Introduction
Toxoplasmosis is caused by the obligate intracellular Apicomplexan parasite, Toxoplasma gondii (Wolf et al., Reference Wolf, Cowen and Paige1939). Toxoplasma gondii stands as one of the most prevalent zoonotic parasites globally, chronically infected one-third of the world’s population(Torgerson and Mastroiacovo, Reference Torgerson and Mastroiacovo2013), making it arguably one of the most successful parasites present in nature. The high prevalence of this parasite can be attributed to its ability to infect any warm-blooded animal, its capacity to persist as a chronic infection, and its multiple modes of transmission which include both horizontal and vertical transmission (Dubey, Reference Dubey1998). Toxoplasmosis is paradigmatic of the One Health concept as it highlights the interconnection of human, animal, and environmental health. Felines, both domestic and wild, act as definitive hosts, responsible for shedding oocysts in their feces, critical for horizontal transmission (Dubey et al., Reference Dubey, Miller and Frenkel1970; Frenkel et al., Reference Frenkel, Dubey and M1970). Similarly, carnivorism among intermediate hosts facilitates parasite transmission through consumption of cysts lodged in muscles and the brain (Webster, 2010). Either parasite source can result in fetal infection if a pregnant human or animal is exposed to the parasite for the first time during gestation. As per its capacity to be vertically transmitted, T. gondii is included as part of the infamous TORCH group of key infectious agents associated with congenital complications in newborns (the ‘T’ in TORCH stands for T. gondii) (Jaan and Rajnik, Reference Jaan and Rajnik2025).
The outcome of acute infection depends on immune status. The most devastating consequences of toxoplasmosis are related to the parasite accessing, infecting, and persisting within immune-privileged sites such as the eye, brain and placenta and immunosuppression (Weiss and Kim, Reference Weiss and Kim2000). During pregnancy, the consequences of an acquired infection vary depending on the gestational period, the foetal age, the inoculum, and both the host’s and the parasite’s genetic background (de Lima Bessa et al., Reference de Lima Bessa, de Almeida Vitor and Dos Santos Martins-duarte2021; Sanchez and Besteiro, Reference Sanchez and Besteiro2021). In general, the earlier the gestational period, the higher the probability of pregnancy loss or fetal damage, while the higher the gestational age, the higher the probability of transmission to the fetus (Weiss and Dubey, Reference Weiss and Dubey2009). Congenitally acquired toxoplasmosis presents itself as broad spectrum of manifestation ranging from premature birth, chorioretinitis or blindness, cognitive deficiencies or impairments in psychomotor development, hydrocephalus, macro- and microcephaly, intracranial calcifications, fetal hydrops and hepatosplenomegaly, among others (Montoya and Remington, Reference Montoya and Remington1996; Montoya et al., Reference Montoya, Jordan, Lingamneni, Berry and Remington1997; Montoya and Liesenfeld, Reference Montoya and Liesenfeld2004).
In most cases, a prior toxoplasmosis infection protects subsequent pregnancies. Nonetheless, this paradigm has been challenged as several studies of pregnant women with chronic infections, this is, acquired prior to pregnancy, report ocular reactivations during the gestational period (I et al.). Likewise, cases of reactivation of latent infections during pregnancy and the consequent vertical transmission (VT), albeit rare, have reported incidence solely in Latin America (Elbez-Rubinstein et al., Reference Elbez-Rubinstein, Ajzenberg, Dardé, Cohen, Dumètre, Yera, Gondon, Janaud and Thulliez2009). Moreover, growing evidence, based not only on murine models of vertical transmission but also on rare clinical cases, suggests that reinfection is possible when a genetically distinct strain reinfects a seemingly ‘immunized’ individual (Nicoll et al., Reference Nicoll, Wright, Maley, Burns and Buxton1997; Dao et al., Reference Dao, Fortier, Soete, Plenat and Dubremetz2001; Elbez-Rubinstein et al., Reference Elbez-Rubinstein, Ajzenberg, Dardé, Cohen, Dumètre, Yera, Gondon, Janaud and Thulliez2009)
The parasite’s genotype plays a role in the severity of toxoplasmosis and is closely linked to its geographical origin. Genetically homogenous populations circulate primarily in North America and Europe, dominated by the presence of Type II and III strains, so-called ‘Archetypal’ (Fernández-Escobar et al., Reference Fernández-Escobar, Schares, Maksimov, Joeres, Ortega-Mora and Calero-Bernal2022). Strains that circulate in South America, Africa and Asia have been termed non-archetypal (Bertranpetit et al., Reference Bertranpetit, Jombart, Paradis, Pena, Dubey, Su, Mercier, Devillard and Ajzenberg2017; Galal et al., Reference Galal, Hamidović, Dardé and Mercier2019, Reference Galal, Ariey, Gouilh, Dardé, Hamidović, Letourneur, Prugnolle and Mercier2022; Hosseini et al., Reference Hosseini, Amouei, Sharif, Sarvi, Galal, Javidnia, Pagheh, Gholami, Mizani and Daryani2019; de Lima Bessa et al., Reference de Lima Bessa, de Almeida Vitor and Dos Santos Martins-duarte2021). Although non-archetypal strains display a variety of phenotypes ranging from attenuated to highly virulent, a number of strains have been shown capable of causing fatal disease in immunocompetent individuals (Bossi et al., Reference Bossi, Caumes, Paris, Dardé and Bricaire1998; De Salvador-Guillouët et al., Reference De Salvador-Guillouët, Ajzenberg, Chaillou-Opitz, Saint-Paul, Dunais, Dellamonica and Marty2006; Leal et al., Reference Leal, Cavazzana, Andrade, Galisteo, Mendonca and Kallas2007; Cuomo et al., Reference Cuomo, D’Abrosca, Rizzo, Nardiello, La Montagna, Gaeta and Valentini2013). It is known that a high frequency of ocular disease is reported in immunocompetent individuals in South America (Vaudaux et al., Reference Vaudaux, Muccioli, James, Silveira, Magargal, Jung, Dubey, Jones, Doymaz, Bruckner, Belfort, Holland and Grigg2010). Consistently, children congenitally infected with these strains tend to develop more severe disease manifestations such as bilateral ocular toxoplasmosis. Moreover, non-archetypal strains have been shown to display natural resistance to the main drugs used for treatment in the clinical settings, even without prior exposure (de Lima Bessa et al., Reference de Lima Bessa, Vitor, Lobo, Rêgo, de Souza, Lopes, Costa and Martins-Duarte2023).
Serological screening for T. gondii in pregnant women in Uruguay is routinely conducted through enzyme immunoassays (EIA) detecting anti-Toxoplasma antibodies. Women resulting in IgG positive and IgM negative results in their initial screening are no longer followed, as pre-existing immunity is generally considered protective. IgG and IgM negative women are screened throughout gestation. Women who seroconvert are derived to specialized, high-risk pregnancy dedicated care. A survey dating from 1996, reported, based on these data, that 53% of pregnant women were chronic carriers of T. gondii (Díaz et al., Reference Díaz, Ismael, Alvaro, Graciela, Claudia, Juliana, Carmen, Berenice, López, Juan and González, A1998a). Additionally, it was determined that up to 3% pregnant women seroconverted during pregnancy. A follow-up study of primary infections during pregnancy reported a vertical transmission efficiency of 9% (Barrios et al., Reference Barrios, Más, Baroloco, Sayaguez and Giachetto2016), a figure that is below the global average of 28% (Torgerson and Mastroiacovo, Reference Torgerson and Mastroiacovo2013). These cumulative figures lead to the estimation that 150 congenitally infected children are born yearly (Amorín et al., Reference Amorín, Pérez and Martínez2015)
However, extensive studies on T. gondii prevalence in the general population have not been carried out recently, nor have there been molecular studies to determine the genetic diversity among circulating strains. CASTELLS, the only Uruguayan-origin strain reported in the literature that has been genotyped, belongs to an genetic clade, which it shares with other non-archetypal isolates (Clade E), and is highly virulent in primary infections (Su et al., Reference Su, Khan, Zhou, Majumdar, Ajzenberg, Dardé, Zhu, Ajioka, Rosenthal, Dubey and Sibley2012; Jensen et al., Reference Jensen, Camejo, Melo, Cordeiro, Julien, Grotenbreg, Frickel, Ploegh, Young and Saeij2015). In addition, Sousa demonstrated by serotyping that most chronic carriers in the country developed immunity to strains exhibiting non-archetypal polymorphic profiles of antigenic GRA proteins (Sousa et al., Reference Sousa, Puime, Costa and Dardé2017). While these findings highlight the importance of genotyping circulating T. gondii strains, as this could directly influence public health policies and decision-making, no previous study has focused on genotyping T. gondii in Uruguay, leaving the full extent of the genetic diversity largely unexplored.
Herein, we update the toxoplasmosis seroprevalence and seroconversion rates of a cohort of patients based on serological surveys done during pregnancy follow-up controls in the Centro Hospitalario Pereira Rossell (CHPR), the major maternity public hospital in Uruguay between years 2019 and 2023. For the mother–baby binomials where congenital toxoplasmosis was confirmed, we analysed these data considering timing of seroconversion and frequency of clinical manifestations present in the newborns. Additionally, molecular detection of T. gondii in a variety of biological fluids and tissue samples enabled us to partially genotype circulating strains, enhancing our understanding of the T. gondii genetic landscape in the country contributing, in turn, to our wider understanding of the parasite’s genetic diversity in the continent.
Materials and methods
Subjects of this study
For this study, three different patient cohorts were selected:
Cohort 1 includes all pregnant women that were followed up at the Centro Hospitalario Pereira Rossell (CHPR) between 2019 and 2023 and had validated anti-T. gondii serological test (23 332). Valid serology tests were those which complied with the quality controls as determined by the test performing laboratory.
Cohort 2 includes all babies born at the CHPR between 2019 and 2023, for which congenital toxoplasmosis (CT) was confirmed (14 cases). CT incidence is expressed as the number of in utero infected newborns every 1000 births at CHPR during the years 2019–2023.
Cohort 3 consists of mother–newborn pairs. Fifty-three patients were recruited from two of the largest public hospitals in Montevideo: The High-Risk Obstetric Clinic ‘Dr. Manuel Quintela’ at the Hospital de Clínicas, and the High-Risk Obstetric Clinic and Department of Neonatology at the CHPR. Patients were recruited into this study, either because the mother presented positive IgM or a serological profile that suggested seroconversion during pregnancy (i.e. low avidity IgG and positive IgM, or persistent IgM despite high avidity IgG, as determined by the VIDAS Avidity test (Biomerieux)). Note that in these cases, mothers were most frequently treated with spiramycin as per the Ministry of Health’s recommendations. Eighty-one samples were collected either from the mother or the newborn at time of birth. In rare occasions, samples were obtained from both the mother and the newborn. These included the mother’s peripheral blood and placenta, umbilical cord blood and the newborn’s peripheral blood. Blood was stored and transported in dry tubes and refrigerated if not processed immediately. All samples were stored at 4°C for less than 24 hours prior to DNA extraction. Only samples from this cohort were subjected to DNA extraction, PCR amplification of parasitic DNA and further analyses. All positive samples from this cohort only were subsequently genotyped as described further.
Seroprevalence and seroconversion determination
Serological results corresponding to cohort 1 were used to determine seroprevalence. Patients were considered incident if they receive a diagnosis during the observational period but not in the pre-observational period. Serostatus was determined using an anti-T. gondii IgG antibody determination kit (automatic commercial quantitative enzyme-linked fluorescence assay ELFA; VIDAS TOXO IgG II;Biomérieux). Results were interpreted as follows: titres of 0–3 were considered negative, 4–7 equivocal and ≥ 8 positive. We excluded all samples with equivocal test results (1.5%) from the analysis. Seroprevalence was defined as the proportion of seropositive women in all women included in the survey with serological information available. Incidence is defined as the number of seroconversions/1000 susceptible women per year. We defined a seroconversion during pregnancy as the presence of IgG- and/or IgM-specific antibodies for toxoplasmosis after negative tests during pregnancy with dates available for the last negative test and the first positive test. Incidence was calculated as the number of new cases of congenital toxoplasmosis per year period of study divided by the number of neonates who are initially disease free.
Congenital toxoplasmosis diagnosis
CT confirmation includes maternal seroconversion, positive serology in the newborn and/or the presence of clinical manifestations consistent with toxoplasmosis, such as chorioretinitis, cerebrospinal fluid abnormalities, brain calcifications, hydrocephalus, seizures, microcephaly, cataracts and microphthalmia, observed after birth (Baquero-Artigao et al., Reference Baquero-Artigao, Del Castillo Martín, Fuentes Corripio, Goncé Mellgren, Fortuny Guasch, de la Calle Fernández-Miranda, González-Tomé, Couceiro Gianzo, Neth and Ramos Amador2013). CT incidence is expressed as the number of in utero infected newborns every 1000 births at CHPR during the years 2019–2023.
DNA extraction
DNA extraction was carried out from tissue and whole-blood samples using the Quick-DNA Tissue/Insect Miniprep Kit (D6016, Zymo Research) and following the manufacturer’s instructions. In the case of placental tissue, efforts were made to identify and purify DNA from regions exhibiting macroscopic lesions suggestive of possible T. gondii presence (see supplementary Figure 2).
PC detection of T. gondii’s DNA
Presence of T. gondii DNA in the samples was assayed by previously published PCR assays (Homan et al., Reference Homan, Vercammen, De Braekeleer and Verschueren2000; Schares et al., Reference Schares, Vrhovec, Pantchev, Herrmann and Conraths2008). In short, specific primer sets were used for T. gondii DNA detection. COC 1–2 (coccidia detection, class of Apicomplexa: F: 5′ AAGTATAAGCTTTTATACGGCT 3′ R: 5′ CACTGCCACGGTAGTCCAATAC 3′ annealing temperature 55°C, amplification product molecular weight 298 base pairs), B1 (the 529 pb repetitive element) and TOX 8-5 using the following primer pairs: B1 Forward 5′ GGACTGCATCCGTTCATGAG 3′ B1 Reverse: 5′ TCTTTAAAGCGTTCGTGGTC 3′ annealing temperature 55°C, amplification product molecular weight 800bp, and Tox8 5′ CCCAGCTGCGTCTGTCGGGAT 3′ Tox5: 5′ CGCTGCAGACACAGTGCATCTGGATT 3′ annealing temperature 60°C, amplification product molecular weight 500 bp, respectively) in combination with Mango Mix (BIO-25033/ meridian BIOSCIENCE) following the manufacturer’s specifications. NPOC (nuclear gene of mammals; NPOC F 5′ GCATCCTTGAGTGTGAAGAGAA 3′ NPOC R: 5′ TGCCTCATAAACTCAGTGAACC 3′ annealing temperature 55°C, amplification product molecular weight 300 bp) was used as control (Reischl et al., Reference Reischl, Bretagne, Krüger, Ernault and Costa2003; Schares et al., Reference Schares, Vrhovec, Pantchev, Herrmann and Conraths2008). PCR products were visualized on a 1% agarose gel. Samples in which PCR amplification was simultaneously successful with two of the primer pairs were considered positive. All samples that tested negative were reassessed with GRA6 and BTUB primer pairs (Su et al., Reference Su, Zhang and Dubey2006).
Genotyping
Toxoplasma gondii genotyping was performed on positive samples by in silico PCR-RFLP. We specifically characterized polymorphisms in the following marker genes: SAG2, SAG3, BTUB, GRA6, c29-2, c22-8, L358, PK1, and ‘Apico’ markers. Subsequently, these loci were amplified by nested PCR following previously published protocols (Su et al., Reference Su, Zhang and Dubey2006). Positive PCR products were sequenced by the Sanger method (MACROGEN, Korea). Once the sequences were obtained, genotyping was performed using online computational tools to analyze restriction profiles following the methods described by Homan et al. (Reference Homan, Vercammen, De Braekeleer and Verschueren2000) and Castro et al. (Reference Castro, Gennari, Lorenzi and Su2020). Each amplified marker sequence was reviewed and refined based on sequencing histogram peaks using the free software ApE Plasmid Editor (Davis and Jorgensen, Reference Davis and Jorgensen2022) . The same tool was then employed to assess restriction digestion patterns for each sequenced marker, as well as for reference strains RH, ME49 and VEG, representing the archetypal T. gondii genotypes. This analysis was conducted using the virtual restriction digest feature of ApE Plasmid Editor, with restriction enzymes selected according to Su et al. (Reference Su, Zhang and Dubey2006). The resulting digestion patterns were compared with those of the reference strains to classify each marker into genotype I, II or III. Any restriction profile deviating from those expected for archetypal was classified as ‘non-archetypal’.
Once sequences were obtained, genotyping was performed using online computational tools to analyse the different restriction profiles according to Homan et al. (Reference Homan, Vercammen, De Braekeleer and Verschueren2000) and Castro et al. (Reference Castro, Gennari, Lorenzi and Su2020). For all restriction profiles which differed from those expected for typical strains (i.e. Types I, II or III), the sample was classified as ‘non-archetypal’. If a combination of typical strain’s genetic loci were detected, the strain was deemed as ‘non-clonal’ as per the nomenclature established by Ajzenberg et al. (Reference Ajzenberg, Bañuls, Su, Dumètre, Demar, Carme and Dardé2004). All new sequences reported in this work have been deposited and are publicly available under GenBank IDs PV564118 to PV564179 (Supplementary T 5).
Phylogenetic inference
Phylogenetic inference was conducted using RAxML v8 (Stamatakis, Reference Stamatakis2014) from a concatenated alignment of all genetic markers. Sequences were aligned using MAFFT (Katoh, Reference Katoh2002) with the L-INS-i algorithm (–localpair and – maxiterate 1000) to ensure high accuracy. Alignment confidence scores were computed with rGUIDANCE (Krah and Heibl, Reference Krah and Heibl2019) and the resulting confidence weights were incorporated into the analysis via a weights file.
The maximum likelihood (ML) tree was constructed using the GTRCAT substitution model. A Neighbor-Joining (NJ) tree based on the TN93 distance was used as the starting tree. The -C and -M options of RAxML were employed to optimize the model and address missing data. Gene-partitioned analyses were specified using a partition file (-q partitions), and the final tree was generated through an exhaustive search (-f I). To evaluate the robustness of the inferred topology, 100 bootstrap replicates were performed using RAxML’s rapid bootstrap algorithm (-f a). The resulting bootstrap support values were mapped onto the ML tree to assess clade reliability. Accession numbers of all sequences used for the construction of the phylogenetic tree can be found in Supplementary T 4.
Results
Seroprevalence and seroconversion (Cohort 1)
Between 2019 and 2023, the country averaged 34 326 births per year, with the CHPR hosting an average of 5751 births annually, accounting for 16.8% of all births in the country during this period. Additionally, CHPR is the largest maternity ward in Montevideo, accounting for approximately 93.4% of the births at public hospitals in the capital city (whereby over 50% of the country’s population resides).
During the years 2019–2023 of the 27 928 pregnancies that were monitored at CHPR, 23 332 presented serological screen results for T. gondii. Over the period analysed, 45.5% (10 611/23 332) of patients had detectable IgG titres, indicating that they had been previously exposed to T. gondii (Figure 1A and Supplementary T 2). Over the 5-year period analysed, this percentage remained relatively stable, showing no significant fluctuation. We also analysed the percentage of patients experiencing seroconversion. On average, we detected a rate of seroconversion of 0.58% during pregnancy, indicating that about 1 in every 172 immune-naive patients were exposed to the parasite for the first time during pregnancy. This translates into 2.6 cases of seroconversion every 1000 births annually (Figure 1A and Supplementary T 2).
Seroprevalence, seroconversion and congenital transmission. (A) Seroprevalence and seroconversion rates are shown as percentages of all serological tests pursued between 2019 and 2023. (B) Seroconversion and CT cases are shown, represented as cases per 1000 births, as indicated. (C). For confirmed congenital cases, the trimester of diagnosis is shown. Grey bars indicate the percentage of cases in which the mother received treatment upon diagnosis. For several cases, the trimester of diagnosed was not registered in the patient’s clinical history and could not be determined (UD). Please note that all data are shown as percentages of the total number of CT cases registered between 2019 and 2023. (D) Quantification of the number of CT cases in which clinical manifestations were observed and its correlation with maternal treatment upon diagnosis. Please note that in most cases in which clinical manifestations were observed, no treatment was indicated. (E). Quantification of the distribution of clinical manifestations, shown as percentages of all CT cases registered between 2019 and 2023.

Congenital toxoplasmosis incidence (Cohort 2)
During the years 2019–2023, 14 cases of congenital toxoplasmosis were reported (Supplementary T 1). Of these, further information was recorded for 13 cases (Supplementary T 2). We note that these data does not include undiagnosed spontaneous pregnancy losses neither voluntarily interrupted pregnancies, which are legal in Uruguay up until week 12 of gestation. CT cases were confirmed by serological detection of increasing titers of IgG and/or presence of IgM in the newborn, and/or clinical manifestations consistent with toxoplasmosis (Baquero-Artigao et al., Reference Baquero-Artigao, Del Castillo Martín, Fuentes Corripio, Goncé Mellgren, Fortuny Guasch, de la Calle Fernández-Miranda, González-Tomé, Couceiro Gianzo, Neth and Ramos Amador2013). Altogether, these data account for a cumulative incidence of CT of 0.5 newborns for every 1000 births tended at CHPR between 2019 and 2023 (14 cases in 23332 controlled pregnancies; Figure 1C and Supplementary T 2).
Forty-five percent (5/13) of the CT cases were diagnosed in utero in the third trimester. Of these, 60% received treatment (3/5, representing a 30% of total cases) (Figure 1B). A total of 25% (3/13) of cases were diagnosed in the second trimester, with 66% receiving treatment (2/3; 15% of total cases). Notably, cases diagnosed in the first trimester did not receive any treatment. Finally, 23.1% of cases (3/13) were detected during pregnancy, however, the trimester of seroconversion was not registered (Undetermined; UD, Figure 1B and Supplementary T 2). One of these cases received treatment. Notably, of the seven treated cases, six received spiramycin.
Among newborns presenting clinical manifestations of CT (60%; 8/13, Figure 1C and D, and Supplementary T 2), the majority (5/7) were born from mothers who did not receive prenatal treatment (Figure 1D and Supplementary T 2). In contrast, the incidence of newborns displaying CT clinical manifestations, among mothers who received treatment decreased to 42% (3/7) (Figure 1D, purple bar). The most prevalent clinical manifestation observed was chorioretinitis (53.9%; 6/13) followed by neurological affectations (38,5%; 5/13) (Figure 1D and Supplementary T 2). No correlation was found between the IgG or IgM titres of these patients and treatment (1).
Genotyping (Cohort 3)
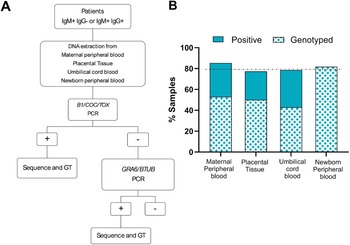
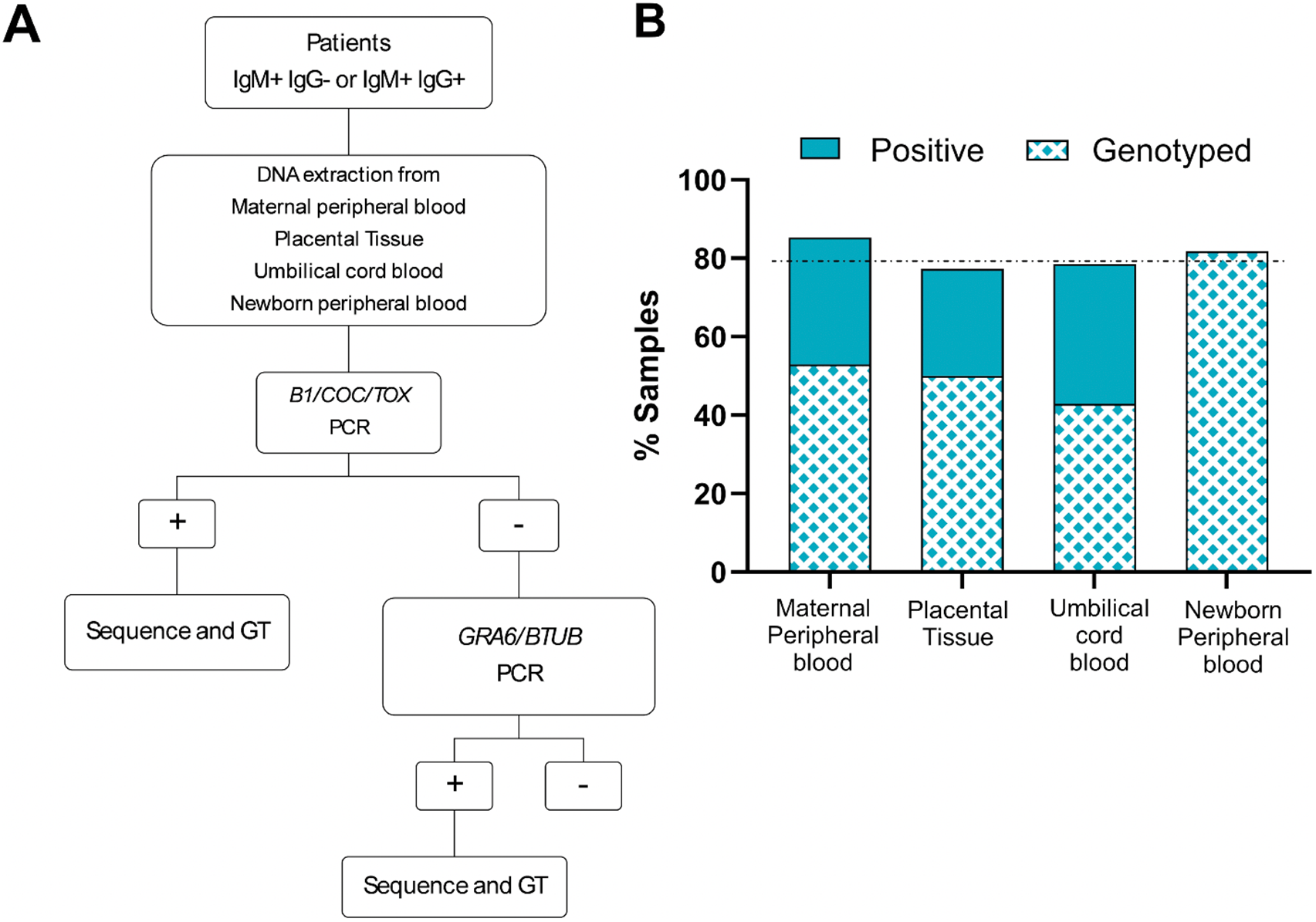
To identify the T. gondii genotypes present in the population of patients who seroconverted during pregnancy, patients who displayed IgG-/IgM + or IgG +/IgM + serologies were enrolled in the study and samples were collected. A total of 81 samples from 53 mother–newborn pairs were received in the laboratory. Samples included maternal peripheral blood (34), placental samples (22), umbilical cord blood samples (14), and peripheral blood samples from the newborns (11). Following DNA extraction, parasite detection was first pursued by PCR. Positive samples were pursued following the procedure shown in Figure 2A
Diagnostic algorithm and sample processing. (A) Patients recruited into this study corresponded to pregnant women seroconverting during pregnancy. Samples obtained were processed as indicated. Genotyping (GT) was pursued for all PCR positive samples, as indicated. (B) Percentages of PCR positive and genotyped samples per sample type are shown. Sample types received and analysed are indicted in the x-axis. The y-axis corresponds to the percentage of samples which resulted positive for T. gondii DNA detection by PCR. From these, only the indicated percentage (patterned) were of sufficient quality to pursue genotyping by in silico RFLP.
Out of all samples analysed, 82% (67/81) were PCR-positive for T. gondii. However, genotyping was only possible for 53% (39/67) of the PCR-positive samples, representative of 32 patients (Figure 2B). Surprisingly, parasite DNA was frequently detectable in maternal peripheral blood (85.3%; 29/34). However, half of these samples rendered insufficient quality DNA to pursue genotyping. In contrast, all newborn’s peripheral blood samples which resulted positive (81.8%; 9/11), could be genotyped. Placental tissue analysed resulted in T. gondii DNA amplification in 77.3% of cases (17/22). Seventy-one percent of umbilical cord blood samples were PCR positive (8/11). In 20 cases, multiple samples were collected for a mother–newborn pair. In these cases, the correlation among samples in terms of PCR detection of the parasite was variable (Supplementary T 3).
Genotyping was pursued by in silico PCR-RFLP using nine different markers according to Su and collaborators (2006) from DNA extracted from samples belonging to 32 patients (Figure 2B). For 12 of these patients, genotyping efforts were pursued starting from more than one sample. In total, 18 maternal peripheral blood samples, 11 placental samples, 6 umbilical cord blood samples, and 10 newborn peripheral blood samples were genotyped.
In 90.6% of cases, we successfully amplified and analysed the BTUB marker, while in 71.9% of cases, we achieved the same for GRA6. Successful PCR amplification of the remaining markers (SAG2, c29-2, c22-8, L358, PK1 and Apico) was on average 25.5%. In all cases, we were unable to amplify all markers simultaneously. In particular, the SAG3 marker, an established component of the commonly used genotyping panel, could not be amplified for any of the samples. Amplicons were sequenced and analysed in silico for polymorphisms affecting restriction sites. In most cases in which more than one sample was analysed for a patient, or a mother–child pair, genotyping results were consistent unless otherwise stated. These results were consolidated and shown as a single line (Figure 3A).
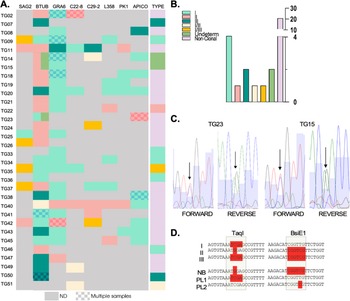
Genotyping of strains circulating in seroconverting pregnant women. (A) Genetic types for each genotyping marker are shown for all indicated samples (‘type’), which resulted from the combination of genetic types obtained per marker, per sample. Note that SAG3 could not be characterized for any of the samples and is hence, not shown. A patterned coloured box indicates that multiple samples were obtained and genotyped for that patient. In all cases in which the block displays a unique colour, the sequence from distinct samples for the same marker matched 100% for those cases in which the same amplified marker from distinct samples resulted in a different genotype, the block is divided in two colours corresponding to the identified genotypes. (B). Quantification of genetic types represented in (A). (C) Sequencing chromatograms. Representative examples of BTUB markers which were deemed undetermined in a and b due to the presence of overlapping peaks in the sequencing chromatogram. Double peaks are indicated by arrows both in the forward and reverse sequences for the indicated samples (tg15 and tg23). (D) The haplotypes obtained for the amplicons sequenced from placental (PL) samples and from newborn blood (NB) are shown in comparison with typical haplotypes (I, II and III) as indicated. Restriction sites, used for haplotype identification in the in silico RFLP analysis, corresponding to regions recognized by the indicated restriction enzymes (taqi and bsie1) are highlighted.
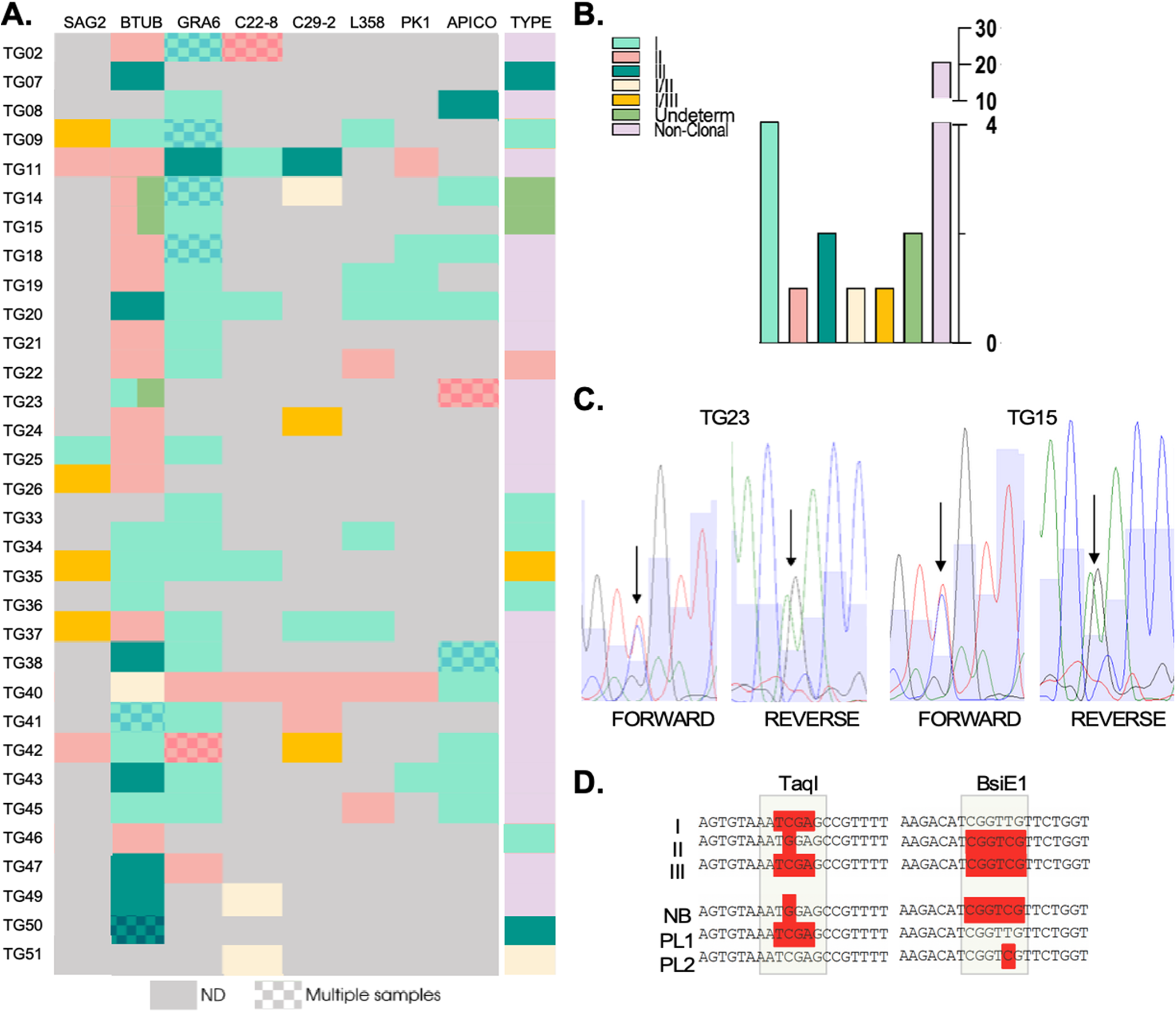
In five cases (corresponding to Tg07, Tg33, Tg36, Tg46 and Tg51), we were able to amplify one marker only out of the nine analysed. Though unlikely, single allele amplification could occur due to stochastic amplification in low concentration DNA samples. We have therefore excluded these samples from further downstream analyses. For Tg34, whereby three markers could be amplified, all corresponded to type I alleles. For the remaining samples corresponding to 26 patients, a combination of restriction profiles corresponding to at least two different archetypal strains, could be detected. The vast majority (17) of these showcased combinations of type I and type II alleles; the resulting genotype obtained for all these samples was denominated ‘non-clonal’ (Figure 3A and B).
Interestingly, for samples corresponding to patients Tg14, Tg15 and Tg23 we could not determine the BTUB allele as no clear base calling could be obtained from the sequencing chromatogram at one specific position which was critical to the genotype determination (Figure 3A and Figure C). For these cases, double peaks with equal representation were present in both the forward and reverse sequences obtained at these specific positions, while all other bases of the allele could be unambiguously called (Figure 3C). In these cases, the BTUB allele was deemed ‘undetermined’. For Tg23, the double peak pattern corresponded to either a Type I or Type III. Sequencing of this marker from DNA isolated from a distinct region of the placenta, resulted in BTUB corresponding to a Type I allele. For these samples, we could additionally amplify and sequence one additional markers (Apico) which resulted, for both samples, in a Type II allele. This combination of markers allowed us to exclude an archetypal genotype. Likewise, for a placental sample corresponding to Tg14, three markers (GRA6, c29-2 y Apico) could be amplified corresponding to Type I and Type I/II alleles.
While in all other cases where multiple samples were analysed for a mother–newborn pair, results were fully consistent (shaded rectangles, Figure 3A). for both Tg14 and Tg15 the alleles amplified from newborn’s blood corresponded to a BTUB Type II alleles in both these cases while the BTUB alelle amplified from a placental sample displayed a similar ‘double peak’ profile whereby the possible genotypes could be attributed to either a Type I or a Type III (Figure 3D).
In case such as those of Tg11, Tg20 and Tg40, among others, we successfully amplified and sequenced a minimum of three alleles. For these samples, we pursued comparison with reported genotypes of isolated strains. While Tg34 partially matches genotype #10, Tg43 partially matches genotype #55. Nonetheless, in both cases not all alleles were amplified, which precludes definitive genotype assignment. Strikingly, all other genotypes (10/12) did not match to any previously reported genotype as obtained by PCR-RFLP, suggesting they correspond to novel genotypes (Figure 4).
Phylogenetic analysis of circulating strains. Maximum likelihood tree reveals the presence of two phylogenetically distinct groups. Markers used for comparison among strains are indicated as shaded boxes for each sample indicated. Samples which lacked sequence information are indicated by clear boxes. Please note that the apico marker was excluded in this analysis. For all samples in which at least three markers could be amplified and sequenced, PCR-RFLP genotype assignment was pursued in toxodb. NR corresponds to genotypes ‘not reported’ previously. Genetic clades to which reference samples belong are indicated. All accession numbers of all sequences used in these analyses can be found in Supplementary T 4 and 5.

To further explore genetic diversity among strains infecting patients who seroconverted during pregnancy, we pursued comparative analyses using fully sequenced amplified alleles. For this, we manually curated all polymorphisms observed, comparing only those strongly supported by sequencing. These analyses consider polymorphisms beyond the restriction sites, further resolving existing variability among infecting strains. Maximum likelihood analysis resolved two distinct groups of strains whose diversity is strongly supported. Both groups contain genotypes displaying a mix of archetypical alleles. However, Group II is characterized by a comparative sub representation of Type II alleles with respect to Group I. In addition, we compared our amplified alleles with both reference archetypal strains and publicly available sequences of regional non-archetypal strains. These analyses revealed that most of the amplified allele combinations belonging to group I segregated with reference strains and isolates belonging most prominently to genetic Clade A and C, while group II alleles segregated with reference strains and isolates belonging in their majority to Clades B and D (Figure 4).
Discussion
Serological studies are included as mandatory controls in pregnancy follow-up routines in Uruguay. Per mandate of the ministry of health, serological determination of T. gondii exposure should be carried out at least twice during pregnancy, once prior to the 18th week of gestation, and once into the second/third trimester. Despite routine follow-up and abundance of publicly available information, seroprevalence for toxoplasmosis in the Uruguayan population has not been updated since 1996. To tackle this, we followed women who monitored their pregnancies between 2019 and 2023 at the public hospital hosting the largest maternity ward in the country; the Centro Hospitalario Pereira Rossell (CHPR). Between 2019 and 2023, the country averaged 34 326 births per year, with CHPR contributing an average of 5751 births annually, accounting for an average of 16.8% of all births in the country during this period. In addition, CHPR is a reference center for complicated pregnancies follow up, hence, receives patients from various regions across the country. Given the high percentage of national births it hosts, the population studies herein are considered representative of national trends.
We found that, over this period 45.5% of patients had detectable IgG titres, indicating that they had been previously exposed to T. gondii. The overall seroprevalence among pregnant women in Europe is estimated at approximately 32.9%, which is notably lower than that observed in our sample population. However, in the Americas, the prevalence is around 45.2%, with notable differences within the continent: Brazil exhibits a higher prevalence (54.4%), while Mexico shows a much lower rate of 7% (Bigna et al., Reference Bigna, Tochie, Tounouga, Bekolo, Ymele, Youda, Sime and Nansseu2020).
Conti and colleagues had reported in 1998 an infection prevalence of 52.7% among 16 936 pregnant women studied between 1991 and 1996, as measured by the presence of antibodies using the latex agglutination test for toxoplasmosis. In contrast, we observed a prevalence of 45.5%, indicating a downward trend. Consistent with this observations, recent surveys highlight an observable trend towards a decrease in human population seroprevalence of toxoplasmosis worldwide (Milne et al., Reference Milne, Webster and Walker2023). For instance, comparative studies carried out in Serbia, highlighted significant reductions, with prevalence dropping from 86% in 1988 to approximately 32.5% in 2007 (Marković-Denić et al., Reference Marković-Denić, Stopić, Bobić, Nikolić, Djilas, Srzentić and Štajner2023). This decline is attributed to several factors, including improvements in prenatal care, diagnostic methods and public education about transmission routes. Enhanced awareness regarding safe practices (such as handling raw meat properly and managing cat litter) has played a significant role in reducing exposure to the parasite, particularly among vulnerable populations like pregnant women (Marković-Denić et al., Reference Marković-Denić, Stopić, Bobić, Nikolić, Djilas, Srzentić and Štajner2023).
In this study, all patients who were IgG positive/IgM positive with low avidity test results or IgG negative/IgM positive were considered to have seroconverted during pregnancy; we determined that seroconversion rate was of 0.58% in the studied population. This figure is also a drastic reduction from previous reports, which placed the rates of seroconversion in 1998 closer to 1 out of 100 immune-naive pregnant women (0.82%) (Díaz et al., Reference Díaz, Ismael, Alvaro, Graciela, Claudia, Juliana, Carmen, Acosta, López, Juan and González, A1998b). Nonetheless, this reduced rate of seroconversion is contrasted with a higher incidence rate of congenital toxoplasmosis, indicating that this rate of seroconversion is likely an underestimation. While pregnancy routine checkups are in place to detect seroconversion, poorly followed pregnancies and untimely detected seroconversions could account for a higher rate of congenital toxoplasmosis cases than that estimated by the observable seroconversion rates. Conspicuously, of the 13 cases of confirmed congenital transmission of T. gondii analysed, only a skim percentage of patients had received treatment during pregnancy. Most newborns exhibited severe clinical manifestations. These data indicate a significant gap in the medical system’s ability to meet the treatment directives given by the ministry of health, which may in turn be attributed to difficulties posed by the very health care system in terms of difficult access to the drugs, voluntary treatment interruptions by the patients and underdiagnosis, among others.
In this study, we pursued PCR detection of T. gondii’s DNA in 81 samples from 53 patients. We serendipitously observed that peripheral blood from women who had recently seroconverted proved an additional valuable sample for detection, and that samples resulting negative for B1 and Tox5-8, could be positively identified using either the BTUB or GRA6 genotyping oligos. This expanded the breath of positively identified samples in our study allowing us in turn to genotype additional samples. These results prompt further studies aiming at evaluating the value of including GRA6 in diagnostic PCRs.
Recent work has demonstrated that GRA7, a gene that is normally not included in the widely used typing panel, outperformed GRA6 both in its diagnostic and genotyping potential in cerebrospinal fluid from AIDS patients suffering from toxoplasmic encephalitis, presenting additional opportunities to optimize both diagnostic PCRs and genotyping algorithms (Harminarti et al., Reference Harminarti, Sari, Artama, Imran and Kurniawan2024). Among 32 sampled variants, we detected a minor fraction (5) of possible archetypal strains, with a preponderance of potential Type I strains. Many samples, (Tg42, Tg34, Tg9, Tg41, Tg45, Tg35), albeit non-clonal, clustered with the Type I strains (GT1, RH88). Tg40 closely related to Type II strain ME49 and Tg49 closely related to Type III strain VEG. Tg21 to Tg37 form a unique cluster. For the majority of samples we could not identify previously reported genotypes in ToxoDB (with only two samples partially matching genotypes #10 and #55). These findings highlight the ample genetic diversity of T. gondii strains circulating in the human population in the country. Prior efforts to characterize genetic variation of T. gondii’s strains in humans in Uruguay, by serotyping, also identified a preponderance of potential non-archetypal strains (Sousa et al., Reference Sousa, Puime, Costa and Dardé2017). However, serotyping has not been validated for use in the identification of non-canonical genetic variants and its value in this context remains to be proven. Our genotyping efforts by molecular methods herein further support these earlier results.
The expansion of epidemiological studies over many different geographical regions and host species has allowed the identification of hundreds of genotypes of Toxoplasma gondii. We now appreciate the intricate population structure of this organism and its connection to geographical origin. Though genetic variants, including non-clonal and non-archetypal strains can be identified, it is well established that the majority of strains infecting humans in Europe and North America are of the Type II and III background (Hosseini et al., Reference Hosseini, Amouei, Sharif, Sarvi, Galal, Javidnia, Pagheh, Gholami, Mizani and Daryani2019). South America is currently regarded as the probable origin of the T. gondii species, mapping its origin to Colombia (Cañón-Franco et al., Reference Cañón-Franco, López-Orozco, Gómez-Marín and Dubey2014). In addition, the population structure in South America has been extensively studied, particularly in Brazil, Colombia and Argentina (Ajzenberg et al., Reference Ajzenberg, Bañuls, Tibayrenc and Dardé2002; Ferreira et al., Reference Ferreira, Vidal, de Mattos, de Mattos, Qu, Su and Pereira-Chioccola2011; Pardini et al., Reference Pardini, Carral, Bernstein, Gos, Olejnik, Unzaga, Kaufer, Freuler, Durlach and Venturini2014, Reference Pardini, Bernstein, Carral, Kaufer, Dellarupe, Gos, Campero, Moré, Messina, Schneider, Freuler, Durlach, Unzaga and Venturini2019; de Lima Bessa et al., Reference de Lima Bessa, de Almeida Vitor and Dos Santos Martins-duarte2021; Galal et al., Reference Galal, Ariey, Gouilh, Dardé, Hamidović, Letourneur, Prugnolle and Mercier2022). Collectively, these studies have determined that strains infecting humans are most often non-clonal and non-archetypal.
Toxoplasma gondii genotypes previously identified in Argentina by Pardini and colleagues, who isolated six strains from cases of acute infection during pregnancy corresponding to genotypes #138, #132 and #14 in Argentina (Pardini et al., Reference Pardini, Bernstein, Carral, Kaufer, Dellarupe, Gos, Campero, Moré, Messina, Schneider, Freuler, Durlach, Unzaga and Venturini2019), did not match our identified genotypes. Genotype #10, which partially corresponds to one of the genotyped samples in our cohort, has been rarely reported infecting birds in Brazil (Brito et al., Reference Brito, de Lima Bessa, Bastilho, Dantas-Torres, de Andrade-neto, Bueno, Fujiwara and Magalhães2023). Likewise, genotype #55 has only been found in DNA coming from cats in Brazil (TgCatBr79 and TgCatBr80) and rarely reported (Su et al., Reference Su, Shwab, Zhou, Zhu and Dubey2010). This is consistent with our amplified alleles being placed among strains that belong to genetic Clade A, whereby strains from Brazilian origin are under-represented in comparison to samples segregating in Group II. Interestingly, a recent study analyzing 156 genomes of T. gondii strains of diverse geographical origins, established that CASTELLS, a Uruguayan T. gondii isolated from an aborted sheep, is the only strain showing an Amazonian exclusive ancestry (vs. other regional strains which seem to be hybrids derived from mixtures of regional and European-ancestry strains) (Galal et al., Reference Galal, Ariey, Gouilh, Dardé, Hamidović, Letourneur, Prugnolle and Mercier2022) supporting the notion that the T. gondii diversity found in the country might differ from that of the region.
Interestingly, we also detected three cases – which make up 10% of our sampled population – in which a mix of alleles was identified for the same markers within the same sample, suggesting the presence of at least two genetically distinct strains. However, sequencing artefacts due to low-quality DNA amplification cannot be formally excluded. Nonetheless, coinfection and mixed infections have been previously reported (Aspinall et al., Reference Aspinall, Guy, Roberts, Joynson, Hyde and Sims2003; Boughattas et al., Reference Boughattas, Ben-Abdallah, Siala, Souissi, Aoun and Bouratbine2010; Carneiro et al., Reference Carneiro, Andrade, Costa, Pinheiro, Vasconcelos-Santos, Ferreira, Su, Januário and Vitor2013). Additionally, sequential infections with distinct strains have been reported in the context of pregnancy, whereby previously immunized mothers with one strain passed onto their fetus a new strain upon reinfection during pregnancy (Elbez-Rubinstein et al., Reference Elbez-Rubinstein, Ajzenberg, Dardé, Cohen, Dumètre, Yera, Gondon, Janaud and Thulliez2009). Albeit rare, these cases should be taken into consideration, especially in the context of the massively diverse parasite populations in circulation among humans detected in many countries of South America.
While the virulence of the three clonal genotypes is well-understood in animal models, non-archetypal natural recombinant strains are not as well understood. Nonetheless, it is well established that the variant genetic makeup of T. gondii corresponds with amped disease severity in South America, particularly pertaining to ocular toxoplasmosis in immunocompetent individuals(de Lima Bessa et al., Reference de Lima Bessa, de Almeida Vitor and Dos Santos Martins-duarte2021; Sanchez and Besteiro, Reference Sanchez and Besteiro2021; Brito et al., Reference Brito, de Lima Bessa, Bastilho, Dantas-Torres, de Andrade-neto, Bueno, Fujiwara and Magalhães2023). Conspicuously, we observed that the most prevalent manifestation in congenitally infected newborns is ocular (chorioretinitis). Further studies would be required, both to establish genotype–phenotype correlations and to further understand whether ample genetic variability impacts the detection efficiency of currently used serological survey tools. Further delving into these questions would require massive strain isolation efforts, both to determine serological response to individual strains, their underlying phenotypic variations, as well as their behavior in pregnant animal models.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182025100334.
Data availability
All sequencing data generated herein is publicly available through the GeneBank public repository, or upon request to the corresponding author via email.
Acknowledgements
The Apicomplexan Biology Lab is funded by a G4 installment grant from the Pasteur Network. The Institut Pasteur de Montevideo is funded by FOCEM – Fondo para la Convergencia Estructural del Mercosur (Grant COF 03/11). P.F.T. and M.E.F. are researchers of PEDECIBA and the national system of researchers (SNI-ANII). We are grateful to all the participating patients.
Author’s contribution
M.E.F., J.G., A.V.-D., L.T.-H. and P.F.-T. conceived and designed the study. M.G., M.R.-R., G.D., V.G., G.G. and F.N. conducted data gathering. P.F. performed bioinformatics analyses. M.E.F., J.G., A.V.-D., L.T.-H. and P.F.-T. wrote the article. M.E.F., J.G. and A.V.-D. revised the article.
Financial support
This research received no specific grant from any funding agency, commercial or not-for-profit sectors.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
All data acquisition involving patients was conducted in accordance with a protocol approved by the research ethics committees of both Centro Hospitalario Pereira Rossell and Hospital de Clínicas. Patients were fully informed about the study’s objectives and scope, and they provided written informed consent prior to participation. All patient data were handled confidentially to ensure their anonymity.