Non-alcoholic fatty liver disease (NAFLD) is one of the most important liver diseases worldwide and refers to a clinical pathological syndrome(Reference Kleiner, Brunt and Van Natta1). It may become the main cause of end-stage liver disease in the coming decades, affecting adults and children(Reference Famouri, Shariat and Hashemipour2,Reference Younossi, Anstee and Marietti3) .
The pathogenesis of NAFLD is mainly due to lipid accumulation in the liver and insulin resistance caused by multiple reasons, such as obesity, type 2 diabetes and lipid metabolism disorders, resulting in hepatocyte steatosis(Reference Buzzetti, Pinzani and Tsochatzis4). In addition, oxidative stress, lipid peroxidation and proinflammatory cytokines also lead to hepatocyte infiltration and necrosis, resulting in NAFLD(Reference Henaomejia, Elinav and Jin5). Fatty liver is usually a reversible disease, and early prevention and diagnosis can normalise the liver(Reference Lee, Han and Kang6). However, several drugs, such as insulin sensitisers, particularly thiazolidinediones, show inconsistent results during treatment. In addition, many of these drugs may have safety issues due to their short duration(Reference Musso, Cassader and Rosina7,Reference Musso, Gambino and Cassader8) .
A studies has shown that bile salt hydrolase (BSH) mediates host–micro-organism balance, functionally regulates host lipid metabolism and plays an important role in cholesterol metabolism(Reference Lye, Kato and Low9). In addition, BSH promotes bile salt excretion in the body, which is the main pathway to clear cholesterol in the body(Reference Jones, Martoni and Parent10). NAFLD presents with lipid accumulation in the liver, which results from an imbalance in the acquisition and clearance of lipids. Several studies have shown that probiotics prevent and alleviate NAFLD by regulating lipid metabolism and improving liver function(Reference Fabbrini, Sullivan and Klein11,Reference Ma, Li and Yu12) . A recent study showed that oral administration of Bifidobacterium and Lactobacillus reduced cholesterol and TAG levels and achieved the goal of relieving NAFLD(Reference Famouri, Shariat and Hashemipour2). Therefore, probiotic strains with high BSH activity can hydrolyse bile acid conjugates, trigger the consumption of liver cholesterol and reduce the cholesterol content in the body.
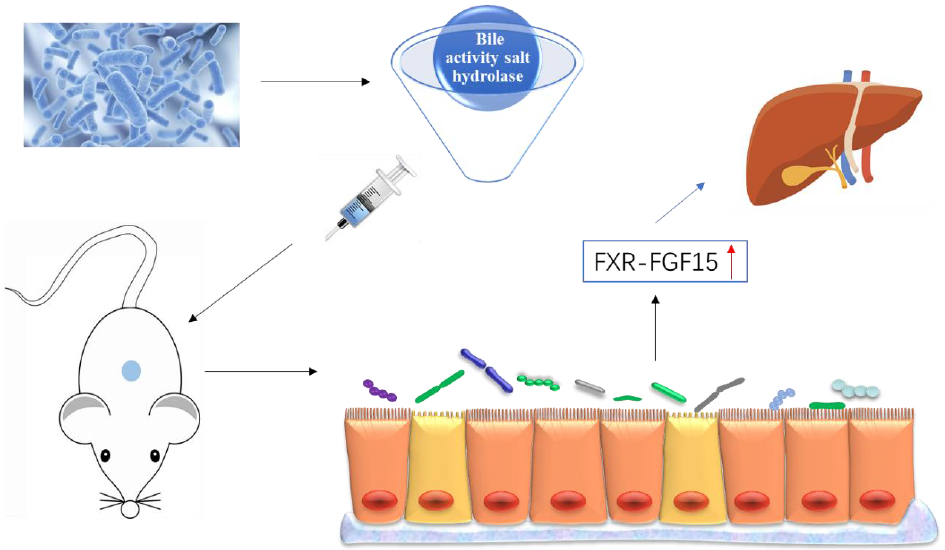
In the enterohepatic circulation of bile acids, bound bile acids are dissociated by the BSH of intestinal microorganisms, and BSH activity is one of the main characteristics of probiotics(Reference Choi, Lew and Yeo13). Regulating the intestinal microbiota changes the composition and structure of bile acids in the body, by affecting the reabsorption of bile acids and activating the farnesoid X receptor (FXR)(Reference Hartmann, Hochrath and Horvath14). FXR activation significantly increased the secretion of fibroblast growth factor 15 (FGF15) in the small intestine of mice, triggering the intestinal–liver signalling pathway, thereby regulating bile acid synthesis(Reference Inagaki, Choi and Moschetta15). A study has shown that the up-regulation of FXR–FGF15 ameliorates NAFLD by improving dyslipidaemia and liver steatosis and reducing the weight of mice(Reference Adorini, Pruzanski and Shapiro16).
In this study, we screened probiotic strains with high BSH activity in vivo and explored their relieving effect on NAFLD in mice. Then, the potential mechanism of probiotics was explored through the genes in the intestinal bile acid pathway.
Experimental methods
Experimental strains and cultivation
The thirty-four strains were stored at the Functional Dairy and Probiotics Engineering Laboratory of Ocean University of China. Before the experiment, all strains were inoculated in MRS liquid medium and cultured in a 37°C incubator for 24 h. The fermentation broth to be used was centrifuged at 8000 rpm for 5 min at 4°C. The bacteria cells were collected, washed twice with sterile PBS buffer and then resuspended in PBS. The final concentration of the strains was 108 colony-forming units/ml, and the plate counting method was used to determine the concentration of the strains(Reference Liang, Lv and Zhang17).
Bile salt hydrolase activity of probiotics in vitro
Qualitative determination of bile salt hydrolase activity
For the qualitative determination of BSH activity, 0·3 % taurine deoxycholate (TDCA), 0·2 % sodium thioglycolate, 0·37 g/l CaCl2 and 1·5 % agar were added to MRS liquid medium that was then sterilised at 121°C for 15 min and poured into plates. A sterile filter paper piece was evenly put into the plate, and 10 μl of the bacterial solution was added dropwise to the filter paper. The plate was then placed in an incubator at 37°C for 48 h. The presence of a white precipitate around the filter paper was indicative of BSH activity(Reference Allain, Chaouch and Thomas18).
Quantitative determination of bile salt hydrolase activity
According to Xu et al.(Reference Xu, Li and Zalzala19), the ninhydrin method was used to quantify the BSH activity of strains. Briefly, 10 μl of completely broken bacterial cells were mixed with 180 μl of PBS buffer (pH 6·0), 10 μl of 200 mm TDCA or glycine deoxycholate and 10 μl of liquid paraffin. Then, 200 μl of TCA (15 %) was added to stop the enzymatic hydrolysis reaction. After centrifugation at 12 000 rpm for 10 min at 4°C, 0·1 ml of the supernatant was taken, to which 1·9 ml of ninhydrin colour solution (Solarbio) was added and mixed. The solution was boiled in a boiling water bath for 14 min and then cooled to ambient temperature. The absorbance of the solution was measured at 570 nm. Glycine was used as the standard for preparing the standard curve. Protein concentration was measured using the BCA analysis kit (Beyotime Biotechnology).
Animal models and experimental groups
All experiments were performed in accordance with the British Animal (Scientific Procedures) Act 1986 (PPL 70/7652) and were approved by the Laboratory Animal Ethics Committee of College of Food Science and Engineering of Ocean University of China (approval number: SPXY2019041501). Six-week-old male C57BL/6 mice purchased from Pengyue Co., Ltd were used as the experimental mice. They were adaptively fed for 7 d (SPF facility, 12-h light/dark cycle, 20–24°C temperature, 40–60 % relative humidity) with ad libitum access to food and water.
In the experiments, we first weighed the animals and numbered them according to their weight. And then the mice were randomly numbered and assigned to five groups using the SPSS 22.0 software: control group (CON), high-fat diet group (HFD), positive control group (simvastatin (SV): positive control for the inhibition of cholesterol biosynthesis), L. casei YRL577 group and L. paracasei X11 group. All investigators were blinded to the treatment. The NAFLD model was constructed according to a previously described method(Reference Wang, Cao and Fu20). The control group was fed a normal diet, and the other groups were fed a HFD (20 % protein composed of casein and L-cystine; 35 % carbohydrate composed of maize starch, maltodextrin and sucrose; 45 % fat composed of soybean oil and lard) for 8 weeks(Reference De Minicis, Rychlicki and Agostinelli21). Mice were intraperitoneally injected with a carbon tetrachloride (CCl4)–vegetable oil (v/v) solution at a dose of 0·72 ml/100 g for the first week, followed by the intraperitoneal injection of 40 % CCl4 solution at a dose of 0·42 ml/100 g for 3 weeks. From weeks 9 to 17, mice in the intervention group began to receive the corresponding reagent. Probiotics were administered at a dose of 10 ml/kg body weight, and simvastatin was administered at 3 mg/kg body weight. Each experimental group was gavaged with probiotics or SV once a day. We used the following formula to determine the human equivalent dose (HED): HED (mg/kg) = animal (mg/kg) × (weight animal (kg)/weight human (kg))(1–0·67)(Reference Nair and Jacob22). Factor method for body surface area was applied to calculate the dose converting between animals and humans. Thus, probiotics = 10 ml/kg × (0·022 kg/70 kg)0·33 = 0·69 ml/kg or 48·86 ml for a 70 kg human and SV = 3 mg/kg × (0·022 kg/70 kg)0·33 = 0·21 mg/kg or 14·67 mg for a 70 kg human. Mice were killed 8 weeks after gavage, which were allowed ad libitum access to drinking water and were subjected to fasting 12 h before killing. Blood samples were collected from the eyeball veins of the mice. Hepatic and intestinal tissues were collected and stored in a refrigerator at –80°C until further use.
Measurement of body weight and liver index
The weight of mice was recorded every week, and weight changes in the mice of each group were monitored. The weight of the whole liver was also recorded, and the liver index of mice was calculated according to the following formula(Reference Kleiner, Brunt and Van Natta1).
 $${\rm{Liver\;index\;}}\left( {\rm{\% }} \right) = {{{\rm{Liver\ weight }}\over {{\rm{Body\ weight}}}}\times 100 \,%.$$
$${\rm{Liver\;index\;}}\left( {\rm{\% }} \right) = {{{\rm{Liver\ weight }}\over {{\rm{Body\ weight}}}}\times 100 \,%.$$
Detection of lipid levels
Liver tissue samples were homogenised in 0·9 % cold saline and then centrifuged at 3000 rpm for 10 min to obtain the supernatant. The levels of total cholesterol (TC) and TAG in the liver and TC, TAG, HDL-cholesterol and LDL-cholesterol levels in the serum were detected using commercial kits (Nanjing Jiancheng) according to the manufacturer’s instructions.
Histopathological analysis of the liver
According to a previous method(Reference Ritze, Bardos and Hubert23), 5 mm × 5 mm mice liver tissue samples were fixed in 4 % paraformaldehyde and then embedded in paraffin. One piece was cut from the paraffin block (4-μm thick) and haematoxylin–eosin-stained. The stained sections were observed under an optical microscope (E100, Nikon) and photographed. The histological assessments were carried out by an independent researcher.
Detection of serum biochemical indicators
The serum levels of alanine aminotransferase (ALT), aspartate aminotransferase (AST), superoxide dismutase (SOD), glutathione peroxidase (GSH-PX) and malondialdehyde (MDA) were measured using commercial kits (Nanjing Jiancheng) according to the manufacturer’s instructions. Commercially available ELISA kits (Nanjing Jiancheng) were used to measure the serum levels of TNF-α and IL-6.
Real-time PCR assay
The mRNA expression levels of FXR, FGF15 and apical Na-dependent bile acid transporter (ASBT) were evaluated via RT-PCR. According to the manufacturer’s instructions, an appropriate amount of liver and ileum tissues were weighed, followed by the addition of Trizol (Invertrogen) lysate to extract the total RNA from the tissues. Then, cDNA was synthesised using the ReverTra Ace® qPCR RT Master Mix reverse transcription reaction kit (TOYOBO). Forward and reverse primer sequences are shown in Table 1. After adding SYBR Green, the reaction was performed in a real-time PCR apparatus (ABI). The ratio of the detection value of each target gene to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) represented the expression level of each target gene. ΔΔCt was used to calculate the fold difference in gene expression(Reference Li, Ikaga and Yamazaki24).
Target gene primer sequences

FXR, farnesoid X receptor; FGF15, fibroblast growth factor 15; ASBT, apical Na-dependent bile acid transporter; GAPDH, glyceraldehyde 3-phosphate dehydrogenase.
Statistical analysis
All data are expressed as mean values and standard deviations. Statistical differences in data amongst groups were determined using one-way ANOVA, and post hoc analysis through the Student–Newman–Keuls difference test using SPSS 22.0 software. P < 0·05 was considered statistically significant. The sample size was chosen based on our previous preliminary experiment. The levels of TC and TAG in the liver were important indicators for NAFLD. So we mainly analysed the TC and TAG in the liver using one-way ANOVA. The F value of TC and TAG is 9·214 and 27·523, and the effect size of TC and TAG is 0·617 and 0·756. We have done an analysis using G* Power (v 3.1.9.4). With a P = 0·05 and a power of 0·80, we got a total sample size of 40 and 30. So we chose ten mice in each group to obtain more valid statistical data.
Results
Analysis of bile salt hydrolase activity of strains
Qualitative determination of the BSH activity of thirty-four strains revealed that the precipitation circle appeared in eleven strains, indicating that these strains may have BSH activity (Table 2). Next, we performed the quantitative determination of the BSH activity of eleven strains. The results are shown in Table 3. All strains had BSH activity on both substrates, and there were significant differences in BSH activity between TDCA and glycine deoxycholate. Among the different strains, L. paracasei X11 and L. casei YRL577 had the highest BSH activity for TDCA and glycine deoxycholate (P < 0·05).
Qualitative determination of bile salt hydrolase (BSH) activity of the strains*

TDCA, taurine deoxycholate.
* BSH activity was determined on the halo zone (diameter); ‘+’ positive response: halo >8 mm; ‘−’ negative response: no detected halo.
Quantitative determination of bile salt hydrolase (BSH) activity of the strains
(Mean values and standard deviations)

TDCA, taurine deoxycholate; GDCA, glycine deoxycholate.
a,b,c,d,e,f Unlike letters in the same column represent significant differences (P < 0·05).
Effects of probiotics on body weight, liver weight and liver index in mice
According to the qualitative and quantitative experimental results of BSH activity, the best effect was observed for L. paracasei X11 and L. casei YRL577 strains. Therefore, they were selected to explore the relieving effect on NAFLD in vivo. As shown in Fig. 1, at baseline, the weight of mice fed with a HFD in each group was not statistically different. However, the body weight of mice changed after 8 weeks of intervention in each experimental group. The weight gain of the HFD group was significantly higher than that of the CON group by the eighth week (P < 0·05). The experimental groups did not show statistically different body weights but showed significantly decreased body weight in the last 2 weeks compared with the HFD group (P < 0·05).
Changes in the body weight of mice in different treatment groups. Data are presented as mean values and standard deviations (n 10 per group). a,b Unlike letters in the same column represent significant differences (P < 0·05). ![]() , Control;
, Control; ![]() , high-fat diet;
, high-fat diet; ![]() , simvastatin;
, simvastatin; ![]() , Lactobacillus paracasei X11;
, Lactobacillus paracasei X11; ![]() , L. casei YRL577.
, L. casei YRL577.

Table 4 shows that the liver weight and liver index of the HFD group were significantly higher than those of the CON and experimental groups (P < 0·05). The liver index of the HFD group increased by 30·88 % compared with the CON group. Compared with the HFD group, the liver index of the L. paracasei X11 and L. casei YRL577 groups decreased by 10·19 and 17·69 %, respectively (P < 0·05). The L. casei YRL577 group showed a better effect, but it was not significantly different from the SV group.
Liver weight and liver index of mice after 8 weeks of intervention
(Mean values and standard deviations)

CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.
a,b,c,d Unlike letters in the same column represent significant differences (P < 0·05).
Effects of probiotics on lipids indicators in mice
According to the analysis of mouse lipid-related indicators (Table 5), compared with other groups, the HFD group had the higher serum TAG, TC and LDL-cholesterol levels (P < 0·05). In contrast, the HFD group had the lower serum HDL-cholesterol level (P < 0·05). Compared with the HFD group, the TC, TAG and LDL-cholesterol levels in the L. paracasei X11 group were reduced by 36·78, 23·67 and 69·28 %, respectively, whereas HDL-cholesterol levels increased by 43·28 % (P < 0·05). TC, TAG and LDL-cholesterol levels in the L. casei YRL577 group decreased by 46·53, 31·88 and 75·16 %, respectively, whereas HDL-cholesterol levels increased by 56·72 % (P < 0·05). The L. casei YRL577 group showed a better effect than the L. paracasei X11 group, which did not have any significant difference compared with the SV group.
Biochemical parameters of lipid metabolism for non-alcoholic fatty liver disease in the serum
(Mean values and standard deviations)

TC, total cholesterol; CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.
a,b,c,d,e Unlike letters in the same column represent significant differences (P < 0·05).
After 8 weeks of continuous intervention, the results of TC and TAG levels in mice liver are shown in Table 6. The TC and TAG levels were the higher in the HFD group, compared with other groups. Compared with the HFD group, the liver TC content in the L. paracasei X11 group decreased by 20·48 % and that of TAG decreased by 11·73 % (P < 0·05). The liver TC and TAG levels in the L. paracasei YRL577 group decreased by 22·85 and 20·35 %, respectively (P < 0·05). L. casei YRL577 and L. paracasei X11 showed similar lipid-lowering effects (P > 0·05).
Contents of total cholesterol (TC) and TAG in liver
(Mean values and standard deviations)

CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.
a,b,c Unlike letters in the same column represent significant differences (P < 0·05).
Histopathological analysis of the liver
As shown in Fig. 2, the liver cells in the CON group had normal morphology and did not have fat accumulation. On the other hand, lipid infiltration was severe and fatty vacuole deposition was most obvious in the HFD group, with a proportion of approximately 10–20 %. The L. casei YRL577 and L. paracasei X11 groups achieved different degrees of improvement, alleviated lipid infiltration and reduced the number of fat vacuoles, even though they did not return to the levels in the CON or SV group.
Histopathological analyses of haematoxylin–eosin-stained liver sections from mice. Arrows indicate the areas of fat accumulation; 400× magnification. CON, control; HFD, high-fat diet; SV, simvastatin; YRL577, Lactobacillus casei YRL577; X11, L. paracasei X11.

Effects of probiotics on serum biochemical indicators
The serum AST and ALT activities are shown in Table 7. Compared with the CON group, the serum AST level in the HFD group increased 3·21-fold and that of ALT increased 2·76-fold. AST and ALT levels were significantly reduced after L. paracasei X11 and L. casei YRL577 supplementation (P < 0·05). The effects of L. casei YRL557 were better than those of L. paracasei X11 in maintaining AST and ALT levels. However, it was inferior to those of SV (P < 0·05).
Serum alanine aminotransferase (ALT) and aspartate aminotransferase (AST) levels of mice
(Mean values and standard deviations)

CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.
a,b,c,d Unlike letters in the same column represent significant differences (P < 0·05).
The results of serum antioxidant biomarkers are shown in Table 8. Compared with other groups, serum SOD and GSH-PX levels were lower in the HFD group. In contrast, serum MDA levels were higher than those of other groups. Compared with the HFD group, the L. paracasei X11 group only decreased the MDA content. However, the SOD and GSH-PX levels of the L. casei YRL577 group increased by 55·23 and 53·17 %, respectively (P < 0·05) and the MDA content decreased by 37·33 % (P < 0·05). The results show that L. casei YRL577 is remarkably better than L. paracasei X11 in terms of antioxidant activities and basically restores them to the levels in the CON group (P < 0·05).
Biochemical parameters of the antioxidant properties in serum
(Mean values and standard deviations)

SOD, superoxide dismutase; GSH-PX, glutathione peroxidase; MDA, malondialdehyde; CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.
a,b,c,d Unlike letters in the same column represent significant differences (P < 0·05).
Effects of probiotics on proinflammatory factors in the non-alcoholic fatty liver disease mouse model
Serum TNF-α and IL-6 levels were higher in the HFD group than in the other groups (P < 0·05) (Fig. 3). Compared with the HFD group, the TNF-α and IL-6 content in the L. casei YRL577 group decreased by 11·24 and 41·28 %, respectively (P < 0·05), which were consistent with those of the SV group. The IL-6 content in the L. paracasei X11 group decreased by 28·33 % without any significant difference in TNF-α content compared with the HFD group, which differed with the results of the L. casei YRL577 group in terms of regulating these proinflammatory factors (P < 0·05).
(A) Serum TNF-α levels in mice. (B) Serum IL-6 levels in mice. Data are presented as mean values and standard deviations (n 8–10 per group). a,b,c,d Unlike letters represent significant differences (P < 0·05). CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.

Effects of probiotics on expression of farnesoid X receptor, fibroblast growth factor 15 and apical sodium-dependent bile acid transporter in the non-alcoholic fatty liver disease mouse model
After 8 weeks of intervention, the HFD group had no significant changes in the expression of FXR and FGF15 compared with CON group (Fig. 4). However, the ASBT level significantly increased in the HFD group (2·3-fold) compared with the CON group (P < 0·05). Compared with the HFD group, the levels of FXR and FGF15 in the SV and L. casei YRL577 groups increased by almost 2·3-fold, whereas those in the L. paracasei X11 group increased by almost 1·8-fold. In addition, the L. casei YRL577 group significantly downregulated the ASBT level more than the L. paracasei X11 group (P < 0·05), which was close to the CON group.
(A) Intestinal mRNA expression level of farnesoid X receptor (FXR). (B) Intestinal mRNA expression level of fibroblast growth factor 15 (FGF15). (C) Intestinal mRNA expression level of apical sodium-dependent bile acid transporter (ASBT). Data are presented as mean values and standard deviations (n 6 per group). a,b,c,d Unlike letters represent significant differences (P < 0·05). CON, control; HFD, high-fat diet; SV, simvastatin; X11, Lactobacillus paracasei X11; YRL577, L. casei YRL577.

Discussion
In this study, we found that L. casei YRL577 had BSH activity and could regulate lipid metabolism, oxidative stress and proinflammatory cytokine levels. Furthermore, it had an effect on the regulation of the mRNA expression levels of FXR, FGF15 and ASBT. These findings suggested that L. casei YRL577 could alleviate NAFLD.
NAFLD covers a range of liver diseases, which are characterised by abnormal liver fat accumulation, inflammation and abnormal liver cell function(Reference Haas, Francque and Staels25). Simvastatin reduces cholesterol synthesis by accelerating bile acid synthesis and reducing cholesterol levels(Reference Bader26), which was used as positive control(Reference Jian, Ao and Wu27). BSH activity is known to be related to the ability to decrease cholesterol. A study has reported that probiotics with BSH activity can improve lipid metabolism by reducing serum cholesterol levels(Reference Kim, Park and Sin28). Cholesterol is converted to bile acids in the liver and excreted towards the intestine, which is the main pathway for cholesterol excretion in the body(Reference Mudaliar, Henry and Sanyal29). In this study, L. casei YRL577 and L. paracasei X11 with high BSH activity were selected from the thirty-four strains that exhibited good probiotic performance. L. casei YRL577 effectively reduced abnormal liver enlargement and blood lipid index, as well as ameliorated the occurrence and development of NAFLD more than L. paracasei X11. Compared with SV, L. casei YRL577 had an effect that was similar to the inhibition of cholesterol absorption by SV. They all reduced serum lipid index and liver fat accumulation.
Long-term HFD were found to cause liver lipid accumulation, hepatomegaly and impaired liver functions. ALT and AST are mainly distributed in the hepatocytes(Reference Al Zarzour, Ahmad and Asmawi30). A study has found that L. fermentum and L. plantarum can improve liver damage and reduce the serum levels of AST and ALT(Reference Chen, Zhang and Yi31). In our experiment, we found that the application of L. casei YRL577 and L. paracasei X11 strains could significantly reduce the serum levels of AST and ALT. The effects of L. casei YRL577 were better than those of L. paracasei X11, indicating that it can improve liver damage caused by consuming a HFD. SOD and GSH-PX are important antioxidant biomarkers, whereas MDA is commonly used as an indicator of peroxidation. Changes in the levels of SOD, GSH-PX and MDA can reflect the function of the body’s antioxidant system(Reference Porras, Nistal and Martínez-Flórez32). Our results showed that the use of L. casei YRL577 improved the antioxidant activity of the mouse NAFLD model. Lipid accumulation in the cytoplasm of the liver leads to an inflammatory response in the liver(Reference Henao-Mejia, Elinav and Jin33). NAFLD causes an increase in proinflammatory factors, such as IL-6 and TNF-α (Reference Minxuan, Sun and Dai34,Reference Mridha, Wree and Robertson35) . In this study, the application of L. casei YRL577 alleviated abnormalities in IL-6 and TNF-α levels, relieved proinflammatory symptoms and further attenuated NAFLD. In addition, L. casei YRL577 had an effect that was more similar to SV in terms of alleviating oxidative stress and inflammatory response.
In the enterohepatic circulation, 95 % of the bile acids generated by the liver are reabsorbed in the terminal ileum mainly by ASBT. Approximately 5 % enter the colon and are partially reabsorbed, whereas some are metabolised by intestinal microorganisms before being excreted out of the body(Reference Thomas, Pellicciari and Pruzanski36). Studies have shown that the colonisation of lactobacilli and bifidobacteria in the intestinal tract of mice changed the structure of the intestinal microbiota, increasing the acidification of bound bile acids to free bile acids as well as increasing their excretion via defecation(Reference Degirolamo, Rainaldi and Bovenga37). In our experiment, we found that HFD increased ASBT expression and that the application of L. casei YRL577 decreased its expression better than L. paracasei X11. This suggests that intestinal bile acid reabsorption increases in a HFD and that L. casei YRL577 intervention reduces intestinal bile acid reabsorption and increases excretion, so that the liver synthesises more bile acids to compensate for intestinal bile acid loss. Other studies have confirmed that decreased ASBT expression regulates blood lipid levels and improves insulin resistance(Reference Wu, Aquino and Cowan38). Furthermore, intestinal microbiota-mediated intestinal bile acid biotransformation has been found to regulate metabolism homoeostasis by reducing intestinal ASBT expression levels(Reference Miyata, Yamakawa and Hamatsu39). These findings are consistent with those of our research.
FXR is highly expressed in the intestine and liver and is a modulatory receptor for bile acids(Reference Vavassori, Mencarelli and Renga40). Bile acids are found to be reabsorbed by FXR in small intestinal epithelial cells, specifically in the ileum, which stimulates the secretion of FGF15 in the intestine, triggers the intestinal–liver signalling pathway and regulates bile acid synthesis(Reference Walters, Johnston and Nolan41). A study has confirmed that by regulating the intestinal microbiota to promote enterohepatic bile acid circulation and FXR expression, cholesterol can be converted to bile acids and excreted in vitro (Reference Gao, Fu and Wang42). In our study, the mRNA expression levels of FXR and FGF15 in the intestine were evaluated. FXR and FGF15 expression increased after intervention with L. casei YRL577 and L. paracasei X11. L. casei YRL577 had a better regulatory effect on the expression of intestinal FXR, suggesting that L. casei YRL577 promotes bile acid production and accelerates cholesterol excretion via defecation. In addition, intestinal FXR and FGF15 activation could be induced by increasing energy consumption and heat production, possibly contributing to improving NAFLD(Reference Fang, Suh and Reilly43,Reference Wang, Liao and Zhou44) .
The strengths of this study are that the use of TDCA and glycine deoxycholate in vitro to estimate the BSH activity of L. casei YRL577 and the effect of L. casei YRL577 on genes in the intestinal bile acid pathway permit us to obtain information of the L. casei YRL577 on relieving NAFLD. However, the weakness of this study is that we did not measure the efflux of cholesterol and bile acids, which might have established a direct effect of the observed changes in gene expression on intestinal bile acid pathway modulation and cholesterol excretion. Therefore, our data only show that there is an association between the changes in gene expression and the observed metabolic effects.
Conclusion
This study screened L. casei YRL577 and L. paracasei X11 with high BSH activity and verified their role in alleviating liver injury based on the basic indicators of NAFLD related to mouse models. It was confirmed that L. casei YRL577 had a potential reparative effect on NAFLD in mice. In addition, L. casei YRL577 alleviated the inflammatory response to a certain extent and improved antioxidant activity. Moreover, L. casei YRL577 activated FXR and FGF15 and decreased ASBT expression in the intestine to improve NAFLD. However, the related mechanism of this effect on NAFLD still needs further research.
Acknowledgements
The authors would like to thank the Marine Bioactive Substances Laboratory of Ocean University of China, Qingdao, Shandong, China.
This study was financially supported by National Key R & D of China (2018YFC0311201) and Heilongjiang province Key Sci. & Techn. Plan (GA16B201-2).
L. Z. designed and guided the experiments; Z. Z. and H. Z. wrote the original draft; Z. Z., H. Z., X. Z., J. S. and J. W. performed the experiments; X. L., Y. L., L. B. and J. Z. analysed the data. P. G., T. L. and H. Y. revised the manuscript.
No conflict of interest exits in the submission of this manuscript, and manuscript is approved by all authors for publication.