Introduction
Nematodes of the genus Syphacia Seurat, 1916 (Oxyuridae Cobbold, 1864: Syphaciinae Railliet, 1916) are among the most common members of the helminth communities in wild rodents worldwide (Roman, Reference Roman1951), but perhaps best known are the two species that parasitize laboratory rodents and their free-living conspecifics: S. obvelata (Rudolphi, 1802) in house mice (Mus spp.) and S. muris Yamaguti, 1935 in rats (Rattus spp.). In the western Palearctic wood mice (also referred to as long-tailed field mice, Apodemus sylvaticus) and yellow-necked field mice (A. flavicollis) are parasitized by two species, S. stroma (Linstow, 1884) Morgan, Reference Morgan1932 and S. frederici Roman, Reference Roman1945. Sympatric voles are also typically parasitized by three species, S. petrusewiczi Bernard, Reference Bernard1966 in bank voles (Myodes glareolus), S. montana Yamaguti, 1943 in European snow voles (Chionomys nivalis) and European pine voles (Microtus subterraneus; see Tenora et al., Reference Tenora, Quentin and Durette-Desset1974) and S. nigeriana Baylis, Reference Baylis1928 in other Microtus spp. [M. agrestis (northern short-tailed field vole), M. rozianus (Portuguese field vole), M. lavernedii (Mediterranean field vole), M. arvalis (common vole) and M. oeconomus (root or tundra vole)]. In addition to the seven listed here, four other species of Syphacia have also been recorded from European rodents, namely S. arvicolae Sharpilo, Reference Sharpilo1973, S. vandenbrueli Bernard, Reference Bernard1966, S. agraria, Sharpilo, Reference Sharpilo1973 and the fourth S. baylisi, which is a synonym of S. muris (see Tenora and Mészáros, Reference Tenora and Mészáros1975). The hosts of S. vandenbrueli and S. arvicolae (the Eurasian harvest mouse, Micromys minutus and the European water vole, Arvicola spp., respectively), are endemic in the British Isles but to the best of our knowledge their Syphacia spp. have never been recorded locally, although it is worth noting that Tenora et al. (Reference Tenora, Barus, Wiger and Genov1979a) recommended that this latter species should be synonymized with S. nigeriana. The striped field mouse, Apodemus agrarius (host of S. agraria) does not exist in the wild in the British Isles.
Syphacia nigeriana was first described by Baylis (Reference Baylis1928) on the basis of worms recovered from five species of West African rodents Taterillus gracilis (slender gerbil from the district of Kano, Nigeria), Gerbilliscus kempi (northern savanna gerbil from Ibadan, Nigeria as Taterona kempi), Praomys tulbergi (Tullberg's soft-furred mouse from Adu, Nigeria), Mastomys erythroleucus (Guinea multimammate mouse from Ife, Nigeria) and Lemniscomys striatus (typical striped grass mouse from Adu, Nigeria). Although he did not designate types in his paper, specimens of Syphacia in Baylis' material labelled as from T. kempi from Ibadan, Nigeria registered in the Natural History Museum London (NHML) as 129.1.24. 26, 27, 31 are labelled paratypes (specimens additional to the holotype on which the description is based).
Subsequently, after studying specimens from a wide range of hosts, and extending the host record list to Microtus spp. from North America and adding six additional rodent hosts from Africa, Quentin (Reference Quentin1971) wrote: ‘As a consequence we think that the Syphacia of holarctic Microtus, which present the cephalic and genital structures morphologically identical to those from Syphacia of Gerbillidae and African Muridae, belong to the same species, Syphacia nigeriana’ (Translation from the original in French, Quentin, Reference Quentin1971, p. 32).
Thereafter, European parasitologists considered the Syphacia species parasitizing European rodents of the genus Microtus to be mainly S. nigeriana (see Sharpilo, Reference Sharpilo1973; Tenora and Mészáros, Reference Tenora and Mészáros1975; Mészáros, Reference Mészáros1977; Tenora et al., Reference Tenora, Wiger and Baruš1978; Mészáros and Murai, Reference Mészáros and Murai1979; Tenora et al., Reference Tenora, Andreassen, Hindsbo and Lodal1991; Tenora and Staněk, Reference Tenora and Staněk1994, Reference Tenora and Staněk1995). Accordingly, S. nigeriana has been recorded in M. agrestis in Denmark (Tenora et al., Reference Tenora, Andreassen, Hindsbo and Lodal1991), Norway (Wiger et al., Reference Wiger, Barus and Tenora1978; Tenora et al., Reference Tenora, Wiger and Barus1979b), Finland (Tenora et al., Reference Tenora, Henttonen and Haukisalmi1983; Haukisalmi et al., Reference Haukisalmi, Henttonene and Pietiäinen1994), in M. rozianus or M. lavernedii in Spain (as M. agrestis, Mas-Coma et al., Reference Mas-Coma, Tenora and Racomara1978; Feliu et al., Reference Feliu, Renaud, Catzeflis, Hugot, Durand and Morand1997), in M. arvalis in Romania (Mészáros and Murai, Reference Mészáros and Murai1979), in M. oeconomus in Norway (Tenora et al., Reference Tenora, Mészáros and Wiger1977), in C. nivalis in Spain (Mas-Coma et al., Reference Mas-Coma, Tenora and Racomara1978) and as an occasional parasite of M. glareolus in Norway (Tenora et al., Reference Tenora, Mészáros and Wiger1977, Reference Tenora, Wiger and Baruš1978; Tenora and Mészáros, Reference Tenora and Mészáros1975; Wiger et al., Reference Wiger, Lien and Tenora1976) and Hungary (Mészáros, Reference Mészáros1977).
Like most oxyurid nematodes, species of Syphacia are believed to show strong co-evolutionary relationships with their hosts, and hence are considered to be generally host-specific (Adamson, Reference Adamson1989; Hugot, Reference Hugot1999; Garcia et al., Reference Garcia, Melin, Aureli and de Leon GP2018). It was our view, given the distance and the intervening terrain involved, that it was highly unlikely that the parasite in European Microtus spp. was panmictic with those parasitizing wild rodents of quite different host genera in Africa. On these grounds, we questioned whether the species of Syphacia found in European Microtus spp. had been erroneously assigned as S. nigeriana and considered that a genetic analysis was warranted.
Syphacia petrusewiczi was described by Bernard (Reference Bernard1966) from bank voles, M. glareolus, trapped near Mikołajki in the Mazury Lake District region of north-eastern Poland, where long-term surveys of bank vole helminth communities have been conducted in more recent years (Grzybek et al., Reference Grzybek, Bajer, Bednarska, Alsarraf, Behnke-Borowczyk, Harris, Price, Brown, Osborne, Siński and Behnke2015). Specimens of S. petrusewiczi in bank voles from this region of Poland therefore may be regarded as having come from close to the type locality (Bernard, Reference Bernard1966). Syphacia petrusewiczi is regarded as a Myodes specialist (Tenora and Mészáros, Reference Tenora and Mészáros1975), as emphasized, for example, by Mészáros (Reference Mészáros1978). Reported locations of S. petrusewiczi occurring in bank voles include Finland (Haukisalmi and Henttonen, Reference Haukisalmi and Henttonen1993), Lithuania (Mažeika et al., Reference Mažeika, Pailauskas and Balčiauskas2003; Skyrienė et al., Reference Skyrienė, Ulevičius and Samas2011), northern France (Ribas Salvador et al., Reference Ribas Salvador, Guivier, Xuéreb, Chavel, Cadet, Poulle, Sironen, Voutilainen, Henttonen, Cosson and Charbonnel2011), southern Italy (Milazzo et al., Reference Milazzo, Casanova, Aloise, Ribas and Cagnin2003), Serbia (Bjelić-Čabrilo et al., Reference Bjelić-Čabrilo, Popović, Šimić and Kostić2009, Reference Bjelic-Čabrilo, Kostic, Popovic, Cirkovic, Aleksic and Lujic2011), Spain (Feliu et al., Reference Feliu, Renaud, Catzeflis, Hugot, Durand and Morand1997; Ribas et al., Reference Ribas Salvador, Torre, Feliu, Arrizabalaga and Casanova2009) and both European and Asian Russia (Gorelysheva et al., Reference Gorelysheva, Efeykin, Yakushov and Sheftel2020). Shortly after Bernard's publication (Bernard, Reference Bernard1966), Quentin (Reference Quentin1969) described a variant of this species, which he regarded as sufficiently distinct to be considered a subspecies, named S. petrusewiczi rauschi, a parasite of Myodes rutilus dawsoni from Anchorage in Alaska, USA.
Female nematodes of the genus Syphacia are character poor, and species identity is largely based on male worms, which, although more character rich than females, are rare in most species and in some cases still unknown (Morgan, Reference Morgan1932; Ogden, Reference Ogden1971). Male worms are thought to be short-lived, inseminating females before they reach full size and then passing out of the host (Morgan, Reference Morgan1932; Adamson, Reference Adamson1994). Species-specific diagnostic characters in female worms have been described for some species (Stewart et al., Reference Stewart, Lowe, Smales, Bajer, Bradley, Dwużnik, Franssen, Griffith, Stuart, Turner, Zleśny and Behnke2018) but most are unsuitable for quantitative studies and some are difficult to detect with certainty, especially in preserved specimens, particularly if they have been fixed in 100% ethanol. Such specimens are likely to be only partially dehydrated and/or the tissues may be inadequately preserved and therefore deformed to varying extents. Hence, the literature is likely to contain cases of misidentification of species of Syphacia parasitizing particular hosts, and S. obvelata appears to be the most commonly misidentified species in this context, possibly because as a parasite of Mus spp. it is so well known [e.g. Sharpe (Reference Sharpe1964) and Kisielewska and Zubczewska (Reference Kisielewska and Zubczewska1973) both recorded S. obvelata in M. agrestis; Lewis (Reference Lewis1968) recorded S. obvelata in M. glareolus]. For these reasons, published host lists, relying primarily on morphological characters, have to be treated with some caution until all the species have been fully characterized genetically and carefully re-examined morphologically.
The helminth community of M. agrestis in the British Isles has been only poorly studied, and as far as we are aware, to date there is only one record of S. nigeriana from a population of M. agrestis in Kielder (Turner et al., Reference Turner, Beldomenico, Bown, Burthe, Jackson, Lamblin and Begon2014). Interestingly, there are no wild Microtus spp. in Ireland and bank voles were detected for the first time only in 1964, although they are thought to have been introduced from Germany in 1926 (White et al., Reference White, Perkins, Heckel and Searle2013). It is pertinent, however, that recently Loxton et al. (Reference Loxton, Lawton, Stafford and Holland2016) did not recover any Syphacia worms from bank voles from sites in Galway in Ireland. There are few records of the helminths of M. glareolus from the British Isles (Elton et al., Reference Elton, Ford, Baker and Gardiner1931; Sharpe, Reference Sharpe1964; Lewis, Reference Lewis1968; Canning et al., Reference Canning, Cox, Croll and Lyons1973; Loxton et al., Reference Loxton, Lawton, Stafford and Holland2016), and to date all those authors that have recorded Syphacia in bank voles from sites on the British mainland have assigned them to S. obvelata, some even after Bernard's (Reference Bernard1966) paper describing S. petrusewiczi (Lewis, Reference Lewis1968; Canning et al., Reference Canning, Cox, Croll and Lyons1973). There are also several reports of S. obvelata from bank voles (Tenora and Zejda, Reference Tenora and Zejda1974) and Microtus spp. on the European mainland, which in the light of our current understanding of this species are likely also to be misidentifications (Tenora, Reference Tenora1972; Tenora et al., Reference Tenora, Murai and Mészáros1973 and citations therein; Prokopič, Reference Prokopič1973).
In this paper, we examine morphologically Syphacia species isolated from both M. glareolus and Microtus spp. voles, from several disparate sites in the British Isles, and we provide the genetic signature of each isolate that we encountered, based primarily on the rDNA internal transcribed spacers (ITS1-5.8S-ITS2) region and for representative isolates also on the 18S rDNA (SSU) region and the mitochondrial cytochrome c oxidase subunit 1 (cox1) gene locus. We also include genetic data on worms recovered from rodents of the same genera from the European mainland, and from other species from North America and West Africa to help clarify the species status of Syphacia in voles from the British Isles. We test Baylis’ original finding (Baylis, Reference Baylis1928) and Quentin's subsequent work (Quentin, Reference Quentin1971) indicating that S. nigeriana is a generalist species capable of infecting a range of different rodent genera, as widely distributed as from West Africa, Europe and even from North America (Quentin, Reference Quentin1971).
Materials and methods
Host nomenclature
We refer to M. glareolus rather than Clethrionomys glareolus, following Wilson and Reeder (Reference Wilson and Reeder2005) and Carleton et al. (Reference Carleton, Musser, Pavlinov, Averianov and Abramson2003), and despite the recent proposal that Clethrionomys has priority for the generic name of all red-backed voles (Kryštufek et al., Reference Kryštufek, Tesakov, Lebedev, Bannikova, Abramson and Georgy Shenbrot2020) to maintain parity with recent publications in the field of parasite ecology. We also refer to wood mice, rather than long-tailed field mice for A. sylvaticus.
Sources of isolates
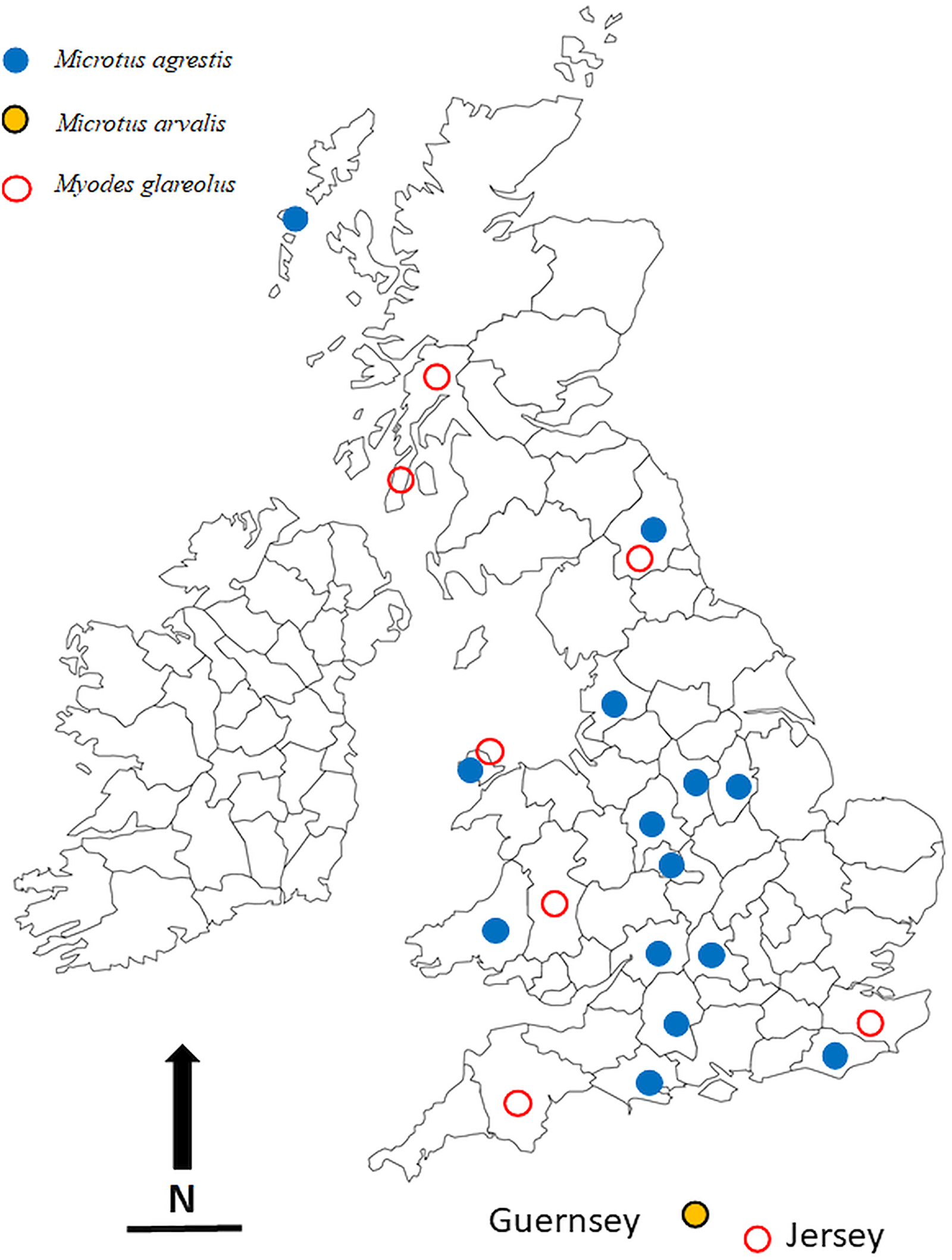
The locations in which voles were trapped in the British Isles are given in Table 1 and are illustrated in Fig. 1. Most of these were obtained through field trips across the British Isles, but we also made use of a large collection of voles, both M. agrestis and M. glareolus (preserved in 100% ethanol) held at the National Museums Collection Centre, Edinburgh, UK. In addition to the voles that were infected with Syphacia, we also processed bank voles from 22 other locations across the country but found these to be not infected with Syphacia. Regardless of the source of the voles, all were examined for nematodes using the same dissection protocol and any oxyurids recovered were fixed in 100% ethanol and stored in 80–100% ethanol, and in some cases frozen, for future study. Although fixation in 100% ethanol is not ideal for morphological examination, it does allow the use of worms obtained from each host individual to be separated into two samples, one for morphological examination and one for molecular analysis. In addition, all the material registered in the Natural History Museum, London, as S. nigeriana, consisting of material from several hosts from African localities, was examined for comparative purposes.
The locations in the British Isles in which rodents for this study were trapped. Symbols are plotted on the outlines of the relevant administrative counties in the British Isles in which the animals were trapped. Blue filled in circles = Microtus agrestis; red open circles = Myodes glareolus; black, yellow filled circles = Microtus arvalis. Scale bar = 100 km, and N = North.
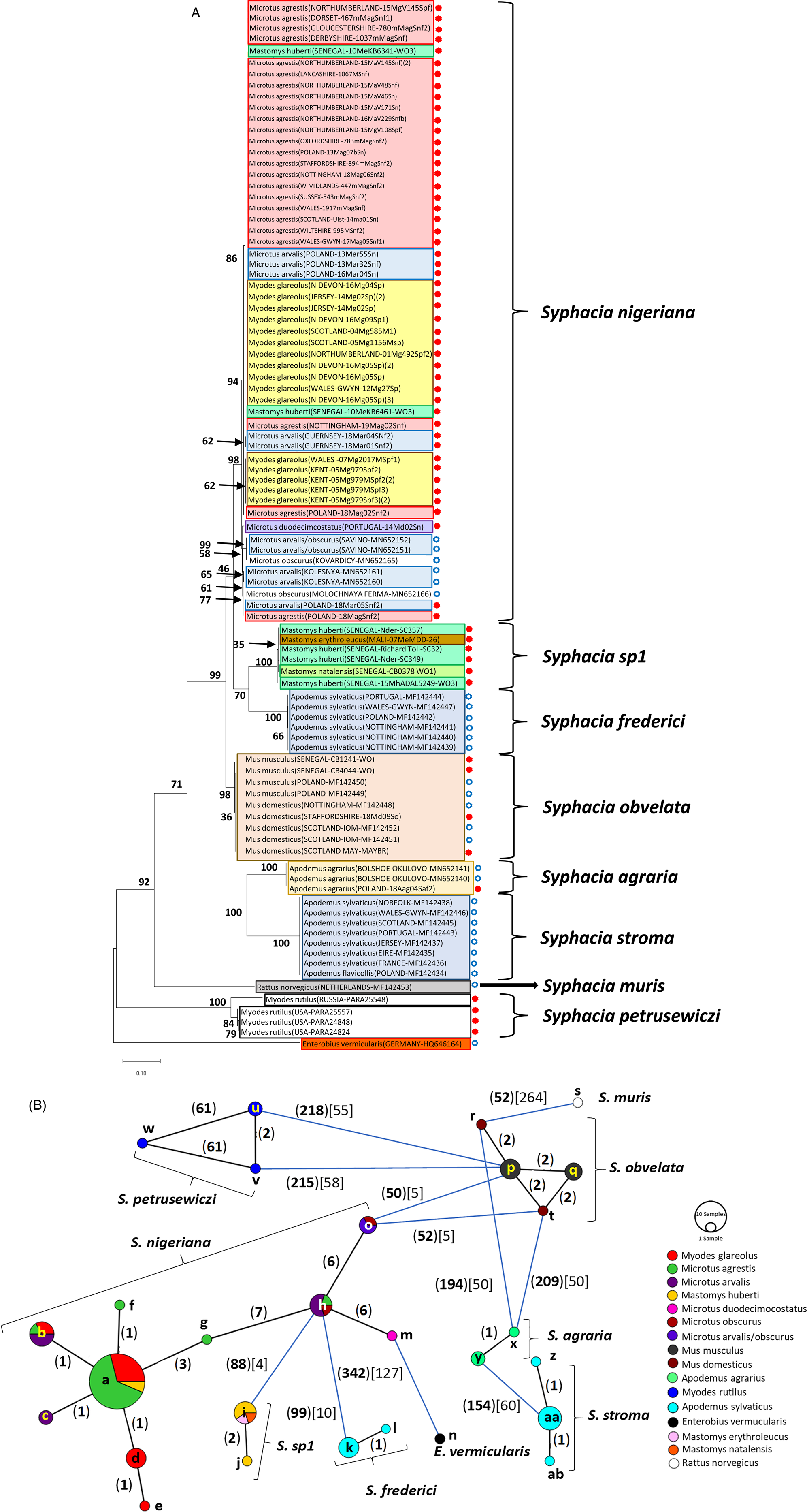
Molecular phylogenetic analysis of rDNA (ITS-1-5.8S-ITS2). (A) Molecular phylogenetic tree of Syphacia from murid and cricetid hosts following maximum likelihood (ML) analysis with 1000 bootstrap replicates. Scores at junctions represent bootstrap support for that junction. Scale bar is proportional to the genetic distance in substitutions per site. New sequences are marked by a red filled in circle and those taken from GenBank in blue open circles. (B) Minimum spanning network analysis. Bold numbers in round brackets on the lines show the number of single nucleotide polymorphisms between nodes and those in regular font and square brackets give the number of indels. Colours represent different rodent host species, and the newly genotyped isolates in each lettered node are listed in Table 1.
Molecular phylogenetic analysis of 18S rDNA (SSU). (A) Molecular phylogenetic tree of Syphacia from murid and cricetid hosts following maximum-likelihood (ML) analysis with 1000 bootstrap replicates. Scores at junctions represent bootstrap support for that junction. Scale bar is proportional to the genetic distance in substitutions per site. (B) Minimum spanning network analysis. Bold numbers in round brackets on the lines show the number of single nucleotide polymorphisms between nodes and those in regular font and square brackets give the number of indels. Colours represent different rodent host species (see legend to Fig. 2B for key), and the newly genotyped isolates in each lettered node are listed in Table 1.
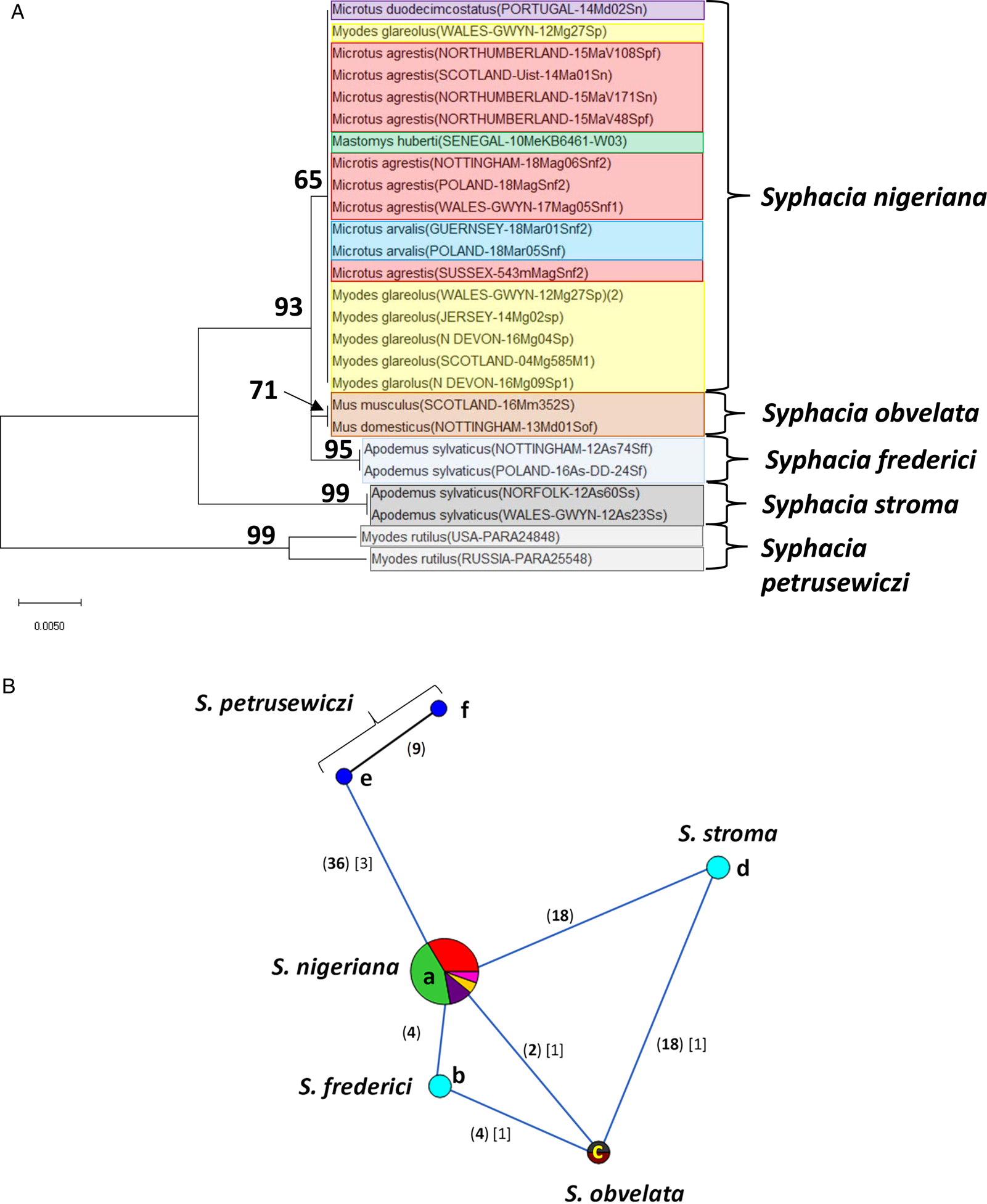
Molecular phylogenetic analysis of the cox-1 gene. (A) Molecular phylogenetic tree of Syphacia from murid and cricetid hosts following maximum-likelihood (ML) analysis with 1000 bootstrap replicates. Scores at junctions represent bootstrap support for that junction. Scale bar is proportional to the genetic distance in substitutions per site. (B) Minimum spanning network analysis. Bold numbers in round brackets on the lines show the number of single nucleotide polymorphisms between nodes and those in regular font and square brackets give the number of indels. Colours represent different rodent host species (see legend to Fig. 2B for key), and the newly genotyped isolates in each lettered node are listed in Table 1.
Molecular genetic comparison of isolates
DNA was isolated from individual worms using Extracta™ DNA prep kits (Quantabio) or DirectPCR lysis buffer (Viagen Biotech) according to the manufacturer's instructions. The cox-1 gene was amplified using primers (Forward: 5′-TGGTCTGGTTTTGTTGGTAGTT-3′, Reverse: 5′-AACCACCCAACGTAAACATAAA-3′; Okamoto et al., Reference Okamoto, Urushima, Iwasa and Hasegawa2007) spanning part of the coding sequence and yielding up to 684 bp amplicons, although some were truncated because of PCR amplification issues. The rDNA region consisting of ITS-1, 5.8S gene and ITS-2 (~700–750 bp) was amplified using the universal NC5 forward (5′-GTAGGTGAACCTGCGGAAGGATCATT-3′) and NC2 reverse primers (5′-TTAGTTTCTTTTCCTCCGCT-3′; Newton et al., Reference Newton, Chilton, Beveridge, Hoste, Nansen and Gasser1998, Table 2). rDNA samples that failed to amplify were further analysed with nested forward (5′-GCTGTTTTCTTACATGCTATAAACG-3′) and reverse (5′-TATCATTACGTGGTTGACAGACA-3′) primers. The 18S rDNA (SSU) region was amplified using primers nSSU_F_03 (5′-GCTTGTCTCAAAGATTAAGCCATGC-3′) and nSSU_R_24 (5′-CCCCRRTCCAAGAATTTCACCTC3′; Blaxter et al., Reference Blaxter, De Ley, Garey, Liu, Scheldeman, Vierstraete, Vanfleteren, Mackey, Dorris, Frisse, Vida and Thomas1998 and http://www.nematodes.org/research/barcoding/sourhope/nemoprimers.shtml), resulting in a maximum amplicon length of 781 bp.
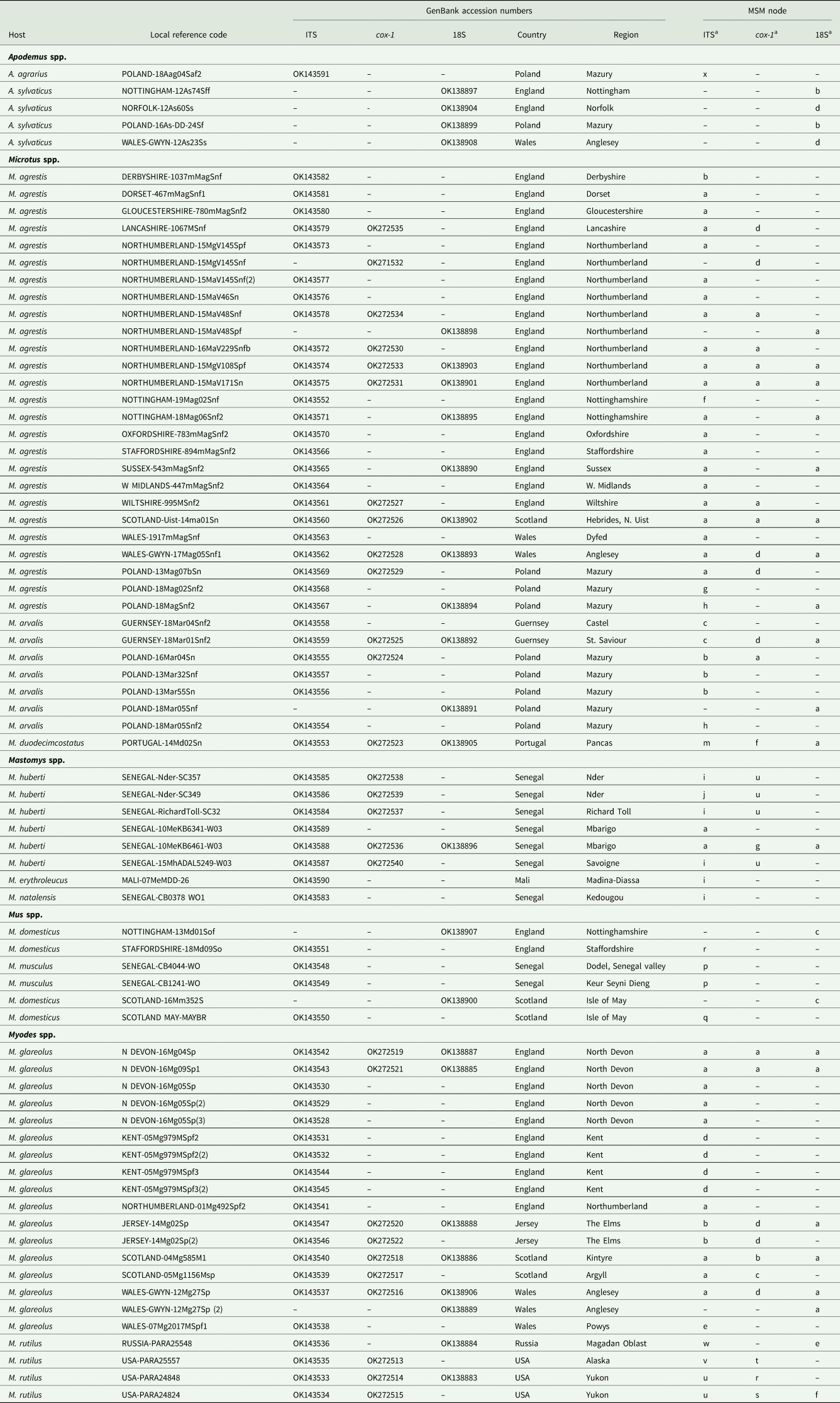
Sequences from GenBank included in this study
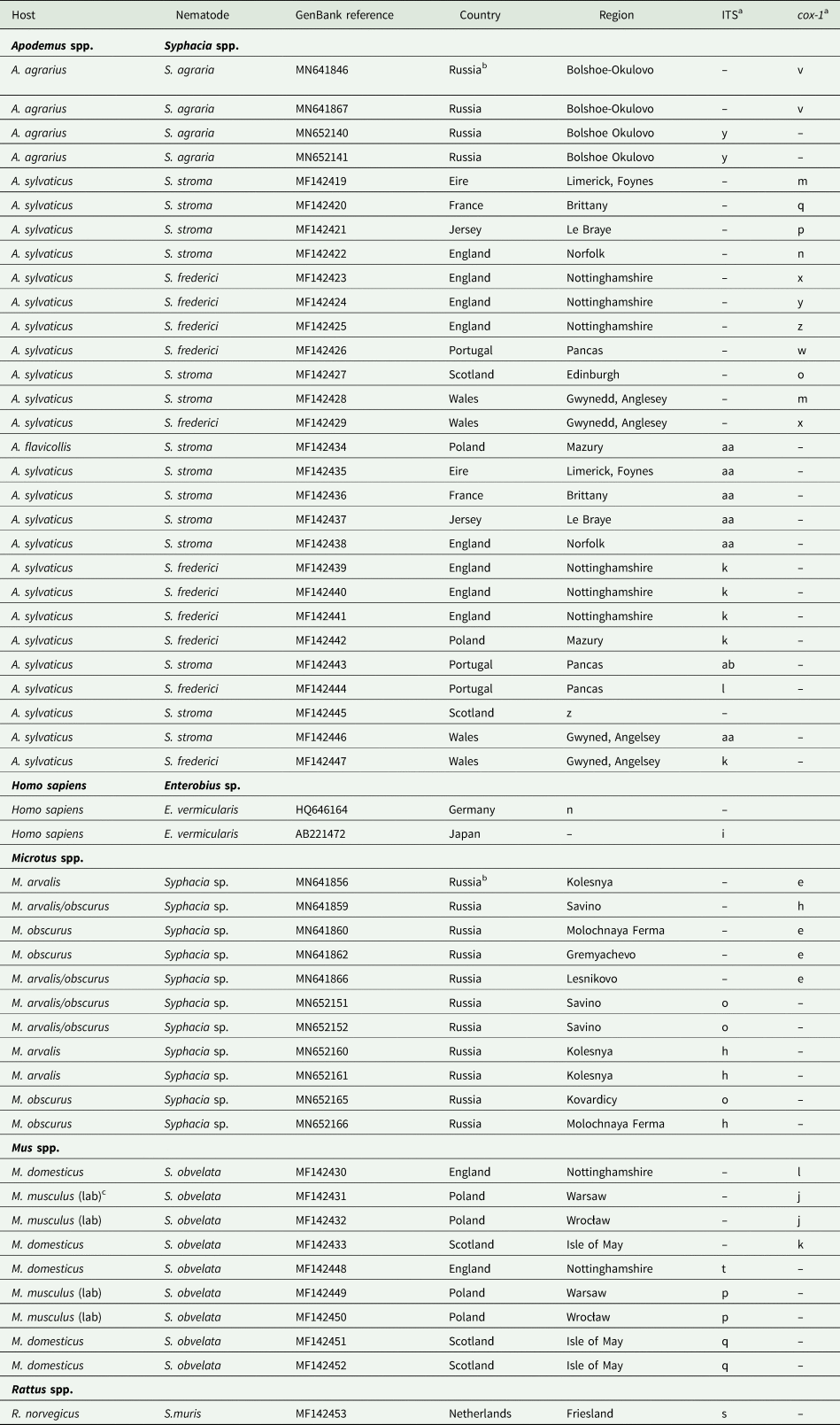
a Letters correspond to nodes in Figs 2B and 4B. Within each column isolates with the same letter have identical sequences.
b For further details of isolates from Russia recorded in GenBank, see Gorelysheva et al. (Reference Gorelysheva, Gromov, Lavrenchenko and Spiridonov2021).
c (lab) These were BALB/c mice from the Universities of Warsaw and Wrocław
PCR reactions used Phusion Taq Polymerase according to manufacturer's instructions with 0.5 μ m of the forward and reverse primers, <250 ng of template DNA and nuclease-free water to a total volume of 25 μL. Thermal cycling conditions for cox-1 were: denaturation for 30 s at 98°C, then 35 cycles of 30 s at 98°C, 1 min at 52°C and 1 min 30 s at 72°C, with a final extension time of 5 min at 72°C before being held at 4°C. Thermal cycling conditions for the rDNA, rDNA-Nested and SSU regions were identical to cox-1 with the exception of an annealing temperature of 60°C in both cases. Amplification in all PCR reactions was confirmed by visualization on a 1x SYBRSafe™ stained 1.5% agarose gel. PCR products were purified using AMPureXP beads according to the manufacturer's instructions and the final DNA concentration was estimated by Nanodrop before dilution with nuclease-free water to the required concentration for sequencing. Sequencing primers, identical to the amplification primers, were diluted to the required concentration with nuclease-free water and supplied to Source Bioscience or Eurofins, along with PCR products, for Sanger sequencing. Chromatograms were inspected visually for ambiguities and sequences trimmed to the 3′ end of the primers.
Sequence alignments were prepared using ClustalW within the MEGA X package (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018) followed by visual inspection. Phylogenetic analysis was performed in MEGA X using maximum likelihood (ML) with a partial deletion threshold set at 95% in MEGA (v10). Intraspecies (each separate parasite species based on phylogenetic analysis) and interspecies (all sequences) minimum spanning network (MSN) plots were produced in PopART (Leigh and Bryant, Reference Leigh and Bryant2015) with an epsilon of zero (Bandelt et al., Reference Bandelt, Forster and Röhl1999). Intraspecies MSNs were linked together based on the interspecies MSN, but indel and single nucleotide polymorphism (SNP) numbers were added in manually given that sites with indels are ignored in MSN model calculations.
In addition to the novel sequences generated in this project, we exploited genetic records of rDNA and cox-1 from Syphacia in GenBank. Some of these were from our own earlier work (Stewart et al., Reference Stewart, Lowe, Smales, Bajer, Bradley, Dwużnik, Franssen, Griffith, Stuart, Turner, Zleśny and Behnke2018), and those for S. agraria and isolates from Microtus spp. from Russia were from the work of Gorelysheva et al. (Reference Gorelysheva, Gromov, Lavrenchenko and Spiridonov2021). Since the focus of the current study was on Syphacia spp. in Microtus spp., we used only representative sequences from Apodemus and Mus spp. but more information on how other isolates from these hosts fit into the phylogenetic trees can be found in Stewart et al. (Reference Stewart, Lowe, Smales, Bajer, Bradley, Dwużnik, Franssen, Griffith, Stuart, Turner, Zleśny and Behnke2018). Because of the large number of identical sequences deposited by Gorelysheva et al. (Reference Gorelysheva, Gromov, Lavrenchenko and Spiridonov2021) for isolates from Russian voles, we selected for the current analyses only representative sequences for the clades referred to in Gorelysheva et al. (Reference Gorelysheva, Gromov, Lavrenchenko and Spiridonov2021), in order to show how our novel sequences related to those from Russian rodents.
Voucher sequences, including all sequences generated by this study and included in the current paper, have been deposited in GenBank (Table 1). Voucher numbers for accession to worms from M. rutilus held at the Museum Southwestern Biology, University of New Mexico, are as follows: Isolate from Russia, MSB:Para:25548; isolate from Alaska, USA, MSB:Para 25557; isolates from the Yukon, USA, MSB:Para:24824 and 24848.
Microscopical analysis of isolates
All the specimens from hosts collected for this study and selected for morphological examination were cleared in lactophenol and viewed as temporary wet mounts using an Olympus BH-2 compound microscope. Light micrographs were taken using the same microscope. Measurements in micrometres were taken with the aid of an ocular micrometer and are given as the range where more than two measurements were taken. En face and transverse sections of the anterior body of representative specimens were prepared using a cataract scalpel and mounted in polyvinyl lactophenol for examination. All such specimens examined morphologically for this study, except those used for sectioning, were deposited in the South Australian Museum (SAMA) Adelaide, South Australia (Voucher numbers for specimens from Senegal are AHC 48827–48830, and all others AHC 48791–48826 and AHC 48831–48837).
Results
Molecular genetic comparison of worms
In total, we provide 118 new sequences [64 for rDNA (ITS-1-5.8S-ITS-2), 28 for cox-1 and 26 for 18S rDNA (SSU)], which we have combined with 52 sequences from GenBank [29 for rDNA (ITS-1-5.8S-ITS-2) and 23 for cox-1].
rDNA (ITS-1-5.8S-ITS-2) region
The ML phylogenetic tree for rDNA (ITS-1-5.8S-ITS-2), illustrated in Fig. 2A, shows that Syphacia isolates from rodents trapped throughout the British Isles had mostly identical genetic sequences, irrespective of whether the worms were isolated from field, bank or common voles (the latter only from Guernsey). A large proportion of the isolates from voles from the British Isles were identical, and these included isolates from the island of N. Uist in the Outer Hebrides, the most northerly location, to those from Devon in the south west of the British Isles, the island of Anglesey in Gwynedd, Wales in the west and Sussex in the southeast (Node a in Fig. 2B, and Table 1). This particular genotype was also identified in field voles from Poland and somewhat unexpectedly in two mice (Mastomys huberti) from Senegal in West Africa (Table 1). The latter was scrutinized very carefully to ensure that there was no possibility of any ambiguity in this finding.
Our data also show that there were some close variants of the node a sequence (Fig. 2B), including isolates from Kent (node d), Nottingham (node f), Jersey, Derbyshire, Poland (node b) and Guernsey (node c), differing by just one SNP in each, but a different locus, and from Powys in Wales [differing by an additional SNP (node e, Fig. 2B)]. Not surprisingly, some S. nigeriana from more distant locations showed greater SNP variation when compared to the main S. nigeriana clade [e.g. some worms from Poland showed more SNPs (up to 3, node g), Russia (10-16 SNPs, nodes o and h) and Microtus duodecimcostatus from Portugal (16 SNPs, node m)]. However, given the length of the amplicon (860 bp), these were all relatively minor variations within what appears to be a single species of S. nigeriana, when compared, for example, to S. muris which differed from the main S. nigeriana clade (node a) by 93 indels and 240 SNPs.
Syphacia isolates from North American and Russian M. rutilus formed their own distinct and distant clade (node u, v and w in Fig. 2B), differing by 215 SNPs and 58 indels from that representing S. obvelata (node p). All the worms from M. rutilus had been identified previously as S. petrusewiczi (New Mexico Museum). Isolates from other specimens of Mastomys formed their own distinct clade (Fig. 2A) and branch (Fig. 2B, nodes i and j) and likely represent a new, as yet undescribed Syphacia species from West African Mastomys spp. (referred to hereafter as Syphacia sp1). We recovered these genotypes of Syphacia from three different species of Mastomys. As Fig. 2A and B shows other recognized species of Syphacia (S. stroma, S. frederici, S. obvelata, S. muris and S. agraria) all formed their own distinct clades (Fig. 2A) and nodes (Fig. 2B) differing from node a isolates mostly by over 50 SNPs and in all cases also by indels. The only additional novel sequence in the current work was for S. agraria from a striped field mouse from Poland (Fig. 2B, node x), which proved to have a sequence that differed by just one SNP from that reported from the same host species from Russia (node y), and for an S. obvelata from the Senegal (Fig. 2B, node p) which had an identical sequence to isolates from laboratory mice from Poland but differed only by 2 SNPs from S. obvelata isolates from Scotland (node q) and by a different set of 2 SNPs from an isolate from Nottingham (node t).
18S rDNA (SSU)
Our analysis of the 18S rDNA region generated a phylogenetic tree that was largely congruent with that for the rDNA (ITS-1-5.8S-ITS-2) region (Fig. 3A). All the sequences we obtained from field, common (only on Guernsey) and bank voles, from locations in the British Isles were identical (Fig. 3B, node a). No variants were identified (Fig. 3A), pointing to this clade being a single species, i.e. S. nigeriana. Moreover, the 18S rDNA sequence from the isolate from M. huberti from the Senegal [that had an identical rDNA (ITS-1-5.8S-ITS-2) sequence to S. nigeriana] was clearly assigned to node a in Fig. 3B (and Table 2). The representative sequences from other recognized Syphacia spp. that we included in this analysis formed their own distinct clades with very high bootstrap values for junctions in the tree. Although sequence variation compared to S. nigeriana (node a) ranged from just 2 SNPs and 1 indel to S. obvelata, and then 18 SNPs and 1 indel to S. stroma (Fig. 3B), the greatest comparative sequence variation was for S. petrusewiczi from M. rutilus from North America (36 SNPs) and Russia (45 SNPs) and 3 indels in both cases compared to S. nigeriana.
cox-1
Analysis of the cox-1 locus produced a more complex picture (Fig. 4A and B). Again, it is clearly apparent that sequences from worms from many of the field, common and bank voles from the British Isles were identical or differed by just 1 SNP (nodes a and d in Fig. 4B). These included worms from M. agrestis and M. arvalis from Poland. In this analysis, the isolate from M. huberti (SENEGAL-10MeKB6461-W03), that had an identical rDNA (ITS-1-5.8S-ITS2) sequence to the S. nigeriana in node a (Fig. 2B), differed in its cox-1 sequence from the main S. nigeriana clade (node g in Fig. 4B) by 4 SNPs. The cox-1 sequence of the isolate from M. duodecimcostatus from Portugal (node f) differed from those in node a by 8 SNPs and those in node d by 5 SNPs. As expected, all the other recognized Syphacia spp. showed greater discrepancies in SNPs, each species forming a distinct clade in Fig. 4A, although some of the bootstrap values were low. Consistent with their status as a new species, the four isolates from M. huberti that had differed from S. nigeriana in rDNA sequences also had their own distinct sequence for the cox-1 locus (node u in Fig. 4B). As in the rDNA, S. petrusewiczi formed a distinct distant clade of its own (Fig. 4A), although the isolate from Alaska (USA-PARA25557, node t) differed by 35 SNPs from the Yukon isolate (USA-PARA24824; Fig. 4B, node s) and there was a single SNP distinguishing the two Yukon isolates (USA-PARA24824 vs USA-PARA24848).
Morphological comparison of specimens
All the worms examined from British and European localities, regardless of host species and locality, were identified as S. nigeriana on the basis of their morphology and measurements of key morphological features (Table 3). No consistent differences were found in the morphometrics of males or females between samples, as shown in Table 3. Characters which showed variability between individual samples of females included egg size and shape but such differences could not be related to a particular host species. For example, the longest eggs were found in an individual of M. arvalis from Guernsey, up to 128 μm and an individual of M. agrestis from the West Midlands of England, up to 132 μm long. The shortest eggs measured were from an individual of M. agrestis from Nottingham, 99 μm. All other egg lengths varied within these limits, showing no particular size pattern. Similarly, female tail length varied between 390 and 900 μm long, shorter tail lengths being found in immature females regardless of the origin of the sample. The same variability was seen in measurements of the distance between the anterior end and the vulva. By contrast, the morphometrics of the male samples showed less variability across locality and host species (Table 3).
Isolates of Syphacia species examined microscopically
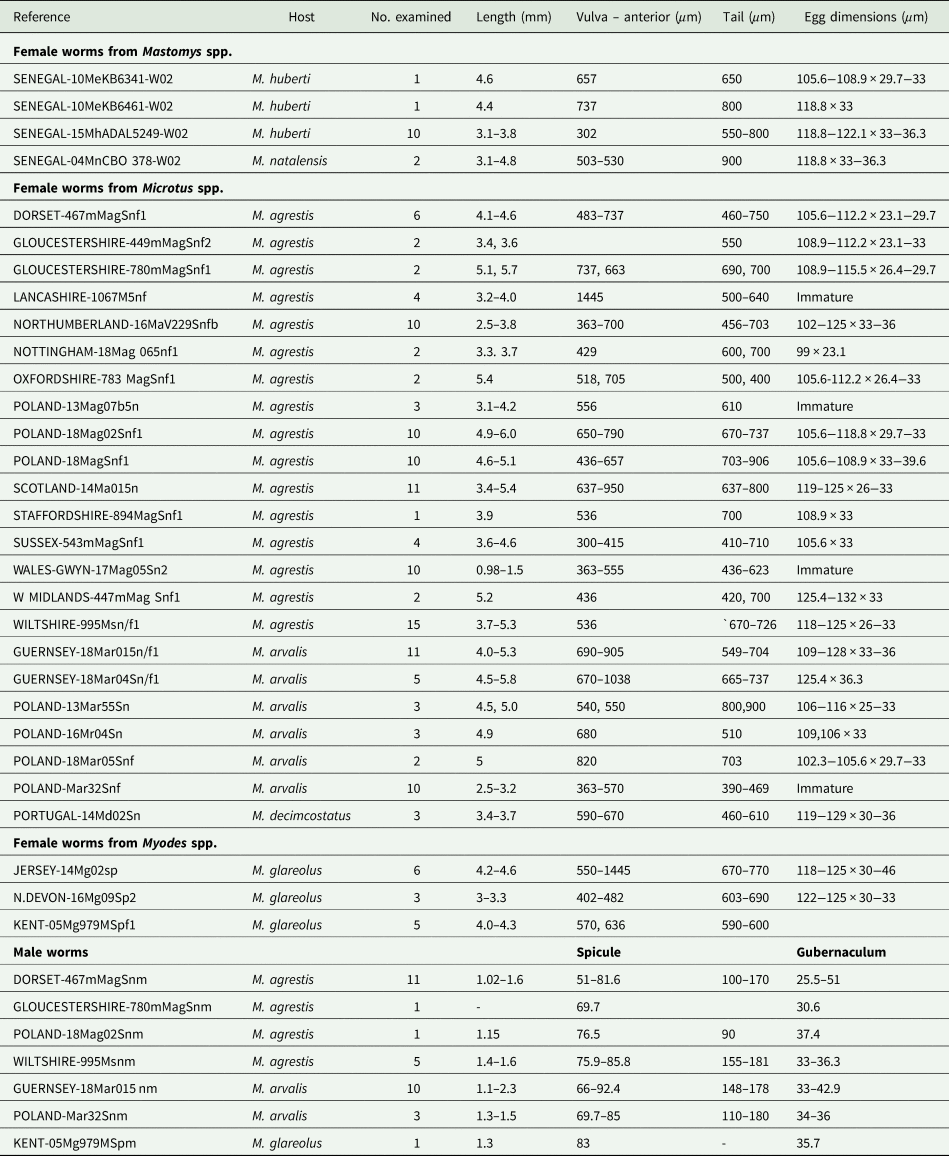
Nor could any consistent morphological differences be found between samples. All of the specimens had the typical key characters of S. nigeriana, including the cephalic structures, absence of cervical alae and presence of narrow lateral alae, that are used to differentiate both males and females of the species. The cephalic vesicle was a prominent feature of most but not all specimens although it should be noted that this character may be adversely affected by the fixation protocol used (Fig. 5A and B). The only character that showed some morphological variation was the shape of the vulva (Fig. 5B and C). Some Polish specimens from M. arvalis had an ornamented vulva, some Polish specimens from M. agrestis had a flap covering the vulva, and some specimens from M. glareolus from North Devon had a protruding vulva.
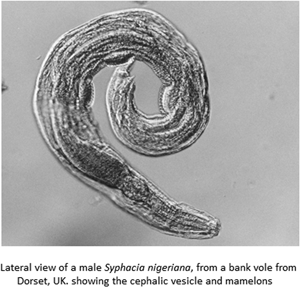
(A) Lateral view of the anterior end of female Syphacia nigeriana from Microtus agrestis (NORTHUMBERLAND 16MaV229Snfb), showing the cephalic vesicle (arrow). (B) Lateral view of the anterior end of a female S. nigeriana from M. agrestis (Wales GWYNLLANB 17Mag05Sn2), showing the cephalic vesicle (large arrow) and non-protruding vulva (small arrow). (C) Lateral view of the anterior end of a female S. nigeriana from M. agrestis (POLAND – 18MagSnf1), showing the cephalic vesicle (large arrow) and protruding vulva (small arrow). (D) Lateral view of male S. nigeriana (DORSET-467mMagSnm), showing the cephalic vesicle (large arrow) and mamelons (thin arrow). Scale bars A and C = 75 μm; B = 150 μm; D = 120 μm.
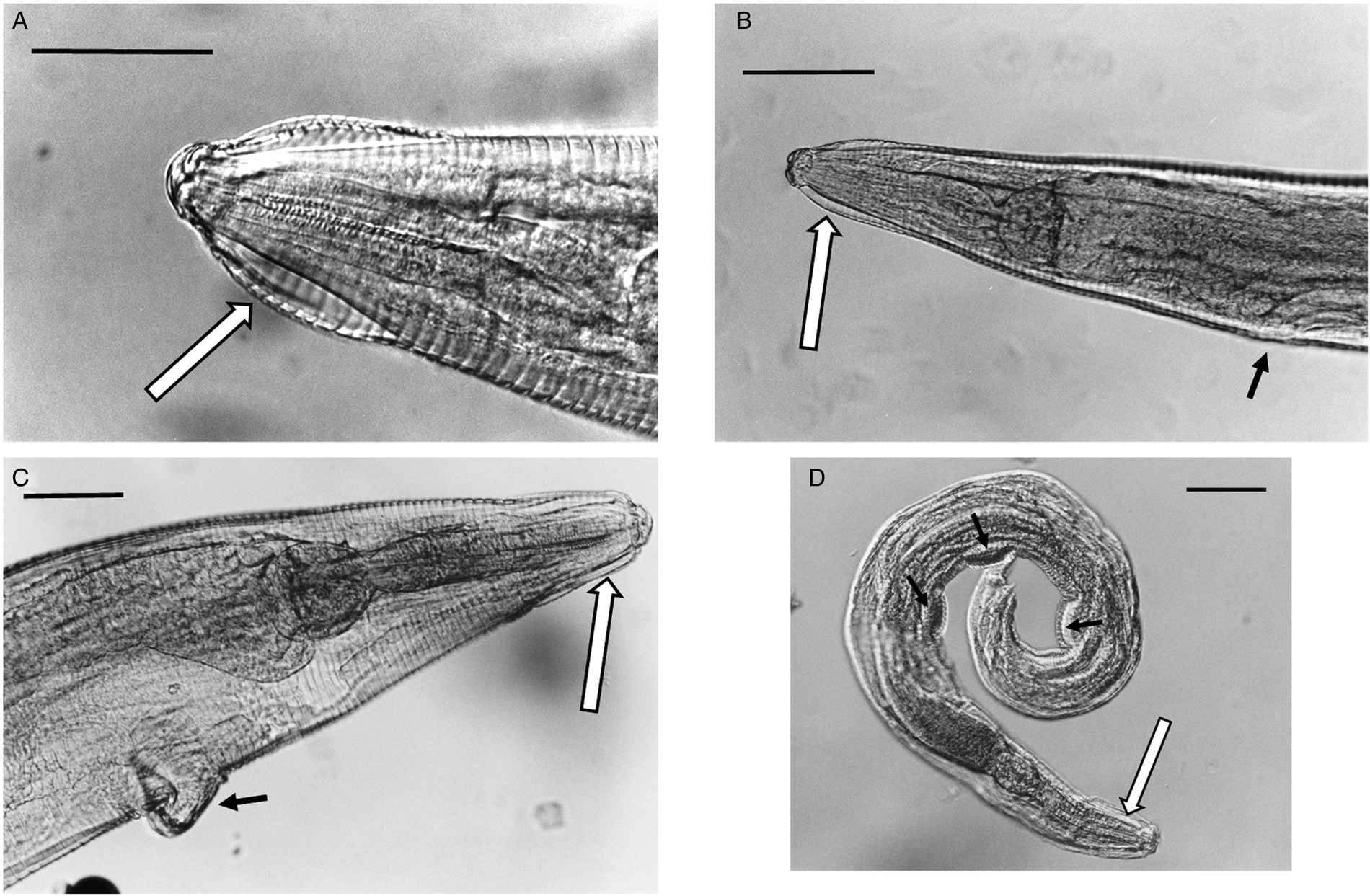
None of the specimens had ornamented cervical alae or lacked lateral alae, key characters for S. petrusewiczi. Males of S. nigeriana from this study had spicules up to 92 μm long, compared with up to 56 μm, a tail up to 180 μm long compared with up to 66 μm long and the first mamelon up to 550 μm from the anterior end compared with up to 300 μm for S. petrusewiczi (Tenora and Mészáros, Reference Tenora and Mészáros1975).
Single specimens were available from each of two M. huberti SENEGAL-10MeKB6341-WO2 and SENEGAL-10MeKB6461-WO2 and no significant differences in the morphometrics between these samples and S. nigeriana could be found (Fig. 6A and B), so from morphometrics and microscopical examination of visible features, both were congruent with S. nigeriana. However, morphometric examination of the females from a single individual of Mastomys natalensis (2 specimens; SENEGAL-04Mn CB0378-W02) and three individuals of M. huberti (12 specimens; SENEGAL-15MhADAL5249-W02; Table 3), showed that whilst having morphometrics consistent with S. nigeriana, these worms differed in not having a cephalic vesicle, and alae that could be interpreted as either no cervical alae, lateral alae beginning anteriorly or no break or differentiation between lateral and cervical alae. The vulva of these females was unobtrusive and not ornamented. Some specimens had the vulva slightly protruding and some resembled the vulva of specimens of S. nigeriana from the Welsh collection of worms.
Photomicrograhs of Syphacia nigeriana and Syphacia sp.1 from Mastomys huberti from Senegal. (A and B) S. nigeriana (10MeKB6341-W02); arrow in A indicates the cephalic vesicle and in B the dome-shaped vulva. (C and D) Syphacia sp.1 (15MhADAL5249-W02); arrows in C show the lack of a cephalic vesicle and in D the non-ornamented vulva. Scale bars are 100 μm.
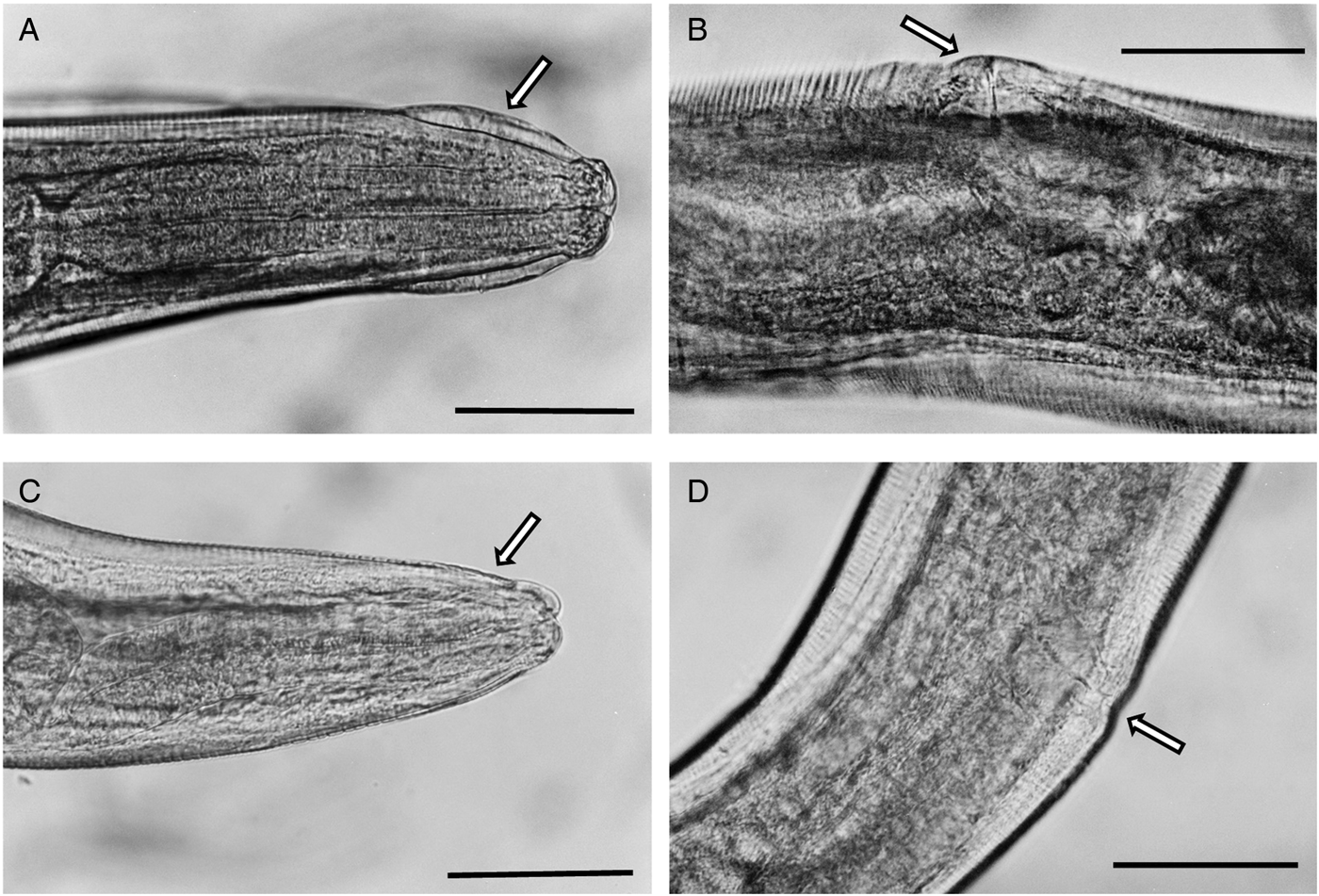
Comparisons between an en face preparations of a specimen from an individual of M. huberti (SENEGAL-15MhADAL5249-WO2) from Senegal, and M. agrestis from Northumberland (NORTHUMBERLAND-16MaV229Snfb) and from Wiltshire (WILTSHIRE-995Msn/f1) indicated that this M. huberti was harbouring a different species. All the specimens had an oval cephalic plate, extended laterally with submedian papillae and a mouth with three lips, and could therefore be placed in Group V of Quentin (Reference Quentin1971). This group includes species of Syphacia from rodents from central and north Africa and the Holarctic. The cephalic plate of the specimen from M. huberti was shaped differently from that typical for S. nigeriana. The distance between the amphids was smaller (35 as compared to 50, 55 μm for the 2 specimens from M. agrestis) and the lips were also proportionally smaller.
Discussion
When we began this project, informed by the robust and often repeated statements of earlier workers that in Europe S. petrusewiczi is the dominant Syphacia infecting rodents of the genus Myodes (e.g. M. glareolus) and S. nigeriana those of the genus Microtus (Tenora and Mészáros, Reference Tenora and Mészáros1975; Mészáros, Reference Mészáros1978), we had expected to see on our phylogenetic trees at least two clearly separated genetic clades for each of the genetic loci we have been sequencing, one associated with worms from bank voles and the other for worms from field and common voles of the genus Microtus. Moreover, given the well-established co-evolution of Syphacia spp. with their hosts (Hugot, Reference Hugot1988), and the enormous geographical barriers to gene flow between Nigeria in West Africa and Europe, we were also sceptical about the likelihood that a species of Syphacia originally described from five species of murid rodents from Nigeria (two species in the subfamily Gerbillinae and three in the subfamily Murinae) could be the same or even very closely related to a common species infecting voles (Cricetidae, Arvicolinae) in Europe. Our data show, however, that on both counts we were wrong.
Our phylogenetic analysis shows clearly that at the genetic level, based on two genetic regions [rDNA (ITS-1-5.8S-ITS2) and 18S rDNA (SSU)] and one locus (cox-1) Syphacia worms recovered from bank voles and from field and common voles from the British Isles, all formed one genetic clade and not two distinct clades as we had expected. Irrespective of the host, or the site from which the hosts had been sampled throughout the British Isles and parts of Europe, and implementing Occam's razor, we conclude that the worms whose genes we sequenced were sufficiently close genetically to be considered just one species and not two. Moreover, all measurements of the specimens of S. nigeriana from both Europe and the British Isles were congruent with those previously reported in descriptions of S. nigeriana, both of the Type (Baylis, Reference Baylis1928) and other material from African hosts, and material from European hosts (see e.g. Tenora and Mészáros, Reference Tenora and Mészáros1975). Their cephalic structures and key morphological characters were all congruent with the analyses of Quentin (Reference Quentin1971) and Hugot (Reference Hugot1988) for S. nigeriana.
Unexpectedly, the gene sequences of two worms, each from a different individual of M. huberti, a species that is closely related to one of the hosts from which Baylis (Reference Baylis1928) had originally described S. nigeriana in Nigeria (M. erythroleucus), proved to be virtually identical or very similar to those from our Myodes/Microtus isolates from the British Isles. This finding lends support to Quentin's (Reference Quentin1971) conclusion that indeed S. nigeriana, as described by Baylis (Reference Baylis1928), is also a parasite of European Microtus spp., and based on our results and those of earlier morphology-based studies, also that of Myodes spp.
Our results raise several relevant issues. Firstly, earlier authors have reported what they considered to be morphological features that differ between S. petrusewiczi and S. nigeriana. As emphasized by Wiger et al. (Reference Wiger, Barus and Tenora1978), S. petrusewiczi has prominent cervical alae, whereas S. nigeriana does not. This was also underlined by Hugot (Reference Hugot1988) whose tables indicate that S. petrusewiczi is typified by prominent and ornamented cervical alae. Careful microscopical examination of representative worms from our collection did not reveal evidence of prominent cervical alae on any of the worms from British Myodes or Microtus. Furthermore, S. petrusewiczi does not have lateral alae whereas S. nigeriana has narrow lateral alae and males of S. petrusewiczi have a short tail compared to the males of S. nigeriana which have a longer tapering tail (Tenora and Mészáros, Reference Tenora and Mészáros1975). Since males, however, are usually not found at dissection, the opportunity to distinguish throughout between the two species based on male characters was not possible. Several of the features that distinguish the two species are based on scanning electron microscopical (SEM) studies (a row of denticles on each of the three main teeth of S. nigeriana and their lack in S. petrusewiczi (Wiger et al., Reference Wiger, Barus and Tenora1978); differences in the topographical surface of the eggs, especially in the shape and outline of the operculum (Barus et al., Reference Barus, Tenora and Wiger1979)]. It will be apparent that features that can only be identified through SEM do not lend themselves well to quantitative studies of parasite burdens, and it is therefore not surprising that identification of worms by previous authors relied largely on the host rather than on specific morphological features that differ between worm species.
Secondly, although we failed to find any worms from bank voles from the British Isles and Europe that differed markedly from those we had sequenced from Microtus spp., we were able to include in our genetic analysis worms that had been clearly identified as S. petrusewiczi rauschi, derived from eastern Russian and North American M. rutilus. In accordance with Okamoto et al. (Reference Okamoto, Urushima and Hasegawa2009), these had genetic sequences that were markedly different from those we had obtained from worms from European voles, and formed their own distinct genetic clade, with only minor SNP variation between isolates from Russia and from North America. Isolates of S. petrusewiczi from Russia have also been found recently to form their own distinct clade based on sequences of the rDNA (ITS-1-5.8S-ITS-2) and the large subunit 28S regions (Gorelysheva et al., Reference Gorelysheva, Efeykin, Yakushov and Sheftel2020), although to date the sequences are not available in GenBank. Our isolates from Russia and North America had been carefully examined by one of us (JMK) and designated as S. petrusewiczi rauschi and deposited as such in the Museum Southwestern Biology, University of New Mexico. It is interesting that this S. petrusewiczi rauschi clade was quite distant to that of our worms from European voles with a deep split in the phylogenetic tree of each of the three genes in our study. This is exactly consistent with the cladistic tree published by Hugot (Reference Hugot1988) in which S. petrusewiczi and S. petrusewiczi rauschi were illustrated as sister species on a branch deeply separated from that on which S. nigeriana and S. obvelata were placed. It is also consistent with Gorelysheva et al. (Reference Gorelysheva, Efeykin, Yakushov and Sheftel2020) and Okamoto et al. (Reference Okamoto, Urushima and Hasegawa2009), the latter concluding that since S. petrusewiczi formed a deep split in the phylogenetic tree, it diverged much earlier than the other rodent pinworms examined in their study. Hugot (Reference Hugot1988) erected three subgenera within the genus based on the morphology he had characterized. As a result, S. nigeriana was placed in the subgenus Syphacia and S. petrusewiczi in the subgenus Seuratoxyuris, thus reinforcing the morphological separation of the two species, and early divergence of S. petrusewiczi from other rodent species (Okamoto et al., Reference Okamoto, Urushima and Hasegawa2009). The close relationship between S. nigeriana and S. obvelata, that was apparent in Hugot'as (Reference Hugot1988) cladistic tree, was also clearly evident in our results. In Hugot's (Reference Hugot1988) cladistic study, specimens of S. nigeriana were not derived from European Microtus spp., but from Hylomyscus stella (Thomas, 1911), the Stella wood mouse from the Republic of Central Africa. So it is possible that the worms he examined may have been the original species that Baylis (Reference Baylis1928) described as S. nigeriana, or a close relative.
Thirdly, nematodes of the genus Syphacia are among the most widespread helminth species infecting wild rodents (Roman, Reference Roman1951), but the different species are also difficult to distinguish, based on morphological characters as traditionally applied. This is partly because much of the taxonomy is based on male worms which are infrequently encountered, or even extremely rare in some species (see e.g. Mészáros, Reference Mészáros1978, who found only females of S. petrusewiczi), and are very delicate and small, but nevertheless are character richer than females. Differences between female worms, which dominate parasite burdens, are mostly minor and difficult to recognize. The difficulties inherent in distinguishing between species of Syphacia are reflected in the literature in studies where worms were first ascribed to S. petrusewiczi (Wiger et al., Reference Wiger, Lien and Tenora1976), followed by recantation of their identity, and subsequently re-identification as S. nigeriana (e.g. Tenora et al., Reference Tenora, Mészáros and Wiger1977). Where parasite burdens are heavy (Syphacia worm burdens in mice can exceed several thousand/host), examination of each individual is an onerous task, and we suspect that many authors may have relied simply on the host species as the key determinant of the Syphacia they may have been infected with. Moreover, in many reports, it is just not clear what specific characters were used to distinguish between S. petrusewiczi and S. nigeriana (e.g. Tenora et al., Reference Tenora, Andreassen, Hindsbo and Lodal1991) and how reliable these may have been. Although we do not know of any studies quantifying variation in morphological characters between individuals of a specific Syphacia sp., it is well established that morphological characters vary in size, shape and even presence or absence within species of animals, and no less so among nematodes [see e.g. Le Jambre (Reference Le Jambre1977) and Hunt et al. (Reference Hunt, Knox, Le Jambre, McNally and Anderson2008), and their studies on variation in the morphology of the vulvar flaps of Haemonchus contortus].
Fourth, we were surprised that gene sequences from worms from M. huberti fell clearly within our clade from European Microtus. Whilst this is consistent with the idea that this West African rodent shares the same parasite as European Microtus and Myodes, we were not able to extend this to the other species of rodents in Baylis' study. As our project progressed we were able to sample worms from a range of other West African rodents and our results will be reported elsewhere, as they raise yet more issues.
Fifthly, despite extensive efforts during this work to detect some evidence that S. petrusewiczi exists in British rodents, we were unable to find this species. The genetic sequences that we obtained from Russian M. rutilus and the recent study by Gorelysheva et al. (Reference Gorelysheva, Efeykin, Yakushov and Sheftel2020) indicate that it does exist in Eurasian Myodes spp., but we were unable to detect its genetic signature in any of the worms we sampled from bank voles from the British Isles. It is instructive that the worms from bank voles on the island of Jersey (which does not have any Microtus spp.) also proved to have the genetic signature of S. nigeriana. In contrast on the island of Guernsey there are no wild bank voles, the only species of vole is M. arvalis, and the Syphacia from these hosts differed from those on Jersey by just a single SNP. It may be that we were just unlucky in failing to find S. petrusewiczi in British bank voles, and that an even more extensive trapping and sampling campaign may have found the species, but given our widespread sampling, this does not sit easily with Tenora and Mészáros (Reference Tenora and Mészáros1975), who emphasized that S. petrusewiczi is a dominant parasite of M. glareolus. They also concluded that S. nigeriana is a parasite of both Microtus spp. and more rarely M. glareolus, which is consistent with our study. In further studies, Tenora et al. (Reference Tenora, Mészáros and Wiger1977, Reference Tenora, Wiger and Barus1979b) reported S. nigeriana from Norwegian bank voles but found only very low abundance with this species. These authors emphasized again that S. nigeriana is essentially a parasite of Microtus spp., but may infect bank voles in particular cases.
Finally, our results lead us to conclude that S. nigeriana is indeed a species which is less host-specific than, for example, S. stroma (only found in Apodemus spp.) or S. obvelata (only found in Mus spp.). Syphacia nigeriana has been recorded from a range of Microtus spp., not just in Europe but also in North America (Quentin, Reference Quentin1971), as well as in other rodent species in Africa (Baylis, Reference Baylis1928; Quentin, Reference Quentin1971). Based on our genetic analysis, currently restricted to British and West African rodents, it appears to exist among these hosts as a species complex with several genetic variants that differ in just a few SNPs from each other but not sufficiently to merit different species status. Its host range in Africa may be even wider, but that remains to be confirmed. On this basis, S. nigeriana must be considered a generalist that has switched hosts on more than one occasion, spreading to new rodent hosts within the cricetid and murid families, and has become the dominant species in European voles. We found no evidence that S. petrusewiczi exists in bank vole populations living in the British Isles but we can now confirm that the only report of S. nigeriana in field voles from England (Turner et al., Reference Turner, Beldomenico, Bown, Burthe, Jackson, Lamblin and Begon2014) was correct, since our sequences from worms from field voles from Northumberland (including from Kielder Forest) were identical to those in clades that we concluded were S. nigeriana.
Acknowledgements
We are grateful to Olivia G. Branston for assistance with fieldwork. We thank Carine Brouat, Christophe A. Diagne and the CBGP Small Mammal Collection for provision of samples from West African rodents (Centre de Biologie pour la Gestion des Population, l'Université de Montpellier, CIRAD, INRAE, Institut Agro, IRD, Montpellier, France, https://doi.org/10.15454/WWNUPO: KB6461, KB6341, ADAL5249, MDD-26 and CB0378). Additional samples from W. Africa were provided by Stefano Catalano and Joanne Webster. We thank Sara Brant (Museum of Southwestern Biology, Division of Parasites, University of New Mexico) for provision of Syphacia isolates from Russian and North American Myodes rutilus, through the efforts of the Beringia Coevolution Project that collects and permanently archives holistic specimens at the Museum of Southwestern Biology (University of New Mexico) under US National Science Foundation grants NSF-DEB 1258010, 0415668; Denise McGowan (Government of Jersey, Natural Environment) for providing bank voles from Jersey in the British Isles, Julia Henney (Biodiversity Education Officer in Guernsey) for provision of common voles from Guernsey for this study, and Jan Bradley for advice.
Author contributions
J.M.B. conceived and designed the study. J.M.K., A.B., D.D.S., J.H., S.C., C.A.D. and J.P.W. provided specimens from fieldwork; A.S., G.C., J.F. and A.L conducted all the genotyping and L.S. the morphological and morphometric comparison of species. The article was written by J.M.B., A.S. and L.S., and then refined by contributions from all the authors. All authors approved the final version.
Financial support
This research received no specific grant from any funding agency, commercial or not-for-profit sectors.
Conflict of interest
None.
Ethical standards
Rodents sampled in the UK were killed according to the ‘Code of Practice for the Humane Killing of Animals under Schedule 1 of the Animals (Scientific Procedures) Act 1986’ or died naturally due to domestic cat predation.