Introduction
Renicolidae is a small family of digeneans (Trematoda, Digenea), currently comprising fewer than a hundred species, taking into account the descriptions of the larvae (Sudarikov and Stenko, Reference Sudarikov, Stenko and Sonin1984; Munyer and Holloway, Reference Munyer and Holloway1990; Kharoo, Reference Kharoo2013). Their transmission is implemented in marine and estuarine ecosystems. In the complex life cycle of renicolids, the role of the first intermediate host is played by marine and brackish-water gastropods, while the role of the second intermediate host is mostly played by molluscs and fish. Adult renicolids parasitize kidneys and ureters of marine or aquatic birds, exhibiting a strong pathogenic effect on their hosts (Campbell and Sloan, Reference Campbell and Sloan1943; Hill, Reference Hill1952, Reference Hill1954; Riley and Owen, Reference Riley and Owen1972; Mahdy and Shaheed, Reference Mahdy and Shaheed2001; Jerdy et al., Reference Jerdy, Baldassin, Werneck, Bianchi, Ribeiro and Carvalho2016; Matos et al., Reference Matos, Meira-Filho, Lorenzetti, Lavorente, Caldart, Bizari, Matos, Domit and Bracarense2021). As they grow, their body becomes densely packed with eggs. It is next to impossible to discern diagnostic characters in such a worm. Considering, in addition, that adult worms of closely related species are similar morphologically, differentiating among them is a challenge. It is therefore unsurprising that the systematics of renicolids is poorly developed. Only 2 genera are recognized within the family: Renicola Cohn, 1904 and Nephromonorcha Leonov, 1958. Their adults differ in the number of testes: 2 separate testes in the former and 1 testis (resulting from merging of the 2) in the latter genus (Sudarikov and Stenko, Reference Sudarikov, Stenko and Sonin1984; Gibson, Reference Gibson, Gibson, Bray and Jones2008). Attempts to elaborate the classification of renicolids (e.g. Wright, Reference Wright1957; Odening, Reference Odening1962; Riley and Owen, Reference Riley and Owen1972) have not gained general recognition (reviewed in Gibson, Reference Gibson, Gibson, Bray and Jones2008; Kharoo, Reference Kharoo2013).
Intramolluscan stages of renicolids are represented by mother sporocysts, which look like small membrane-enveloped aggregations of cells, and cercariae-producing sac-like daughter sporocysts, parasitizing the molluscan gonad and digestive gland (Wright, Reference Wright1956; James, Reference James1969). Life cycles of only a few renicolid species have been elucidated (Stunkard, Reference Stunkard1964; Werding, Reference Werding1969; Prevot and Bartoli, Reference Prevot and Bartoli1978). At the same time, several cercariae whose descriptions are present in the literature are considered as renicolid larvae (e.g. Martin and Gregory, Reference Martin and Gregory1951; Cable, Reference Cable and Miner1956, Reference Cable1963; James, Reference James1968a, Reference James1969; Martin, Reference Martin1971; Sannia and James, Reference Sannia and James1977; Cannon, Reference Cannon1978, Reference Cannon1979; Hechinger, Reference Hechinger2007, Reference Hechinger2019; Martorelli et al., Reference Martorelli, Fredensborg, Leung and Poulin2008; Flores et al., Reference Flores, López, Levicoy, Muñoz-Ramírez, González-Wevar, Oliva and Cárdenas2019). There are among them cercariae with contrasting morphotypes: from typical xiphidiocercariae (small styleted cercariae with a simple tail) to large non-styleted larvae of the Rhodometopa group with tail fins (Wright, Reference Wright1953, Reference Wright1956; Odening, Reference Odening1962; Cable, Reference Cable1963; Stunkard, Reference Stunkard1971; Prevot and Bartoli, Reference Prevot and Bartoli1978). Such a high diversity of cercarial morphotypes within a small family is unusual for trematodes (Galaktionov and Dobrovolskij, Reference Galaktionov and Dobrovolskij2003). This matter apparently requires clarification, all the more so, as the results of molecular studies are ambiguous: some molecular data confirm that the larvae of the Rhodometopa group belong to renicolids (Matos et al., Reference Matos, Lavorente, Lorenzetti, Meira-Filho, Nóbrega, Chryssafidis, Oliveira, Domit and Bracarense2019), while other data indicate the opposite (Heneberg et al., Reference Heneberg, Sitko, Bizos and Horne2016).
Despite the contrasting differences in the morphotype, species identification of cercariae is problematic because they are morphologically very similar in closely related species. This is the case, in particular, of renicolid intramolluscan stages from intertidal snails Littorina spp. in the North Atlantic (NA). Stunkard and Shaw (Reference Stunkard and Shaw1931) and Stunkard (Reference Stunkard1932) described cercariae Cercaria parvicaudata Stunkard and Shaw, Reference Stunkard and Shaw1931 and Cercaria roscovita Stunkard, Reference Stunkard1932 from these molluscs, but they are extremely difficult to differentiate (Stunkard, Reference Stunkard1950; Galaktionov and Skírnisson, Reference Galaktionov and Skírnisson2000). After the life cycle of, presumably, C. roscovita was elucidated and the species was named Renicola roscovitus (Stunkard, Reference Stunkard1932) Werding, Reference Werding1969, it has been generally assumed that this is the dominant renicolid species using periwinkles as the first intermediate hosts in NA (Lauckner, Reference Lauckner and Kinne1980, Reference Lauckner and Kinne1983). In a study of cercariae from intertidal molluscs in Iceland, Galaktionov and Skírnisson (Reference Galaktionov and Skírnisson2000) recorded only larvae corresponding to C. parvicaudata described by Stunkard and Shaw (Reference Stunkard and Shaw1931) and Stunkard (Reference Stunkard1950). No cercariae matching the description of C. roscovita have been found during long-term studies of the fauna of digenean intramolluscan stages associated with Littorina spp. at the coasts of NA and the North Pacific (NP) (K. V. Galaktionov, personal observation). All these observations indicate that the question of the species composition of renicolids in NA and NP should be revisited.
The aim of this study was to ascertain the species composition of renicolids using periwinkles as the first intermediate hosts and to determine their transmission routes into NA and NP. We used an integrative approach, combining the analysis of morphological and molecular data, which has been shown to be the most effective in addressing taxonomy, phylogeny and elucidation of digenean life cycles (Blasco-Costa and Poulin, Reference Blasco-Costa and Poulin2017). Relatively few studies on renicolids have employed this approach (Skírnisson et al., Reference Skírnisson, Guðmundsdóttir, Andrésdóttir and Galaktionov2002–2003–2003; Hechinger and Miura, Reference Hechinger and Miura2014; O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014, Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015; Patitucci et al., Reference Patitucci, Kudlai and Tkach2015; Heneberg et al., Reference Heneberg, Sitko, Bizos and Horne2016; Flores et al., Reference Flores, López, Levicoy, Muñoz-Ramírez, González-Wevar, Oliva and Cárdenas2019; Matos et al., Reference Matos, Lavorente, Lorenzetti, Meira-Filho, Nóbrega, Chryssafidis, Oliveira, Domit and Bracarense2019), and our study is an addition to their number. In the course of our research on trematodes from the nearshore areas of NA and NP seas, we have collected and analysed extensive material on both intramolluscan stages and adults of renicolids from coastal birds, including gulls. Based on this material, we ascertained the species composition of renicolids from gulls in NA and NP and outlined the ways towards the elucidation of some aspects of their classification, evolution and ways of host colonization and geographical expansion. In addition, we confirmed that the larvae of Rhodometopa group belonged to renicolids and suggested how the cercariae of this type could have originated during the evolution of the taxon.
Material and methods
Material collection and treatment
The material presented in this study was collected from definitive and intermediate hosts (birds and molluscs) in 2002–2021 on the Atlantic coasts of Europe and North America and Pacific coast of North Asia (Table 1). Gastropod molluscs Littorina saxatilis (Olivi, 1792) and Littorina obtusata (Linnaeus, 1758) were collected in the intertidal zone of the White Sea, Barents Sea (Eastern Murman and Finmark) and Iceland, Littorina sitkana Philippi, 1846, in the Sea of Okhotsk (Magadan region) and Littorina littorea (Linnaeus, 1758), in the White Sea and the North Sea (Texel, the Netherlands) (Table 1). We also included in the molecular analysis C. parvicaudata isolates from L. littorea collected at the coasts of North East Atlantic (NEA) and North West Atlantic (NWA) during the study by Blakeslee and Byers (Reference Blakeslee and Byers2008), with the sequence data reported in Blakeslee and Fowler (Reference Blakeslee, Fowler and Caliskan2012), in the summer months between 2002 and 2005 (Table 1). Herring gull Larus argentatus Pontoppidan, 1763 and slaty-backed gull Larus schistisagus Stejneger, 1884 were obtained by shooting in accordance with local regulations in South-West Iceland (Reykjavik region) and the Sea of Okhotsk (Magadan region), correspondingly.
List of samples used in this study and corresponding GenBank accession numbers

L. littorea, Littorina littorea; L. saxatilis, Littorina saxatilis; L. argentatus, Larus argentatus; L. schistisagus, Larus schistisagus; N. lapillus, Nucella lapillus
The molluscs were dissected under a stereomicroscope to identify those infected with renicolid intramolluscan stages. Some snails were placed in plastic jars filled with seawater (1 snail per jar) and exposed to sunlight or direct artificial light for 1 h. The jars were examined under a stereomicroscope and the individuals that had shed cercariae of Renicola spp. were selected. These snails, kept in the refrigerator under 4°C, were used as a source of cercariae, which were obtained when required following the same procedure as in case of freshly collected snails.
The species of renicolid intramolluscan stages was identified on the basis of the original descriptions by Stunkard and Shaw (Reference Stunkard and Shaw1931) and Stunkard (Reference Stunkard1932, Reference Stunkard1950). Live sporocysts and cercariae were observed, measured and photographed using Olympus CH40 compound microscope equipped with an Olympus XC-30 digital camera at the ‘Kartesh’ White Sea Biological Station of the Zoological Institute of the Russian Academy of Science (ZIN RAN); Leica compound microscope in the Institute of Pathology (Keldur, Iceland) and Leitz Dialux 20B compound microscope in the Institute of Biological Problems of the North (Magadan, Russia). Only newly shed cercariae were used for morphometric studies and scanning electron microscopy (SEM). Cercariae to be measured were fixed by heating in a drop of seawater on the object slide (until the water started to evaporate), and then gently pressed with a coverslip. Sporocysts and encysted metacercariae were measured in vivo. For SEM, we used cercariae C. parvicaudata newly shed from the White Sea L. littorea. Cercariae fixation procedure and treatment before SEM examination were done as described in Galaktionov et al. (Reference Galaktionov, Solovyeva and Miroliubov2021). The treated cercariae were viewed under a FEI Quanta 250 scanning electron microscope in ‘Taxon’ Research Resource Center (http://www.ckp-rf.ru/ckp/3038/) of ZIN RAS. For molecular studies, we used renicolid intramolluscan stages whose species had been tentatively identified based on morphological criteria. This material was fixed in 95% ethanol.
Gulls were dissected and the renicolid individuals were extracted from the kidney. These adults were fixed in 70% ethanol under a slight pressure of a coverslip. Samples of adults were stored in 70 and 95% ethanol for further morphological and molecular analysis, correspondingly. Carmine-stained whole mounts were used for morphological studies, to make drawings and photographs using Leica DM2500 compound microscope with camera lucida and ToupCam UCMOS14000 digital camera. All measurements presented in the paper are in micrometres, with the mean in parentheses. Drawings were made with the aid of camera lucida.
DNA extraction, amplification and sequencing
We determined the sequences of 28S ribosomal RNA (rRNA) and cox1 mitochondrial genes for rediae and cercariae of Renicola spp. from infected periwinkles and birds (Table 1). Genomic DNA was extracted with cetrimonium bromide (CTAB) detergent according to the published protocol with modifications (Winnepenninckx et al., Reference Winnepenninckx, Backeljau and De Wachter1993) from ethanol-fixed isolates. Fixed specimens were rinsed in 1× phosphate-buffered saline for 15 min before extraction. The D1–D3 fragment of 28S rRNA gene was amplified with primers ZX-1 (5′-ACCCGCTGAATTTAAGCATAT-3′) (Palm et al., Reference Palm, Waeschenbach, Olson and Littlewood2009) and 1500R (5′-GCTATCCTGAGGGA AACTTCG-3′) (Olson et al., Reference Olson, Cribb, Tkach, Bray and Littlewood2003) according to the following temperature profile: initial DNA denaturation at 95°C for 5 min, then 30 cycles (95°C for 1 min; 55°C for 30 s; 72°C for 1 min) and a final elongation step at 72°C for 5 min. The cox1 gene fragments were amplified with primers JB3 (5′-TTTTTTGGGCATCCTGAGGTTTAT-3′) и JB4-5 (5′-TAAAGAAAGAACATAATGAAAATG-3′) (Bowles et al., Reference Bowles, Blair and McManus1992) with the following conditions: initial DNA denaturation at 95°C for 5 min, then 30 cycles (95°C for 1 min; 53°C for 30 s; 72°C for 45 s) and a final elongation step at 72°C for 5 min. PCR reactions were run on the Mastercycler personal 5332 (Eppendorf, USA) thermal cycler. ITS2 fragment was amplified with NC13(ITS2)/F(5′-ATC GAT GAA GAA CGC AGC-3′) и Dd28SR1(5′-ACA AAC AAC CCG ACT CCA AG-3′) primers according to Heneberg et al. (Reference Heneberg, Sitko, Bizos and Horne2016). PCR products were purified following a modified protocol (Dyachenko et al., Reference Dyachenko, Beck, Pantchev and Bauer2008; Galaktionov et al., Reference Galaktionov, Solovyeva and Miroliubov2021). DNA sequencing was performed at the Development of Molecular and Cellular Technologies Resource Centre at St. Petersburg State University and the University of New Hampshire (Durham, New Hampshire, USA). Two cox1 gene sequences of samples from NWA L. littorea recognized as R. roscovita were kindly provided by Natalia Ershova (University of Chicago). All the sequences obtained in this study were deposited in GenBank (Table 1).
Alignments and phylogenetic analyses
We performed alignment, trimming and basic analyses in Geneious 7.1.4 http://www.geneious.com (Kearse et al., Reference Kearse, Moir, Wilson, Stones-Havas, Cheung, Sturrock, Buxton, Cooper, Markowitz, Duran, Thierer, Ashton, Meintjes and Drummond2012) of the newly generated sequences together with 28S rRNA gene and cox1 partial sequences retrieved from GenBank for other Renicola spp. Genetic divergences among taxa were calculated as uncorrected p-distances for each gene region using MEGA v. X (Tamura et al., Reference Tamura, Stecher, Peterson, Filipski and Kumar2013). Phylogenetic relationships were reconstructed using Bayesian inference (BI) on MrBayes v. 3.2.6 (Ronquist et al., Reference Ronquist, Teslenko, van der Mark, Ayres, Darling, Höhna, Larget, Liu, Suchard and Huelsenbeck2012) and maximum likelihood (ML) on MEGA X (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018). The most suitable evolutionary models were determined by the corrected Akaike information criterion in the PartitionFinder program (https://github.com/brettc/partitionfinder). The Hasegawa–Kishino–Yano model with estimates of gamma-distributed among-site rate variation (HKY + G) was chosen as best fitted for cox1 gene. Kimura 2-parameter model with estimates of gamma-distributed among-site rate variation was chosen for fragments of 28S rRNA genes. Genetic divergences among taxa were calculated as uncorrected p-distances for each gene region using MEGA X (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018). Mismatch distribution and Tajima's D neutrality test were calculated in DNASP 6 program (Rozas et al., Reference Rozas, Ferrer-Mata, Sánchez-DelBarrio, Guirao-Rico, Librado, Ramos-Onsins and Sánchez-Gracia2017). We also performed the species partitioning with clustering algorithm implemented in ASAP tool (Puillandre et al., Reference Puillandre, Brouillet and Achaz2020). Haplotype network was reconstructed with PopArt tool (Leigh and Bryant, Reference Leigh and Bryant2015).
Results
Molecular results showed that renicolid intramolluscan stages from L. littorea and L. sitkana and most isolates from L. obtusata and L. saxatilis, identified as C. parvicaudata based on morphological criteria, belonged to one and the same species. Their sequences also matched that of the adult from the Icelandic herring gull, which made it possible to complete the life cycle of this species. We named it Renicola parvicaudatus (Stunkard and Shaw, Reference Stunkard and Shaw1931) nov. comb. (see Molecular results and Remarks for details). Among the isolates from L. obtusata and L. saxatilis, initially identified as C. parvicaudata, the analysis of molecular markers made it possible to differentiate intramolluscan stages of the cryptic species, which we named Cercaria littorinae saxatilis VIII larva nov. In slaty-backed gulls of the Sea of Okhotsk, besides R. parvicaudatus, we found the adults of one more Renicola species, which we identified as Renicola keimahuri Yamaguti, Reference Yamaguti1939.
Description
Family Renicolidae Dollfus, 1939
Renicola parvicaudatus (Stunkard and Shaw, Reference Stunkard and Shaw1931) nov. comb.
[syn. C. parvicaudata Stunkard and Shaw, Reference Stunkard and Shaw1931, R. roscovitus (Stunkard, Reference Stunkard1932) Werding, Reference Werding1969; sexual adults of Renicola thaidus Stunkard, Reference Stunkard1964].
ZooBank LSID: urn:lsid:zoobank.org:pub:86EDD019-DF69-487C-A6C9-DF790F43966D
Type host (definitive): herring gull L. argentatus Pontoppidan, 1763, slaty-backed gull L. schistisagus Stejneger, 1884 (Laridae).
Site in definitive host: kidney.
Type-locality: South-West Iceland.
Other localities (in definitive host): Nagaeva Bay, Sea of Okhotsk.
Type material: 11 syntypes (on slides 3732-1, 3732-2, 3733-1, 3733-2, 3734-1 and 3734-2), deposited in the Collection of Helminths, section Trematoda, of the Zoological Institute of the Russian Academy of Sciences, St. Petersburg, Russia. This material represents paragenophores.
First intermediate host: L. littorea (Linnaeus, 1758), L. saxatilis (Olivi, 1792), L. obtusata (Linnaeus, 1758) and L. sitkana Philippi, 1846 (Caenogastropoda: Littorinimorpha: Littorinidae) (natural).
Site in first intermediate host: gonad.
Localities (in first intermediate host): NEA, NWA, NP.
Second intermediate host: Mytilus edulis (Linnaeus, 1758), Cerastoderma edule (Linnaeus, 1758), Argopecten irradians irradians (Lamarck, 1819), occasionally L. littorea, L. saxatilis and L. obtusata.
Representative DNA sequences: 28S rDNA (ON650718, ON650721, ON650723, ON650724, ON650726), cox1 (ON652703, ON652704, ON652707–ON652709, ON652711–ON652713, ON652718, ON652636–ON652702) and ITS2 rDNA (ON667891–ON667893, ON667895) (according to Table 1).
Sexual adults (Table 2, Figs 1 and 2)
Morphometric parameters of adults of Renicola spp. parasitizing gulls
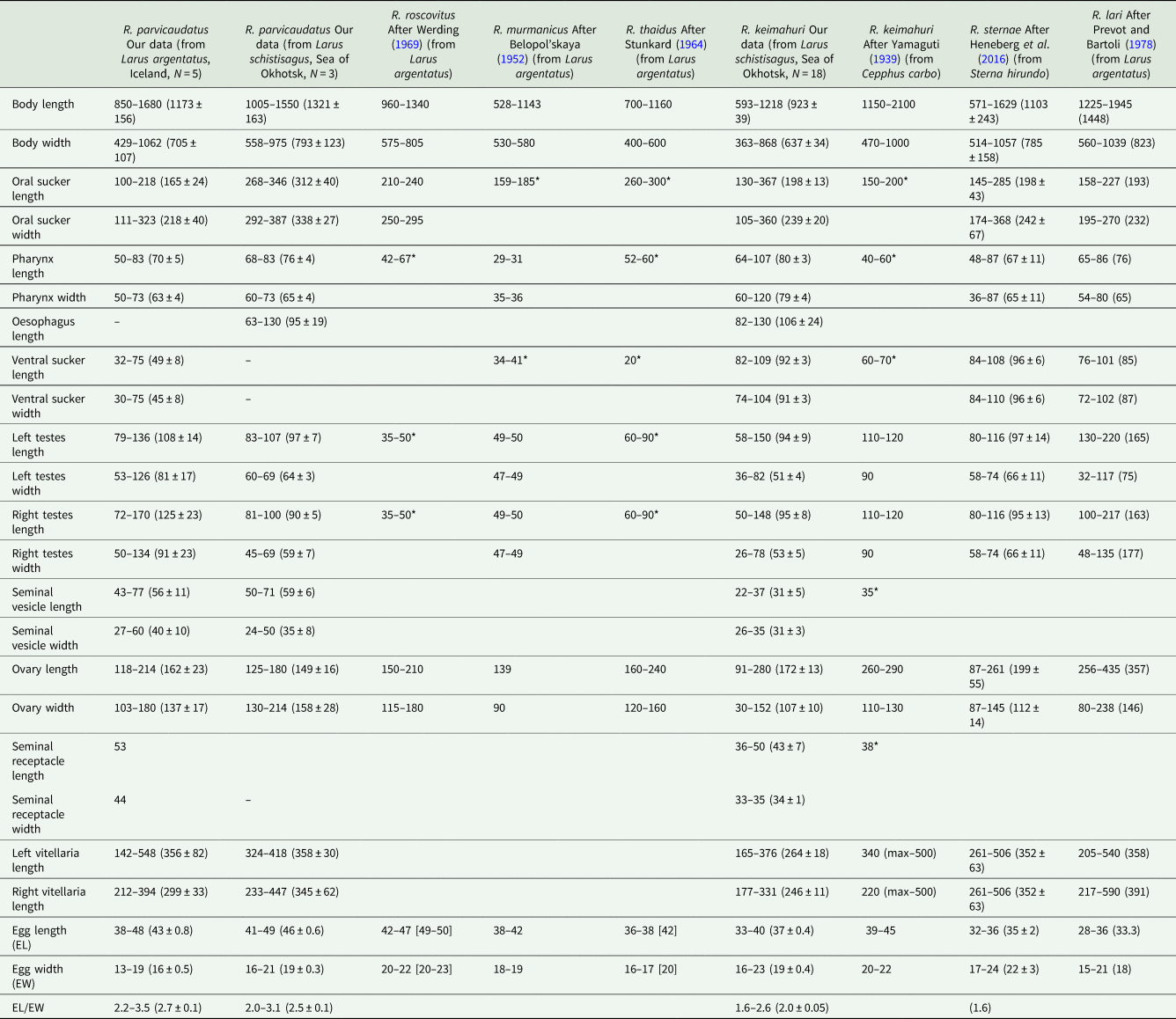
Host species is indicated in brackets in the column heads; N, number of measured individuals; * diameter of organs; measurements of live worms are given in square brackets.
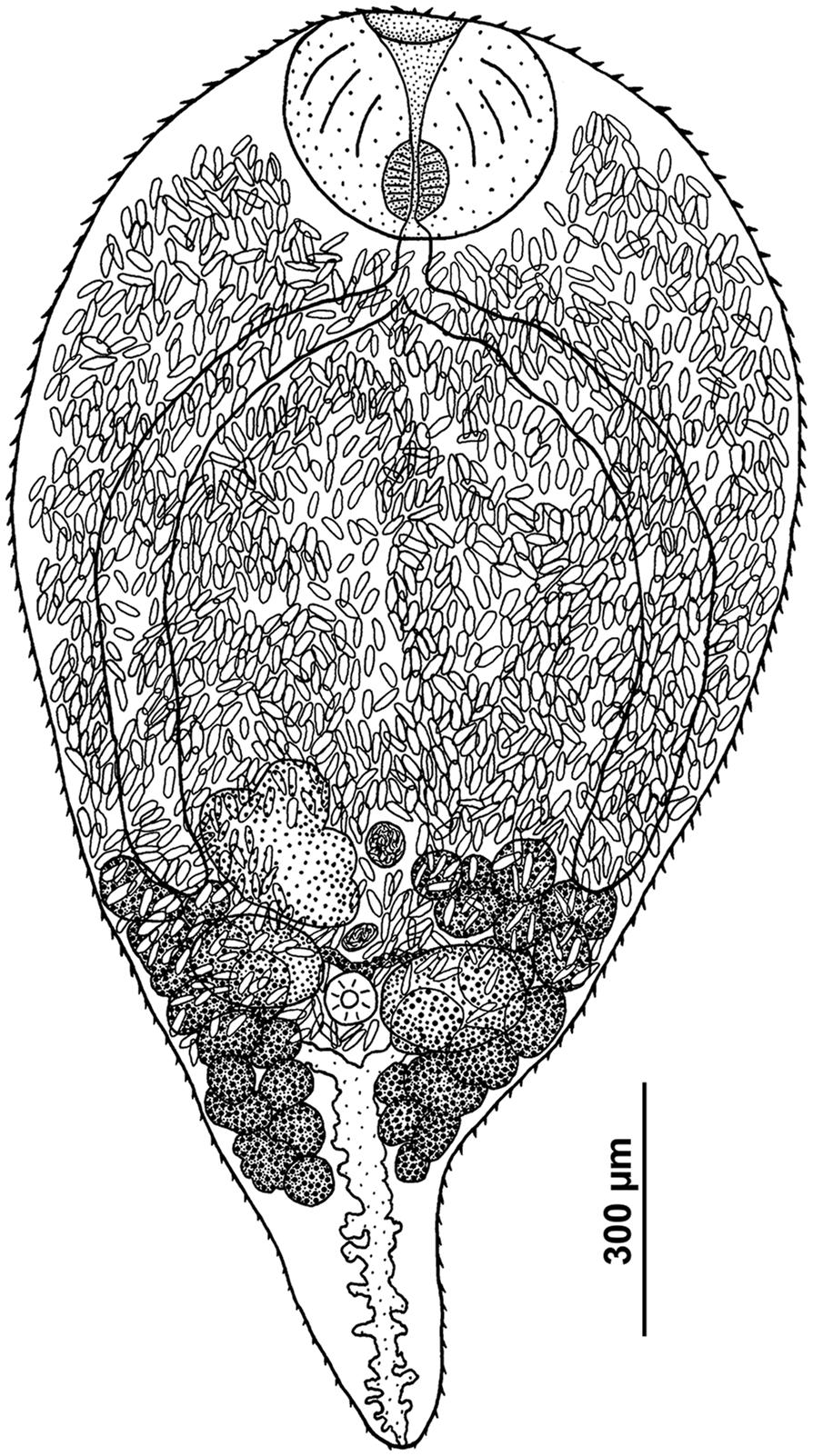
Sexual adult of R. parvicaudatus (ventral view).
Representative microphotographs of sexual adults of Renicola spp. analysed in this study (ventral view): R. parvicaudatus from the Icelandic Larus argentatus (A); R. parvicaudatus (specimen heavily pressed by cover glass) from L. schistisagus of the Sea of Okhotsk (B); R. keimahuri from L. schistisagus of the Sea of Okhotsk (C).

The description is based on morphologically identical adults from herring gull obtained in South-West Iceland. One of the adult worms matched intramolluscan stages of C. parvicaudata in the marker DNA sequences.
Body ovoid, rounded anteriorly and attenuated posteriorly. Size of worms varying greatly depending on number of eggs in uterus. Oral sucker subterminal to terminal, transversely elongated-oval. Ventral sucker 3–5 times smaller than oral sucker, in posterior third of body. Ventral sucker poorly discernible in large worms with numerous eggs. Prepharynx absent; pharynx small, often deeply embedded in wall of oral sucker. Oesophagus short, caeca 2, extending into posterior third of body. Testes oval, lying laterally of the ventral sucker, more or less opposite to each other. Left testis somewhat larger than right testis. Seminal vesicle lying anteriorly of ventral sucker approximately at level of middle to anterior part of ovary, median or lightly dextral of body midline. Ovary dextral (rarely sinistral), pretesticular, larger than testes, variously lobed. Seminal receptacle median or lightly dextral, just anterior to ventral sucker. Uterus strongly developed, occupying most of body. Eggs numerous, elongated (length about 3 times greater than width), operculate, with thin eggshell. Vitellarium follicular; follicles in 2 lateral fields in posterior third of body extended from the base of attenuated posterior part of body to level of middle or anterior border of ovary; consisting of 10–18 large follicles on ovarian side of body and 13–18 on opposite side; follicles most often fusing together. Excretory bladder Y-shaped, with distinct lateral diverticula; bifurcates just posterior to the ventral sucker, arms extending into forebody up to level of oral sucker.
Intramolluscan stages
The description is based on examination of intramolluscan stages from L. littorea collected in Texel (the Netherlands) and in the White Sea, from L. saxatilis and L. obtusata collected in Iceland (Reykjavik region) and in the White Sea, and from L. sitkana collected in the Sea of Okhotsk (Nagaeva Bay). Intramolluscan stages isolated from each snail were conspecific, as confirmed by the analysis of the molecular markers.
Sporocyst [measurements based on 30 live specimens]
Sporocyst (Fig. 3A) elongate oval, 437–876 × 213–444 (641 ± 25 × 345 ± 9), containing 1–12 (4) motile cercariae and numerous embryos. Sporocysts occupy the molluscan gonad tissue forming a tumour-like structure. The pseudo-tumour, milky white in case of early infection, becomes lemon-yellow or orange as cercariae mature in the sporocysts. The pigment responsible for the colour of the tumour is mostly concentrated in the surrounding host tissue, not in the sporocyst wall.
Microphotographs of the intramolluscan stages and cercaria of R. parvicaudatus: daughter sporocysts in the gonad of Littorina littorea (A); cercaria (B) and metacercariae encysted in the same molluscan host where daughter sporocysts develop (C). mt, Molluscan tissue; sp, daughter sporocysts.

Cercaria (Table 3, Figs 3B and 4)
Morphometric parameters of cercariae of R. parvicaudatus and closely related species (‘Parvicaudata’ group)

N, number of measured individuals; * diameter; measurements of live worms are given in square brackets. Stylet width 1 – width in the broad part of the spearhead; stylet width 2 – width of the handle.
Cercaria of R. parvicaudatus: drawings of cercaria (ventral view) (A) and stylet (ventral view) (B); SEM microphotographs showing spines in oral (C) and ventral sucker (D). sn, Spines; sne, spines of the external row; sni, spines of the internal row; sp, uniciliated sensory papillae surrounded by wide convex tegumental collars.

Cercariae small, body oval, highly contractile, body length more than 1.5 times greater than tail length. Oral sucker ventro-subterminal, muscular, approximately the same size as ventral sucker. Oral sucker armed with a single row of 38–43 spines (Fig. 4C). Stylet spear-shaped with a weakly expressed light-refracting spearhead, dorsal to mouth opening (Fig. 4B). Ventral sucker equatorial, armed with 2 alternating rows of spines of 38–40 (Fig. 4D). Anteriorly to external row of spines, ventral sucker bears 6 characteristic short sensory papillae surrounded by wide convex tegumental collars (2 anterior and 4 posterior) (Fig. 4D).
Penetration gland cells numbering 6 pairs. Their nucleated bodies arranged symmetrically on either side of oesophagus approximately at level of its middle and posteriorly. Ducts skirting oral sucker dorsally and opening at each side with common bundle near external opening of stylet pocket. Anterior parts of ducts forming pronounced curve near anterior end of oral sucker (Fig. 4A). Contents of penetration gland cells finely granular, stained with neutral red.
Entire body of larva densely packed with tegumental cystogenous gland cells. Two types of these cells distinctly seen: cells with coarsely granular contents staining with neutral red and cells with granular unstaining contents. Cells of first type with distinct nuclei, nuclei in cells of second type indistinguishable. At final stages of larva formation gland cells apparently discharging some of contents into tegument, granular material being visible throughout body and not only in cells.
Prepharynx not pronounced, pharynx rounded, intestine short, bifurcating anteriorly of ventral sucker. Excretory bladder Y-shaped, its arms skirting the ventral sucker posteriorly. Main collecting tubes opening at either side into unpaired part of bladder close to its bifurcation. Excretory formula 2[(3 + 3 + 3) + (3 + 3 + 3)] = 36.
Metacercaria (Fig. 3C)
Metacercariae are enclosed in a spherical cyst 150–180 μm in diameter; cyst wall is 10–20 μm thick. The preferred second intermediate host is the mussel M. edulis. In mussels, the cysts with metacercariae are located in the hepatopancreas and, more rarely, in the tentacles at the mantle edge. The cercariae may also encyst in the same individuals of Littorina spp. that harbour daughter sporocysts. In this case they are located in the host tissues between the sporocysts. Encystment in periwinkles is more common during the cold season, after the arrest of cercarial emergence.
Cercaria littorinae saxatilis VIII larva nov. (Table 3)
First intermediate host: L. saxatilis and L. obtusata (Caenogastropoda: Littorinimorpha: Littorinidae) (natural).
Site in first intermediate host: gonad.
Localities (in first intermediate host): Dalniye Zelentsy, Barents Sea, Grindavik, South-West Iceland.
Representative DNA sequences: 28S rDNA (ON650719, ON650722, ON650725), cox1 (ON652705, ON652710, ON652714–ON652717) and ITS2 rDNA (ON667894) (according to Table 1).
Etymology: the name of the intramolluscan stages continues the tradition of the classification of cercariae and parthenitae developing in molluscs Littorina spp., introduced by Lebour (Reference Lebour1911) and continued by James (Reference James1968b, Reference James1969), Sannia and James (Reference Sannia and James1977) and Newell (Reference Newell1986).
The species was identified based on the analysis of molecular markers of intramolluscan stages from snails L. saxatilis collected in Iceland (Reykjavik region) and the Barents Sea (coast of the Kola Peninsula) (see molecular results). Daughter sporocysts and cercariae of C. littorina saxatilis VIII are morphologically and morphometrically identical to the intramolluscan stages of R. parvicaudatus described above (Table 3).
Renicola keimahuri Yamaguti, Reference Yamaguti1939 (Table 2, Figs 2C and 5)
Sexual adult of R. keimahuri: general view from ventral side (A) and arrangement of ovary, vitellaria, genital complex and testes in relation to ventral sucker (ventral view) (B).

Representative slides: 47 individuals on slides 3735-1–3735-10, deposited in the Collection of Helminths, section Trematoda, of the Zoological Institute of the Russian Academy of Sciences, St. Petersburg, Russia. This material represents paragenophores.
Representative DNA sequences: 28S rDNA (ON650720) and cox1 (ON652706) (according to Table 1).
This species has been described by Yamaguti (Reference Yamaguti1939) based on individuals from spectacled guillemot (Cepphus carbo Pallas, 1811) obtained in Japan. In our material, R. keimahuri was represented by adults from slaty-backed gull from the northern part of the Sea of Okhotsk. Considering the differences in the hosts and the geographic sites, we provide the description of the adult worms found in our study.
Worms small, drop-shaped. Oral sucker rounded, subterminal. Ventral sucker subequatorial, approximately 2–3 times smaller than the oral sucker. Prepharynx absent; pharynx small, overlapped anteriorly by oral sucker. Oesophagus short, caeca 2, extending into posterior third of body. Testes longitudinally oval, close together, sometimes partly overlapping, dorsal from ventral sucker. Vasa efferentia start from the anterior part of each testis, pass anteriorly and fuse to form a short vas deferens just before opening into the seminal vesicle (Fig. 5B). Seminal vesicle anterior to ventral sucker, median or slightly dextral of body midline. Ejaculatory duct short, opening into genital atrium. Seminal vesicle, a few prostatic gland cells and ejaculatory duct enveloped by fine membranous structure. Genital atrium slightly sinistral of seminal vesicle, opens ventrally with genital pore.
Ovary dextral (rarely sinistral), pretesticular, deeply lobbed. Oviduct starting from posterior part of ovary, receiving first seminal receptacle and then duct of vitelline reservoir. Ootype weakly developed, tubular, surrounded by Mehlis' gland cells. All ducts of female reproductive system mentioned above as well as seminal receptacle located dorsally of ventral sucker, at level of its anterior part or somewhat anteriorly. Laurer's canal absent. Ootype passing into uterus, which forms numerous ascending and descending loops and opens into genital atrium from behind. In mature worms uterus loops are densely packed with eggs and occupy almost all body volume except caudal end. Eggs operculate, elongate, their length approximately twice greater than width. Vitellarium lateral to caeca in middle third of body, consisting of 6–8 large follicles on ovarian side of body and 7–10 on opposite side. Transverse yolk ducts originating on each side as pair of ducts filled with yolk, fusing into single duct before joining with each other to form vitelline reservoir. Vitelline reservoir dorsal at the level of anterior part of ventral sucker or pre-acetabular. Excretory bladder Y-shaped with short stem in caudal end of body and 2 arms extending to level of pharynx. Stem and branches with distinct lateral diverticula.
Molecular results
Our study generated 9 partial D1–D3 fragments of 28S rDNA (1160 bp) and 82 new mitochondrial DNA cox1 gene sequences (313 bp) for Renicola spp. (Table 1). Both ML and BI analyses resulted in consensus trees with similar topologies (Figs 6–8). In addition, we obtained 6 ITS2 sequences (354–374 bp) for several isolates: 7saxIP, 10nIR, 13saxWSP, 14obtWSP, 26saxBP and 27litHR (Table 1).
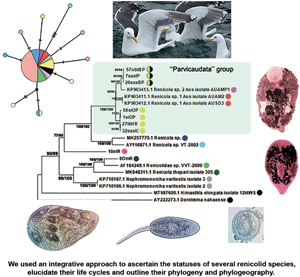
Phylogenetic relationships between Renicola spp. based on maximum-likelihood and Bayesian inference (BI) analyses of the D1–D3 fragment of 28S rRNA genes dataset: phylogenetic tree reconstructed with D1–D3 fragments of 28S rRNA genes (A); phylogenetic tree reconstructed with D3 fragment of 28S rRNA genes (B). Maximum-likelihood bootstrap support values inferred from 1000 replicates are followed by posterior probabilities from BI analysis. Bootstrap values followed by posterior probabilities are shown in nodes. Asterisk indicates posterior probabilities. Coloured circles indicate groups detected by ASAP tool. Yellow circles indicate R. parvicaudatus; yellow/black circles indicate C. littorinae saxatilis VIII. Light-blue ellipses indicate ‘Parvicaudata’ group.

Phylogenetic relationships between Renicola spp. based on maximum-likelihood and Bayesian inference (BI) analyses of cox1 gene dataset. Maximum-likelihood bootstrap support values inferred from 1000 replicates are followed by posterior probabilities from BI analysis. Asterisks indicate only bootstrap values. Coloured circles show groups detected by ASAP tool.

Phylogenetic relationships between Renicola spp. based on maximum-likelihood analysis of ITS2 dataset. Maximum-likelihood bootstrap support values inferred from 1000 replicates are followed by posterior probabilities from BI analysis. Coloured circles show groups detected by ASAP tool.

In all our trees, Renicola spp. involved in the analysis were mostly distributed across 2 large clades (I and II). Renicola somateriae Belopol'skaya, Reference Belopol'skaya1952 (10nIR) formed a separate branch (Figs 6 and 7), which was sister to clade I in the tree based on D1–D3 fragment of 28S rRNA (Fig. 6) and sister to clade I + II in the cox1 tree (Fig. 7). In clade I, isolates morphologically identified as C. parvicaudata grouped with Australian isolates of Renicola sp. 1 Aus O'Dwyer et al., Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015 and Renicola sp. 2 Aus O'Dwyer et al., Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015 into one and the same cluster, which we will refer to as the ‘Parvicaudata’ group (Figs 6 and 7). In this group, isolates tentatively identified as C. parvicaudata were distributed across 2 separate branches. One of the branches comprised isolates of R. parvicaudatus sensu stricto, and the other comprised several isolates from Iceland (7saxIP) and the Barents Sea (26saxBP, 41–43saxBP and 57obtBP), which we referred to as C. littorinae saxatilis VIII (see above) (Figs 6 and 7). Isolates 1siOP and 58siOP in the tree based on partial D1–D3 fragments of 28S rDNA were separate from the samples of R. parvicaudatus from the Netherlands (27litHR) and Iceland (32saxIC). However, genetic distances between the latter 2 samples on the one hand and 1siOP and 58siOP on the other made up 0.003 ± 0.002 and 0.004 ± 0.002, respectively, and were indistinguishable from the distance within the pooled group of these isolates, 0.003 ± 0.001 (Table S1b and S1d). The average interspecific genetic divergence amongst Renicola spp. ranged from 0.015 ± 0.005 (C. littorinae saxatilis VIII/Renicola sp. 2 Aus) to 0.166 ± 0.011 (R. keimahuri/Renicola sp. 2 Aus) (Table S1a).
Renicola sp. 2 Aus was a sister species to C. littorinae saxatilis VIII, while Renicola sp. 1 Aus was closer to R. parvicaudatus (Fig. 6). The genetic distance between the group of isolates of R. parvicaudatus and C. littorinae saxatilis VIII, calculated based on partial D1–D3 fragments of 28S rDNA, made up 0.028 ± 0.005, which corresponds to the interspecific level for Renicola (Table S1a). An analysis in ASAP also showed that the differences between R. parvicaudatus and C. littorinae saxatilis VIII corresponded to the interspecies level (Fig. 6, coloured circles). Thus, C. littorinae saxatilis VIII should be considered as a cryptic species relative to R. parvicaudatus. Our analysis also confirmed that Renicola sp. 1 Aus and Renicola sp. 2 Aus were independent species. Isolates of intramolluscan stage Renicola sp. Huston et al., Reference Huston, Cutmore and Cribb2018 (MH257770.1) found in the cerithiid gastropod Clypeomorus batillariaeformis Habe and Kosuge, 1966 (see Huston et al., Reference Huston, Cutmore and Cribb2018) and Renicola sp. VT-2002 (AY116871.1) from Eurasian curlew Numenius arquata (Linnaeus, 1758) (see Olson et al., Reference Olson, Cribb, Tkach, Bray and Littlewood2003) (Fig. 6) formed a sister branch to the ‘Parvicaudata’ group in the tree based on partial D1–D3 fragments of 28S rDNA.
Renicola keimahuri (8OmR) was placed in clade II. Within this clade, it was a sister taxon to Renicolidae sp. VVT-2000 (AF184249), the sequence of renicolid intramolluscan stages from the marine gastropod Cerithium vulgatum Bruguière, 1792 sampled near Corsica (Tkach et al., Reference Tkach, Pawlowski, Mariaux, Swiderski, Littlewood and Bray2001), in the tree based on partial D1–D3 fragments of 28S rDNA (Fig. 6). These 2 species together with Renicola thapari Caballero, Reference Caballero, Dayal and Singh1953 formed, within clade II, a sister group to Nephromonorcha varitestes Patitucci et al., Reference Patitucci, Kudlai and Tkach2015, the only member of the genus Nephromonorcha represented in GenBank.
We involved in the analysis of a short fragment of 28S rRNA gene obtained from the isolate identified by Litvaitis and Rohde (Reference Litvaitis and Rohde1999) as R. roscovitus (AF023113), as it was the only marker available in GenBank for this species (Fig. 6B). The support of the branches decreased, and ASAP analysis became impossible because the branches were too short and the programme sorted all the samples into 2 groups only. However, the main clades remained unchanged in the resulting tree. The genetic distance between R. roscovitus (AF023113) and isolates of R. parvicaudatus by the shortened fragment of 28S rRNA gene made up 0.018 ± 0.007, which is equivalent to the intraspecific level (0.011 ± 0.004) (Table S1c and S1e).
In contrast to 28S rRNA gene, there are numerous nucleotide sequences of renicolids for cox1 in GenBank. In our cox1 phylogenetic tree, the species of the ‘Parvicaudata’ group formed a separate branch within clade I. A sister branch was represented by renicolid xiphidiocercaria species from New Zealand [Renicola sp. Martorelli et al., Reference Martorelli, Fredensborg, Leung and Poulin2008 (FJ765490–FJ765493)] and North America (Renicola sp. ‘martini’ Hechinger and Miura, Reference Hechinger and Miura2014 and Renicola sp. ‘polychaetophila’ Hechinger and Miura, Reference Hechinger and Miura2014) (Fig. 7).
The phylogenetic reconstruction and the analysis in ASAP showed that groups of isolates of R. parvicaudatus and C. littorinae saxatilis VIII diverged (Fig. 7). Intragroup p-distances in these 2 groups varied from 0.003 ± 0.003 to 0.016 ± 0.007, while the intergroup distance made up 0.106 ± 0.016. This corresponds to the interspecific genetic divergence, which, as estimated by cox1, ranged amongst Renicola spp. from 0.094 ± 0.016 (R. parvicaudatus/Renicola sp. 1 Aus) to 0.291 ± 0.025 (R. somateriae/Renicola sternae Heneberg et al., Reference Heneberg, Sitko, Bizos and Horne2016) (Table S2a). The group of R. parvicaudatus contained all samples from NEA, NWA and NP, including those tentatively identified (based on the colour of sporocysts) as R. roscovitus (11 Ersh and 12 Ersh).
Similarly to the tree based on D1–D3 fragment of 28S rRNA, the Australian species Renicola sp. 1 Aus in the cox1 tree appeared as a sister to R. parvicaudatus, while C. littorinae saxatilis VIII together with Renicola sp. 2 Aus and Renicola sp. NZ O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014 formed a sister clade to them. P-distances between Renicola sp. 2 Aus and Renicola sp. NZ (0.035 ± 0.01) corresponded to intraspecific genetic diversity (Table S3, pair distances, Table S2b), and an analysis in ASAP did not show them to be separate species, either. Within clade II, R. keimahuri (8OmR) was closest to R. sternae and Renicola lari Timon-David, 1933, but p-distance between the former species and the latter 2 species (0.121 ± 0.018 and 0.125 ± 0.018, respectively, Table S2a) corresponded to the interspecific level. These 3 species were also distinct based on ASAP (Fig. N2, coloured circles).
In the tree based on ITS2 fragment of 28S rRNA (Fig. 8), Cercaria doricha Rothschild, Reference Rothschild1935 and Cercaria pythionike Rothschild, Reference Rothschild1935 belonged to the Renicolidae, grouping with representatives of clade II according to D1–D3 28S rDNA and cox1 phylogenetic trees. The analysis in ASAP showed that C. doricha and C. pythionike were separate species, closest to Renicola sloanei but distinct from it. Genetic distances between C. doricha and C. pythionike also corresponded to the interspecific level (0.026 ± 0.009, Table S4). Renicola parvicaudatus and C. littorina saxatilis VIII diverged in the ITS2 tree, while the genetic distance between them based on this rDNA fragment made up 0.044 ± 0.010, which corresponds to the interspecific level (Table S4).
To study the history and the structure of R. parvicaudatus population, we calculated the mismatch distribution and constructed a haplotype network (Fig. 9). Mismatch distribution showed low pairwise differences and was skewed unimodal (Fig. 9A). We detected 10 haplotypes, which were arranged in a ‘star’ network (Fig. 9B). Most isolates represented the main haplotype, except the isolates from the White Sea and one of the Sea of Okhotsk isolates (Fig. 9B). The latter was separated from the dominant haplotype by 4 substitutions. An additional haplotype is conjectured to be present between the White Sea haplotypes and the dominant one (black dot, Fig. 9B). The Tajima's D neutrality test resulted in –2.239 (P < 0.01).
Mismatch distribution based on cox1 haplotypes (A) and a median joining haplotype network for R. parvicaudatus (B) based on cox1 gene sequences. Solid lines in mismatch distribution graph show observed frequencies, dashed lines show expected frequencies. Colours in haplotype network indicate sampling regions, circle size is proportional to sample size. Hatch marks represent nucleotide substitutions. Black dots represent missing haplotypes.

Remarks
Adults of R. parvicaudatus
Adult Renicola from gulls in Iceland matched in all the molecular markers used in our study the intramolluscan stages of C. parvicaudata both from periwinkles collected at the Atlantic coast of Europe and North America and from periwinkles collected in the Sea of Okhotsk (Figs 6–8). In particular, they matched the cercariae C. parvicaudata collected from snails L. littorea in the Woods Hole region (11G′ Ersh and 12D′ Ersh), that is, from the same place and the same molluscan host as in the first description by Stunkard and Shaw (Reference Stunkard and Shaw1931). Morphologically these adults matched the adults of R. roscovitus described by Werding (Reference Werding1969) (Table 2). Since the description of C. parvicaudata Stunkard and Shaw, Reference Stunkard and Shaw1931 was published before that of C. roscovita Stunkard, Reference Stunkard1932, in accordance with the International Code of Zoological Nomenclature (ICZN, Reference Ride, Cogger, Dupuis, Kraus, Minelli, Thompson and Tubbs1999) (23.1. Statement of the Principle of Priority) a valid species name would be R. parvicaudatus (Stunkard and Shaw, Reference Stunkard and Shaw1931) nov. comb. The use of the name R. roscovitus should be discontinued until the clarification of the status of C. roscovita (see Remarks on cercariae). Three adult worms found in a slaty-backed gull L. schistisagus obtained in the north of the Sea of Okhotsk also agreed with the description of adult R. parvicaudatus (Табл. 2, Fig. 2A and B). In addition, snails L. sitkana from that area (Nagaeva Bay) harboured sporocysts and cercariae (isolates 1siOP and 58sitO) that corresponded to C. parvicaudata from the European periwinkles based on molecular and morphological characters.
The adults (but not the cercariae!) described by Stunkard (Reference Stunkard1964) as R. thaidus should also be synonymized with R. parvicaudatus. To note, the synonymy of R. thaidus and R. roscovitus (=R. parvicaudatus) has been suggested by Werding (Reference Werding1969). The adults of R. parvicaudatus and those of R. thaidus are similar morphometrically (Table 2). Other similarities are the location of vitellaria in the body of the worms (in 2 symmetrical lateral groups in the posterior body part, from the base of the tail to approximately the level of anterior border of the ovary) as well as a lobate ovary and 2 oval testes.
The cercaria described by Stunkard (Reference Stunkard1964) from Woods Hole region as the larva of R. thaidus is common at the European coast of the NA in molluscs Nucella lapillus (Linnaeus, 1758) (see Galaktionov and Skírnisson, Reference Galaktionov and Skírnisson2000). This larva is markedly different from the species of the ‘Parvicaudata’ group both morphologically (Galaktionov and Skírnisson, Reference Galaktionov and Skírnisson2000) and in molecular markers used in this study (Figs 6–8). Skírnisson et al. (Reference Skírnisson, Guðmundsdóttir, Andrésdóttir and Galaktionov2002–2003–2003) showed with the use of the ITS1 rDNA marker sequence that this cercaria is the larva of adult worms R. somateriae parasitizing in the kidneys of common eider (Somateria mollissima Linnaeus, 1758) (this conclusion was supported by the analysis of 28S rDNA and cox1 mitochondrial DNA sequences, our unpublished data). The adults of R. somateriae are strikingly different from those of R. parvicaudata as their vitellaria stretch laterally in 2 symmetrical rows from approximately the level of the pharynx to the tail.
Stunkard (Reference Stunkard1964) raised adults of R. thaidus in a young herring gull by feeding it on mussels experimentally infected with metacercariae of R. thaidus, but failed to raise them in ducklings of common eider. Apparently, the experiments went wrong at some stage (most probably, some of the mussels had natural infection with metacercariae of R. roscovita). Later Stunkard (Reference Stunkard1971) repeatedly tried to infect gulls L. argentatus, cormorants, chicks, ducklings and laboratory mammals with metacercariae of C. parvicaudata from periwinkles and mussels collected near Woods Hole, but unsuccessfully. Therefore, he assumed that R. roscovitus and C. parvicaudata were different species and that while the former, in accordance with Werding (Reference Werding1969), used gulls as the definitive host, the latter could use some sandpipers (Stunkard, Reference Stunkard1971). It is difficult to say why Stunkard's experiments with infection of the birds were unsuccessful, but our results unequivocally indicate that the definitive hosts of R. parvicaudatus are gulls.
In principle, there may be another species that should be made synonymous with R. parvicaudatus: Renicola murmanicus Belopol'skaya, Reference Belopol'skaya1952 described by Belopol'skaya (Reference Belopol'skaya1952) from gulls in Eastern Murman (Barents Sea). The adults of these 2 species are morphologically and morphometrically identical (Table 2). We do not synonymize them yet because the adults of R. murmanicus have been registered in the same region (Barents Sea, Eastern Murman) where, according to our observations, the periwinkles are infected only with intramolluscan stages of the cryptic species C. littorinae saxatilis VIII. It is possible that they are the life-cycle stages of one and the same species. In that case, it should be referred to as R. murmanicus and considered a cryptic species relative to R. parvicaudatus.
Cercariae of R. parvicaudatus and C. littorinae saxatilis VIII
Morphologically, the cercaria R. parvicaudatus described in our study completely matches the cercariae of C. parvicaudata, whose intramolluscan stages have been described by Stunkard and Shaw (Reference Stunkard and Shaw1931) from snails L. littorea in the Woods Hole region (north of the USA East Coast). Later, Stunkard (Reference Stunkard1950) supplemented the description and added L. saxatilis and L. obtusata, also from the Woods Hole region, to the list of the first intermediate hosts. Cercaria roscovita has been described by Stunkard (Reference Stunkard1932) from L. saxatilis from Roscoff (France, Atlantic coast). Intramolluscan stages of C. parvicaudata and C. roscovita are barely distinguishable from each other. Stunkard (Reference Stunkard1950), when differentiating between these 2 species, noted that ‘except for the difference in colour of the daughter sporocysts, the 2 species are almost identical’. However, the colour of the parthenitae cannot be considered as a reliable character for species differentiation (Werding, Reference Werding1969; Galaktionov and Skírnisson, Reference Galaktionov and Skírnisson2000). It depends on the infection age: young groups of sporocysts of R. parvicaudatus (infection of the current year) in periwinkles at the White Sea are white, while old groups that have overwintered in the molluscan host are lemon yellow (Nikolaev et al., Reference Nikolaev, Levakin and Galaktionov2021). Nadakal (Reference Nadakal1960) has shown that daughter sporocyst and redial colour is determined by the presence of β-carotene accumulated both in the molluscan tissues and in the parasites. The source of carotenoids in the molluscan organism is the alga the molluscs feed on. In case of the renicolids in our material, it was not so much the sporocysts that were coloured but the layers of molluscan tissues between them. Our analysis of cox1 sequences of the sporocysts from lemon-coloured pseudo-tumours [C. roscovita in accordance with Stunkard (Reference Stunkard1950)] and from orange-coloured ones [C. parvicaudata in accordance with Stunkard (Reference Stunkard1950)] showed that they belonged to the same species, which we refer to as R. parvicaudatus. To conclude, differences in the colour of the sporocysts (or, rather, in the colour of the surrounding host tissues) cannot be considered as a diagnostic character.
There is 1 character that remains to be discussed, and it is the number of penetration gland cells. Stunkard (Reference Stunkard1950) indicated that the cercariae of C. parvicaudata had 6 pairs of penetration gland cells, while cercaria of C. roscovita had ‘several’ (Stunkard, Reference Stunkard1932, Reference Stunkard1950). This character was later used for differentiating the cercariae of these 2 species by James (Reference James1968a, Reference James1969). It is difficult to count the penetration gland cells in renicolid cercariae, because the distal parts of their ducts are extremely narrow while the nuclei-containing cell bodies are obscured by numerous cystogenous gland cells. Werding (Reference Werding1969) noted that the number and the exact location of penetration gland cells in the cercariae described by him as R. roscovitus could not be determined. This may be the reason why Stunkard (Reference Stunkard1950) did not include this character into the list of characters differentiating the 2 species of cercariae under consideration.
However, the number of penetration gland cells is mentioned in the identification keys by James (Reference James1968a), who differentiated the cercariae of C. parvicaudata and those of C. roscovita based on sporocyst colour and the number of penetration gland cells. It is noteworthy that the cercaria of C. roscovita is said to have ‘numerous’ gland cells. In the drawing of a cercaria of this species from L. saxatilis in Cardigan Bay, Wales (UK), 15–17 pairs of penetration gland cells can be counted, whose external pores form 2 longitudinal rows on either side of the stylet (Fig. 77, p. 301, James, Reference James1969). This drawing disagrees with our data and with the drawing of a cercaria of R. roscovitus in Werding (Reference Werding1969), in which the ducts of the penetration gland cells open in 2 compact groups near the anterior edge of the oral sucker in the area of the stylet, that is, exactly as they do in C. parvicaudata.
In our opinion, it was C. parvicaudata that Werding (Reference Werding1969) studied, not C. roscovita described by James (Reference James1969). This opinion is supported by the fact that Werding (Reference Werding1969) worked with intramolluscan stages from L. littorea, while C. roscovita has been reported only from L. saxatilis and Melarhaphe neritoides (Linnaeus, 1758) (Syn. Littorina neritides) (Stunkard, Reference Stunkard1932, Reference Stunkard1971; James, Reference James1968b, Reference James1969). The region where Werding (Reference Werding1969) collected his material is the same as the region where Litvaitis and Rohde (Reference Litvaitis and Rohde1999) worked: the coast of Germany, including Isle of Sylt (Wadden Sea). Moreover, the sequence of short 28S DNA fragment of R. roscovitus (AF023113) from Litvaitis and Rohde (Reference Litvaitis and Rohde1999) matched the sequences that we obtained for C. parvicaudata (Fig. 6B). Snails L. littorea from the North Sea coast (Texel Island, the Netherlands) surveyed in our study were infected only with intramolluscan stages of C. parvicaudata, as supported by molecular data (Figs 6A and 7).
In addition to C. parvicaudata and C. roscovita, cercariae of 3 other renicolid species are recorded in periwinkles in NA: Cercaria emasculans Pelseneer, 1906, Cercaria brevicauda Pelseneer, 1906 and C. littorinae saxatilis VI Sannia and James, Reference Sannia and James1977 (James, Reference James1968a; Sannia and James, Reference Sannia and James1977). They differ from C. parvicaudata and C. roscovita in morphometric characteristics, the shape of the stylet, the number of penetration gland cells and the position of their ducts in the larval body. Cercaria littorinae saxatilis VI, which has been described from L. saxatilis in the north of Iceland (Eyjafjördur) (Sannia and James, Reference Sannia and James1977), is strikingly different from the larvae of the other species, because it has only 1 pair of penetration gland cells. We did not find any cercariae of this species in the south-western Iceland though we dissected more than 10 000 individuals of L. saxatilis and L. obtusata in the course of our surveys; we registered only intramolluscan stages of C. parvicaudata (Galaktionov and Skírnisson, Reference Galaktionov and Skírnisson2000; Skírnisson and Galaktionov, Reference Skírnisson and Galaktionov2002; K. V. Galaktionov, personal observations) and, as molecular analysis showed, those of a cryptic species C. littorinae saxatilis VIII.
Cercariae of Renicola sp. NZ, Renicola sp. 1 Aus and Renicola sp. 2 Aus from Australian and New Zealand Austrolittorina spp., which make up the ‘Parvicaudata’ group together with R. parvicaudatus and C. littorinae saxatilis VIII, differ from the latter 2 species genetically as well as in the number of penetration gland cells (5 pairs), number and position of large spines in the suckers and of sensory papillae on the body surface (chaetotaxy) (O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014, Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015; Denisova and Shchenkov, Reference Denisova and Shchenkov2020).
Summing up, our molecular and morphological studies indicate that R. parvicaudatus is the most common species among the renicolid intramolluscan stages in snails Littorina spp. at the Atlantic coast of Europe and North America. There are no credible findings of C. roscovita in this area. Werding (Reference Werding1969) suggested to synonymize these 2 species under the name of R. roscovitus (as noted before, he dealt with R. parvicaudatus). Nevertheless, it is premature to synonymize C. roscovita Stunkard, Reference Stunkard1932 with R. parvicaudatus because: (1) a cercaria with numerous penetration gland cells, minutely described by James (Reference James1969), should be attributed to C. roscovita Stunkard, Reference Stunkard1932 and (2) Denisova and Shchenkov (Reference Denisova and Shchenkov2020) found that the number and position of the sensory receptors on the body of cercariae C. parvicaudata from L. littorea at the White Sea were different from those of C. roscovita from L. saxatilis near Roscoff (Richard, Reference Richard1971), that is, the same snail species and the same site from which this larva was first described by Stunkard (Reference Stunkard1932). It cannot be ruled out that the species R. roscovitus does exist, and its transmission is implemented further southwards in the Atlantic (e.g. British Isles, France). At the same time, intramolluscan stages of R. roscovitus parasitize only snails L. saxatilis and M. neritoides, while R. parvicaudatus is found in L. littorea and, more rarely, in L. saxatilis and L. obtusata. To note, the analysis of cox1 sequences of the isolates from the Atlantic coast of France including the vicinity of Roscoff (EUCPTROUV1, EUCPTROUV2, EUCPTROUV3, EUCPMINDI1, EUCPMINDI2, EUCPMINDI3, EUCPMINDI4 and EUCPMINDI5 – Table 1) did not reveal the presence of any species different from R. parvicaudatus. Whether or not R. roscovitus is a true species can only be established in integrative morphological and molecular studies of intramolluscan stages of renicolids in periwinkles from the British Isles and the Atlantic coast of France.
Renicola keimahuri
Adult worms of the second species isolated from gulls L. schistisagus from the Sea of Okhotsk in our study (isolate 8OmR) morphologically correspond to R. keimahuri described by Yamaguti (Reference Yamaguti1939) from spectacled guillemot C. carbo in Japan. They are somewhat smaller than the worms described by Yamaguti (Reference Yamaguti1939) (Table 2), which may be associated with the host-induced variability. Leonov et al. (Reference Leonov, Belogurov, Shagvaleeva, Bondarenko, Leonov, Mamaev and Oshmarin1963) recorded R. keimahuri in larids in Kamchatka: slaty-backed gull (L. schistisagus), black-legged kittiwake [Rissa tridactyla (Linnaeus, 1758)], common tern [Sterna hirundo (Linnaeus, 1758)] and Aleutian tern (Onychoprion aleuticus Baird, 1869). This broad range of hosts may indicate that we deal with a complex of close or cryptic species. Detailed morphological and molecular studies are needed to prove or disprove this hypothesis. To note, this hypothesis is also supported by some morphological differences of R. keimahuri in Leonov et al. (Reference Leonov, Belogurov, Shagvaleeva, Bondarenko, Leonov, Mamaev and Oshmarin1963) from the first description by Yamaguti (Reference Yamaguti1939) and the description given above: fewer vitelline follicles (4–5) and testes that do not touch each other but are spaced apart [Fig. 20, p. 151 from Leonov et al. (Reference Leonov, Belogurov, Shagvaleeva, Bondarenko, Leonov, Mamaev and Oshmarin1963)]. At the same time, we examined mounted specimens of R. keimahuri from gulls L. schistisagus of Kamchatka (mounts ## 2439/Tr–2441/Tr, col. & det. Leonov) deposited in the collection of the Centre of Parasitology of the Russian Academy of Sciences and found that they fully corresponded to those described in this study.
Both by the molecular marker cox1 and by morphological criteria R. keimahuri is closest to the European species R. sternae described by Heneberg et al. (Reference Heneberg, Sitko, Bizos and Horne2016) from common tern (S. hirundo) and to R. lari from the herring gull (L. argentatus) and black-headed gull (L. ridibundus Linnaeus, 1766) (Prevot and Bartoli, Reference Prevot and Bartoli1978). These 3 species are similar in size and morphology (Table 2). Renicola sternae differs from R. keimahuri in having separate testes lying beside the ventral sucker (but see remark in Discussion) and in somewhat greater number of follicles in the vitellaria: 6–12 on the ovarian side of body and 9–13 on the opposite side. Renicola lari is slightly larger than R. keimahuri from our material, but corresponds to the size characteristics of this species given by Yamaguti (Reference Yamaguti1939). Despite their morphological similarity, R. keimahuri, R. sternae and R. lari are quite distinct genetically.
Discussion
It was shown for the first time that out of all Renicola spp. using snails Littorina spp. as the first intermediate hosts in the nearshore areas of NA seas the dominant species is C. parvicaudata, as identified based on the combination of morphological characters. The analysis of molecular markers and morphology showed that the adults of this species are found in gulls from Iceland and the Sea of Okhotsk. This means that we successfully elucidated the life cycle of this species. According to the Code of Zoological Nomenclature (ICZN, Reference Ride, Cogger, Dupuis, Kraus, Minelli, Thompson and Tubbs1999), we name it R. parvicaudatus (Stunkard and Shaw, Reference Stunkard and Shaw1931) nov. comb. The name R. roscovitus (Stunkard, Reference Stunkard1932) Werding, Reference Werding1969 and the name R. thaidus Stunkard, Reference Stunkard1964 used by Stunkard (Reference Stunkard1964) for the adult worms should be considered as its synonyms.
The cercariae of C. parvicaudata and those of C. roscovita are difficult to differentiate. This circumstance gave rise to a long-lasting confusion. It started with an experimental study of Werding (Reference Werding1969), who identified the cercaria of the renicolid species whose life cycle he studied as C. roscovita and named the species R. roscovitus. As explained in the Remarks section above, Werding (Reference Werding1969) actually studied the cercariae of R. parvicaudatus. This means that intramolluscan stages from Littorina spp. identified in numerous ecological and faunistic studies as R. roscovitus (e.g. Lauckner, Reference Lauckner1987; Granovitch and Johannesson, Reference Granovitch and Johannesson2000; Thieltges, Reference Thieltges2006; Thieltges and Rick, Reference Thieltges and Rick2006; Mouritsen and Elkjær, Reference Mouritsen and Elkjær2020) actually belong to R. parvicaudatus or to its cryptic species C. littorinae saxatils VIII first described in this study.
‘Parvicaudata’ species complex: composition and phylogeography
The species of the ‘Parvicaudata’ group form a separate clade on phylograms constructed on the basis of molecular markers used in our study. All these species use intertidal snails Littorina spp. and Austrolittorina spp. (Littorinoidea, Littorinidae) as the first intermediate host. The definitive host is known only for R. parvicaudatus, but other species of the ‘Parvicaudata’ group probably also use gulls or other birds, such as sandpipers, that feed on nearshore invertebrates. A sister clade of the ‘Parvicaudata’ group is formed by species whose first intermediate hosts are various molluscs from the superfamily Cerithioidea, some of which belong to the family Cerithiidae (C. batillariaeformis Habe and Kosuge, 1966) (Cannon, Reference Cannon1978, Reference Cannon1979), some to the Batillariidae [Zeacumantus subcarinatus (G. B. Sowerby II, 1855)] (Leung et al., Reference Leung, Donald, Keeney, Koehler, Peoples and Poulin2009) and some to the Potamididae [Cerithideopsis californica (Haldeman, 1840)] (Hechinger and Miura, Reference Hechinger and Miura2014). This observation suggests that the formation of the ‘Parvicaudata’ group was associated with the colonization of periwinkles as the first intermediate host.
The only morphological differences between cercariae of R. parvicaudatus and C. littorinae saxatilis VIII on the one hand and the larvae of Australian-New Zealand species on the other are the number of penetration gland cells and the number and position of large spines in the suckers and the sensory papillae on the body surface (see Remarks). Renicola sp. NZ and Renicola sp. 2 Aus have some differences in the latter 2 characters and the size (O'Dwyer et al., Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015), but genetic divergence between them was within the species level (Tables S2b and S3, Figs 6A and 7), which means that they are likely to be morphs of the same species. At the same time, genetic differences between morphologically indistinguishable cercariae of R. parvicaudatus and C. littorinae saxatilis VIII corresponded to those between different species (Tables S1a, S2a and S4), which suggests that they are cryptic species. At the same time, C. littorinae saxatilis VIII is genetically closer to Australian-New Zealand species that to R. parvicaudatus (Tables S1a, S2a and S4, Figs 6 and 7).
To sum up, morphological differences between the renicolid cercariae may not necessarily mean that they belong to different species. By the same token, the absence of morphological differences does not prove that the cercariae are conspecific. The case of Renicola sp. NZ and Renicola sp. 2 Aus shows that subtle differences in cercarial morphology and chaetotaxy revealed with the use of SEM (O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014, Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015) are not always reliable criteria for species differentiation. These considerations strongly indicate that an integrative approach is the key to ascertaining the species status of digeneans. This approach should involve the analysis of morphological characters (preferably, of all life-cycle stages), molecular markers and the data on the larval and adult biology, host range, transmission pathways and geographic distribution (Blasco-Costa et al., Reference Blasco-Costa, Cutmore, Miller and Nolan2016; Blasco-Costa and Poulin, Reference Blasco-Costa and Poulin2017; Gonchar and Galaktionov, Reference Gonchar and Galaktionov2021, Reference Gonchar and Galaktionov2022).
Small genetic distances between the species of the ‘Parvicaudata’ group (Tables S1a, S2a and S4) indicate its relatively recent formation. The differentiation of R. parvicaudatus could be associated with the colonization of a new first intermediate host, the snail L. (Littorina) littorea, the only Atlantic species of the subgenus Littorina. Its ancestor split from its NP sister species L. (Littorina) squalida Broderip and G. B. Sowerby I, 1829 and colonized the NA via the Arctic route ca. 5.5–2.4 million years ago (Reid, Reference Reid1996; Reid et al., Reference Reid, Rumbak and Thomas1996, Reference Reid, Dyal and Williams2012). This assumption is supported by the facts that R. parvicaudatus is the only renicolid parasitizing L. (L.) littorea and that it occurs in the latter more frequently than in the Atlantic periwinkles of the subgenus Neritrema, i.e. L. (N.) saxatilis and L. (N.) obtusata (our data). Intramolluscan stages of R. parvicaudatus have never been registered in L. (L.) squalida in NP (Tsimbaljuk et al., Reference Tsimbaljuk, Kulikov, Ardasheva, Tsimbaljuk and Kusakin1978; Rybakov, Reference Rybakov1983; our data), and out of all the Pacific Neritrema, only L. (N.) sitkana serves as their host, and only rarely (Tsimbaljuk et al., Reference Tsimbaljuk, Kulikov, Ardasheva, Tsimbaljuk and Kusakin1978; our data). Intramolluscan stages of the other species of the ‘Parvicaudata’ group develop in Atlantic periwinkles of the subgenus Neritrema (C. littorinae saxatilis VIII as well as C. emasculans, C. brevicauda and C. littorinae saxatilis VI, which most likely also belong to this group) or in Australian Austrolittorina spp. (James, Reference James1968a, Reference James1969; Sannia and James, Reference Sannia and James1977; O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014, Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015; our data).
The star-like patterns in cox1 haplotype network for R. parvicaudatus suggest a low geographic structure (Fig. 9B). Thus, we may conclude that R. parvicaudatus is represented by a single population throughout its Holarctic range (so far this species has not been detected at the Pacific coast of North America). The widespread haplotype positioned at the centre of the network can be considered as the ancestral one (Jenkins et al., Reference Jenkins, Castilho and Stevens2018). The other haplotypes, which are linked to this dominant haplotype by a single mutational step or a few steps, are the result of recent mutation events. Unimodal mismatch distribution (Fig. 9A) and significant negative value of the Tajima's D indicate a bottleneck event, possibly dating from the last glacial maximum (LGM). During LGM, the transmission of R. parvicaudatus may have persisted in one of the NEA glacial refugia, where periwinkles, including L. littorea, and seabirds, including gulls, were concentrated. The refugium in question could be one of the southern refugia near the Iberian Peninsula and the British Isles, where L. littorea survived during LGM (Maggs et al., Reference Maggs, Castilho, Foltz, Henzler, Jolly, Kelly, Olsen, Perez, Stam, Vainola, Viard and Wares2008; Blakeslee et al., Reference Blakeslee, Miller, Ruiz, Johannesson, André and Panova2021). Thus, contraction into a single refugium appears to have resulted in a strong bottleneck for both the ancestral L. littorea (Blakeslee et al., Reference Blakeslee, Miller, Ruiz, Johannesson, André and Panova2021) and its parasite, R. parvicaudatus.
If that is the case, the post-LGM expansion of R. parvicaudatus proceeded along the Atlantic coast of Europe, following the advance of their first intermediate hosts. The incursion into the NWA may have occurred after L. littorea was introduced into this region from the NEA. This introduction was human-mediated and tentatively dates back to the 19th century (Blakeslee et al., Reference Blakeslee, Byers and Lesser2008, Reference Blakeslee, Miller, Ruiz, Johannesson, André and Panova2021). The fact that almost all NWA haplotypes of R. parvicaudatus coincide with the dominant haplotype (Fig. 9B) can be interpreted as a further evidence of a fairly recent introduction of L. littorea into the NWA. Alternatively, natural trans-Atlantic migration with another first-intermediate periwinkle, L. obtusata, may have transported R. parvicaudatus to the NWA. This is because this periwinkle appears to have recolonized the NWA following glacial retreat via a stepping stone migration across NA islands (Wares and Cunningham, Reference Wares and Cunningham2001).
Two circumstances explain the fact that R. parvicaudatus has a broad geographical distribution and, at the same time, its cox1 haplotypes are identical or very similar in different parts of its range. Firstly, the definitive hosts of this parasite are highly mobile migrating birds such as gulls, and secondly, the life span of adult worms in them is very long. Gulls that breed at high latitudes (e.g. L. argentatus, Larus fuscus Linnaeus, 1758, Larus canus Linnaeus, 1758, Larus glaucescens Naumann, 1840, Larus glaucoides Meyer, 1822, L. schistisagus) make long seasonal migrations along the coasts of Europe and North America (Helberg et al., Reference Helberg, Systad, Birkeland, Lorentzen and Bustnes2009; Newton, Reference Newton2010; Hallgrimsson et al., Reference Hallgrimsson, Gunnarsson, Torfason, Buijs and Camphuysen2012; Klaassen et al., Reference Klaassen, Ens, Shamoun-Baranes, Exo and Bairleind2012; Davis et al., Reference Davis, Maftei and Mallory2016; Anderson et al., Reference Anderson, Gilchrist, Ronconi, Shlepr, Clark, Fifield, Robertson and Mallory2020). White-headed gulls associated with coastal habitats, such as L. argentatus and L. canus, which have a circumpolar distribution, were shown to have a limited population genetic subdivision among northern Arctic populations (Sonsthagen et al., Reference Sonsthagen, Chesser, Bell and Dove2012). This observation indicates that there is an intense genetic exchange between the populations of these birds owing to their migratory activity. Some individuals, apparently, are even capable of making trans-Arctic flights. Otherwise, the coincidence of R. parvicaudatus haplotypes in NA and NP would be difficult to explain. Trans-Arctic flights are known for Arctic-breeding seabirds (Clairbaux et al., Reference Clairbaux, Fort, Mathewson, Porter, Strøm and Grémillet2019) and have recently been reported for a larid bird, the black-legged kittiwake (R. tridactyla) (Ezhov et al., Reference Ezhov, Gavrilo, Krasnov, Bråthen, Moe, Baranskaya and Strøm2021). Another option is the transfer of the parasite by birds from the Atlantic coast of North America to the Pacific coast, and from there to the coast of North Asia.
The snag of both hypothetical variants of the trans-Arctic transfer of R. parvicaudatus is the absence of its first intermediate hosts, the periwinkles, at the coasts of the Siberian seas and at the Arctic coast of North America (Arctic coast of Alaska and the Canadian Arctic Archipelago) (Reid, Reference Reid1996). Only long-living helminths such as renicolids can endure such a long flight. According to Werding (Reference Werding1969), the lifespan of R. parvicaudatus (R. roscovitus in Werding's article) in the final host is at least 7 months. The fact that one of the haplotypes from the Sea of Okhotsk coincides with the dominant one indicates that there is an ongoing exchange between NA and NP parts of the R. parvicaudatus population (Fig. 9B). It may be associated with the warming of the Arctic, which opens opportunities for trans-Arctic bird migrations (Clairbaux et al., Reference Clairbaux, Fort, Mathewson, Porter, Strøm and Grémillet2019; Ezhov et al., Reference Ezhov, Gavrilo, Krasnov, Bråthen, Moe, Baranskaya and Strøm2021). Another haplotype of R. parvicaudatus from the Sea of Okhotsk is significantly different from the Atlantic one (Fig. 9B), possibly indicating some degree of isolation between the NP and the NA part of the parasite's population. Another evidence of the possibility of some local differentiation within the population of R. parvicaudatus is the fact that all the haplotypes from the White Sea are different from the dominant one (Fig. 9B).
In our opinion, it is premature to hypothesize about the ways of geographical expansion of other species of the ‘Parvicaudata’ group since genetic data are limited and we do not know the actual number of species constituting the group, the array of their second intermediate and definitive hosts and their ranges. The establishment of the Australian-New Zealand species was probably associated with the colonization of the local Austrolittorina spp., the incursion of the ancestral species being ensured by migrating birds. Considering that in our phylograms the Australian species Renicola sp. 1 Aus is sister to the Holarctic R. parvicaudatus, while the group comprising Australian-New Zealand Renicola sp. 2 Aus and Renicola sp. NZ is sister to the NA C. littorinae saxatilis VIII, we can assume that Australia and New Zealand were colonized as a result of 2 putative independent events.
Notes on taxonomy and phylogeny of renicolids
Two large clusters can be seen in the phylograms constructed on the basis of the molecular markers used in our study (Figs 6 and 7). Renicola parvicaudatus falls into cluster I, while R. keimahuri falls into cluster II. There are considerable morphological differences between species in cluster I and those in cluster II. Moreover, these differences are pronounced at all life-cycle stages. The adults differ in the position of vitellaria and testes, which is considered as an important taxonomical character in renicolids (Wright, Reference Wright1956, Reference Wright1957; Odening, Reference Odening1962; Sudarikov and Stenko, Reference Sudarikov, Stenko and Sonin1984; Gibson, Reference Gibson, Gibson, Bray and Jones2008). The adults of R. parvicaudatus (the only species from cluster I for which adults have been described) have vitellaria in the posterior body part and separate, non-contiguous testes. At the same time, in all adults from cluster II described so far vitellaria are located lateral to the caeca in the middle third of the body, which is a characteristic feature of species from the pinguis-group in accordance with Wright (Reference Wright1957) and Odening (Reference Odening1962). Testes are contiguous [R. thapari, R. sloanei Wright, Reference Wright1954, Renicola pinguis (Mehlis in Creplin, 1846) Cohn, 1904, R. lari and R. keimahuri (Yamaguti, Reference Yamaguti1939; Caballero, Reference Caballero, Dayal and Singh1953; Wright, Reference Wright1954; Prevot and Bartoli, Reference Prevot and Bartoli1978; Rubio-Godoy et al., Reference Rubio-Godoy, Pérez-Ponce de León, Mendoza-Garfias, Carmona-Isunzaf, la Moral and Drummondt2011; Matos et al., Reference Matos, Lavorente, Lorenzetti, Meira-Filho, Nóbrega, Chryssafidis, Oliveira, Domit and Bracarense2019; this study)] or fused to form a single mass, which is a diagnostic character of Nephromonorcha (Gibson, Reference Gibson, Gibson, Bray and Jones2008). Renicola sternae is an exception: Fig. 2b (p. 1601) in Heneberg et al. (Reference Heneberg, Sitko, Bizos and Horne2016) shows testes lying laterally of the ventral sucker. To note, however, Fig. 2a (p. 1601) in Heneberg et al. (Reference Heneberg, Sitko, Bizos and Horne2016) shows a similar position of testes in R. lari, which is at odds with the description of Prevot and Bartoli (Reference Prevot and Bartoli1978). This means that the position of testes in R. sternae and R. lari should be verified.
Nephromonorcha varitestis, the only species of the genus for which molecular data are available (Patitucci et al., Reference Patitucci, Kudlai and Tkach2015), forms an independent branch within cluster II in our phylograms. This finding supports the validity of the genus Nephromonorcha. The tendency towards contingence and subsequent merging of the testes seems to be characteristic of species in cluster II. Partial merging of contiguous testes is noted for R. lari (Prevot and Bartoli, Reference Prevot and Bartoli1978), and we observed similar pictures in R. keimahuri. At the same time, an incomplete merging of the testes into a common mass has been described in some individuals of Nephromonorcha ralli Byrd and Heard, Reference Byrd and Heard1970 and N. varitestis (Byrd and Heard, Reference Byrd and Heard1970; Patitucci et al., Reference Patitucci, Kudlai and Tkach2015). In addition to the species involved in our analysis, testes contiguous near the ventral sucker have been noted in some other renicolids, e.g. Renicola wright Odening, Reference Odening1962, Renicola pelecani Wright, Reference Wright1954, Renicola fischeri Odening, Reference Odening1962, Renicola pseudosloanei Odening, Reference Odening1962, Renicola hayesannieae Byrd and Kellogg, Reference Byrd and Kellogg1972, Renicola pollaris Kontrimavitschus and Bachmet'eva, Reference Kontrimavitschus and Bachmet'eva1960, Renicola glacialis Riley and Owen, Reference Riley and Owen1972 and Renicola williamsi Munyer and Holloway, Reference Munyer and Holloway1990 (Wright, Reference Wright1954, Reference Wright1956; Kontrimavitschus and Bachmet'eva, Reference Kontrimavitschus and Bachmet'eva1960; Odening, Reference Odening1962; Byrd and Kellogg, Reference Byrd and Kellogg1972; Riley and Owen, Reference Riley and Owen1972; Munyer and Holloway, Reference Munyer and Holloway1990). The emergence of this character in the course of morphological evolution of adult renicolids could be determined by the pressure on the testes from the eggs in the uterus loops expanding in posterior–lateral direction and could occur independently in different phylogenetic lineages of this taxon. For instance, contiguous or even merging testes are noted in Renicola philippensis Stunkard et al., Reference Stunkard, Nigrelli and Gandal1958 (Stunkard et al., Reference Stunkard, Nigrelli and Gandal1958) and R. hayesannieae (Byrd and Kellogg, Reference Byrd and Kellogg1972). At the same time, vitellaria in these species are located in the posterior-lateral part of the body, as in R. parvicaudatus from cluster I. Apparently, the determination of the taxonomic ‘weight’ of all these characters requires a more complete molecular phylogeny of renicolids involving a greater number of species differing in the structure and position of testes and vitellaria.
Structural differences between the species from the 2 clusters identified in our molecular phylogenies concern not only adults but also cercariae. Cercariae of all species from cluster I look like typical xiphidiocercariae: small size (body and tail each approximately 150–250 μm long), stylet, 1–6 pairs of penetration gland cells (rarely more), excretory formula 2[(3 + 3 + 3) + (3 + 3 + 3)] = 36, main collecting tubes join the stem of the excretory bladder, simple tail (Hechinger and Miura, Reference Hechinger and Miura2014; O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014, Reference O'Dwyer, Faltýnková, Georgieva and Kostadinova2015). Xiphidiocercariae are also known for some renicolids for which molecular data are lacking, e.g. Cercaria opaca Holliman, Reference Holliman1961, Cercaria caribbea XXXII Cable, Reference Cable and Miner1956, C. caribbea XXXIII Cable, Reference Cable and Miner1956 (Cable, Reference Cable and Miner1956; Holliman, Reference Holliman1961). The cercaria of R. somateriae (isolate 10nIR, syn. R. thaidus Stunkard, Reference Stunkard1964), which is sister to I or I + II in the phylogenetic trees (Figs 6 and 7), also looks like a typical xiphidiocercaria (Stunkard, Reference Stunkard1964).
Cercariae of Renicola buchanani (Martin and Gregory, Reference Martin and Gregory1951) and Renicola cerithidicola Martin, Reference Martin1971 in clade II have the same general appearance but lack the stylet (Martin and Gregory, Reference Martin and Gregory1951; Martin, Reference Martin1971). In R. lari, which is similar to R. keimahuri, cercaria, besides lacking the stylet, also have a well-developed excretory bladder with a short stem and arms extending to the anterior end of the body and carrying numerous lateral diverticula (Prevot and Bartoli, Reference Prevot and Bartoli1978). In addition, though the excretory formula remains the same, the main collecting tubes join not the stem but the arms of the excretory bladder. Prevot and Bartoli (Reference Prevot and Bartoli1978) considered cercariae R. lari and a similar C. caribbea VIII Cable, Reference Cable and Miner1956 as a transitional morphotype to the typical cercariae of Rhodometopa group. The latter are large (body up to 2 mm), have a long tail with fin-folds, numerous penetration gland cells that form 1–3 groups in the anterior part of the body, a well-developed Y-shaped excretory bladder with lateral diverticula in the stem and the arms and numerous flame cells (Stunkard, Reference Stunkard1932; Rothschild, Reference Rothschild1935; Wright, Reference Wright1956).
Cable (Reference Cable1963) noted that excretory system of cercariae of Rhodometopa group was organized similarly to that in the adults. In the course of development of adult renicolids, the excretory bladder expands considerably and forms lateral diverticula, as it does in Rhodometopa cercariae. The number of flame cells also increases in the course of development, which is a characteristic of trematodes (Galaktionov and Dobrovolskij, Reference Galaktionov and Dobrovolskij2003). On the basis of these observations, Cable (Reference Cable1963) suggested that Rhodometopa cercariae were more advanced than xiphidiocercariae and had certain traits of adult organization, particularly pronounced in the structure of their excretory system.
A series of transition forms from renicolid xiphidiocercariae to the cercariae of Rhodometopa group can be arranged. In cercariae of R. buchanani and R. cerithidicola, the site where the main collecting tube leaves the stem of the excretory bladder is shifted forwards; in R. buchanani it is located just before the bifurcation (Martin and Gregory, Reference Martin and Gregory1951; Martin, Reference Martin1971). Cercariae of C. caribbea VII Cable, Reference Cable and Miner1956, C. caribbea VIII, C. caribbea IX Cable, Reference Cable and Miner1956 and R. lari not only lack the stylet, but also have a well-developed excretory bladder with lateral diverticula; the main collecting tube starts not from the stem but from the arms (Cable, Reference Cable and Miner1956; Prevot and Bartoli, Reference Prevot and Bartoli1978). In drawings showing successive stages of embryogenesis in cercariae of C. caribbea VII (Cable, Reference Cable and Miner1956; Plate 3, Fig. 16, p. 550) one can see that the site of the origin of main collecting tube, which in early embryos is located at the site of bifurcation of the excretory bladder, is shifted forwards together with the outgrown branches of the excretory bladder. This also seems to be the case during the ontogenesis of adults in renicolids with xiphidiocercariae, since their adults also have outgrown branches of the excretory bladder with numerous diverticula.
Another morphological character shared by the renicolid xiphidiocercariae and the Rhodometopa cercariae is the organization of surface structure in the oral and the ventral sucker. SEM studies of xiphidiocercariae of the ‘Parvicaudata’ group have revealed 1–2 rows of large spines in the suckers and 6 large uniciliated sensory papillae (2 anterior and 4 posterior) with a wide convex tegumental collars in the ventral sucker (Fig. 4D) (O'Dwyer et al., Reference O'Dwyer, Blasco-Costa, Poulin and Faltýnková2014; Denisova and Shchenkov, Reference Denisova and Shchenkov2020; this study and our unpublished data). Rothschild (Reference Rothschild1935) noted a circle of spines and 6 large cuticular tubercles outside of them in the ventral sucker of a typical Rhodometopa cercaria C. pythionike. These ‘cuticular tubercles’ are arranged in the same manner as the sensory papillae in renicolid xiphidiocercariae and are, undoubtedly, sensory papillae, too.
The final evidence that Rhodometopa cercariae belong to renicolids came from the analysis of sequences of ITS2 rDNA of 2 typical larvae of Rhodometopa group: C. pythionike and C. doricha (Matos et al., Reference Matos, Lavorente, Lorenzetti, Meira-Filho, Nóbrega, Chryssafidis, Oliveira, Domit and Bracarense2019). This conclusion was supported by our analysis based on ITS2 rDNA sequences for a greater number of renicolid species (Fig. 8). To note, C. pythionike and C. doricha did not group with renicolids in the NCBI Blast analysis by Heneberg et al. (Reference Heneberg, Sitko, Bizos and Horne2016), which now seems to have been an error associated with the scarcity of the relevant sequences in the GenBank at the time of the analysis. In our phylogenetic tree both larvae of Rhodometopa group grouped with the species that belonged to clade II in cox1-based tree (Fig. 7). These 2 larvae clearly belong to different species. Cercaria pythionike is close to R. sloanei, but ASAP analysis convincingly shows that it is a distinct species.
It has been suggested that the formation of Rhodometopa cercariae in renicolids was associated with the colonization of plankton-eating fish as the second intermediate host and through them, of fish-eating seabirds such as alcids, penguins, petrels, pelicans, etc. (Wright, Reference Wright1956; Odening, Reference Odening1962; Cable, Reference Cable1963; Prevot and Bartoli, Reference Prevot and Bartoli1978). This hypothesis is supported by the fact that C. doricha and C. pythionike group in the phylogenetic tree together with R. sloanei, a parasite of several species of penguins and alcids (Matos et al., Reference Matos, Lavorente, Lorenzetti, Meira-Filho, Nóbrega, Chryssafidis, Oliveira, Domit and Bracarense2019, Reference Matos, Meira-Filho, Lorenzetti, Lavorente, Caldart, Bizari, Matos, Domit and Bracarense2021).
Renicolidae belong to the superfamily Microphalloidea Ward, 1901 (suborder Xiphidiata) (Cribb et al., Reference Cribb, Bray, Olson and Littlewood2003; Olson et al., Reference Olson, Cribb, Tkach, Bray and Littlewood2003; Pérez-Ponce de León and Hernández-Mena, Reference Pérez-Ponce de León and Hernández-Mena2019), whose cercariae possess the stylet. Its origin is thought to be associated with the involvement of arthropods as the second intermediate host into the life cycle of the ancestral microphalloideans (Cribb et al., Reference Cribb, Bray, Olson and Littlewood2003). The cercaria uses the stylet to penetrate the arthropod cuticle or arthrodial membranes. In renicolid cercariae the stylet is reduced to some degree or even absent, as in the larvae of Rhodometopa group and ‘transitional morphotypes’. The reduction of the stylet is associated with the transition to the use of organisms without rigid cuticular covers, such as molluscs and fish, as the second intermediate host. Only a few of the stylet-bearing renicolid cercariae penetrate polychaetes (Hechinger and Miura, Reference Hechinger and Miura2014) and occasionally crabs (Robson and Williams, Reference Robson and Williams1970) alongside with molluscs.
Metacercariae of species with xiphidiocercariae develop in invertebrates inhabiting nearshore areas, usually the intertidal zone. Therefore, the range of their definitive hosts is limited by the birds feeding on these invertebrates such as gulls, terns and sandpipers. Colonization of fish-eating seabirds became possible after renicolids began to use fish, especially planktonic fish, as the second intermediate host. This transition called for new adaptations to the infection of second intermediate host by cercariae, and finally resulted in the evolution of the larvae of Rhodometopa type.
Morphological changes of the cercariae were also accompanied by the changes in their behavioural strategies. Stylet-bearing renicolid larvae, as most xiphidiocercariae, demonstrate an active searching strategy. They are constantly moving, searching and infecting animals with low mobility (in case of renicolids, mostly molluscs) (Prokofiev and Galaktionov, Reference Prokofiev and Galaktionov2009; Nikolaev et al., Reference Nikolaev, Prokofiev, Levakin and Galaktionov2017). In contrast, renicolid cercariae of ‘transitional morphotype’ and of Rhodometopa group exhibit intermittent swimming, alternating periods of active swimming with passive floating in the water column (Combes et al., Reference Combes, Fournier, Moné and Théron1994). In the passive phase, the cercariae acquire a characteristic resting pose, bending the tail to enhance the ‘parachuting’ effect and slow down the sinking (Cable, Reference Cable and Miner1956, Reference Cable1963; Prevot and Bartoli, Reference Prevot and Bartoli1978). An enlarged tail and the development of fin-folds, characteristic of Rhodometopa cercariae, serve the same aim. This behaviour corresponds to the active waiting strategy, characteristic of the cercariae infecting actively moving hosts such as fish (Prokofiev and Galaktionov, Reference Prokofiev and Galaktionov2009). Cercaria buchanani unite by the proximal portions of their tails forming aggregations (Martin and Gregory, Reference Martin and Gregory1951), which corresponds to the prey mimetism strategy (Combes et al., Reference Combes, Fournier, Moné and Théron1994; Prokofiev and Galaktionov, Reference Prokofiev and Galaktionov2009). Some elements of this strategy also seem to be characteristic of large Rhodometopa cercariae, which might be taken by the fish for food objects, e.g. small pelagic polychaetes. The example of renicolids illustrates a high plasticity of the structure of cercariae, which limits the use of the cercarial morphotype as a character for the establishment of taxa of high taxonomical level.
Conclusions
We showed that the use of morphological criteria alone is insufficient for a revision of the Renicolidae. Characters such as the location of testes and vitellaria appear to have been evolving in a convergent manner in different phylogenetic branches of these digeneans. Based on our molecular analyses, we outlined 3 main branches of renicolids for which molecular data are available. Although our data are incomplete, we can tentatively suggest that the first branch (clade I) is characterized by parasitism of adults in gulls (possibly also in sandpipers) and by the presence of the xiphidiocercaria stage in the life cycle. Renicolid species from the second branch (clade II) use sea birds, including gulls, as the definitive host, and their cercariae belong to the Rhodometopa group or to ‘transitional morphotype’. The third branch is represented for now by 1 species, R. somateriae, a typical parasite of sea ducks, with xiphidiocercaria in the life cycle.
In our opinion, it is premature to attempt a thorough taxonomic revision of the renicolids. This task would be meaningful after the accumulation of molecular data, especially on morphologically contrasting species, the elucidation of life cycles of a greater number of species and the determination of the range of their hosts. A detailed analysis of the morphological features of adults and cercariae is also necessary.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182022001500.
Data
Data available on request from the authors.
Acknowledgement
The authors are grateful to the White Sea Biological Station of the Zoological Institute of the Russian Academy of Sciences (ZIN RAS) for providing fieldwork infrastructure. We thank Dr Gennady Atrashkevich, Dr Kira Regel, Dr Kirill Nikolaev and Dr Ivan Levakin for their help with sampling and primary treatment of the material, Dr A. Miroliubov for his help with SEM and Dr Natalia Ershova who provided us with 2 cox1 gene sequences of Renicola cf. roscovita. We also thank Dr D. I. Gibson (NHM, London) for his assistance in taxonomical questions. We wish to acknowledge the ‘Taxon’ Research Resource Center (http://www.ckp-rf.ru/ckp/3038/) of ZIN RAS and the research resource centre ‘Molecular and Cell Technologies’ of St. Petersburg State University for granting access to their facilities. We are grateful to Natalia Lentsman for her help with the manuscript preparation. We thank the anonymous reviewers for their well-considered comments on an earlier draft of the manuscript.
Author's contributions
Contributions are addressed as follows: conceptualization (K. V. G., A. I. S.); data collection (all authors); data analysis and interpretation (K. V. G., A. I. S., A. M. H. B.); writing (K. V. G., A. I. S.).
Financial support
The fieldwork at the White Sea Biological Station and at the ‘Taxon’ Research Resource Center was partly financed by the research programme of Zoological Institute RAS, project number 122031100260-0. The treatment and analysis of the accumulated data were supported by the Russian Science Foundation (Grant No. 18-14-00170-P).
Conflict of interest
None.
Ethical standards
Not applicable.