Introduction
The order Mugiliformes comprises of a single family, Mugilidae Cuvier, 1829, which contains about 70 species, distributed worldwide, commonly known as mullets. The great majority of mullets are highly euryahaline, inhabiting tropical and temperate habitats that include rivers, estuaries, coastal areas and seas (Hotos and Vlahos, Reference Hotos and Vlahos1998; Cardona, Reference Cardona2001; Durand et al., Reference Durand, Shen, Chen, Jamandre, Blel, Diop, Nirchio, Garcia de Leon, Whitfield, Chang and Borsa2012). Due to their omnivorous nature and benthic feeding strategy, mullets are able to feed on a great variety of materials including epiphytic algae, insects, annelids, crustaceans, mollusks and even detritus. As a result of their ecological plasticity, this family plays an important role in the ecosystem, namely by contributing to the flow of energy and matter from the lower to the upper levels (Cardona, Reference Cardona2001; Laffaille et al., Reference Laffaille, Feunteun, Lefebvre, Radureau, Sagan and Lefeuvre2002; Almeida, Reference Almeida2003; Zetina-Rejón et al., Reference Zetina-Rejón, Arreguín-Sánchez and Chávez2003). At the commercial level, the importance of mugilids depends on the geographic region and whether they are cultured for gathering roe or for food consumption. Nonetheless, the world production of mullets is an increasing trend, both in fishery and aquaculture industries (Crosetti and Cataudella, Reference Crosetti, Cataudella and Nash1995; Saleh, Reference Saleh2006).
Several studies have been conducted on the protozoan and metazoan microorganisms parasitizing mullets worldwide (e.g. Merella and Garippa, Reference Merella and Garippa2001; Bahri et al., Reference Bahri, Andree and Hedrick2003; Fioravanti et al., Reference Fioravanti, Caffara, Florio, Gustinelli, Marcer and Quaglio2006; Yurakhno and Ovcharenko, Reference Yurakhno and Ovcharenko2014; Özer and Kirca, Reference Özer and Kirca2015; Sarabeev, Reference Sarabeev2015). According to Yurakhno and Ovcharenko (Reference Yurakhno and Ovcharenko2014), this fish group accounts for the description of about 64 myxosporean species from the genera of the families: Sphaeromyxidae Lom and Noble, 1984; Sphaerosporidae Davis, 1917; Myxidiidae Thélohan, 1892; Myxobilatidae Shulman, 1953 [the genus Ortholinea has been recently transferred to this family, and the family Ortholineidae dismantled (Karlsbakk et al., Reference Karlsbakk, Kristmundsson, Albano, Brown and Freeman2017)]; Sinuolineidae Shulman, 1959; Alatosporidae Schulman, Kovaleva and Dubina, 1979; Chloromyxidae Thélohan, 1892; Kudoidae Meglitsch, 1960 and Myxobolidae Thélohan, 1892. The latter family is the largest within the subclass Myxozoa Grassé, 1970, namely due to the species-richness of the genera Myxobolus Bütschli, 1882 and Henneguya Thélohan, 1892. Worldwide, the genus Myxobolus comprises over 850 species, the majority of which are histozoic in freshwater fish, less frequently infecting hosts from estuarine and marine environments. Few species present coelozoic development and even fewer have been reported to occur in amphibian hosts. On its turn, the genus Henneguya comprises about 200 species that mostly infect freshwater fish, with the exception of ca. 35 species that are known to occur in marine hosts (e.g. Lom and Dyková, Reference Lom and Dyková1992, Reference Lom and Dyková2006; Eiras, Reference Eiras2002; Eiras et al., Reference Eiras, Molnár and Lu2005, Reference Eiras, Zhang and Molnár2014; Eiras and Adriano, Reference Eiras and Adriano2012; Khlifa et al., Reference Khlifa, Miller, Adlard, Faye and Sasal2012; Li et al., Reference Li, Sato, Kamata, Ohnishi and Sugita-Konishi2012; Azevedo et al., Reference Azevedo, Rocha, Matos, Matos, Oliveira, Al-Quraishy and Casal2014; Rocha et al., Reference Rocha, Casal, Garcia, Matos, Al-Quraishy and Azevedo2014a; Özer et al., Reference Özer, Özkan, Gurkanli, Yurakhno and Ciftci2016a).
Traditionally, the taxonomy of myxosporeans was mainly based on spore morphology, and its association with a particular host and organ of infection. However, molecular analyses have been shown that the comparison of spore morphological traits is insufficient for classifying myxosporeans, both at the genus and species level (e.g. Fiala, Reference Fiala2006; Bartošová et al., Reference Bartošová, Fiala and Hypša2009; Fiala and Bartošová, Reference Fiala and Bartošová2010; Liu et al., Reference Liu, Whipps, Gu and Zeng2010). In the case of myxobolids, the artificiality of the morphological criterion hampers identification at the species level, since many species share similar spore shape and size and others present significant intraspecific variations (Lom, Reference Lom1987; Mitchell, Reference Mitchell1989; El-Matbouli et al., Reference El-Matbouli, Fischer-Scherl and Hoffmann1992). Also, although most species are acknowledged to be host and tissue restricted (Molnár, Reference Molnár1994), others have been reported to indiscriminately infect a wide range of hosts and tissues (e.g. Forró and Ezsterbauer, Reference Forró and Eszterbauer2016). Taxonomic comparisons are further challenged by the paucity of reliable data from most original descriptions, which relied solely on light microscopy and schematic line drawings (e.g. Lubat et al., Reference Lubat, Radujkovic, Marques and Bouix1989), with few studies using transmission electron microscopy for ultrastructural characterization. Consequently, several species have been identified as potentially cryptic (Easy et al., Reference Easy, Johnson and Cone2005; Ferguson et al., Reference Ferguson, Atkinson, Whipps and Kent2008; Atkinson et al., Reference Atkinson, Bartošová-Sojková, Whipps, Bartholomew, Okamura, Gruhl and Bartholomew2015), thus warranting authentication through the use of currently accepted taxonomic criteria, i.e. combined spore morphology, host specificity, tissue specificity and molecular data. Considering all of the above, this study provides, for the first time, a detailed and critical revision of the data available for myxobolids reported in mullets, evaluating the reliability of these reports through the comprehensive and careful analysis of original species descriptions, available molecular data and currently accepted taxonomic and phylogenetic criteria. Myxobolids, which occurrence is considered to be reliable in mullets, are herein referred to as having bona fide mugiliform fish hosts. A morphological, ultrastructural and molecular redescription is further performed for the cryptic species Myxobolus exiguus Thélohan, Reference Thélohan1895 from infections in the visceral peritoneum of the thinlip-grey mullet Chelon ramada in the River Minho, Portugal.
Materials and methods
Fish and parasite sampling
Between March 2015 and January 2018, 18 specimens of the thinlip-grey mullet C. ramada (Risso, 1827) (Teleostei and Mugiliformes) were captured from the River Minho (41°56′N, 08°45′W), Vila Nova de Cerveira, Portugal. Specimens were transported live to the laboratory and, prior to dissection, anesthetized with ethylene glycol monophenyl ether until dead. The parasitological survey of several organs and tissues was performed at both the macro and microscopic levels. Cysts and parasitized tissues were prepared for light microscopy, transmission electron microscopy and molecular procedures.
Light microscopy and morphological examination
Parasitized tissues were examined and photographed using a Leitz-Dialux 20 microscope, equipped with a differential interference contrast (DIC) optics. Morphometry was determined from fresh material (Lom and Arthur, Reference Lom and Arthur1989). All measurements include the mean value ± standard deviation (s.d.), range of variation and number of spores measured (range, n).
Transmission electron microscopy
Fragments of parasitized tissue were fixed in 5% glutaraldehyde buffered in 0.2 m sodium cacodylate (pH 7.2) for 20–24 h, washed in the same buffer and postfixed in 2% osmium tetroxide also buffered with 0.2 m sodium cacodylate (pH 7.4) for 3–4 h. All these steps were performed at 4 °C. The samples were then dehydrated in an ascending graded series of ethanol, followed by embedding using a series of oxide propylene and Epon mixtures, ending in EPON. Semithin sections were stained with methylene blue-Azure II. Ultrathin sections were double-contrasted with uranyl acetate and lead citrate, and then examined and photographed using a JEOL 100 CXII TEM (JEOL Optical, Tokyo, Japan), operating at 60 kV.
DNA extraction, amplification and sequencing
Fragments of parasitized tissue were obtained from three fish specimens and preserved in 80% ethanol at 4 °C. Genomic DNA extraction was performed using a GenElute™ Mammalian Genomic DNA Miniprep Kit (Sigma-Aldrich, St Louis, USA), following the manufacturer's instructions. The DNA was stored in 50 µL of TE buffer at −20 °C until further use.
The SSU rRNA gene was amplified and sequenced using both universal and myxosporean-specific primers (Table 1). Polymerase chain reactions (PCRs) were performed in 50 µL reactions using 10 pmol of each primer, 10 nmol of dNTPs, 2 mm MgCl2, 5 µL 10× Taq polymerase buffer, 2.5 units Taq DNA polymerase (Nzytech, Lisbon, Portugal) and approximately 50–100 ng of genomic DNA. The reactions were run on a Hybaid PxE Thermocycler (Thermo Electron Corporation, Milford, Massachusetts), with initial denaturation at 95 °C for 3 min, followed by 35 cycles of 94 °C for 45 s, 53 °C for 45 s and 72 °C for 90 s. The final elongation step was performed at 72 °C for 7 min. Five microlitre aliquots of the PCR products were electrophoresed through a 1% agarose 1× tris-acetate-EDTA buffer (TAE) gel stained with ethidium bromide. PCR products were purified using the ExoFast method, in which an enzymatic clean-up that eliminates unincorporated primers and dNTPs is performed with Exonuclease I (Escherichia coli) and FastAP Thermosensitive (SAP).
PCR primers used for the amplification and sequencing of the SSU rRNA gene.

The PCR products from different regions of the SSU rRNA gene were sequenced directly. The sequencing reactions were performed using BigDye Terminator v1.1 from the Applied Biosystems Kit (Applied Biosystems, Carlsbad, California), and were run on an ABI3700 DNA analyser (Perkin-Elmer, Applied Biosystems, Stabvida, Oeiras, Portugal).
Distance estimation and phylogenetic analysis
The partial sequences obtained for the case isolate were aligned in MEGA 6.06, allowing the construction of the parasite's assembled SSU rRNA sequence, with a total of 2013 bp. In order to calculate distance estimation, a dataset was created solely for the SSU rRNA sequence of the case isolate and all other SSU rRNA sequences available for Myxobolus spp. that have been reported from mullets. This includes M. bramae Reuss, 1906, M. branchialis (Markevitsch, 1932) Landsberg and Lom, Reference Landsberg and Lom1991 and M. rotundus Nemeczek, 1911, despite these species having not been sequenced from hosts of the order Mugiliformes. Of the several SSU rRNA sequences available in the GenBank for M. rotundus, only one (FJ851447) was chosen to represent this species in the dataset, considering that all sequences were identical among each other, with the exception of one (AY165179) that is misidentified (Zhang et al., Reference Zhang, Wang, Li and Gong2010) and, therefore, was not included. Similarly, two SSU rRNA sequences of M. bramae (AF085177 and AF507968) are available in the GenBank but, despite having both been obtained from infections occurring in the cypriniform host Abramis brama Linnaeus, 1758, only one is considered valid (AF507968), thus having been included in the dataset. Sequences were aligned using software MAFFT version 7 available online, and distance estimation was performed in MEGA 6.06, with the p-distance model and all ambiguous positions were removed for each sequence pair.
For phylogenetic analyses, the dataset was widened to encompass other representatives of the clade of myxobolids. The final dataset comprised of a total of 93 SSU rRNA sequences, and included Chloromyxum riorajum (FJ2624481), Myxidium lieberkuehni (X76638) and Sphaerospora oncorhynchi (AF201373) as the outgroup. Sequences were aligned using software MAFFT version 7 available online, and posteriorly manually edited in MEGA 6.06. Phylogenetic trees were calculated from the sequence alignments using maximum-likelihood (ML) and Bayesian inference (BI). Models of nucleotide substitution were evaluated using MEGA 6.06. The general time reversible substitution model with estimates of invariant sites and gamma distributed among site rate variation (GTR + I + Γ) was chosen as the best suited model for the dataset, and was used in both ML and BI analyses. ML analyses were also conducted in MEGA 6.06 (Tamura et al., Reference Tamura, Stecher, Peterson, Filipski and Kumar2013), with bootstrap confidence values calculated from 1000 replicates. BI analyses were performed using MrBayes v3.2.6 (Ronquist and Huelsenbeck, Reference Ronquist and Huelsenbeck2003), with posterior probability distributions generated using the Markov Chain Monte Carlo method, with four chains running, simultaneously, for 500 000 generations, and every 100th tree sampled.
Results
Revised description and taxonomic summary of M. exiguus
Diagnosis
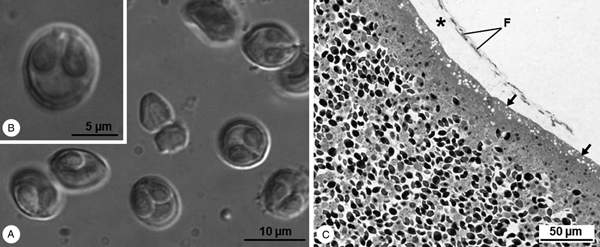
Cysts whitish and spherical, about 1 mm in diameter, located adjacent to the peritoneum lining the viscera. Mature myxospores subspherical in valvular view and ellipsoidal in sutural view, measuring 9.3 ± 0.6 (8.4–10.7) μm in length and 8.2 ± 0.5 (7.6–8.9) μm in width (n = 25). Valves are smooth presenting several markings near the suture line. Two pyriform equal-sized polar capsules located side by side at the myxospores’ anterior pole, 4.8 ± 0.2 (4.4–5.3) μm long and 2.8 ± 0.3 (2.2–3.1) μm wide (n = 25), each containing an isofilar polar filament forming five coils (Fig. 1A and B).
Light micrographs of Myxobolus exiguus infecting the peritoneum of Chelon ramada in the River Minho. (A) DIC micrograph showing some free fresh mature myxospores, subspherical in valvular view and ellipsoidal in sutural view, and containing two polar capsules. (B) Free fresh mature myxospore displaying several markings near the suture line. (C) Semithin section of the periphery of a cyst evidencing the vacuolated ectoplasm (arrow) adhering to loose connective tissue (*), where some fibroblasts (F) are observed.

Ultrastructural description
Cysts’ wall with cytoplasmic expansions forming ladder-like junctions that strongly adhere to mesothelial cells of the peritoneum. Detachment of cytoplasmic portions of mesothelial cells by insertion of the cysts’ wall expansions into connective tissue. Several fibroblasts widely separated by bundles of collagen fibres in loose connective tissue (Figs 1C and 2A–C). Cysts’ ectoplasm highly vacuolated and devoid of cytoplasmic organelles (Figs 1C and 2C); endoplasm rich in mitochondria, vegetative nuclei and containing all sporogonic stages. Sporogony asynchronous and centripetal: generative cells and young sporoblasts located at the cysts’ periphery; immature and mature myxospores in the centre, each within a vacuole-like structure (Figs 1C and 2B, D). Myxospores wall thin and smooth, comprised of two symmetrical valves adhering together along a straight suture line. Polar capsules with a double-layered wall formed by an outer thin electron-dense layer and an inner thick electron-lucent layer. Polar filament coils in an electron-dense homogenous matrix (Fig. 2D–F). Cap-like structure at the apex of polar capsule, directed at the corresponding extrusion pore. Extrusion pores near the suture line, corresponding to the portions of the valves with diminished thickness (Fig. 2G). Sporoplasm at the myxospores’ posterior pole, with two nuclei and several sporoplasmosomes randomly distributed in a heterogeneous matrix (Fig. 2D and E).
Transmission electron micrographs of Myxobolus exiguus infecting the peritoneum of Chelon ramada in the River Minho. (A) Periphery of a cyst (C) adhering to a mesothelial cell near its nucleus (N), and reaching the loose connective tissue (LCT), where fibroblasts (F) are observed widely separated by bundles of collagen fibres. (B) Periphery of a cyst displaying numerous generative nuclei (Gn) and forming cytoplasmic expansions (arrows) that strongly adhere to the mesothelial cells (Mc) and reach the LCT. (C) Detailed aspect of the cytoplasmic expansions forming ladder-like junctions (arrows) that connect the cyst (C) to the mesothelial cells (Mc). Notice the numerous vacuoles (Vs) occupying the cyst's ectoplasm. (D) Longitudinal section of a myxospore in valvular view, located within a vacuole-like structure (*), and displaying its two polar capsules (PC) and binucleate sporoplasm (Sp). E. Longitudinal section of a myxospore in sutural view, depicting the number of polar filament (PF) coils, as well as some sporoplasmosomes (Sps) randomly distributed in the sporoplasm. (F) Transverse section of a myxospore showing its two valves united along a straight suture line (arrowheads), and its two PCs presenting a double-layered wall (arrow) that surrounds an electron-dense matrix (*) and a coiled polar filament (PF). (G) Longitudinal oblique section of a polar capsule displaying its cap-like structure (arrow) in continuity with the valve's extrusion pore (*), located near the suture line (arrowheads).

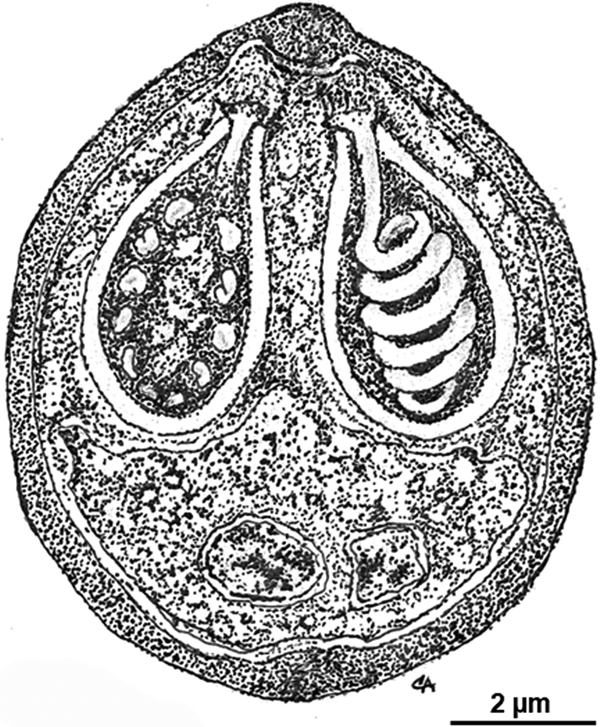
Morphology of the myxospores is represented in a schematic drawing (Fig. 3) depicting the ultrastructural features here described.
Schematic drawing depicting the ultrastructural organization of a myxospore of Myxobolus exiguus in sutural view.

Type host: the thinlip-grey mullet C. ramada (Risso, 1827) (Teleostei, Mugiliformes).
Site of infection: the visceral peritoneum.
Prevalence: three infected in 18 specimens analysed (16.7%).
Type locality: France (Vivier-sur-Mer, Marseille, Banyuls).
Other localities: Tunisia (Ichkeul Lake); Portugal (River Minho).
Pathogenicity: long-term pathological assessments were not performed, but collected and analysed fish did not present evident external symptoms of infection or disease.
Vouchers: one glass slide containing semi-thin sections of the hapantotype was deposited in the Type Slide Collection of the Laboratory of Animal Pathology at the Interdisciplinary Centre of Marine and Environmental Research, Porto, Portugal, reference CIIMAR 2018.19.
Sequences: one assembled SSU rRNA gene sequence with a total of 2013 bp and GenBank accession no. MH236070.
Molecular comparison of the case isolate to other Myxobolus spp. reported from mullets
Pairwise comparisons between the SSU rRNA sequence here obtained and all others available in the GenBank for Myxobolus spp. reported from mugiliform fish hosts (Table 2) revealed the case isolate presenting 100% of similarity to two sequences: one identified as M. exiguus (AY129317), and the other identified as M. muelleri Bütschli, 1882 (AY129314), with these also sharing 100% of similarity between each other. The other SSU rRNA sequences of M. exiguus (AY129316) and M. muelleri (AY129313) obtained from infections in mugiliform fish hosts followed, with 99.4% of similarity to the case isolate, and 100% of similarity between each other. The two remaining available sequences of M. muelleri (AY325284 and DQ439806), which correspond to infections of this species in the cypriniform fish host Squalius cephalus (Linnaeus, 1758), shared 97.0% of similarity between each other, but differed significantly from its conspecific sequences available from mugiliform fish hosts, as well as from those of M. exiguus and the sequence in study, with percentages of identity that varied between 72.2 and 73.4%. All other species resulted in percentages of similarity lower than 90%.
Comparison between the SSU rRNA sequences of the case isolate and all other Myxobolus spp. infecting mugilids: percentage of identity (top diagonal) and nucleotide difference (bottom diagonal)

Phylogenetic positioning of M. exiguus and other myxobolids reported from mugiliform fish hosts
BI and ML analyses resulted in similar topologies, with some entropy in the middle of the tree, namely due to the unstable positioning of the subclade comprising SSU rRNA sequences from myxobolids that infect marine perciform fish hosts. The phylogenetic analyses here performed revealed the case isolate clustering to form a well-supported clade together with most of the SSU rRNA sequences available for myxobolid species described from mugiliform fish hosts: M. exiguus (AY129316 and AY129317); M. muelleri (AY129313 and AY129314); M. parvus Schulman, 1962 (KX242161); M. episquamalis Egusa, Maeno and Sorimachi, 1990 (KC733437); M. bizerti Bahri and Marques, Reference Bahri and Marques1996 (AY129318); M. ichkeulensis Bahri and Marques, Reference Bahri and Marques1996 (AY129315); Myxobolus sp. Kim, Kim and Oh, 2013 (KC733438); M. spinacurvatura Maeno, Sorimachi, Ogawa and Egusa, 1990 (AF378341); and also with members of the sphaeractinomyxon, endocapsa and triactinomyxon collective groups. Exceptions to the mugiliform-infecting clade are the other two SSU rRNA sequences available for M. muelleri (DQ439806 and AY352284), as well as those of M. bramae (AF507968), M. branchialis (JQ388887) and M. rotundus (FJ851447), which cluster among cypriniform-infecting myxobolids (Fig. 4).
Tree topology resulting from the Bayesian analysis of 93 SSU rRNA sequences representative of the clade of myxobolids. Numbers at the nodes are Bayesian posterior probabilities/ML bootstrap values; asterisks represent full support in both methodologies; dashes represent a different branching for the ML tree or a bootstrap support value under 50. Bold taxa correspond to species that have been reported from mugiliform fish hosts, with the invalid sequences of M. muelleri contained within square brackets. The SSU rRNA sequence obtained in this study for M. exiguus is marked with a dark grey box. Final host groups are indicated by vertical lines.

Discussion
Overview of mugiliform-infecting myxobolids
In this paper, we summarize the data available for myxobolids that have mugiliforms as bona fide fish hosts (Tables 3 and 4), with measurements from the original descriptions (whenever given), and updated scientific names for host species (according to FishBase). The vast majority of mugiliform-infecting Myxobolus species are registered from the genus Mugil Linnaeus, 1758, with the flathead grey mullet M. cephalus Linnaeus, 1758 accounting for an astonishing number of 19 species infecting several of its organs in coastal waters of the Mediterranean Sea, Atlantic Ocean, Indian Ocean and North Pacific Ocean; while white mullet M. curema Valenciennes, 1836 accounts for two species from the Atlantic coast off Senegal, and lebranche mullet M. liza Valenciennes, 1836 for one species in Brazilian waters. The genus Chelon Artedi, 1793 accounts for 12 species: four from several organs of golden grey mullet C. auratus (Risso, 1810) (syn. Liza aurata) in the Mediterranean Sea and North Atlantic Ocean; three from thicklip grey mullet C. labrosus in European and North African waters; two from several organs of the leaping mullet C. saliens (Risso, 1810) [syn. L. saliens (Risso, 1810)] in Eurasian coastal waters; one from goldspot mullet C. parsia (Hamilton, 1822) in India; one from tade grey mullet C. planiceps (Valenciennes, 1836), also in India; and one from C. ramada (Risso, 1827) [syn. L. ramada (Risso, 1827)] in Europe and North Africa. Seven species have been reported from hosts of the genus Planiliza Whitley, 1945: four from the gills and gut of large-scale mullet P. macrolepis (Smith, 1846) [syn. C. macrolepis (Smith, 1846)] in India; and three from several organs of so-iuy mullet P. haematocheila (Temminck and Schlegel, 1845) [syn. L. haematocheila (Temminck and Schlegel, 1845)] in Eurasian coastal waters. Other hosts accounting for a single species are: the corsula Rhinomugil corsula (Hamilton, 1822), the yellowtail mullet Sicamugil cascasia (Hamilton, 1822) and the squaretail mullet Ellochelon vaigiensis (Quoy and Gaimard, 1825), all from in India; the hornlip mullet Oedalechilus labiosus (Valenciennes, 1836) in the Red Sea off Egypt; and the bluespot mullet Moolgarda seheli (Forsskål, 1775) off Thailand. On its turn, only two species of Henneguya have been reported from the gills and brain of M. cephalus off Senegal (Table 4). Some of these parasite species have been reported, and even originally described, from more than one mugiliform fish host; it is the case of M. achmerovi Schulman, Reference Schulman1966, M. anili Sarkar, Reference Sarkar1989, M. cheni Schulman, 1962, M. exiguus and M. parvus. Also, some have been indiscriminately reported from several organs and tissues; it is the case of M. achmerovi, M. adeli, M. cephalis, M. dasguptai, M. parvus and M. spinacurvatura (Table 3). The great biodiversity of Myxobolus spp. parasitizing mugiliform fish hosts reflects not only the species-richness of this myxosporean genus, which is the most common in freshwater environments, but probably correlates with the migratory patterns and feeding strategies of mullets. The catadromous nature of mullets allows these fish species to move into freshwater and brackish environments, thus increasing risk of exposure to typically freshwater parasites, such as Myxobolus spp. Also, being benthic feeders, mullets have increased proximity to infected annelids and, therefore, are more prone to contact with waterborne actinosporean stages. The high number of Myxobolus spp. parasitizing M. cephalus in particular, might suggest that this species possesses higher susceptibility than other mullets to myxosporean infection, but most likely simply reflects the higher number of parasitological surveys that have been conducted in this fish species, as a result of its economic importance in fisheries and aquaculture.
Summary of data available for Myxobolus spp. with bona fide mugiliform fish hosts
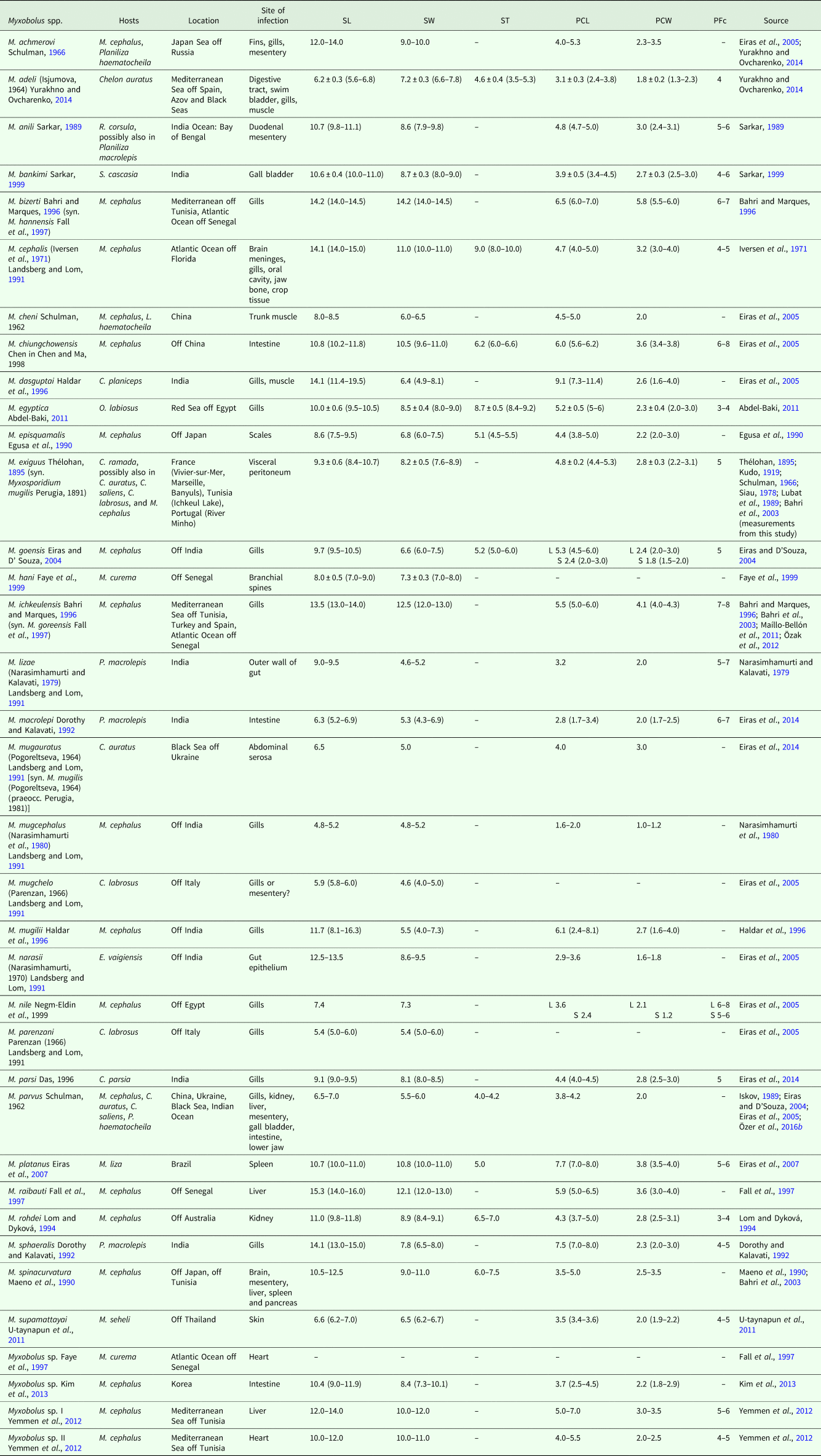
SL, myxospore length; SW, myxospore width; ST, myxospore thickness; PCL, polar capsule length; PCW, polar capsule width; PFc, number of polar filament coils; S, smaller; L, larger. Measurements are means ± s.d. (range) (when available), given in μm.
Summary of data available for Henneguya spp. with bona fide mugiliform fish hosts

SBL, myxospore body length; SBW, myxospore body width; SBT, myxospore body thickness; STL, total length of myxospore; LCA, length of caudal appendages; PCL, polar capsule length; PCW, polar capsule width; PFc, number of polar filament coils; S, smaller; L, larger. Measurements are means ± s.d. (range) (when available), given in μm.
Most mugiliform-infecting species are without molecular data, so that their reports (original and subsequent) have been solely based on morphological traits, which molecular-based systematics reveal are artificial for the reliable description of myxobolids (e.g. Fiala, Reference Fiala2006; Bartošová et al., Reference Bartošová, Fiala and Hypša2009; Fiala and Bartošová, Reference Fiala and Bartošová2010; Liu et al., Reference Liu, Whipps, Gu and Zeng2010). Thus, the legitimacy of these species, and their occurrence in the several reported sites of infection and hosts, must be evaluated through the use of molecular tools; a task that it might prove more difficult than expected, not only due to the frequent occurrence of co-infections (e.g. Molnár et al., Reference Molnár, Marton, Eszterbauer and Székely2006), but also due to the vague boundary between intraspecific and interspecific variability of myxobolids. For instance, our molecular analysis shows that the SSU rRNA sequence provided by Molnár et al. (Reference Molnár, Marton, Eszterbauer and Székely2006) for M. muelleri displays only 97% of similarity to its conspecific sequence by Eszterbauer (Reference Eszterbauer2004), while it also shares a similar percentage of identity (97.0–97.5%) to the SSU rRNA sequences of M. arrabonensis (KP025680), M. bliccae (HM138772) and M. bramae (AF507968). Similarly, high values of intraspecific variability have been reported for different isolates of M. koi (3.0%), M. flavus (1.9%), H. corruscans (2.3%) and H. maculosus (1.9%) (Camus and Griffin, Reference Camus and Griffin2010; Carriero et al., Reference Carriero, Adriano, Silva, Ceccarelli and Maia2013). On the other hand, very low interspecific variability has been reported between M. pseudodispar, M. musculi and M. cyprini (0.3–0.6%); M. pendula and M. pellicides (0.4%); M. fryeri and M. insidiosus (0.5%); M. intramusculi and M. procerus (2.1%); M. paksensis and M. cycloides (2.4%); and M. szentendrensis and M. intimus (2.8%) (Kent et al., Reference Kent, Andree, Bartholomew, El-Matbouli, Desser, Devlin, Feist, Hedrick, Hoffmann, Khattra, Hallett, Lester, Longshaw, Palenzeula, Siddall and Xiao2001; Molnár et al., Reference Molnár, Eszterbauer, Székely, Dán and Harrach2002; Easy et al., Reference Easy, Johnson and Cone2005; Ferguson et al., Reference Ferguson, Atkinson, Whipps and Kent2008; Cech et al., Reference Cech, Borzák, Molnár and Székely2015). Considering all of the above, it is clear that the reliable classification of myxobolids can only result from the comprehensive evaluation of biological, morphological and molecular features. Another problem that researchers face when studying myxobolids, as well as myxosporeans in general, is the amount of unpublished, incomplete, erroneous and confusing data in the GenBank. Thus, it is important to recognize and avoid the use of poor records. In fact, Molnár (Reference Molnár2011) identified and deemed invalid several SSU rRNA sequences of Myxobolus spp. In this study, it is further acknowledged that some SSU rRNA sequences of M. exiguus are erroneously attributed to M. muelleri (AY129313 and AY129314), as are the cases of the SSU rRNA sequences of M. turpisrotundus, M. toyamai and M. cutanei, erroneously designated as M. rotundus (AY165179), T. toyamai (HQ338729) and U. caudatus (JQ388890), respectively.
Assessment of the legitimacy of mugiliform fish as hosts for cryptic species, with the redescription of M. exiguus
The great majority of myxobolids are host specific (Molnár, Reference Molnár1994), with few having been recognized to infect a wide range of hosts belonging to the same taxonomic family or order. For instance, M. pseudodispar has been shown to infect a wide range of cypriniforms (Molnár et al., Reference Molnár, Eszterbauer, Székely, Dán and Harrach2002; Forró and Ezsterbauer, Reference Forró and Eszterbauer2016) and, in the same manner, M. cerebralis is known to parasitize several species of salmonids (El-Matbouli et al., Reference El-Matbouli, Hoffmann, Schoel, McDowell and Hedrick1999; Hoffman, Reference Hoffman1999; Hedrick et al., Reference Hedrick, McDowell, Mukkatira, Georgiadis and MacConnell2001; Ferguson et al., Reference Ferguson, Atkinson, Whipps and Kent2008). Similarly, most myxobolids have well-defined sites of infection (Molnár, Reference Molnár1994; Reference Molnár2002; Molnár et al., Reference Molnár, Marton, Eszterbauer and Székely2006), but several species have been indiscriminately reported from multiple organs, either due to misidentifications, or to the parasite's specificity to a given tissue. For instance, the plasmodia of M. diaphanus develop in the connective tissue of several different organs of the banded killifish Fundulus diaphanus (Lesueur, 1817) (Cone and Easy, Reference Cone and Easy2005). As such, indicating specific tissue tropism, rather than just the organ of infection, is necessary for the correct characterization of myxobolids, and myxosporeans in general. Furthermore, recent phylogenetic studies have consistently shown vertebrate host group as the strongest evolutionary signal for myxobolids (e.g. Ferguson et al., Reference Ferguson, Atkinson, Whipps and Kent2008; Carriero et al., Reference Carriero, Adriano, Silva, Ceccarelli and Maia2013; Rocha et al., Reference Rocha, Casal, Garcia, Matos, Al-Quraishy and Azevedo2014a), followed by the aquatic environment of the host species and tissue tropism (Kent et al., Reference Kent, Andree, Bartholomew, El-Matbouli, Desser, Devlin, Feist, Hedrick, Hoffmann, Khattra, Hallett, Lester, Longshaw, Palenzeula, Siddall and Xiao2001; Eszterbauer, Reference Eszterbauer2004; Holzer et al., Reference Holzer, Sommerville and Wootten2004; Fiala, Reference Fiala2006).
Overall, about 38 species and four records of Myxobolus have been performed from mugiliform fish hosts worldwide (Table 3; Naidenova et al., Reference Naidenova, Schulman and Donets1975; Donets, Reference Donets1979; Ibragimov, Reference Ibragimov1987; Yurakhno and Maltsev, Reference Yurakhno and Maltsev2002; Yurakhno, Reference Yurakhno and Nigmatullin2004; Eiras et al., Reference Eiras, Abreu, Robaldo and Pereira Júnior2007, Reference Eiras, Zhang and Molnár2014), with C. ramada being host to M. exiguus and M. muelleri (Siau, Reference Siau1978; Lubat et al., Reference Lubat, Radujkovic, Marques and Bouix1989; Thélohan, Reference Thélohan1895; Bahri et al., Reference Bahri, Andree and Hedrick2003). Both these species are cryptic, having been described from a wide range of tissues and hosts. The original description of M. exiguus simultaneously reported the parasite from the stomach epithelium, pyloric caeca, kidney and spleen of the mugilids C. labrosus and C. ramada in France, and from the gills of the cypriniform fish host A. brama (Linnaeus, 1758), with basis on a single schematic line drawing and some spores’ measurements (Thélohan, Reference Thélohan1895). Since then, this parasite was reported from several other cypriniform and mugiliform fish hosts inhabiting freshwater across Europe and the Mediterranean Sea, including: Alburnus alburnus (Linnaeus, 1758), Leuciscus aspius (Linnaeus, 1758), Blicca bjoerkna (Linnaeus, 1758), Chondrostoma nasus (Linnaeus, 1758), L. idus (Linnaeus, 1758), Pelecus cultratus (Linnaeus, 1758), Rutilus rutilus (Linnaeus, 1758), Scardinius erythrophthalmus (Linnaeus, 1758), C. auratus, C. saliens and Mugil cephalus (Kudo, Reference Kudo1919; Siau, Reference Siau1978; Lubat et al., Reference Lubat, Radujkovic, Marques and Bouix1989; Bahri et al., Reference Bahri, Andree and Hedrick2003). Bahri et al. (Reference Bahri, Andree and Hedrick2003) sequenced the SSU rRNA gene of this parasite (AY129316, AY129317) using samples obtained from the intestine of C. ramada from the Ichkeul Lake, Tunisia. However, these authors also provided two SSU rRNA sequences for M. muelleri (AY129313 and AY129314) obtained from the mesenteric vessels of C. ramada, which our analysis shows, are equal to the ones provided for M. exiguus. According to the phylogenetic analysis performed by Bahri et al. (Reference Bahri, Andree and Hedrick2003), the sequences obtained for M. muelleri and M. exiguus differed solely by three nucleotide substitutions, with the myxospores exhibiting subtle morphological differences and undergoing sporogony in different organs (M. muelleri in mesenteric vessels and M. exiguus in the intestine). However, acknowledging that C. ramada should not be considered as a bona fide host for M. muelleri, it seems probable that Bahri et al. (Reference Bahri, Andree and Hedrick2003) sequenced the SSU rRNA gene of the same parasite, M. exiguus, from the visceral peritoneum, which lines the intestine and double-folds to form the mesentery attaching to the gastrointestinal organs. Molecular comparison of the SSU rRNA sequence here obtained to those obtained by Bahri et al. (Reference Bahri, Andree and Hedrick2003) identified the parasite in study as M. exiguus. This identification is corroborated by the specificity of the site of infection and host species, but also by the morphological characters of the myxospores, which dimensions are congruent with those provided in the original description of M. exiguus by Thélohan (Reference Thélohan1895), as well as those provided by Bahri et al. (Reference Bahri, Andree and Hedrick2003) (albeit being slightly bigger, and presenting fewer polar filament coils (five) than those (six to seven) described for the myxospores collected from the mesenteric vessels and misidentified as M. muelleri). Bahri et al. (Reference Bahri, Andree and Hedrick2003) further described the valves as smooth, with 10–12 markings appearing near the suture line, as it was also observed in this study. Considering that M. exiguus was originally described on the basis of a single schematic line drawing, and that its morphometrics were obtained from myxospores that most probably belong to different Myxobolus spp., as shown by the several tissues of infection and hosts accounted for in the original description, this paper aims to present a comprehensive morphological and molecular redescription of this myxobolid species. Chelon ramada is here suggested as type host for M. exiguus, not only because it is among the host species included in the original description, but also because the parasite gained its molecular identity from infections in the visceral peritoneum of this mullet species. Thus, other fish species, namely cypriniforms, should be disregarded as bona fide hosts for M. exiguus. Similarly, the visceral peritoneum is suggested as type tissue, so that other tissues and organs of infection, such as the gills, should also be disregarded as sites of infection for M. exiguus.
On its turn, M. muelleri, type species of the genus, was originally described from the gills of several cypriniforms, without indication of a type host, and since then reported from several different tissues and organs in a great number of fish hosts from Eurasia and North America, including: the kidney and ovaries of Phoxinus phoxinus (Linnaeus, 1758); the eyes of Symphodus melops (Linnaeus, 1758) and A. alburnus; the gills of Zingel asper (Linnaeus, 1758), Barbus barbus (Linnaeus, 1758), R. rutilus and Lota lota (Linnaeus, 1758); the pseudobranches of Cottus gobio (Linnaeus, 1758); and the gills, fins, eyes, mesentery, intestine, gall bladder, urinary bladder, liver, kidney, spleen, gonads, heart and muscle of M. cephalus, C. auratus, C. saliens and C. ramada (Kudo, Reference Kudo1919; Shulman, Reference Shulman and Bauer1984; Lom and Dyková, 1992; Bahri et al., Reference Bahri, Andree and Hedrick2003; Molnár et al., Reference Molnár, Marton, Eszterbauer and Székely2006; Umur et al., Reference Umur, Pekmezci, Beyhan, Gurler and Acici2010; Yurakhno and Ovcharenko, Reference Yurakhno and Ovcharenko2014). Considering the currently accepted evolutionary signals of Myxosporea, namely the importance of host affinity for myxobolids (Carriero et al., Reference Carriero, Adriano, Silva, Ceccarelli and Maia2013), it is clear that M. muelleri constitutes a species-complex, comprising several species that are phenotypically similar and, therefore, have been misidentified. This is further supported by the astonishing variation in the shape and size of the myxospores of M. muelleri between reports. In their taxonomic revision of the genus Myxobolus, Landsberg and Lom (Reference Landsberg and Lom1991) suggested settling S. cephalus as the type host for M. muelleri. Eszterbauer (Reference Eszterbauer2004) provided a SSU rRNA sequence (AY325284) for the parasite sampled from the gills of the chub S. cephalus from the River Danube, Hungary; but it was Molnár et al. (Reference Molnár, Marton, Eszterbauer and Székely2006) who characterized M. muelleri by providing a comprehensive morphological and molecular redescription of the species from samples obtained from the gills, as well as from the swimbladder of S. cephalus in Hungary (DQ439806). Bahri et al. (Reference Bahri, Andree and Hedrick2003) had supposedly sequenced M. muelleri from the mesenteric vessels of C. ramada from the Ichkeul Lake, Tunisia. Nonetheless, our molecular analyses show that the sequences obtained from the mugilid fish host (AY129313 and AY129314) display lower percentage of identity (72.2–73.4%) than those obtained by Eszterbauer (Reference Eszterbauer2004) and Molnár et al. (Reference Molnár, Marton, Eszterbauer and Székely2006), revealing that the parasite infecting C. ramada is not M. muelleri, but M. exiguus, as previously stated. Thus, our study agrees with Molnár (Reference Molnár2011) in that the SSU rRNA sequences obtained by Bahri et al. (Reference Bahri, Andree and Hedrick2003) for M. muelleri should be deemed invalid. We further suggest disregarding C. ramada and other mugilids has bona fide hosts for M. muelleri, as well as gadids, percids and scorpaenids.
Analysing the legitimacy of mugiliform fish as hosts for other Myxobolus spp. reported from mullets, some species require obvious attention, as their original descriptions were performed from cypriniforms. Myxobolus acutus (Fujita, 1912) Landsberg and Lom, Reference Landsberg and Lom1991, originally S. acuta, was first described from the gills of Carassius auratus gibelio in Japan, and later reported from the scales of M. cephalus and P. haematocheila from several Russian Rivers, and from the Sea of Japan (Landsberg and Lom, Reference Landsberg and Lom1991; Eiras et al., Reference Eiras, Molnár and Lu2005; Yurakhno and Ovcharenko, Reference Yurakhno and Ovcharenko2014). Myxobolus bramae was originally described from the gills of A. brama in Russia, and later reported from a wide range of organs and tissues of M. cephalus in the Black Sea, including the gills, skin, fins, heart, muscle, mouth, oesophagus, intestine, swim bladder, liver, gall bladder, spleen and kidney (Iskov, Reference Iskov, Markevitch and Schulman SS1989; Eiras et al., Reference Eiras, Molnár and Lu2005; Yurakhno and Ovcharenko, Reference Yurakhno and Ovcharenko2014). Both Andree et al. (Reference Andree, Székely, Molnár, Gresoviac and Hedrick1999) and Eszterbauer (Reference Eszterbauer2004) deposited an SSU rRNA sequence for this parasite obtained from the gills of its type host in Hungary (AF085177 and AF507968, respectively), which turned out shared very low percentage of similarity between each other. Considering that the common bream is the host for several other gill-infecting Myxobolus spp. in Hungary (Molnár and Székely, Reference Molnár and Székely1999), Eszterbauer (Reference Eszterbauer2004) and Ferguson et al. (Reference Ferguson, Atkinson, Whipps and Kent2008) suggested that the samples used by Andree et al. (Reference Andree, Székely, Molnár, Gresoviac and Hedrick1999) were probably contaminated by myxospores of another species and the corresponding SSU rRNA sequence was ultimately deemed invalid (Molnár, Reference Molnár2011). Myxobolus rotundus, which was also originally described from the gills of A. brama in Germany, as well as from the gudgeon Gobio gobio (Linnaeus, 1758), was later reported from the gills, heart and other internal organs of C. auratus in the Black Sea (Donets, Reference Donets1979; Iskov, Reference Iskov, Markevitch and Schulman SS1989; Eiras et al., Reference Eiras, Molnár and Lu2005). This parasite ultimately had its SSU rRNA gene characterized from infections in its type tissue and host (Székely et al., Reference Székely, Hallett, Atkinson and Molnár2009). Myxobolus branchialis, originally Myxosoma branchialis, was first described from the gills of B. barbus in Ukraine, but then reported from the gills, kidney and spleen of M. cephalus, C. auratus and C. saliens in the Caspian Sea and Black Sea (Schulman, Reference Schulman1966; Ibragimov, Reference Ibragimov1987; Iskov, Reference Iskov, Markevitch and Schulman SS1989; Eiras et al., Reference Eiras, Molnár and Lu2005). Molnár et al. (Reference Molnár, Eszterbauer, Marton, Székely and Eiras2012) gave molecular identity to the parasite upon its redescription from the gills of common barbel and Iberian barbel Luciobarbus bocagei (Steindachner, 1864) in Hungary and Portugal. Finally, M. circulus (Achmerov, 1960) Landsberg and Lom, Reference Landsberg and Lom1991, originally M. circulus, was first described from the gills of Cyprinus carpio Linnaeus, 1758 in Russia, being later reported from the gills, fins, muscle and kidney of M. cephalus in the Black Sea (Naidenova et al., Reference Naidenova, Schulman and Donets1975; Iskov, Reference Iskov, Markevitch and Schulman SS1989; Yurakhno, Reference Yurakhno and Nigmatullin2004; Eiras et al., Reference Eiras, Molnár and Lu2005). Given the molecular trends accepted for myxobolids (Andree et al., Reference Andree, Székely, Molnár, Gresoviac and Hedrick1999; Kent et al., Reference Kent, Andree, Bartholomew, El-Matbouli, Desser, Devlin, Feist, Hedrick, Hoffmann, Khattra, Hallett, Lester, Longshaw, Palenzeula, Siddall and Xiao2001; Eszterbauer, Reference Eszterbauer2004; Fiala, Reference Fiala2006; Carriero et al., Reference Carriero, Adriano, Silva, Ceccarelli and Maia2013), it is highly unlikely for a cypriniform-infecting species to parasitize members of the order Mugiliformes. Thus, we suggest disregarding mullets as legitimate hosts for all these species, which should be considered restricted to their original hosts, and others proven through means of molecular tools (as is the case of M. branchialis) (Table 5). For the same reason, we suggest disregarding the cyprinid C. carpio haematopterus as a bona fide host for M. achmerovi Schulman, Reference Schulman1966, which original description was performed from the gills, fins and mesentery of M. cephalus and P. haematocheila (Eiras et al., Reference Eiras, Molnár and Lu2005; Yurakhno and Ovcharenko, Reference Yurakhno and Ovcharenko2014).
Summary of valid data for Myxobolus spp. erroneously reported from mugiliform fish hosts
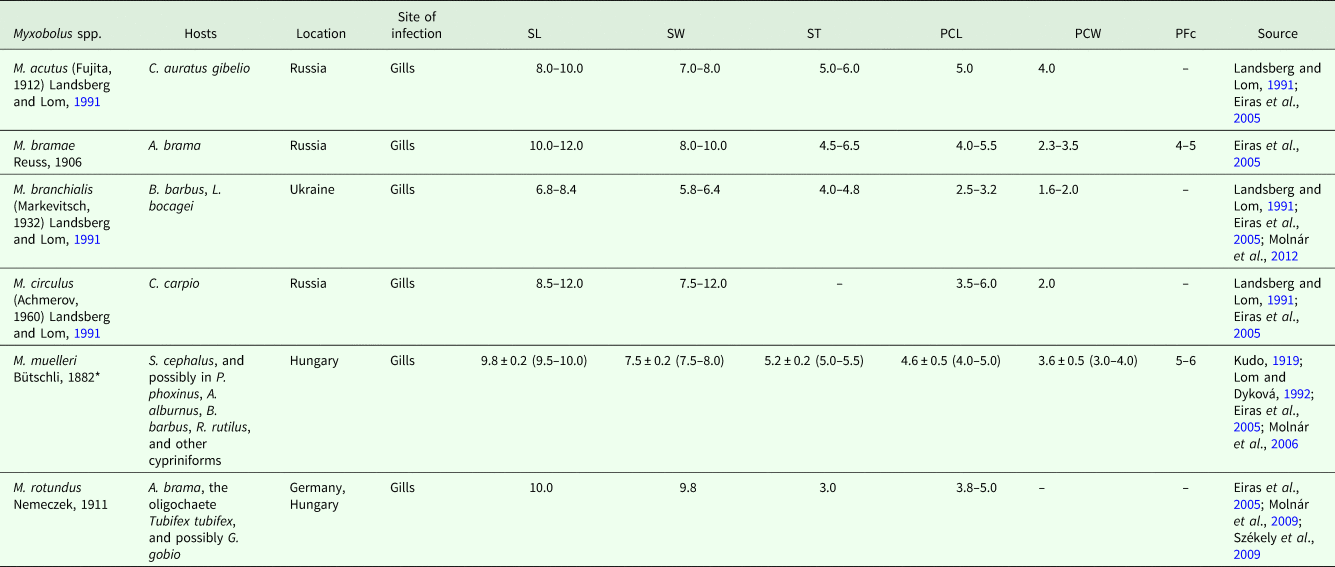
SL, myxospore length; SW, myxospore width; ST, myxospore thickness; PCL, polar capsule length; PCW, polar capsule width; PFc, number of polar filament coils; S, smaller; L, larger. Measurements are means ± s.d. (range) (when available), given in μm.
*Data from the morphological redescription and molecular identification of the parasite from its original site of infection and host species in Hungary (Molnár et al., Reference Molnár, Marton, Eszterbauer and Székely2006).
Ultrastructure of M. exiguus
Ultrastructural studies of the plasmodial and sporogonic development can provide valuable supplementary information for the distinction of individual species and, more importantly, for understanding host–parasite interactions (Hallett and Diamant, Reference Hallett and Diamant2001; Rocha et al., Reference Rocha, Casal, Rangel, Severino, Castro, Azevedo and Santos2013; Reference Rocha, Casal, Al-Quraishy and Azevedo2014b). Current (Reference Current1979) further suggested that certain ultrastructural differences in the plasmodium wall may partially correlate with the degree of pathogenicity of the parasite.
In general, the wall of cyst-forming myxosporeans is structured similarly between different species and genera, with few variations that probably result from the physical and biological conditions of the tissue/organ of infection, as well as the host immune response (Lom and Dyková, 1992; Hallett and Diamant, Reference Hallett and Diamant2001; Rocha et al., Reference Rocha, Casal, Rangel, Severino, Castro, Azevedo and Santos2013). Most histozoic species present a smooth plasmodial wall, with pinocytosis or phagocytosis being widely accepted as the processes supplying the nutrients necessary for the parasite's development (Current and Janovy, Reference Current and Janovy1976; Mitchell, Reference Mitchell and Krier1977; Current, Reference Current1979; Current et al., Reference Current, Janovy and Knight1979; Casal et al., Reference Casal, Matos and Azevedo2006; Azevedo et al., Reference Azevedo, Casal, Marques, Silva and Matos2011). The ultrastructural features of the plasmodial development of M. exiguus is similar to that of other histozoic myxosporeans only in that pinocytotic activity is evidenced by the large number of vacuoles occupying the ectoplasmic layer. Its plasmodial wall, however, displays an irregular outline, with peripheral projections expanding the parasite–host interface, probably for optimizing nutrient intake. This feature has been reported for few other histozoic species, e.g. M. insignis (Azevedo et al., Reference Azevedo, São Clemente, Casal, Matos, Oliveira, Al-Quraishy and Matos2013) and M. filamentum (Naldoni et al., Reference Naldoni, Zatti, Capodifoglio, Milanin, Maia, Silva and Adriano2015), since the differentiation of peripheral projections is common to the plasmodial development of coelozoic species (Sitjà-Bobadilla and Alvarez-Pellitero, Reference Sitjà-Bobadilla and Alvarez-Pellitero1993; Reference Sitjà-Bobadilla and Alvarez-Pellitero2001; Rocha et al., Reference Rocha, Casal, Matos, Matos, Dkhil and Azevedo2011). Overall, the ultrastructural study performed revealed significant unique features of the plasmodial development of M. exiguus, namely in its attachment to the mesothelial cells. In turn, the sporogony of M. exiguus is essentially similar to that of other myxobolids with centric asynchronous development (e.g. Current, Reference Current1979; Current et al., Reference Current, Janovy and Knight1979; Naldoni et al., Reference Naldoni, Zatti, Capodifoglio, Milanin, Maia, Silva and Adriano2015), in that the ectoplasm appears highly vacuolated and devoid of cytoplasmic organelles, while the endoplasm is riddled with organelles and the parasite's different developmental stages: generative cells and developing sporoblasts at the periphery, and immature myxospores at the centre.
Phylogenetic analysis
The phylogenetic analysis here performed is congruent with previously published cladograms (e.g. Kent et al., Reference Kent, Andree, Bartholomew, El-Matbouli, Desser, Devlin, Feist, Hedrick, Hoffmann, Khattra, Hallett, Lester, Longshaw, Palenzeula, Siddall and Xiao2001; Fiala, Reference Fiala2006; Ferguson et al., Reference Ferguson, Atkinson, Whipps and Kent2008; Carriero et al., Reference Carriero, Adriano, Silva, Ceccarelli and Maia2013; Rocha et al., Reference Rocha, Casal, Garcia, Matos, Al-Quraishy and Azevedo2014a), in that it shows the vertebrate host group as the most relevant evolutionary signal for myxobolids, with tissue tropism and aquatic environment playing less conspicuous roles. Accordingly, all SSU rRNA sequences available for myxobolids with legitimate mugiliform fish hosts are here shown clustering together to form a well-supported subclade of the clade of myxobolids. Members of the sphaeractinomyxon, endocapsa and triactinomyxon collective groups that probably play a role in the life cycles of myxobolid species from mullet hosts also cluster within this subclade. In turn, the valid SSU rRNA sequences available for M. bramae, M. branchialis, M. muelleri and M. rotundus, which were obtained from their cypriniform type hosts, all cluster within the clade comprising cypriniform-infecting myxobolids. This emphasizes the incongruence of reporting these species from mugiliform fish hosts, as well as the artificiality of using morphological characters for species identification. The fallibility of morphology as an evolutionary signal for myxobolids has been well reported in several studies (e.g. Fiala, Reference Fiala2006; Bartošová et al., Reference Bartošová, Fiala and Hypša2009; Fiala and Bartošová, Reference Fiala and Bartošová2010; Liu et al., Reference Liu, Whipps, Gu and Zeng2010). For instance, traditional taxonomy separates the genera Myxobolus and Henneguya according to the absence or presence of caudal appendages, respectively (Lom and Dyková, 2006). Nonetheless, molecular-based taxonomy has consistently shown the convergent evolution of caudal appendages (Fiala and Bartošová, Reference Fiala and Bartošová2010; Liu et al., Reference Liu, Whipps, Gu and Zeng2010), revealing that this morphological trait bares little insight into the relationships of myxobolids. In fact, abnormal spore extensions have been reported for some Myxobolus spp. (Mitchell, Reference Mitchell1989; Cone and Overstreet, Reference Cone and Overstreet1997; Bahri, Reference Bahri2008; Liu et al., Reference Liu, Whipps, Gu and Zeng2010, Reference Liu, Whipps, Gu, Huang, He, Yang and Molnár2013, Reference Liu, Whipps, Nie and Gu2014, Reference Liu, Zhang, Batueva and Voronin2015; Zhang et al., Reference Zhang, Al-Quraishy and Abdel-Baki2014), including M. bizerti from the gills of M. cephalus, and M. exiguus (misidentified as M. muelleri, as previously mentioned) from the mesenteric vessels of C. ramada (e.g. Longshaw et al., Reference Longshaw, Frear and Feist2003; Eiras et al., Reference Eiras, Molnár and Lu2005; Kaur and Singh, Reference Kaur and Singh2010; Camus et al., Reference Camus, Dill, Rosser, Pote and Griffin2017).
It has been suggested that the origins and radiations of myxosporean parasites probably reflect the evolution of their fish hosts (e.g. Carriero et al., Reference Carriero, Adriano, Silva, Ceccarelli and Maia2013; Kodádková et al., Reference Kodádková, Bartošová-Sojková, Holzer and Fiala2015). Evolutionary phylogenies of fish reveal that the order Mugiliformes is monophyletic in relation to its sister taxa, despite the polyphyly and/or paraphyly that takes place at the genera-level, due to systematics based in poorly informative anatomical characters (Durand et al., Reference Durand, Shen, Chen, Jamandre, Blel, Diop, Nirchio, Garcia de Leon, Whitfield, Chang and Borsa2012). The phylogenetic analyses here performed supports the coevolutionary history of myxosporeans and their vertebrate hosts, as it shows all legitimate mugiliform-infecting myxobolids clustering together to form a monophyletic well-supported subclade within the clade of myxobolids. In the future, it would be interesting to unravel the significance that this coevolutionary history had in the adaptive strategies of myxosporeans to different micro- and macroenvironments.
Financial support
This work was financially supported by FCT (Lisbon, Portugal) within the scope of the Ph.D. fellowship grant attributed to S. Rocha (SFRH/BD/92661/2013) through the programme QREN-POPH/FSE; and the Eng° António de Almeida Foundation (Porto, Portugal).
Conflicts of interest
None.
Ethical standards
The work developed in this study was performed in accordance with European ethical standards.