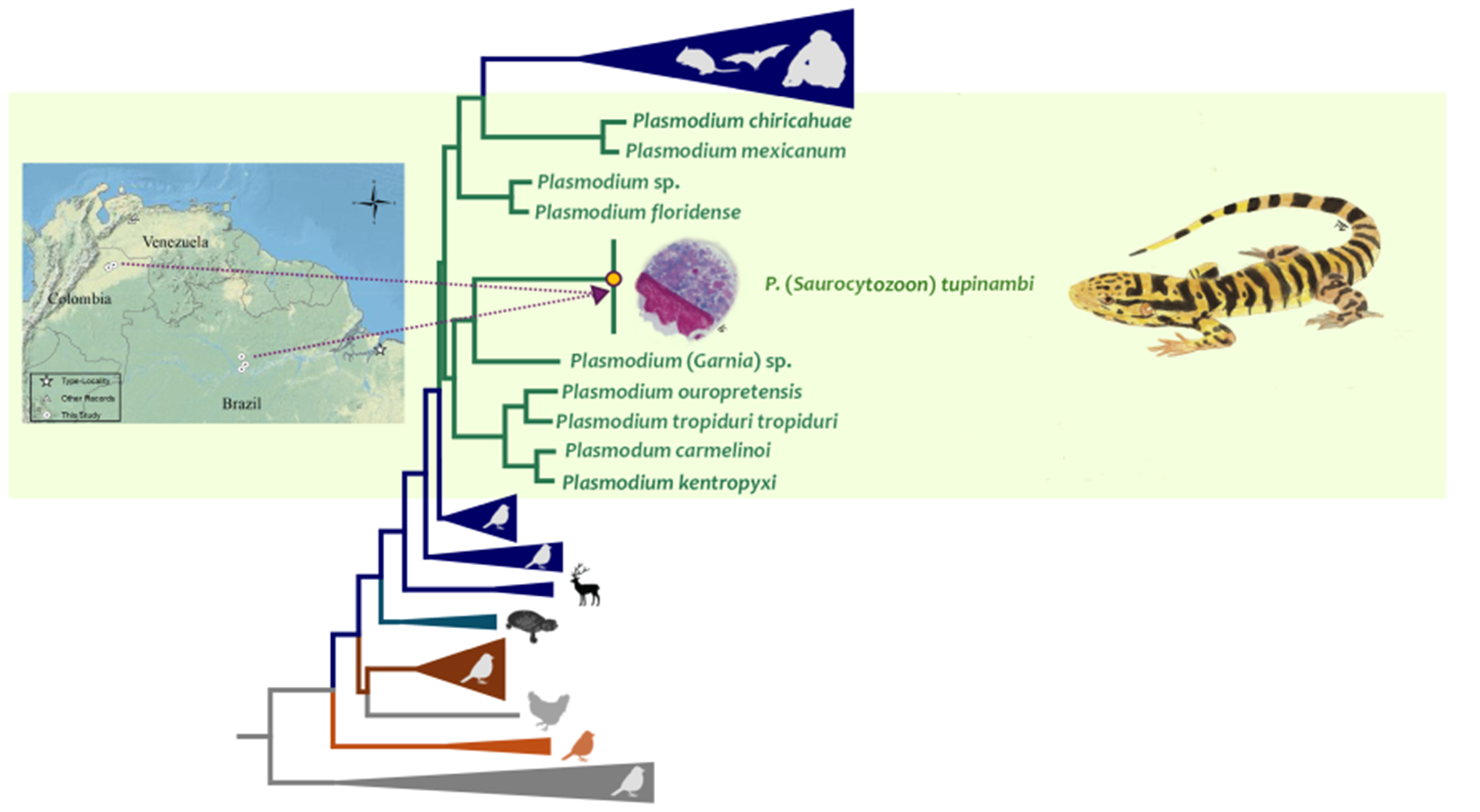
Seventeen out of 26 samples from Brazil (65%) and eight out of 13 (61%) from Colombia were positive by PCR. Both parasite partial cytb gene and the nearly complete parasite mtDNA genome sequences obtained here showed three haplotypes, one from Brazilian and Colombian (H1, 15/17 = 88% and 8/8 = 100%, respectively) samples, one (H2, 2/17 = 11%) only present in Brazil, and one only present in Colombia in only one sample (H3, 1/17 = 6%) (Supplementary Figure S1). Bayesian phylogenetic trees, using both the parasite partial cytb gene and the nearly complete parasite mtDNA genome, show that these three haplotypes are closely related (Figure 6, Supplementary Fig. S1, and Figure 7, respectively) and nested within the Plasmodium species group. Indeed, a very low genetic distance between H1 and H2 was found using both the cytb gene (0.0015 ± 0.00133, N sites = 745) and parasite mtDNA genome sequences (0.00046 ± 0.00022, N sites = 5494), with only one synonymous substitution in the cytb gene.

Figure 6. Bayesian phylogenetic hypothesis of Plasmodium (Saurocytozoon) tupinambi comb. nov. based on cytb gene (383 bp excluding gaps). branch colours indicate different genera/hosts. GenBank accession numbers for all parasite sequences used in this analysis are provided in parentheses. Lizard image credit: Robson Ávila.
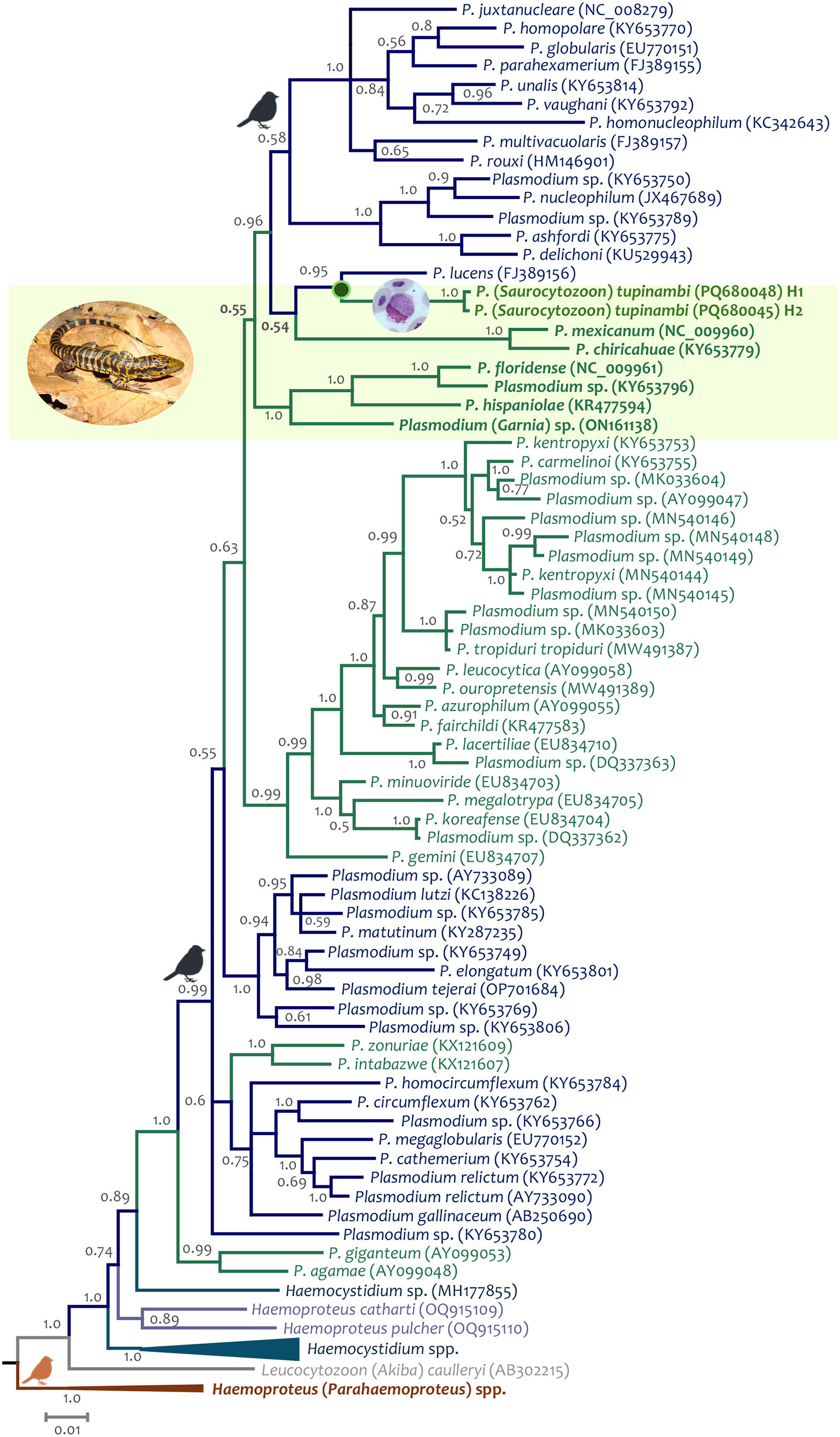
Figure 7. Bayesian phylogenetic hypothesis of Plasmodium (Saurocytozoon) tupinambi comb. nov. based on mtDNA genome (5081 bp excluding gaps). branch colours indicate different genera/hosts. GenBank accession numbers for all parasite sequences used in this analysis are provided in parentheses. Lizard image credit: Robson Ávila.
Unfortunately, given the few informative sites from the partial cytb gene sequences (N = 383 bp), the phylogenetic relationships between both parasite haplotypes detected here and haemosporidian parasites with data available could not be resolved (many nodes have posterior probabilities less than 0.85) using this fragment. However, given the data, the closest taxon is Plasmodium (Novyella) lucens Valkiūnas et al., Reference Valkiūnas, Iezhova, Loiseau, Smith and Sehgal2009 (FJ389156), a parasite described in Olive Sunbird Cyanomitra olivacea (Smith, 1840) from Africa (Valkiūnas et al., Reference Valkiūnas, Iezhova, Loiseau, Smith and Sehgal2009) (Figure 6, Supplementary Fig. S1). Although the dataset with parasite mtDNA genomes has more informative sites (N = 5081 bp, Figure 7), fewer sequences are available for lizard parasites compared to the cytb dataset. However, both haplotypes from T. teguixin appear to share a common ancestor (PP = 1) with Plasmodium (Garnia) sp. (ON1611381), a non-pigmented parasite found in turnip-tailed gecko Thecadactylus rapicauda (Houttuyn, 1782) from Colombia (Matta et al., Reference Matta, González, Vargas-Ramírez, Valkiūnas, Escalante and Pacheco2023). Both parasites share a common ancestor (posterior probability PP = 1) with a parasite clade that contains Plasmodum carmelinoi Lainson et al., 2010 (KY653755), Plasmodium kentropyxi Lainson et al., 2001 (KY653753), Plasmodium ouropretensis Córdoba et al., 2021 (MW491389), and Plasmodium tropiduri tropiduri (Aragão and Neiva, 1909) (MW491387), parasites reported recently in Colombia and Brazil (Matta et al., Reference Matta, González, Pacheco, Escalante, Moreno, González and Calderón-Espinosa2018; Ferreira et al., Reference Ferreira, Alves, Jager, Franzini, Mesquita, Díaz-Delgado, Catão-Dias and Braga2020; Córdoba et al., Reference Córdoba, Ferreira, Pacheco, Escalante and Braga2021).
Thus, the molecular evidence presented here indicates that haplotypes infecting T. teguixin belong to the same species of non-erythrocytic parasites previously identified as S. tupinambi and support classifying this parasite as a Plasmodium taxon. Therefore, we propose placing the non-erythrocytic parasites detected in golden tegu lizards in Plasmodium (new genus combination).
Species morphological characterization
Plasmodium (Saurocytozoon) tupinambi (Lainson and Shaw, Reference Lainson and Shaw1969) comb. nov. (Figures 2 and 3; Table 3).
Type Host: Golden tegu Tupinambis teguixin (Linnaeus, 1758) (Teiidae).
Other Hosts: Probably Crocodilurus amazonicus Spix, 1825 (Teiidae) (Lainson et al. Reference Lainson, Landau and Shaw1974a).
Vectors: Natural vectors are unknown. Sporogony completed in experimentally exposed mosquito Culex quinquefasciatus Say, 1823 (Landau et al., Reference Landau, Lainson, Boulard and Shaw1973).
Type Locality: Forest edge bordering Lake Água Preta, Utinga State Park, municipality of Belém, Pará State, Brazil (Lainson and Shaw, Reference Lainson and Shaw1969).
Other Localities: Brazil – Barcarena, Pará State (Lainson and Shaw, Reference Lainson and Shaw1969); Biological Dynamics of Forest Fragments Project (BDFFP) campsites in municipalities of Rio Preto da Eva and Manaus, and Rio Pardo Rural Settlement (RPRS) in the municipality of Presidente Figueiredo, Amazonas State (Table 2; this study). Colombia – Fincas Buena Vista and Chaviripa in Paz de Ariporo, Reserva Puro Llano in the municipality of Yopal, Casanare Department (Table 2; this study). Venezuela – Portuguesa and Cojedes states (Telford, Reference Telford1978, Reference Telford1980).
Parasite frequency and distribution: Brazil – 4 of 5 (80%) in Belém, and a single specimen from Barcarena, Pará State (Lainson and Shaw, Reference Lainson and Shaw1969); 4 of 7 (57%) lizards in BDFFP, and 13 of 19 (68%) from RPRS, Amazonas State (this study). Colombia – 4 of 6 (80%) in Paz de Ariporo, and 5 of 7 (71%) in Yopal, Casanare Department (this study). Venezuela – 24 of 81 (29%) golden tegus (Telford, Reference Telford1978).
Site of infection: Leukocytes and occasionally immature erythrocytes were speculated to be infected (Lainson and Shaw, Reference Lainson and Shaw1969). In this study, blood cell origin could not be identified due to the marked deformation caused by P. (S.) tupinambi comb. nov. gametocytes.
Parasitemia (this study): Brazil – mean of 5.62 parasites/10000 erythrocytes (0.06%; ±0.08), ranging from 1 to 25 parasites per 10000 erythrocytes; Colombia – mean of 8.5 parasites/10000 erythrocytes (0.08%; ±0.08), ranging from 1 to 20 parasites per 10000 erythrocytes.
Type material: Hapantotypes (nos. 949, 950, 951) from the type host and the type locality are deposited in the Natural History Museum, London (Garnham and Duggan, Reference Garnham and Duggan1986).
Additional material: Voucher blood smears (UFMG31 and UFMG197), parasitemia are 0.02% and 0.08%, respectively. Collected by A. M. Picelli (Manaus and Presidente Figueiredo municipalities, Brazil) and deposited in the Institute of Biological Sciences (UFMG, Belo Horizonte, Brazil). Voucher blood smears (GERPH:CAH311, GERPH:CAH359 and GERPH:CAH765), parasitemia are 0.06%, 0.2%, and 0.01% respectively. Collected by L. M. Velandia (Paz de Ariporo, Colombia); and (GERPH:CAH400) parasitemia 0.07% Collected by L. M. Velandia (Yopal, Colombia); were deposited in the Biological Collection GERPH at (Universidad Nacional de Colombia, Bogotá, Colombia).
DNA Sequences: Haplotype H1, partial cytb gene: PQ680048 – PQ680068, and mtDNA genones: PQ680072 – PQ680073. Haplotype H2, partial cytb gene: PQ680045 – PQ680047 and mtDNA genomes: PQ680070 – PQ680071, Haplotype H3, only partial cytb gene: PQ680069.
ZooBank registration: The Life Science Identifier for P. (S) tupinambi comb. nov. is urn:lsid:zoobank.org:act:C4380897-05DA-43CC-8E50-9D2F54C3781E.
Diagnosis: Gametocytes develop in white blood cells; host-parasite complexes are indistinguishable from those of avian Leucocytozoon species developing in roundish host cells. Nuclei of host cells are appressed to gametocytes and extend less than ½ of the circumferences of gametocytes.
Mature gametocytes are round to oval and occupy almost their entire host cell, which often results in significant deformation of the host cell. Male and female mature gametocytes can be readily distinguished due to sexual dimorphic characters that are typical for haemosporidian parasites – paler staining cytoplasm and the large diffuse nuclei in microgametocytes (Figure 2j-l and Figure 3g-l) in comparison to macrogametocytes (Figure 2a-i and Figure 3a-f). Hemozoin pigment is absent. Only mature gametocytes found and no merogonyic cycle observed.
Macrogametocyte (Figure 2a-i and Figure 3a-f; Table 3) cytoplasm is vacuolated, has scattered and abundant dark (red to pinkish) volutin granules, and bluish-grey staining with less evident azurophilic granules. The nucleus of the macrogametocytes is oval or irregular in shape, stains pinkish, lies at a central to peripheral location, and typically exhibits a darker stained round nucleolus (Figure 2f and Figure 3c).
Microgametocyte (Figure 2j-l and Figure 3g-l; Table 3) exhibit the cytoplasm stained lightly in purple and less vacuolated than the macrogametocyte. Because of these characteristics, volutin granules are readily visible. Parasite nuclei are inconspicuous, often pale-pink stained and sometimes hardly distinguishable from the cytoplasm. Occasionally, a small pink round mass, of unclear origin can be observed in the cytoplasm (Figure 2k-l).
Effects on host cells: Host cells became hypertrophied and rounded as gametocytes grow and, in some cases, the cytoplasm is distended, forming a thin layer around the parasite (Figure 2d and Figure 3d). The host cell nuclei are distorted, pushed aside, and extend to less than half of the circumference of the gametocyte, to which the nucleus is typically appressed, and in rare instances. The nucleus appears to be in contact with the parasite and, in certain regions, there is a thin and delicate layer of the host nucleus overlapping the parasite’s cytoplasm (Figure 2l). The part of the host cell nucleus that is opposite to gametocyte may exhibit an irregular or wavy outline (Figure 3j).
Remarks: Lainson and Shaw (Reference Lainson and Shaw1969) described this large parasite in T. teguixin from the Brazilian Amazonia; it was present mostly in non-erythrocytic cells, lacked a merogonic cycle in peripheral blood, and did not exhibit hemozoin pigment. Therefore, the authors created a new genus for this haemosporidian within the Family Leucocytozoidae and described a new species, S. tupinambi, the first species of this family identified in reptiles. Here, DNA sequences recovered for this parasite species infecting T. teguixin in Brazil and Colombia clustered within the genus Plasmodium in the Plasmodiidae, with strong support at both mtDNA and cytb level. This provides phylogenetic support for a new combination, Plasmodium (Saurocytozoon) tupinambi comb. nov. We suggest keeping Saurocytozoon as a subgenus due to the taxonomic tradition of subgeneric classification of haemosporidian parasites as well as the distinct morphology of the parasite and its influence on the host cell nucleus, which is deformed and enlarged.
Two other species have been described in Saurocytozoon based on morphological characteristics: Saurocytozoon mabuyi Lainson et al., 1974 in the black-spotted skink Copeoglossum nigropunctatum (Spix, 1825) from the same region as P. (S) tupinambi comb. nov. (Lainson et al., Reference Lainson, Landau and Shaw1974a); and Saurocytozoon agamidorum Telford, Reference Telford2013 in the Caucasian agama Paralaudakia caucasia (Eichwald, 1831) from Pakistan (Telford, Reference Telford2013). Telford (Reference Telford1978, Reference Telford1983) also reported S. mabuyi in the common sun skink Eutropis multifasciata (Kuhl, 1820) from Thailand and Singapore, and P. (S) tupinambi comb. nov. in its type host species from Venezuela. These three species share similar morphologies. However, they can be distinguished from each other by the size of their gametocytes, where S. agamidorum [8.2 ± 1.7 × 6.2 ± 0.8 μm (Telford, Reference Telford2013)] and S. mabuyi [11.3 ± 1.5 × 8.7 ± 1.0 μm (Lainson et al., Reference Lainson, Landau and Shaw1974a; Telford, Reference Telford1983)] are smaller than P. (S.) tupinambi comb. nov. (Table 3). Furthermore, S. agamidorum gametocytes sometimes exhibit a distinct single vacuole, severely distorting the host cells by stretching out their nuclei (Telford, Reference Telford2013).
Plasmodium (Saurocytozoon) tubinambi comb. nov., exhibits morphological variations in mature gametocyte dimensions across different geographic areas (Table 3). Macrogametocytes found in Brazil (Lainson and Shaw, Reference Lainson and Shaw1969; this study) showed slightly larger sizes compared to those from Venezuela (Telford, Reference Telford1978) and Colombia (this study, Table 3). However, in all locations, no differences were observed between microgametocyte dimensions. According to the species diagnosis made by Telford (Reference Telford2009), gametocytes have dimensions ranging between 13–17 × 10–16 μm, encompassing the larger dimensions found in Brazil. Nevertheless, this range of dimensions is larger than the size of the parasites (see Table 3) shown in his study in Venezuela (Telford, Reference Telford1978) and the original description (Lainson and Shaw, Reference Lainson and Shaw1969).
Telford (Reference Telford1980, Reference Telford2009) briefly mentioned the presence of parasites in lymphocytes of the giant ameiva Ameiva praesignis (Baird and Girard, 1852) with length and width data (10.9 ± 1.9 × 6.6 ± 1.3 μm; 9–16 × 5–10 μm) like those for P. (S) tupinambi comb. nov. However, this author did not provide a detailed description or images for this haemosporidian. Lainson et al. (Reference Lainson, Landau and Shaw1974a) reported finding P. tubinambi comb. nov. in one crocodile tegu C. amazonicus without providing further information. Although these two additional teiid species may be possible hosts for P. (S) tupinambi comb. nov., the lack of thorough morphological data in these reports precludes comparisons with the parasitic forms found here.
Plasmodium (Carinamoeba) sp. (Figure 4)
This haemosporidian parasite is characterized by small forms infecting erythrocytes of T. teguixin in Brazil.
Trophozoites (Figure 4a-b and l) are small, round, or ring-shaped, sometimes displaying short cytoplasmatic outgrowths (Figure 4a). Young trophozoites tend to be positioned close to the host cell nucleus at the lateral or polar region (Figure 4a). Mature trophozoites have the greyish-blue stained cytoplasm with a centrally located vacuole, and small golden pigment granules (Figure 4b and 4l).
Meronts (Figure 4c-e) in young stages are oval or slightly elongated, with the nuclei appearing as thick bands and having one or two large vacuoles (Figure 4c). Mature meronts (Figure 4d-e) are broadly fan-shaped, do not exhibit vacuoles, the cytoplasm stains dark-blue, and the nuclei stains pink. The pigment granules of mature meronts are dark golden, clumped into a mass, which is positioned opposite to the parasite nuclei (Figure 4d). Mature meronts are in a polar position close to the host cell nuclei, which sometimes leads to displacement and slight distortion of these nuclei (Figure 4d-e). Meronts produce an average of 4.5 ± 1.0 merozoites, ranging from 4 to 6 (n = 4). Length and width of mature meronts are 4.9 ± 0.9 × 3.7 ± 0.6 μm (4.1–6 × 3.2–4.5 μm; n = 4), and area 16.2 ± 2.8 μm2 (13–20.2 μm2; n = 4). Mature meront size relative to host cell nucleus size is 0.79 ± 0.35 (0.58–1.33; n = 4), and to non-infected erythrocyte nucleus size is 0.70 ± 0.15 (0.55–0.89; n = 4).
Gametocytes (Figure 4f-k) are roundish and small, nucleophilic, generally occupy a polar position in erythrocytes and often have one or two small vacuoles. They produce few evident effects on the erythrocytes, usually slightly deforming the host cell and its nucleus, and sometimes displacing the latter.
Macrogametocytes (Figure 4f-i) have greyish-blue cytoplasm lightly stained at the central area. Golden pigment granules are grouped on the periphery and opposite to parasite nucleus (Figure 4h). The macrogametocyte nuclei are stained pinkish-red and appear as a thick band or mass along one of the margins (Figure 4h). Macrogametocytes are relatively larger than microgametocytes. Macrogametocytes average dimensions are 5.0 ± 0.8 × 4.2 ± 0.7 μm (3.5–6.6 × 3.1–5.5 μm; n = 20), with area 18.7 ± 5.3 μm2 (10–30.6 μm2; n = 20) and length/width ratio (L/W) 1.14 ± 0.10 (1.00–1.36). Macrogametocyte size relative to host cell nucleus averages 0.87 ± 0.34 (0.40–1.52; n = 20), and to non-infected erythrocyte nucleus size is 0.76 ± 0.31 (0.41–1.50; n = 20). Macrogametocyte nuclei average dimensions 2.6 ± 0.6 × 1.5 ± 0.6 μm (1.5–3.6 × 0.6–2.5 μm; n = 20) and area 3.6 ± 1.4 μm2 (1.3–6.2 μm2; n = 20).
Microgametocyte (Figure 4j-k) cytoplasm stains light pink and there is a thick reddish nuclear mass in a lateral or central position in gametocytes. The golden pigment granules are clumped on the parasite margin (Figure 4g). Microgametocytes average dimensions are 4.3 ± 0.8 × 3.6 ± 0.8 μm (3.4–6.0 × 2.6–5.2 μm; n = 10), with area 15.0 ± 6.5 μm2 (9.5–30.5 μm2; n = 10) and L/W 1.21 ± 0.18 (0.96–1.64). Microgametocyte size relative to host cell nucleus size averages 0.72 ± 0.21 (0.36–1.00; n = 10), and to non-infected erythrocyte nucleus size is 0.65 ± 0.16 (0.39–0.90; n = 10). Microgametocyte nuclei average dimensions 2.1 ± 0.4 × 1.4 ± 0.4 μm (1.7–3.0 × 0.9–2.3 μm; n = 10) and area 3.4 ± 1.1 μm2 (1.8–6.0 μm2; n = 10).
Remarks: The absence of a sequence for the erythrocytic Plasmodium sp. and the need for further morphological data on its asexual stages precludes its formal description. However, it is valuable to compare its morphology to that of P. minasense tegui, another haemosporidian species found in T. teguixin. Plasmodium minasense tegui, one of seven subspecies of P. minasense, was found infecting T. teguixin lizards from Venezuela (Telford, Reference Telford1979, Reference Telford1980). While only blood stages are known for this subspecies, it can be distinguished from the other subspecies by its nucleophilic erythrocytic forms, macrogametocytes with band-like nucleus, similar average sizes between meronts (4.2 ± 0.8 × 3.4 ± 0.5 µm; N = 50) and gametocytes (4.2 ± 0.7 × 3.6 ± 0.5 µm; N = 50), and by having the smallest gametocytes among all subspecies (Telford, Reference Telford2009). Although P. minasense tegui and the parasite identified in the present study share some morphological characteristics, macrogametocytes and microgametocytes of P. minasense tegui were not morphometrically differentiated in previous reports, with their measurements varying broadly, and images available for comparison are of low quality (Telford, Reference Telford1979, Reference Telford1980, Reference Telford2009). Given these discrepancies and the taxonomic challenges surrounding P. minasense and its subspecies, it is premature to confirm that the erythrocytic parasite in our study is the same species as described by Telford (Reference Telford1979). Further morphological and molecular data are necessary to resolve this taxonomic uncertainty. Regarding other records of Plasmodium parasites in T. teguixin, small round gametocytes (5 × 3 μm) like P. minasense tegui were observed in French Guiana (Leger, Reference Leger1919), while in Brazil (Landau et al., Reference Landau, Lainson, Boulard and Shaw1974) and Colombia (Ayala et al., Reference Ayala, D’Alessandro, Mackenzie and Angel1973) no morphological data were provided.
Discussion
The morphology of Saurocytozoon is unique among reptile haemosporidians, but it has remained poorly investigated. The debate that started just a few years after its description in the 1960s was at a stalemate, with some authors assuming that it should be placed in Plasmodiidae, either as Saurocytozoon (Telford, Reference Telford1983, Reference Telford2009, Reference Telford2013; Perkins, Reference Perkins2014) or as Plasmodium (Ayala, Reference Ayala and Kreier1977, Reference Ayala1978; Levine, Reference Levine1988; Schall, Reference Schall1996). Nevertheless, keeping the genus within the Leucocytozoidae was widely preferred, although this classification, or potential reclassification, needed confirmation by molecular methods (Pacheco and Escalante, Reference Pacheco and Escalante2023; Valkiūnas and Iezhova, Reference Valkiūnas and Iezhova2023). Although morphologically similar to gametocytes of avian Leucocytozoon parasites (Leucocytozoidae), genetic data obtained here from natural infections occurring in Brazil and Colombia have placed Saurocytozoon within the Plasmodium clade. Consequently, S. tupinambi – the type species of Saurocytozoon – should be considered Plasmodium (Saurocytozoon) tupinambi comb. nov. It follows that the other two species of Saurocytozoon also should be reclassified as Plasmodium (Saurocytozoon) mabuyi comb. nov. and Plasmodium (Saurocytozoon) agamidorum comb. nov. These three parasites are part of the same subgenus and are similar based on the morphology of blood stages. However, future molecular evidence is needed to corroborate the taxonomic placement of these two parasites.
This finding addresses the use of broader definition of the family Plasmodiidae than the traditional one proposed by Garnham (Reference Garnham1966), which was limited to merogony in circulating blood cells, production of hemozoin and development of gametocytes in erythrocytes. The inclusion of unpigmented parasites developing in both white and red blood cells, an additional morphological character suggested by Telford (Reference Telford1973) and Ayala (Reference Ayala and Kreier1977), has gained solid phylogenetic support (Córdoba et al., Reference Córdoba, Ferreira, Pacheco, Escalante and Braga2021; Matta et al., Reference Matta, González, Vargas-Ramírez, Valkiūnas, Escalante and Pacheco2023; this study). Given the molecular evidence, we suggested the following slightly corrected diagnosis for the family Plasmodiidae: mainly, merogony takes place in cells of fixed tissues (obligatory) and blood cells (with few exceptions) of vertebrate hosts; hemozoin pigment is present in meronts and gametocytes, which develop in mature erythrocytes but during the development in immature erythrocytes and leukocytes, it can be absent in some species; sexual process and sporogony take place predominantly in mosquitoes (Culicidae), but occasionally in sandflies (Phlebotominae), and biting midges (Ceratopogonidae); this family contains one genus, Plasmodium.
The one attempt to understand the P. (S.) tupinambi comb. nov. development in an experimental vector was made by Landau et al. (Reference Landau, Lainson, Boulard and Shaw1973); it was shown that the oocysts developed and sporogony completed in the mosquito C. quinquefasciatus (syn. Culex pipiens fatigans). Interestingly, the oocysts developed inside the epithelial cell layer of the vector stomach, as it occurs in Leucocytozoidae species. Although sporozoites developed in mature oocysts, there was no evidence of their invasion into the salivary glands, indicating that this mosquito species was a non-competent vector (Telford, Reference Telford2009). Interestingly, oocysts of P. (S.) tupinambi comb. nov. developed slowly (over 16 days) in mosquitoes; they contained numerous germinative centres and reached large sizes (up to 62 µm), producing several hundred long, thin sporozoites – the features of malaria parasites of Plasmodium. This is not characteristic of Leucocytozoon parasites, in which oocysts develop faster (usually < 7 days); are small (<20 µm), contain one germinative centre, and produce less than one hundred shorter sporozoites (Valkiūnas, Reference Valkiūnas2005). Therefore, those aspects of P. (S.) tupinambi comb. nov. sporogony within mosquitoes prompted Telford (Reference Telford1978, Reference Telford2009) to classify Saurocytozoon parasites within Plasmodiidae. Leucocytozoidae parasites complete sporogony in black flies (Simuliidae) in all investigated species, except for, Leucocytozoon (Akiba) caulleryi Mathis and Leger, 1909, which develops in biting midges, while Plasmodiidae species complete sporogony in mosquitoes in most examined species (Valkiūnas and Iezhova, Reference Valkiūnas and Iezhova2023).
We did not detect merogony associated with infections by P. (S.) tupinambi comb. nov., and its absence was an important argument for classifying this parasite within Leucocytozoidae (Lainson and Shaw, Reference Lainson and Shaw1969). Indeed, only mature gametocytes were found in all studies, indicating the predominant persistence of the parasite at this stage when merogony is absent/inactive. However, this does not rule out that it cannot exist in this parasite. Despite several attempts, Lainson et al. (Reference Lainson, Landau and Shaw1974a) failed to demonstrate the merogony of this parasite in circulating blood cells; they reported a broken meront in a spleen smear from a lizard. However, meronts in the peripherical blood can occur. Still, they might be overlooked because the asexual reproduction in the blood may be transient or occur at submicroscopic levels (Ayala, Reference Ayala and Kreier1977) and can be restricted to certain stages of infection in lizards (Paperna and Landau, Reference Paperna and Landau1990). Thus, additional observations are needed to confirm if merogony is absent in the blood cells of Saurocytozoon species. It is worth mentioning that Telford (Reference Telford1978) showed the existence of large mature meronts (32–102 nuclei) in lymphocytes from an infected T. teguixin, suggesting that P. (S.) tupinambi comb. nov. was more closely related to Plasmodiidae than to Leucocytozoidae parasites. Nevertheless, this author could not confirm that lymphocytic meronts belonged to P. (S.) tupinambi comb. nov. due to the coinfection with P. minasense tegui in the same host.
Infections with erythrocytic Plasmodium (Carinamoeba) sp. were visualized in blood smears from some (n = 6) positive T. teguixin sampled from Brazil; most of them were coinfected (n = 5) with P. (S.) tupinambi comb. nov., and only one presented a single infection. Unfortunately, the molecular screening did not detect a sequence for this parasite. Interestingly, coinfections involving both erythrocytic and non-erythrocytic haemosporidians are relatively common in lizard hosts, including T. teguixin (Lainson et al., Reference Lainson, Landau and Shaw1974b; Perkins, Reference Perkins2000; Córdoba et al., Reference Córdoba, Ferreira, Pacheco, Escalante and Braga2021). Except for Lainson and Shaw (Reference Lainson and Shaw1969) report, which was unclear regarding the presence of coinfections, and in T. teguixin from Colombia (this study), all other reports on Saurocytozoon have documented higher numbers of coinfections with other erythrocytic haemosporidians (Landau et al., Reference Landau, Lainson, Boulard and Shaw1974; Lainson et al., Reference Lainson, Landau and Shaw1974a; Telford, Reference Telford1978, Reference Telford1980, Reference Telford1983, Reference Telford2013; Picelli et al., Reference Picelli, Ramires, Masseli, Pessoa, Viana and Kaefer2020). Because these parasites belong to the same genus (Plasmodium), their molecular detection using general primers constitutes a challenge, requiring species-specific molecular diagnostics.
The presence of non-erythrocytic and erythrocytic parasites without recovering two divergent genetic lineages may lead to the consideration of three hypotheses: (1) both parasites observed here would constitute the same species infecting different host cells since there are other species displaying similar behaviour (i.e. Plasmodium chiricahuae Telford, 1970 and Plasmodium mexicanum Thompson and Huff, 1944) (Telford, Reference Telford2009); (2) sequences obtained belonged to the erythrocytic parasites instead of the P. (S.) tupinambi comb. nov., because molecular detection is particularly challenging for some lineages of leukocyte-inhabiting parasites of genus Leucocytozoon (Lotta et al., Reference Lotta, Valkiūnas, Pacheco, Escalante, Hernández and Matta2019); and (3) DNA from the erythrocytic parasite was not amplified and sequenced due to low parasitemia and primers affinities (Perkins et al., Reference Perkins, Martinsen and Falk2011).
The first two scenarios seem unlikely. First, coinfections are predominant in wildlife (Pacheco and Escalante, Reference Pacheco and Escalante2023), including in T. teguixin lizards (Landau et al., Reference Landau, Lainson, Boulard and Shaw1974; Lainson et al., Reference Lainson, Landau and Shaw1974a). Furthermore, morphological differences between the erythrocytic parasites (presence of small gametocytes and meronts, production of hemozoin granules) and P. (S.) tupinambi comb. nov. (large parasites distorting host cells and their nuclei) suggest that they likely represent distinct species. Second, within a coinfection, PCR often amplifies sequences from the species with higher parasitemia (Perkins et al., Reference Perkins, Martinsen and Falk2011). Additionally, the amplification is often selective irrespective of the intensity of parasitemia during coinfections of different haemosporidians (Valkiūnas et al., Reference Valkiūnas, Bensch, Iezhova, Križanauskienė, Hellgren and Bolshakov2006; Bernotienė et al., Reference Bernotienė, Palinauskas, Iezhova, Murauskaitė and Valkiūnas2016). Further, some readily visible parasitemia can be non-detectable by established PCR-based protocols, as documented in birds with Plasmodium coinfections (Zehtindjiev et al., Reference Zehtindjiev, Križanauskienė, Bensch, Palinauskas, Asghar, Dimitrov, Scebba and Valkiūnas2012). In our case, we visualized coinfections in only 5 of 26 T. teguixin in Brazil, while all 13 individuals from Colombia exhibited single infections with P. (S) tupinambis comb. nov., suggesting that this parasite does not produce pigmented erythrocytic stages. Whole mtDNAs and cytb gene sequences were identical and almost identical (1 bp difference, synonymous mutation in cytb gene) among all parasites from 5 lizards with coinfections and from 21 harbouring single infections, including the individual with single infection by the erythrocytic Plasmodium (Carinamoeba) sp. In this case, we may have failed to detect a coinfection with P. (S) tupinambi comb. nov. by microscopy screening, because our sequencing results showing the presence of haplotype H1 suggests submicroscopic coinfection with P. (S) tupinambi comb. nov.
Lastly, lizards with single infections with P. (S) tupinambi comb. nov. detected by microscopy and PCR screening were found for both localities, reinforcing that sequences from those samples correspond to P. (S) tupinambi comb. nov. This suggests the validation of our third hypothesis. Mainly, the PCR assays used here amplified DNA from the P. (S) tupinambi comb. nov., which had a higher parasitemia than the erythrocytic parasite in all but one case of mixed infections, and it is also possible that primers do better match the DNA of this parasite without relationship to parasitemia intensity. These assays amplify mtDNAs (≤ 6 kb) of many Haemosporida species belonging to several genera infecting multiple hosts (Pacheco et al., Reference Pacheco, Cepeda, Miller, Beckerman, Oswald, London, Mateus-Pinilla and Escalante2024), including in coinfections with Leucocytozoon and Haemoproteus parasites (Vieira et al., Reference Vieira, Pereira, Vilela, Landau, Pacheco, Escalante, Ferreira and Braga2023), but we were not able to recover sequences from the erythrocytic parasite. Therefore, future studies employing next-generation sequencing tools, such as those developed by Pacheco et al. (Reference Pacheco, Cepeda, Miller, Beckerman, Oswald, London, Mateus-Pinilla and Escalante2024), may be required to uncover molecular data related to the erythrocytic Plasmodium (Carinamoeba) sp. infecting T. teguixin.
A higher frequency of infection by P. (S) tupinambi comb. nov. in T. teguixin using molecular screening (65%) compared to microscopy analysis (43%) was found here. This result is not unexpected, as it is well known that PCR tools tend to detect infections not visible on blood smears from reptiles (Ferreira et al., Reference Ferreira, Alves, Jager, Franzini, Mesquita, Díaz-Delgado, Catão-Dias and Braga2020) and avian hosts (Pacheco et al., Reference Pacheco, Ferreira, Logan, McCune, MacPherson, Albino Miranda, Santiago-Alarcon and Escalante2022). Additionally, our molecular screening showed the frequency of infection of P. (S) tupinambi comb. nov. did not vary between sampling sites, 65% in Brazil and 61% in Colombia, staying in the range observed for this species, which has the highest prevalence among the other Saurocytozoon species. Records before this study showed microscopy prevalences ranging from 20 to 80% for P. (S) tupinambi comb. nov. in teiids from South America (Lainson and Shaw, Reference Lainson and Shaw1969; Telford, Reference Telford1978; Picelli et al., Reference Picelli, Ramires, Masseli, Pessoa, Viana and Kaefer2020), 6 to 14% of P. (S). mabuyi comb. nov. in skinks from South America and Southeast Asia (Lainson et al., Reference Lainson, Landau and Shaw1974a; Telford, Reference Telford1983), and 3% of P. (S). agamidorum comb. nov. in agamids from Southern Asia (Telford, Reference Telford2013).
Parasitemia intensity data is absent in most of the studies with Saurocytozoon. Telford (Reference Telford1978) noted a low parasitemia (below 1%) for P. (S) tupinambi comb. nov., suggesting chronic infections in a T. teguixin from Venezuela, and Telford (Reference Telford1983) marked a parasitemia ranging from 0.2 to 2.0% for P. (S). mabuyi comb. nov. from Southeast Asia. Here, parasitemia values were below 0.2%. Although the sampling sites from Colombia and Brazil were in tropical ecosystems, they were in different landscapes. The Colombian Orinoquia region (Eastern Plains or Llanos Orientales) is a plain lowland area comprising flooded savannas and gallery forests characterized by annual rainfall of 2400–2600 mm and mean temperature of 25°C (Angulo-Silva et al., Reference Angulo-Silva, Castellanos-Domínguez, Flórez-Martínez, Esteban-Adarme, Pérez-Mancipe, Farfán-García and Luna-Marín2016). Whereas in Brazil, the study was in an Amazonian rainforest region, which varies from primary and secondary forests to open areas, characterized by a humid tropical climate with annual rainfall of 2340–2630 mm and mean annual temperature of approximately 26°C with (Laurance et al., Reference Laurance, Camargo, Luizão, Laurance, Pimm, Bruna, Stouffer, Williamson, Benítez-Malvido, Vasconcelos and Van Houtan2011). The ecological differences seem not to affect parasite transmission significantly, probably because T. teguixin is abundant in these regions (Ribeiro-Junior and Amaral, Reference Ribeiro-Junior and Amaral2016), in addition to the presence of suitable vectors in these locations (Almeida et al., Reference Almeida, Belchior, Batista, Guimarães, Maitra, Ríos Velasquez, Izzo and Pessoa2023).
A recent review of leucocytozoids (Valkiūnas and Iezhova, Reference Valkiūnas and Iezhova2023) recognized P. (S) tupinambi comb. nov. and P. (S). mabuyi comb. nov. as valid species, but the validity of P. (S). agamidorum comb. nov. remains unclear. These species are distinguished primarily by the large gametocyte size and details of host cell distortion, aiding microscopic identification (Telford, Reference Telford2013). Lymphocytes, monocytes, and occasionally immature erythrocytes were reported as host cells for these species (Lainson and Shaw, Reference Lainson and Shaw1969; Telford, Reference Telford2013). Here, P. (S.) tupinambi comb. nov. host cell types were not determined. Although most host cells are likely to be leukocytes, the possibility that some are erythrocytes, as only deformed host cells were visualized here, cannot be discarded, hampering the identification of host-cell origin (Valkiūnas and Iezhova, Reference Valkiūnas and Iezhova2023). These features are common in Leucocytozoon parasites because merozoites of leucocytozoids lack a pellicle, making gametocytes fragile and easily deformed in blood films (Valkiūnas and Iezhova, Reference Valkiūnas and Iezhova2023). Thus, it would be relevant to identify further if merozoites of P. (S.) tupinambi comb. nov. also lacks this structure.
While other erythrocytic Plasmodium parasites that infect lizards, like the subgenera Sauramoeba Garnham, 1966 and Lacertamoeba Telford, 1988, are diverse and cosmopolitan (Telford, Reference Telford2009, Reference Telford2013), Saurocytozoon parasites are found in a few lizard host species (teiids, skinks, and agamids) but in locations of South America and Asia (Lainson et al., Reference Lainson, Landau and Shaw1974a; Telford, Reference Telford1978, Reference Telford1983, Reference Telford2013). However, the available distribution data may be due to insufficient sampling efforts rather than other factors since they are based on the scarce microscopy detection of these parasites in a few biogeographic regions. Here, we expand the known geographic range of P. tupinambi comb. nov. by reporting new localities in Brazil and their first occurrence in Colombia. The three countries, Brazil, Colombia, and Venezuela, where P. (S) tupinambi comb. nov. has been recorded to border each other in northern South America, forming a continuous environmental gradient of tropical landscapes where the host species, T. teguixin, is widespread and abundant (Ribeiro-Junior and Amaral, Reference Ribeiro-Junior and Amaral2016). Therefore, P. (S.) tupinambi comb. nov. is probably more widely distributed than current records suggest, and its presence in the entire distribution area of these hosts in South America should be explored.
In addition, our study represents the first genetic assessment of haemoparasites in T. teguixin. Although recognized as hosts for numerous protozoan species (Telford, Reference Telford2009), these teiid lizards have been largely overlooked in modern parasitological research. While previous studies have identified haemosporidians in golden tegus, these assessments have primarily relied on morphological characteristics (Picelli et al., Reference Picelli, Ramires, Masseli, Pessoa, Viana and Kaefer2020). Thus, combining morphological and genetic data provided a more accurate understanding of this haemosporidian parasite in these reptiles.
In conclusion, we provide the first genetic data to elucidate the taxonomic position of P. (S.) tupinambi. comb. nov. This study contributes to a better understanding of the diversity of malaria parasites of the genus Plasmodium by providing molecular evidence that species of the subgenus Saurocytozoon are malaria parasites but not leucocytozoids. Interestingly, the erythrocytic merogony – the typical stage of development of malaria parasites in vertebrates – was not found in lizards naturally infected P. (S.) tupinambi comb. nov. However, the close phylogenetic relationships of P. (S.) tupinambi. comb. nov. with other Plasmodium species indicate that the multiplication in the blood might occur and can be detected if more delicate studies on the life cycle are carried out, which is an important task for current wildlife parasitology research aiming to better understand the diversity of malaria pathogens. Thus, combining morphological and genetic data markedly improves an accurate understanding of haemosporidian parasite diversity in reptiles.
Supplementary material
The supplementary material for this article can be found at https://doi.org/.10.1017/S0031182025000381.
Data availability statement
Sequence data are available at GenBank accessions: PQ680045–PQ680069 (partial cytb); and PQ680070–PQ680073 (mitochondrial genome). Lizard specimen vouchers are deposited in the Zoological Collections of Universidade Federal do Amazonas (CZPB-RP 1051) and Instituto Nacional de Pesquisas da Amazonia (INPA-H037419-22). Haemosporidian vouchers are deposited in the Universidade Federal de Minas Gerais (UFMG31; UFMG197) and at Biological Collection Grupo de Estudio Relación Parásito Hospedero (GERPH) at Universidad Nacional de Colombia-Bogotá (GERPH:CAH311, GERPH:CAH359, GERPH:CAH765 and GERPH:CAH400).
Acknowledgements
We are grateful to Giulliana Appel, Adriane C. Ramires, Gabriel S. Masseli, ‘Juruna’ Ocírio Pereira, and the field team of EDTA for the help with lab and fieldwork in Brazil. To Alejandra Quiroz, Gustavo Rodriguez, and Karly Ojeda from the GERPH for their assistance in the lab. Luz Mary Velandia from the LEMO group thanks her valuable help in the field and in capturing lizards in Casanare, Colombia. We also thank Dr. Alan Warren former NHM curator for providing access to the hapantotypes of S. tupinambi, Dr. Robson Ávila for kindly providing the lizard images used in the figures, Júlia González for the silhouettes design, Paulo Vedovello for generating the map, and Dr. Aaron M. Bauer for his comments on the manuscript.
Author contributions
A.M.P., E.M.B, and N.E.M. conceived and designed the study. A.M.P., O.A.R.F., L.J.C.H., and N.E.M. conducted fieldwork. A.M.P., A.D.G., P.H.O.P., E.M.B., and N.E.M. performed the microscopic analysis. M.A.P. and A.A.E. performed molecular and phylogenetic analysis. A.M.P., M.A.P., F.C.F., A.A.E., E M.B., and N.E.M. processed the data, interpreted the results, and worked on the manuscript. G.V. examined hapantotypes of P. (S.) tupinambi, provided photographs of the parasite, and reviewed the manuscript. I.L.K., F.A.C.P., L.A.V., L.J.C.H., A.A.E., and N.E.M. were responsible for funding acquisition, supervision, and project administration. All authors took part in the preparation, revised, and approved the final version of the manuscript.
Financial support
This study in Brazil was financed in part by the Brazilian National Council for Scientific and Technological Development (CNPq Universal 461.573/2014-8 to I.L.K. and 429.132/2016-6 to L.A.V.) and Excellence Program in Basic and Applied Health Research (PROEP FIOCRUZ FAPEAM 001/2014 to F.A.C.P.). We also thank the Biological Dynamics of Forest Fragments Project (BDFFP) Thomas Lovejoy Research Fellowship Program for fieldwork support to A.M.P. This is the publication number 881 in the BDFFP technical series. In Colombia, this research was funded by SGR Project ‘Investigación para el avance en el conocimiento y desarrollo de capacidades regionales en la vigilancia de agentes potencialmente patógenos de riesgo para la población de Casanare’ [Code BPIN 202100010033]. We are grateful to The National Science Foundation (NSF) (grants NSF 2146653 to A.A.E. and 2146654 postdoctoral fellowship to A.M.P.). F.A.C.P. and I.K. thank to CNPq for their productivity scholarships.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
All procedures involving animals in this study in Brazil were approved by the ethics committee on animal use from Universidade Federal do Amazonas (protocol number 012/2016) and were authorized by the Brazilian Ministry of the Environment (SISBIO number 53851 and SISGEN AA6199D; A814F23). In Colombia, the procedures were approved by the Ethics Committee of the Departamento de Ciencias Básicas of the Universidad Internacional del Trópico Americano, Yopal, Casanare, Colombia by Act 001 of November 16, 2021.