Introduction
Understanding the evolutionary patterns and processes that drive the generation of new phenotypes and occupation of previously unexploited environments (the occurrence of novelty and innovation, sensu Erwin Reference Erwin2017) is a fundamental goal of evolutionary study. The topic of novelty and innovation has most frequently been explored through the lens of adaptive radiation (Losos Reference Losos2010; Yoder et al. Reference Yoder, Clancey, Roches, Eastman, Gentry, Godsoe, Hagey, Jochimsen, Oswald, Robertson, Sarver, Schenks, Spear and Harmon2010), which posits that ecological opportunities combined with chance phenotypic evolution permits exploration of new ecospace (Stroud and Losos Reference Stroud and Losos2016). This model of adaptive radiation has been invoked as the dominant causal factor of major evolutionary events in Earth's history, including the Cambrian explosion (Erwin and Valentine Reference Erwin and Valentine2013), invasion of freshwater ecosystems (Davis et al. Reference Davis, De Grave, Delmer and Wills2018), colonization of land (Benton Reference J2010), and recovery from mass extinction events (Toljagić and Butler Reference Toljagić and Butler2013). In turn, the innovation of phenotypes is recognized to occur through heritable alterations in the timing of an organism's development, a process known as heterochrony (McNamara and McKinney Reference McNamara and McKinney2005; McNamara Reference McNamara2012; Colangelo et al. Reference Colangelo, Ventura, Piras, Bonaiuti and Ardizzone2019). Therefore, heterochronic processes result in new morphological characteristics that can allow organisms to exploit new environments and subsequently diversify.
Natural biological systems are complex entities, in which both environmental interaction and historical components are potentially equally important (Eldredge and Salthe Reference Eldredge and Salthe1984; Vrba and Eldredge Reference Vrba and Eldredge1984; Erwin Reference Erwin2015b; Lamsdell et al. Reference Lamsdell, Congreve, Hopkins, Krug and Patzkowsky2017; Congreve et al. Reference Congreve, Falk and Lamsdell2018). Evolutionary trajectories are defined by the interaction of two semi-independent genealogical and ecological hierarchies (Eldredge and Salthe Reference Eldredge and Salthe1984; Congreve et al. Reference Congreve, Falk and Lamsdell2018); as such, ecological changes have the potential to influence heterochronic processes to the same extent as heterochronic changes might alter the potential for ecological occupation. Therefore, pluralistic approaches are required to determine the relative importance of ecological or evolutionary mechanisms behind biotic responses (Lamsdell et al. Reference Lamsdell, Congreve, Hopkins, Krug and Patzkowsky2017; Tucker et al. Reference Tucker, Cadotte, Carvalho, Davies, Ferrier, Fritz, Grenyer, Helmus, Jin, Mooers, Pavoine, Purschke, Redding, Rosauer, Winter and Mazel2017; Congreve et al. Reference Congreve, Falk and Lamsdell2018), representing a synthesis of macroevolutionary and macroecological topics and methods.
Heterochrony, however, has rarely been studied within an explicitly phylogenetic context (Bardin et al. Reference Bardin, Rouget and Cecca2017). Cranial evolution of anguimorphan lizards (Bhullar Reference Bhullar2012) and theropod dinosaurs (Bhullar et al. Reference Bhullar, Marugán-Lobón, Racimo, Bever, Rowe, Norrell and Abzhanov2012) has been examined by placing heterochronic changes within a phylogenetic hypothesis, but no attempt to incorporate ecological occupation into such investigations has been made. These studies have also focused on only a subset of morphological characters that may show heterochronic trends, which may obscure overall patterns of heterochrony within lineages, as traits display patterns of mosaic evolution (Hopkins and Lidgard Reference Hopkins and Lidgard2012; Hunt et al. Reference Hunt, Hopkins and Lidgard2015), whereby individual traits may exhibit different modes of evolution within a lineage. Recognizing overall heterochronic trends within a lineage beyond the trajectories of individual traits is key to understanding the role of heterochrony in evolution. Furthermore, appropriate methods to quantitatively compare heterochronic trends between lineages have been lacking until now. Here, I use phylogenetic paleoecology to analyze patterns in ecological affinity and heterochronic trends across xiphosuran chelicerates and present a new metric for quantitatively representing heterochronic changes within an evolutionary lineage.
Data and Methods
Quantifying Heterochrony
A novel approach for quantifying heterochrony is proposed herein, whereby a character matrix comprising a series of multistate characters coded for each species within the analysis is developed. Each character represents an aspect of morphology that may exhibit a paedomorphic, peramorphic, or neutral heterochronic expression. The peramorphic and paedomorphic conditions for each character are determined based on a ranked series of criteria:
1. direct observations of ontogenetic changes within the target species;
2. ontogenetic changes observed in closely related species;
3. ontogenetic changes observed in extant relatives;
4. comparison with outgroup juvenile morphology or ontogeny for character polarity; and
5. comparison with outgroup adult morphology for character polarity.
Once the polarity of the morphology of heterochronic expression for each character is established, characters are coded for each species and assigned a score. A paedomorphic condition is assigned a negative (−1) score, a peramorphic condition is assigned a positive (+1) score, and a neutral condition is assigned a score of 0. If a character cannot be coded for a given species, either due to it not being preserved or its condition being otherwise unclear, it is not given a score and does not contribute to the analysis. The total number of characters within the matrix depends on the number of identifiable heterochronically influenced traits within a lineage; as such, organisms with more complex morphologies and better-understood ontogenies will result in larger heterochronic matrices. A larger matrix allows for increased granularity in the heterochronic weighting, and ideally a matrix will comprise 10 or more characters; however, it is possible to generate a heterochronic weighting for a taxon that has as few as three characters coded.
Each of the characters coded for a species in the matrix contributes to the heterochronic weighting of the species, calculated as:
where Hw is the heterochronic weighting of species j, derived from the mean of the combined heterochronic scores (η) of n characters, resulting in a value between 1.00 (more peramorphic) and −1.00 (more paedomorphic). In turn, the heterochronic weighting of a given clade is calculated as:
where [Hw] is the heterochronic weighting of clade k, derived from the mean of the combined heterochronic weightings (Hw) in N species, again resulting in a value between 1.00 and −1.00. The heterochronic weighting of a clade therefore represents the average of the heterochronic weightings of its constituent species; it is explicitly not an ancestral state reconstruction. What heterochronic weighting does ensure is that all taxa within an analysis have a directly comparable metric, permitting analysis of heterochronic trends and correlation with other evolutionary phenomena. While the scale of the heterochrony metric raw values within an analysis is relative and cannot be directly compared between analyses, patterns and timing of heterochronic shifts can be directly compared between datasets in an identical manner to quantitative analyses of morphospace (Korn et al. Reference Korn, Hopkins and Walton2013; Hopkins and Gerber Reference Hopkins, Gerber, de la Rosa and Müller2017).
To determine whether the heterochronic weighting of a clade represents a concerted trend shift distinct from what could be expected from random, nondirectional evolution, the observed clade scores are randomized across the tree topology. In total, 100,000 randomizations are performed, with the distribution of the randomized heterochronic weightings then collated into a histogram for each clade to which the observed clade heterochronic weighting can be directly compared. If the actual weighting score falls within either tail of the distribution, the weighting is significantly different from what would be expected under random (nondirectional) character change. Randomizing the heterochronic weightings across the observed phylogeny accounts for vagaries such as clade size and topological relationships that may impact the distribution of heterochronic weighting scores. The heterochronic weights retrieved for a given clade are therefore compared with a distribution of weight scores over an identical topology; as such, randomized distributions should be generated for each topology used in an analysis. In this way, disparate analyses can determine whether clades fall within the tails of the expected heterochronic weighting distribution given their phylogenetic topology.
Node- versus Tip-based Analyses
Scores for heterochronic weighting can be applied two ways, through either a node-based or a tip-based analysis (Fig. 1). Applying heterochronic weighting through node-based calculations accurately encompasses the fact that whether a condition for a given taxon is peramorphic, paedomorphic, or neutral is dependent on the condition in its ancestor as inferred by ancestral state reconstruction, and so it is possible for the polarity of a character to change over the evolutionary history of a clade. As such, when applying node-based calculations, it is critical to document the polarity of each character for each node in the phylogeny. Heterochronic weights are calculated for nodes and are reset at the base of each branch of the phylogeny, with individual character weights representing the inferred shift or transition from the ancestral condition. Node-based application of heterochronic weighting more accurately reflects the actual process of heterochrony; however, it is sensitive to sampling, as failing to sample a species can result in an incorrect assumption of character polarity for a given node, while large gaps in sampling can result in unnatural stacking of transitions at sampled nodes. The analysis also becomes incredibly sensitive to phylogenetic topology, as the relative order in which nodes occur will have a major impact on character polarity. Therefore, node-based heterochronic weighting ideally requires a well-constrained and dated phylogeny for an evenly sampled group with good ontogenetic data. In practice, only a minority of clades may combine all of these attributes, limiting the extent to which heterochronic weighting can be applied from a node-based perspective—although the promise of phylogenetic advances in a variety of molluscan groups such as ammonoids and gastropods, which regularly preserve details of their ontogeny and have a good fossil record, marks them as potential candidates for node-based heterochronic weighting.
Example of heterochronic weightings calculated from three traits evolving across a lineage comprising taxa A–G. In the top tree, evolution of the three traits is shown with their condition (peramorphic + 1, paedomorphic −1, or neutral 0) for each species and internal node of the phylogeny shown in boxes. Transitions between character states are shown beneath each branch. The polarity of a transition is dependent on the condition of the character at the preceding node; therefore, a transition to 0 from −1 would be positive (a peramorphic transition), while a transition to 0 from + 1 would be negative (a paedomorphic transition). Node-based calculations are shown on the bottom left, where heterochronic weights are derived from the transitions leading to each node or tip species, while tip-based calculations of heterochronic weights derived from the terminal character conditions of tip species are shown on the bottom right. Both analytical variations accurately capture the overall peramorphic trend among species A and B and the paedomorphic trend from species E to G. Notably, the tip-based application of the method fails to recognize the peramorphic reversal in species F; however, tip-based heterochronic weights would recognize the peramorphic influence if this were to develop into a long-term trend. Node- and tip-based calculations of heterochronic weights are therefore both equally accurate with regard to recognizing overall trends, but node-based calculations are more precise.

Alternatively, the tip-based application of heterochronic weighting provides a grand average of heterochronic traits within observed taxa compared with the root character polarity. Rather than tracing the transition of the heterochronic event, tip-based heterochronic weighting quantifies the overall outcome of heterochronic events. This makes the method less precise than its node-based articulation, as it may fail to recognize relative polarity changes in characteristics, but is more broadly applicable to groups with uneven sampling or uncertain phylogenetic topology. Tip-based heterochronic weighting gives an overall indication of peramorphic or paedomorphic trends over the evolutionary history of a clade; in this way, it is similar to the method recently used by Martynov et al. (Reference Martynov, Lundin, Picton, Fletcher, Malmberg and Korshunova2020) to study paedomorphosis in nudibranchs, with the distinction that heterochronic weighting provides a quantitative rather than qualitative assessment of the degree of peramorphic or paedomorphic traits within a species. For the present study, heterochronic weighting was applied using the tip-based procedure, as sampling of Xiphosura is uneven across their evolutionary history, as demonstrated by the lack of Silurian exemplars. This is considered an appropriate field test of the method, as I suspect that tip-based analyses will form the bulk of any subsequent studies that adopt heterochronic weighting.
Heterochronic Character Matrix
A heterochronic character matrix for Xiphosura was assembled comprising 20 characters. Characters and their peramorphic and paedomorphic conditions were developed following the criteria detailed in “Quantifying Heterochrony”. The characters encompass a wide variety of xiphosuran morphology, equally divided between traits of the prosoma (anterior tagma) and traits of the thoracetron (posterior tagma) (Figs. 2, 3, Table 1). The resulting matrix, coded for 54 xiphosurid taxa, is available in the Supplementary Material. While a number of aspects of these characters also appear in the phylogenetic character matrix, several (such as body size) are not considered appropriate phylogenetic characters and are not incorporated into the phylogenetic analysis. Those heterochronic characters that are included comprise only a minority (5%) of the total phylogenetic characters, ensuring a degree of independence between the two character sets.
Heterochronic characters coded for Xiphosura encompassing aspects of overall body size and prosomal morphology, showing paedomorphic (−1), neutral (0), and peramorphic (+1) conditions. A character unavailable for coding in a species is considered missing data (?) and does not contribute to the species score.

Heterochronic characters coded for Xiphosura encompassing aspects of thoracetron and telson morphology, showing paedomorphic (−1), neutral (0), and peramorphic (+1) conditions. A character unavailable for coding in a species is considered missing data (?) and does not contribute to the species score.

Determination of polarity of character morphs for assigning peramorphic and paedomorphic conditions was performed with reference to the observed ontogenetic trajectories of extinct and extant species. The most extensive work on xiphosurid development and ontogeny has focused on the extant American species Limulus polyphemus, with studies of embryonic (Scholl Reference Scholl1977; Sekiguchi et al. Reference Sekiguchi, Yamamichi, Costlow, Bonaventura, Bonaventura and Tesh1982; Sekiguchi Reference Sekiguchi and Sekiguchi1988; Shuster and Sekiguchi Reference Shuster, Sekiguchi, Shuster, Barlow and Brockmann2003; Haug and Rötzer Reference Haug and Rötzer2018a) and postembryonic (Sekiguchi et al. Reference Sekiguchi, Seshimo, Sugita and Sekiguchi1988a,Reference Sekiguchi, Seshimo and Sugitab; Shuster and Sekiguchi Reference Shuster, Sekiguchi, Shuster, Barlow and Brockmann2003) ontogeny supplemented by detailed studies of the early development of the thoracic opercula and book gills (Farley Reference Farley2010) and compound lateral eyes (Harzsch et al. Reference Harzsch, Vilpoux, Blackburn, Platchetzki, Brown, Melzer, Kempler and Battelle2006). Comparatively few studies of the extant Asian species’ ontogenies exist; those few that do focus on Tachypleus tridentatus (Sekiguchi et al. Reference Sekiguchi, Yamamichi, Costlow, Bonaventura, Bonaventura and Tesh1982, Reference Sekiguchi, Seshimo, Sugita and Sekiguchi1988a,Reference Sekiguchi, Seshimo and Sugitab; Sekiguchi Reference Sekiguchi and Sekiguchi1988), often via direct comparison with the development of Limulus.
Ontogenetic data also exist for a number of extinct taxa from across the xiphosurid phylogeny. The bellinurines Euproops danae and Euproops sp., from the Carboniferous of the United States and Germany, respectively, have been subject to detailed study of their postlarval development (Schultka Reference Schultka2000; Haug et al. Reference Haug, Van Roy, Leipner, Funch, Rudkin, Schöllmann and Haug2012; Haug and Rötzer Reference Haug and Rötzer2018b; Tashman et al. Reference Tashman, Feldmann and Schweitzer2019). Ontogenetic data have also been reported from the bellinurine Alanops magnificus, known from the Carboniferous Montceau-les-Mines Konservat-Lagerstätte in France, a species considered to demonstrate a paedomorphically derived adult morphology (Racheboeuf et al. Reference Racheboeuf, Vannier and Anderson2002). Bellinurine embryonic larvae preserved within egg clutches have also been reported from the Carboniferous of Russia and assigned to the newly erected species Xiphosuroides khakassicus (Shpinev and Vasilenko Reference Shpinev and Vasilenko2018), which probably represent the larvae of an existing species of Bellinurus. Outside of the bellinurines, juvenile and adult individuals of the paleolimulid Paleolimulus kunguricus, from the Permian of Russia have been described (Naugolnykh Reference Naugolnykh2017). Subadult and adult stages have also been differentiated among the available material of the Cretaceous tachypleine Tachypleus syriacus from the Hâqel and Hjoûla Konservat-Lagerstätten of Lebanon (Lamsdell and McKenzie Reference Lamsdell and McKenzie2015), while the Cretaceous mesolimulid Mesolimulus tafraoutensis from the Gara Sbaa Konservat-Lagerstätte of Morocco is known from juvenile material (Lamsdell et al. Reference Lamsdell, Tashman, Pasini and Garassino2020).
This diversity in phylogenetic sampling of xiphosurid ontogeny reveals a number of consistent trends in horseshoe crab development across their evolutionary history that can still be observed in the ontogeny of modern species (Fig. 4). These evolutionarily conserved ontogenetic trends include a progressive reduction in expression of the prosomal ophthalmic ridge from the larval form through to the adult, a relative decrease in the length of the prosomal appendages, a reduction in expression of the free lobe segment, an increase in the number and complexity of opisthosomal opercula, and an increase in the relative length of the telson compared with the body. It is these traits, alongside other observed consistent trends in xiphosuran ontogeny, that were incorporated into the heterochronic character matrix.
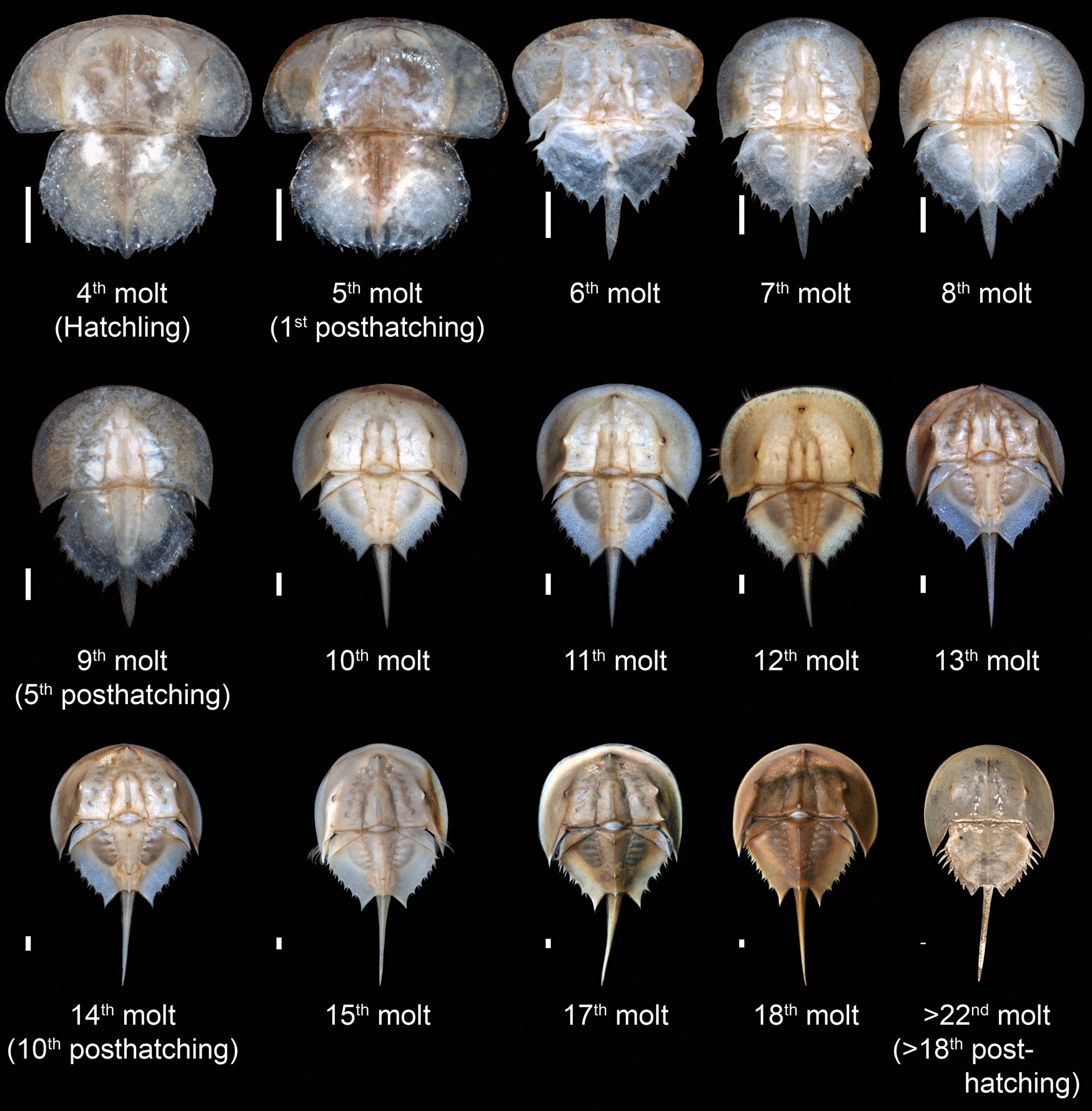
Ontogenetic sequence of Limulus polyphemus from the Yale Peabody Museum teaching collection, beginning with the hatchling (fourth molt) and proceeding to the adult (post–22nd molt, which corresponds to the 18th posthatching molt). The final molt is represented by specimen YPM IZ 070174. The size of each instar has been standardized to more clearly demonstrate changes in relative morphological proportions. Scale bars, 1 mm.
Phylogenetic Analysis
A phylogenetic framework for Xiphosura was constructed using an expanded version of the chelicerate character matrix from Lamsdell (Reference Lamsdell2016), which itself draws on matrices presented in Lamsdell (Reference Korn, Hopkins and Walton2013), Selden et al. (Reference Selden, Lamsdell and Liu2015), Lamsdell and McKenzie (Reference Lamsdell and McKenzie2015), and Lamsdell et al. (Reference Lamsdell, Briggs, Liu, Witzke and McKay2015). The resulting matrix comprises 256 characters coded for 153 taxa and is available in the online MorphoBank database (O'Leary and Kaufman Reference O'Leary and Kaufman2012) under the project code p2606 (accessible from http://morphobank.org/permalink/?P2606) as well as in the Supplementary Material. Sampling of Xiphosurida within the matrix was increased by six, incorporating Limulus woodwardi (Watson Reference Watson1909) and the newly described Limulitella tejraensis (Błażejowski et al. Reference Błażejowski, Niedźwiedzki, Boukhalfa and Soussi2017), M. tafraoutensis (Lamsdell et al. Reference Lamsdell, Tashman, Pasini and Garassino2020), P. kunguricus (Naugolnykh Reference Naugolnykh2017), Paleolimulus woodae (Lerner et al. Reference Lerner, Lucas and Mansky2016), and Vaderlimulus tricki (Lerner et al. Reference Lerner, Lucas and Lockley2017). Three taxa were removed from the matrix: Anacontium brevis, which is a synonym of the co-occurring Anacontium carpenteri (Anderson Reference Anderson1997); Liomesaspis leonardensis, which is poorly preserved and lacks diagnostic characters separating it from Liomesaspis laevis (see Tasch Reference Tasch1961); and Willwerathia laticeps, which may not be a chelicerate (Lamsdell Reference Lamsdell2019). In total, 54 Xiphosura (sensu Lamsdell Reference Lamsdell2013, Reference Lamsdell2016) were included, and it is these taxa that were the focus of subsequent analyses of heterochrony and ecological affinity, with the remaining 99 taxa within the matrix serving to adequately root and demonstrate the monophyly of the xiphosuran clade. Additionally, the character coding for Yunnanolimulus luopingensis was updated to incorporate information from recently described specimens exhibiting exceptional preservation (Hu et al. Reference Hu, Zhang, Feldmann, Benton, Schweitzer, Wen, Zhou, Xie, Lü and Hong2017). Four new characters were also added to the matrix, encompassing cardiac lobe width (character 32), the cardiac lobe bearing a median cardiac ridge with rounded cross section (character 38), the occurrence of a genal groove (character 45), and the course and extent of the genal groove (character 46). Morphological terminology for character definitions follows Selden and Siveter (Reference Selden and Siveter1987) and Lamsdell (Reference Lamsdell2013).
Tree inference was performed using Bayesian statistical analysis, which has been shown to outperform maximum parsimony analyses of simulated data (Wright and Hillis Reference Wright and Hillis2014; O'Reilly et al. Reference O'Reilly, Puttick, Parry, Tanner, Tarver, Fleming, Pisani and Donoghue2016; Puttick et al. Reference Puttick, O'Reilly, Tanner, Fleming, Clark, Holloway, Lozano-Fernandez, Parry, Tarver, Pisani and Donoghue2017, Reference Puttick, O'Reilly, Pisani and Donoghue2019), although tree topologies derived from empirical data analyzed under parsimony methods exhibit higher stratigraphic congruence than those retrieved from Bayesian analysis (Sansom et al. Reference Sansom, Choate, Keating and Randle2018). Bayesian inference was performed using Markov chain Monte Carlo analyses as implemented in MrBayes 3.2.6 (Huelsenbeck and Ronquist Reference Huelsenbeck and Ronquist2001), with four independent runs of 100,000,000 generations and four chains each under the maximum-likelihood model for discrete morphological character data (Mkv + Γ; Lewis Reference Lewis2001), with gamma-distributed rate variation among sites. All characters were treated as unordered and equally weighted (Congreve and Lamsdell Reference Congreve and Lamsdell2016). Trees were sampled with a frequency of every 100 generations, resulting in 1,000,000 trees per run. The first 25,000,000 generations (250,000 sampled trees) of each run were discarded as burn-in, and the 50% majority rule consensus tree was calculated from the remaining 750,000 sampled trees across all four runs; this represents the optimal summary of phylogenetic relationships given the available data (Holder et al. Reference Holder, Sukumaran and Lewis2008). Posterior probabilities were calculated from the frequency at which a clade occurred among the sampled trees included in the consensus tree.
Additionally, maximum parsimony analysis was performed using TNT (Goloboff et al. Reference Goloboff, Farris and Nixon2008) (made available with the sponsorship of the Willi Hennig Society) to test for topological congruence between Bayesian and parsimony methods. The search strategy employed 100,000 random addition sequences with all characters unordered and of equal weight (Congreve and Lamsdell Reference Congreve and Lamsdell2016), each followed by tree bisection-reconnection branch swapping (the mult command in TNT). Jackknife (Farris et al. Reference Farris, Albert, Källersjö, Lipscomb and Kluge1996), bootstrap (Felsenstein Reference Felsenstein1985), and Bremer (Bremer Reference Bremer1994) support values were also calculated in TNT and the ensemble Consistency, Retention and Rescaled Consistency Indices were calculated in Mesquite 3.02 (Maddison and Maddison Reference Maddison and Maddison2018). Bootstrapping was performed with 50% resampling for 1,000 repetitions, while jackknifing was performed using simple addition sequence and tree bisection-reconnection branch swapping for 1,000 repetitions with 33% character deletion.
Ecological Affinity
Previous work by Kiessling and Aberhan (Reference Kiessling and Aberhan2007) and Hopkins (Reference Hopkins2014) has set out a series of standard environmental categories for organisms found in marine environments; latitudinal occupation, substrate preference, bathymetry, and reef association. For the present study, however, a novel characteristic is considered: salinity. Previous work has shown that horseshoe crabs invaded nonmarine environments multiple times over their evolutionary history (Lamsdell Reference Lamsdell2016), a transition associated with a variety of biomechanical and physiological challenges (Lamsdell Reference Lamsdellin press), and it is these macroevolutionary events that comprise the focus of the work herein. As with the characteristics of Kiessling and Aberhan (Reference Kiessling and Aberhan2007) and Hopkins (Reference Hopkins2014), salinity is reduced to a binary variable, and species are assigned either a marine or nonmarine (including freshwater and deltaic/transitional marginal-marine environments with a large amount of continental flora and fauna) affinity. The ecological affinity for each taxon was estimated through the modified method of Miller and Connolly (Reference Miller and Connolly2001). This is a departure from Hopkins (Reference Hopkins2014), who implemented a refined version of the Bayesian estimation of affinity developed by Simpson and Harnik (Reference Simpson and Harnik2009), which has traditionally been applied to higher taxa (e.g., genera). The drawback of applying the Bayesian estimation method to species data is that it requires multiple samples; Simpson and Harnik (Reference Simpson and Harnik2009) only considered genera with at least four occurrences. This is impossible for many species in the fossil record, particularly unmineralized marine organisms (which include horseshoe crabs), the majority of which are known solely from single localities—representing only a single occurrence. In contrast, the method of Miller and Connolly (Reference Miller and Connolly2001) simply calculates the affinity of a taxon for a given environment as the number of occurrences within the environment divided by the total number of occurrences, resulting in a metric ranging from 0 to 1; when combined, the value of the metric for each of the two possible affinities and no clear affinity will always sum to 1. The advantage of this method is that it can operate with only a single observation, with the drawback that lower sample sizes increase the chances of a false positive of ecological affinity. However, species tend to have a narrow range of habitat preferences (Saupe et al. Reference Saupe, Hendricks, Portell, Dowsett, Haywood, Hunter and Lieberman2014, Reference Saupe, Qiao, Hendricks, Portell, Hunter, Soberón and Lieberman2015), thereby reducing the chance of occurrences outside their preferred environments; when species do have broader habitat preferences, they also tend to have broader geographic ranges, increasing the probability of fossilization and thereby increasing the available sample size, reducing the chance of false positives.
The resulting metrics for ecological affinity were interpreted as any single value >0.5 indicating a preference for that environment. When no single environment scored >0.5, the taxon was considered to exhibit no affinity. Ties (an instance where the highest-scoring environment achieved a 0.5) were also considered an indication of no affinity. In practice, as the majority of horseshoe crab species are known from only one or a handful of localities, most species showed a clear ecological affinity (see Supplementary Material).
Phylogenetic Paleoecology
The emergent field of phylogenetic paleoecology leverages phylogenetic information to assess the long-term evolutionary significance of ecological trends within lineages through the synthesis of phylogenetic theory and quantitative paleoecology (Lamsdell et al. Reference Lamsdell, Congreve, Hopkins, Krug and Patzkowsky2017). Using the historical phylogenetic and ecological framework, it is possible to test for phylogenetic or ecological signal in the distribution of heterochronic weightings, thereby accounting for the possible influence of both the genealogical and ecological biological hierarchies (Congreve et al. Reference Congreve, Falk and Lamsdell2018).
The phylogenetic framework also permits for the estimation of the ancestral conditions of both morphological and ecological characteristics. To determine the number of transitions between marine and nonmarine environments, the ecological affinity of each species as determined above was mapped onto the resulting phylogenetic tree and parsimony-based ancestral state reconstruction performed in Mesquite 3.02 (Maddison and Maddison Reference Maddison and Maddison2018). The inferred ancestral ecological affinities were used to polarize the internal nodes of the tree, thereby establishing whether the ecological affinities of tip species represent inherited habitat preferences or are the result of a shift in the occupied niche.
Multivariate statistical tests (permutational multivariate analysis of variance [PERMANOVA] using the Euclidian distance measure) were performed in the statistical software package PAST (Hammer et al. Reference Hammer, Harper and Ryan2001) and using the adonis function in the R (R Core Team 2018) package vegan (Oksanen et al. Reference Oksanen, Blanchet, Friendly, Kindt, Legendre, McGlinn, Michin, O'Hara, Simpson, Solymos, Stevens, Szoecs and Wagner2019) to ascertain the statistical significance of differences in heterochronic weightings across ecological affinities and phylogenetic clades. Several analyses were performed: comparison between marine and nonmarine habitats, comparison between clades, and a test of the interaction between clade and habitat categories. For the comparison between marine and nonmarine habitats, all species were included in the analysis. However, for analyses that required the assignment of species to clades, only species assignable to Bellinurina, Paleolimulidae, Austrolimulidae, or Limulidae were included. This resulted in the exclusion of Lunataspis aurora and Kasibelinurus amoricum (stem Xiphosurida), Bellinuroopsis rossicus and Rolfeia fouldenensis (stem Limulina), and Valloisella lievinensis (stem Limulidae), as their inclusion would necessitate the formulation of unnatural, paraphyletic “groups” that possess no evolutionary cohesion and could serve to bias analyses. Significance was estimated by permutation across groups with 10,000 replicates, including Bonferroni correction to correct for multiple comparisons in the case of comparison between clades.
To test whether the distributions of species’ heterochronic weightings within clades exhibit directional trends, Spearman's (Reference Spearman1904) rank correlation coefficient ρ, calculated using the cor.test function in R (R Core Team 2018), was used to compare the heterochronic weighting of observed taxa within a clade to their distance from the root of the clade. A statistically significant ρ, in combination with a statistically significant clade heterochronic weighting [Hw], was considered evidence of a concerted heterochronic trend within the clade. The distance of a species from the root of the clade was determined using the cladistic rank method (Gauthier et al. Reference Gauthier, Kluge and Rowe1988; Norell and Novacek Reference Norell and Novacek1992a,Reference Norell and Novacekb; Norell Reference Norell1993; Benton and Storrs Reference Benton and Storrs1994; Benton and Hitchin Reference Benton and Hitchin1997; Carrano Reference Carrano2000, Reference Carrano, Carrano, Gaudin, Blob and Wible2006; Wagner and Sidor Reference Wagner and Sidor2000; Poulin Reference Poulin2005; Prado and Alberdi Reference Prado and Alberdi2008; Huttenlocker Reference Huttenlocker2014). Cladistic rank is determined by counting the sequence of primary nodes in a cladogram from the basal node to the ultimate node (Fig. 5). The main axis of the cladogram is defined as the internal branch supporting the most inclusive clade after each bifurcation. Smaller offshoot clades are reduced to polytomies for the purpose of assigning a clade rank, with each constituent species within a polytomy receiving the same rank. The assignment of equal rank to all constituents of an offshoot clade is a conservative approach that ensures any revealed trend is the majority pattern demonstrated by the most inclusive clades within the cladogram, without biasing the analysis through exclusion of taxa located in offshoot clades. This was done for each of the four main xiphosurid clades: Bellinurina, Paleolimulidae, Austrolimulidae, and Limulidae.
Example of the method for assigning taxa to clade ranks for Spearman's rank correlation.

Results
Analysis of the phylogenetic character matrix resulted in Bayesian inference and parsimony optimality criteria retrieving a concordant tree topology, with the strict parsimony consensus of six most parsimonious trees having an identical hypothesis of xiphosuran relationships to the Bayesian majority rule consensus (Fig. 6; for details of the parsimony tree and the full Bayesian tree, see Supplementary Material). The phylogenetic topology, while broadly consistent with those of previous analyses (Lamsdell Reference Lamsdell2013, Reference Lamsdell2016; Lamsdell and McKenzie Reference Lamsdell and McKenzie2015), is more stratigraphically congruent due to movement in the placement of K. amoricum, ‘Kasibelinurus’ randalli, and Casterolimulus kletti. Previously resolving as a paraphyletic grade at the base of Xiphosura, K. amoricum now resolves as the sister taxon to the clade comprising the two suborders Limulina and Bellinurina, while ‘Kasibelinurus’ randalli resolves at the base of Bellinurina. This leaves L. aurora, the sole described Ordovician species, as the sister taxon to all other Xiphosura. Casterolimulus, previously resolved as the sole post-Triassic representative of Austrolimulidae, now forms a clade with Victalimulus within Limulidae. Victalimulus and Casterolimulus are both Cretaceous taxa known from nonmarine environments, and the notion that they share common ancestry has been suggested previously (Lamsdell in press). The internal relationships of the bellinurines and limulids are also better resolved, and a sister clade to the xiphosuran crown group (composed of Tachypleinae and Limulinae) comprising Mesolimulus, ‘Limulitella’ vicensis, ‘Limulus’ woodwardi, Victalimulus, and Casterolimulus is recognized for the first time.
Bayesian phylogeny of xiphosurids showing environmental affinity of salinity and heterochronic weighting mapped onto the tree. Environmental affinity is indicated on the branches (blue, marine; brown, nonmarine), heterochronic weighting is shown at the tips alongside the taxon names through heat-map shading (green, more paedomorphic; orange, more peramorphic). Bayesian posterior probabilities are shown below each node. The clades shown in Figs. 7 and 8 are labeled alongside the tree.

The distribution of environmental affinity across the phylogeny of Xiphosura supports previous assertions that the group has invaded nonmarine environments from the marine realm multiple times over the course of its evolutionary history (Lamsdell Reference Lamsdell2016). However, whereas previous estimates suggested that the transition from a marine to nonmarine environmental affinity had occurred at least five times during xiphosuran evolution, the present results indicate such a transition has occurred only four, or potentially as few as three, times (Fig. 6). Ancestral state reconstruction demonstrates that a marine affinity is plesiomorphic for Xiphosura and that shifts to a nonmarine affinity occurred for the clades Bellinurina and Austrolimulidae, as well as Valloisella and the sister taxa Victalimulus and Casterolimulus. While Bellinurina and the Victalimulus/Casterolimulus clade are clearly derived from marine-dwelling lineages that subsequently transitioned to nonmarine environments, the situation for Valloisella and the austrolimulids is less clear. Due to the tree topology, with Valloisella—a nonmarine xiphosurid—resolving as sister taxon to a large clade divided between the predominantly marine Limulidae and the nonmarine Austrolimulidae, ancestral state reconstruction is unable to determine a clear environmental affinity for the internal nodes between Limulidae, Austrolimulidae, and Valloisella. These results infer a number of potential evolutionary scenarios. One possibility is that Valloisella and Austrolimulidae independently transitioned to nonmarine environments and a marine affinity is plesiomorphic for Limulidae. Alternatively, Valloisella, Austrolimulidae, and Limulidae may all be derived from fully euryhaline ancestors that genuinely have no environmental affinity and occupied both marine and nonmarine environments; in this case, the transition to a nonmarine affinity in Valloisella and Austrolimulidae, and a marine affinity in Limulidae, would be the result of a reduction in their ancestral occupied niche. Finally, it is possible that the transition to nonmarine environments occurred in the ancestors of Valloisella, Austrolimulidae, and Limulidae and that a nonmarine affinity is therefore plesiomorphic for all three taxa, with Limulidae having undergone a subsequent reversal to reoccupy marine environments.
Heterochronic weightings are unevenly distributed across the phylogeny, with more extreme (i.e., higher positive or negative) values occurring among taxa with a nonmarine environmental affinity. This distinction is borne out statistically, with PERMANOVA tests showing that the variance of heterochronic weightings of all species occupying marine environments is significantly distinct from that of those occupying nonmarine environments (Table 2). Statistically significant differences also separate the heterochronic weightings of the xiphosuran clades Bellinurina, Paleolimulidae, Austrolimulidae, and Limulidae (Table 3). Environmental affinity and phylogenetic relatedness therefore both influence heterochronic weightings; however, there appears to be no interaction between the two factors, indicating that they exhibit conflicting signals and do not covary (Table 4).
One-way permutational multivariate analysis of variance (F (1,53) = 4.197, η2 = 0.075, p = 0.0424), 10,000 permutations, Euclidean distance measure. Value in regular font is the p-value, value in italics is the raw F-value. Total sum of squares = 5.120, within-group sum of squares = 4.737, between-group sum of squares = 0.383.

One-way permutational multivariate analysis of variance (F (3,49) = 73.87, η2 = 0.83, p = 0.0001) excluding stem taxa, 10,000 permutations, Euclidean distance measure. Values in regular font are Bonferroni corrected p-values, those in italics are raw F-values. Total sum of squares = 5.088, within-group sum of squares = 0.8588, between-group sum of squares = 4.2292.

Two-way permutational multivariate analysis of variance excluding stem taxa, 10,000 permutations, Euclidean distance measure.

Comparing the heterochronic weightings of Bellinurina, Paleolimulidae, Limulidae, and Austrolimulidae to the distribution of randomized heterochronic weightings reveals the heterochronic weightings of Bellinurina, Austrolimulidae, and Limulidae to be significantly different from what would be expected from random, while that of Paleolimulidae falls within what could be explained from a random distribution (Fig. 7). Interestingly, Spearman's rank correlation (Fig. 8), indicates directional trends in heterochronic weighting within Austrolimulidae and Bellinurina, but no significant trend within Limulidae or Paleolimulidae. The heterochronic weightings of Bellinurina and Austrolimulidae show a consistent directional trend as demonstrated by locally estimated scatterplot smoothing (LOESS) regression (although Bellinurina do undergo a slight shift in trajectory among their highest-ranked clades), while Paleolimulidae and Limulidae exhibit random directional shifts across the phylogeny. This distinction is potentially borne out in the position of the observed heterochronic weightings relative to the randomized distribution of heterochronic weightings: the observed weightings for Bellinurina and Austrolimulidae sit far outside the randomized distribution, expressing a value that does not occur within the randomized weights. In comparison, the observed heterochronic weighting for Limulidae falls within the tail of the randomized distribution, with a value equal to that of a set of those retrieved from the randomizations.
Histograms showing the distribution of randomized heterochronic weightings across 100,000 permutations for Bellinurina, Paleolimulidae, Limulidae, and Austrolimulidae. The actual heterochronic weightings of the clades, derived using characters shown in Figs. 2 and 3, are indicated by the black arrows. Weightings in either tail of the distribution are considered to be more extreme than would be expected from random. The negative tail indicates the occurrence of paedomorphosis, the positive tail indicates peramorphosis.

Graphs showing distribution of heterochronic weightings along clade rank for Bellinurina, Paleolimulidae, Limulidae, and Austrolimulidae. Each plot displays a solid linear regression line and a dashed LOESS regression line. A negative slope indicates a general paedomorphic trend, while a positive slope is representative of a peramorphic trend. The results of Spearman's rank correlation, both in terms of statistical significance and raw ρ, are shown at the top of each graph.

Discussion
Shifts in ecological affinity correlate with changes in evolutionary regime in Xiphosura. Clades that invade nonmarine environments exhibit distinct differences in the prevalence of heterochronic traits in comparison to those that inhabit the marine realm (Fig. 6), with Austrolimulidae demonstrating increased prevalence of peramorphy, while paedomorphy is prevalent among Bellinurina (Fig. 7). Paedomorphic traits have long been recognized in bellinurines (Haug et al. Reference Haug, Van Roy, Leipner, Funch, Rudkin, Schöllmann and Haug2012; Lamsdell in press), including their retention of long, gracile prosomal appendages into adulthood; visible opisthosomal segmentation; and elongated dorsal prosomal shield spines. Austrolimulids, meanwhile, develop elongate and splayed prosomal genal spines; reduce the size of their opisthosomal tergopleura; and exhibit enlarged, posteriorly positioned lateral eyes—all of which are recognized as peramorphic conditions herein. Interestingly, lineages that make the transition to nonmarine environments demonstrate concerted and enduring heterochronic trends (Fig. 8) that persist for millions of years, with species progressively exhibiting an increasingly greater number of paedomorphic (in Bellinurina) or peramorphic (in Austrolimulidae) traits. It seems that shifts in environmental occupation set these lineages along a heterochronic trajectory resulting in a directional bias (Gould Reference Gould and Milkman1982), whereby changes in the timing or rate of development produce innovative morphologies, the selection of which—mediated by the environment—result in increasingly specialized phenotypes. Heterochronic processes are known to be one of the primary ways by which morphological innovation (sensu Erwin Reference Erwin2015a, Reference Erwin2017) occurs (McNamara and McKinney Reference McNamara and McKinney2005; McNamara Reference McNamara2012; Colangelo et al. Reference Colangelo, Ventura, Piras, Bonaiuti and Ardizzone2019), and it is the nonmarine xiphosurans that contribute the most to xiphosuran disparity and occupy novel regions of morphospace (Lamsdell Reference Lamsdell2016).
It is notable that only rarely do either of the heterochronic trends observed here show any indication of heterochronic reversals and that in both cases the trend is associated with a marked shift in diversity dynamics, with Bellinurina increasing in diversity before rapidly going extinct and Austrolimulidae exhibiting decreased rates of speciation and a lower diversity even in relation to other Xiphosura (Lamsdell Reference Lamsdell2016, in press). The combination of limited reversal and shift in diversity dynamics suggests that these directional trends may represent macroevolutionary ratchets (trends where reversals are rare and ultimately result in increased extinction risk or a decrease in rates of origination: Van Valkenburgh Reference Van Valkenburgh1991, Reference Van Valkenburgh1999; Van Valkenburgh et al. Reference Van Valkenburgh, Wang and Damuth2004), although whether this is due to consistent environmental pressure or some inherent property of the developmental processes operating is unclear. Numerous consistent peramorphic and paedomorphic evolutionary trends associated with environmental gradients—termed peramorphoclines and paedomorphoclines (McNamara Reference McNamara1982), or more generally heteroclines (McKinney Reference McKinney1999), although these terms conflate process (heterochrony) and evolutionary outcome (directional bias or macroevolutionary ratcheting)—have been recognized in the fossil record (McNamara Reference McNamara1982, Reference McNamara1986, Reference McNamara1988; Simms Reference Simms1988; Korn Reference Korn1995; Crônier et al. Reference Crônier, Renaud, Feist and Auffray1998; McKinney Reference McKinney1999; Poty Reference Poty2010; Fernandez-Lopez and Pavia Reference Fernandez-Lopez and Pavia2015). Most described heterochronic trends do not exhibit heterochronic reversals, with only a few notable exceptions (Gerber Reference Gerber2011), although it should be noted that none of these studies were performed within a phylogenetic framework. It has previously been suggested that concerted heterochronic trends are always controlled by environmental factors (McKinney Reference McKinney1986, Reference McKinney and McKinney1988); however, subsequent studies have documented heterochronic trends occurring apparently independently of any environmental gradient (Breton Reference Breton1997). In the present study, xiphosurans possibly undergo one reversal in environmental affinity, with the ancestors of the predominantly marine Limulidae potentially having a nonmarine or mixed environmental affinity (Fig. 6). The ramifications of this are twofold. First, it would suggest that xiphosurans survived the end-Permian mass extinction by occupying nonmarine environments and returned to the marine realm during the subsequent recovery, a possibility first suggested by Błażejowski et al. (Reference Błażejowski, Niedźwiedzki, Boukhalfa and Soussi2017). Second, Limulidae display on average more positive heterochronic weightings than other marine taxa (Fig. 7), although these peramorphic traits do not manifest as part of a concerted trend (Fig. 8) as they do in the limulid nonmarine sister group, Austrolimulidae, which is characterized by extreme peramorphism. This opens up the possibility that, if limulids have reoccupied marine environments from an ancestral nonmarine habitat, the elevated occurrence of peramorphic character conditions in the clade may be a relict of the peramorphic trajectory that continued in austrolimulids. If this were to be the case, it would suggest that returning to marine environments stopped the heterochronic bias in limulids, and perhaps most interestingly, that the changes the lineage had undergone while occupying nonmarine environments were not subsequently reversed.
The exact mechanism by which heterochrony operated in the cases observed here is uncertain. It is notoriously difficult (arguably impossible) to discern between changes in timing and changes in rate of development without detailed ontogenetic sequences of consecutive species (Gould Reference Gould and McKinney1988; Jones Reference Jones and McKinney1988; McKinney Reference McKinney and McKinney1988; Allmon Reference Allmon1994; McNamara and McKinney Reference McNamara and McKinney2005; Bardin et al. Reference Bardin, Rouget and Cecca2017). Nevertheless, it is possible to recognize broad peramorphic and paedomorphic trends. Why these processes unfolded in a ratchet-like fashion is also unclear. Classical natural selection (Darwin Reference Darwin1859) may explain the evolution of increasingly unusual morphologies; however, it is unclear whether these morphologies are truly specialized or simply bizarre. Another possibility among the Bellinurina is that physiological changes required in order for xiphosurans to tolerate low-salinity environments for extended periods of time may have been accomplished through the retention of larval physiology into adulthood (Lamsdell in press). The larvae of modern horseshoe crabs are extremely tolerant of salinities lower than 35‰ (Shuster Reference Shuster, Bonaventura, Bonaventura and Tesh1982; Ehlinger and Tankersley Reference Ehlinger and Tankersley2007; Botton et al. Reference Botton, Tankersley and Loveland2010), and paedomorphic processes permitting the maintenance of this tolerance in adults may have also resulted in the retention of larval morphological characteristics. The broad, shallow prosomal carapaces of austrolimulids, meanwhile, may have developed in response to unidirectional hydrodynamic environments that the group encountered as it radiated in lacustrine environments. It is worth considering that macroevolutionary ratchets in even closely related groups may occur through distinct mechanisms.
Ultimately, one of the more interesting outcomes of the study is the support for the quasi-independence of the signal imparted by history and ecology in evolution. History (as represented by phylogeny) and ecology are both significant sources of variation among heterochronic weights but do not interact (Table 4), indicating that although they both exert influence on the distribution of heterochronic weights, they do so with conflicting signals. This conflict is due to the impact of the quasi-independent genealogical and ecological biological hierarchies (Congreve et al. Reference Congreve, Falk and Lamsdell2018), whereby historical contingency limits the morphological framework for subsequent adaptation to ecological pressures (Gould and Lewontin Reference Gould and Lewontin1979; Eldredge and Salthe Reference Eldredge and Salthe1984; Anderson and Allmon Reference Anderson and Allmon2018). Logically, and as hinted at by the data discussed here, this tension between competing hierarchies can also extend to developmental frameworks as the mediating factors by which morphologic phenotypes are expressed.
Conclusions
Applying this new method for quantifying heterochrony, expressed as a heterochronic weighting, within a phylogenetic context reveals concerted independent heterochronic trends in xiphosurans. These trends correlate with, and may be driven by, shifts in environmental occupation from marine to nonmarine habitats, resulting in a macroevolutionary ratchet whereby environmental selective factors result in the preferential retention of phenotypes derived from heterochronic processes, which in turn reinforces directional heterochronic trends and the proliferation of peramorphic or paedomorphic characteristics. Critically, the distribution of heterochronic weightings among species shows evidence of being influenced by both historical, phylogenetic processes and external ecological pressures. This is most clearly demonstrated by the manner in which the independent occupation of nonmarine environments is accompanied by significant heterochronic trends in both Bellinurina and Austrolimulidae, but manifesting as a paedomorphic trend among bellinurines and a peramorphic trend within austrolimulids. Therefore, while the environment can exert a strong pressure on both phenotype (as expected by the fundamental evolutionary process of natural selection) and the underlying developmental processes that govern phenotype, it can only do so utilizing the available morphological and developmental frameworks. The availability and composition of these frameworks is mediated by contingent, historical factors (as expressed by phylogeny) that limit both the potential for adaptation to certain environmental conditions and the structural or developmental outcome of concerted selective pressure. A well-known example of the former is the fact that vertebrates returning to fully aquatic environments are constrained to breathing air, while a structural example of the latter are the hook-like projections of the beak's tomia in mergansers that are used to aid in gripping their fish prey. The serrations perform a function similar to that of the narrow, curved teeth of gars, which also prey upon small fish. Teeth, however, were lost in the avian stem-lineage; lacking the developmental framework to express teeth, mergansers instead developed modifications to the serrated tomia prevalent in ducks and geese. Xiphosurans demonstrate that such contingent processes can also affect the mechanisms by which developmental shifts occur.
Heterochronic weighting has the proven potential to be an effective method to quantify heterochronic trends within a phylogenetic framework. Comparing the observed heterochronic weightings of clades to randomized distributions permits the discrimination of concerted heterochronic trends from what would be expected under random (nondirectional) character change. The method is readily applicable to any group of organisms that have well-defined morphological characteristics, ontogenetic information, and resolved internal relationships; indeed, in order to test the generality of the observations made for xiphosurans, it is imperative that additional studies be performed in disparate clades. Future work should also aim to apply node-based heterochronic weighting in appropriate groups and seek to apply both tip- and node-based calculations to the same data to further explore the behavior and comparability of both. The combination of this heterochronic metric with ecological affinity data affords easy study of the correlation between developmental changes and environmental shifts as a branch of phylogenetic paleoecology and has the potential to open new avenues into studying the relationship between evolutionary developmental processes and external environmental causal factors.
Acknowledgments
I thank E. Lazo-Wasem (Yale Peabody Museum) for providing photographs of Limulus instars and A. Downey (West Virginia University) for coding assistance that aided in collating randomization results. The concepts and perspectives presented in this paper have been refined over the years through in-depth discussion with my colleagues B. Anderson (West Virginia University), C. Congreve (North Carolina State University), A. Falk (Centre College), A. Manafzadeh (Brown University), and J. Miyamae (Yale University). I am especially grateful to A. Whitaker (University of Kansas) who provided graphical design services to aid in the presentation of this research at the Geological Society of America annual conference and possesses invaluable skills in helping to reassemble 3D printed turtle skulls. I thank D. Bapst (Texas A&M University) and an anonymous referee for their detailed and thoughtful reviews that greatly improved the article and prompted me to clarify aspects of the method, as well as encouraging me to devote further page space to explanations of macroevolutionary phenomena.















