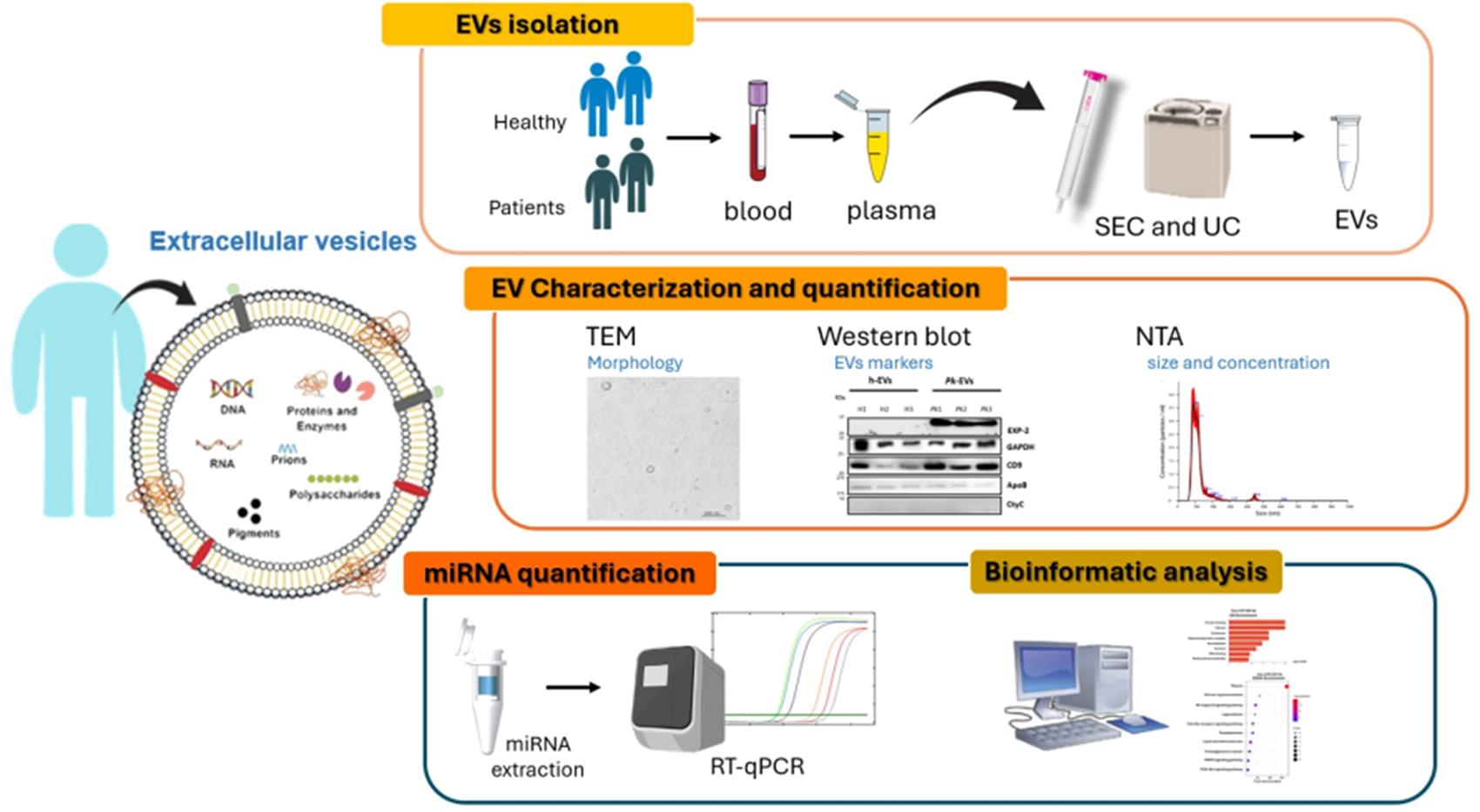
Introduction
Extracellular vesicles (EVs) are small lipid-bound vesicles that cells release either directly from the plasma membrane or through endosomal mechanisms, which merge with the membrane before entering the extracellular space (Théry et al., Reference Théry, Witwer, Aikawa, Alcaraz, Anderson, Andriantsitohaina, Antoniou, Arab, Archer, Atkin-Smith, Ayre, Bach, Bachurski, Baharvand, Balaj, Baldacchino, Bauer, Baxter, Bebawy, Beckham and Zuba-Surma2018). In the context of malaria infection, increased EV levels in the bloodstream have been observed to correlate with disease severity and clinical symptoms. These EVs, sourced from a variety of cells including infected erythrocytes, platelets, leucocytes and endothelial cells, are typically major contributors to malaria-derived EVs, with infected erythrocytes being a significant source (Combes et al., Reference Combes, Taylor, Juhan-Vague, Mège, Mwenechanya, Tembo, Grau and Molyneux2004; Pankoui Mfonkeu et al., Reference Pankoui Mfonkeu, Gouado, Fotso Kuaté, Zambou, Amvam Zollo, Grau and Combes2010; Sahu et al., Reference Sahu, Sahoo, Kar, Mohapatra and Ranjit2013; Abels and Breakefield, Reference Abels and Breakefield2016). Platelets and reticulocytes play prominent roles in Plasmodium vivax infections as the main contributors to EV production (Toda et al., Reference Toda, Diaz-Varela, Segui-Barber, Roobsoong, Baro, Garcia-Silva, Galiano, Gualdrón-López, Almeida, Brito, de Melo, Aparici-Herraiz, Castro-Cavadía, Monteiro, Borràs, Sabidó, Almeida, Chojnacki, Martinez-Picado, Calvo and Del Portillo2020). EVs play a significant role in the biological and pathogenic processes of malaria. EVs, originating from both host and parasite cells, transfer cell-specific biomolecules to immune cells or parasitized cells, influencing the inhibition or promotion of cytokine secretion, thereby inducing inflammation, promoting parasite survival or facilitating gametocyte formation (Mantel et al., Reference Mantel, Hoang, Goldowitz, Potashnikova, Hamza, Vorobjev, Ghiran, Toner, Irimia, Ivanov, Barteneva and Marti2013; Regev-Rudzki et al., Reference Regev-Rudzki, Wilson, Carvalho, Sisquella, Coleman, Rug, Bursac, Angrisano, Gee, Hill, Baum and Cowman2013; Sisquella et al., Reference Sisquella, Ofir-Birin, Pimentel, Cheng, Abou Karam, Sampaio, Penington, Connolly, Giladi, Scicluna, Sharples, Waltmann, Avni, Schwartz, Schofield, Porat, Hansen, Papenfuss, Eriksson, Gerlic, Hill, Bowie and Regev-Rudzki2017).
MicroRNAs (miRNAs) are small non-coding RNA molecules that are secreted from cells into the bloodstream where they can either be bound to proteins or packaged into EVs. miRNAs participate in post-transcriptional gene regulation predominantly by binding to the 3ʹ untranslated region (3ʹUTR) of the target messenger RNA (mRNA), but they can also bindto the 5ʹUTR or even within the coding region of the mRNA, leading to protein translation inhibition or mRNA degradation (Schuster and Hsieh, Reference Schuster and Hsieh2019). P. falciparum is unable to produce miRNAs and lacks genes encoding Argonaute and Dicer (Xue et al., Reference Xue, Zhang, Huang, Feng and Pan2008). In this context, human miRNAs are involved in modulating gene expression in both Plasmodium parasites and human hosts, affecting the host immune response, inflammation, tissue damage, survival and replication of the parasite throughout the life cycle (Lodde et al., Reference Lodde, Floris, Muroni, Cucca and Idda2022). In mild to severe malaria, the dysregulation of gene expression involved in immune regulation is marked by miRNAs such as miRNA-16, miRNA-155, miRNA-150, miRNA-223 and miRNA-451, indicating the potential of these miRNAs as biological markers for malaria infection (Rangel et al., Reference Rangel, Teerawattanapong, Chamnanchanunt, Umemura, Pinyachat and Wanram2019). Children with severe malaria showed increased expression of plasma miR-3158–3p, linked to seizures, and hsa-miR-4497, which is associated with parasite biomass and HRP2 levels. Both miRNAs are associated with acute respiratory distress syndrome (Gupta et al., Reference Gupta, Rubio, Sitoe, Varo, Cisteró, Madrid, Cuamba, Jimenez, Martiáñez-Vendrell, Barrios, Pantano, Brimacombe, Bustamante, Bassat and Mayor2021). In the serum of P. vivax-infected patients, levels of miR-451 and miR-16 are decreased, while the level of miR-223 is not changed (Chamnanchanunt et al., Reference Chamnanchanunt, Kuroki, Desakorn, Enomoto, Thanachartwet, Sahassananda, Sattabongkot, Jenwithisuk, Fucharoen, Svasti and Umemura2015). The reduction was likely due to red blood cell degradation and miRNA clearance by the spleen. In P. vivax patients, other studies have demonstrated notably elevated plasma levels of miR-191, miR-223, miR-145 and miR-155 (Hadighi et al., Reference Hadighi, Heidari, Fallah, Keshavarz, Tavakoli, Mansouri and Sadrkhanloo2022). An increase in miR-4454 and miR-7975 was also observed in patients with severe thrombocytopenia (Santos et al., Reference Santos, Coimbra, Sousa, Guimarães, Gomes, Amaral, Pereira, Fontes, Hawwari, Franklin and Carvalho2021). Human miRNA has been found to be transferred to the intracellular parasite and regulate gene expression, with involvement in pathogenicity and host defence mechanisms (LaMonte et al., Reference LaMonte, Philip, Reardon, Lacsina, Majoros, Chapman, Thornburg, Telen, Ohler, Nicchitta, Haystead and Chi2012; Dandewad et al., Reference Dandewad, Vindu, Joseph and Seshadri2019). The overexpression of miR-451, miR-223 and let-7i in P. falciparum within sickle cell erythrocytes can regulate parasite genes, leading to a reduction in growth (LaMonte et al., Reference LaMonte, Philip, Reardon, Lacsina, Majoros, Chapman, Thornburg, Telen, Ohler, Nicchitta, Haystead and Chi2012). Plasmodium apicortin targeted by miR-150-3p and miR-197-5p also impaired the growth and invasion of the malaria parasite (Chakrabarti et al., Reference Chakrabarti, Garg, Rajagopal, Pati and Singh2020). miRNAs found in both plasma and EVs are important diagnostic markers due to their stability in the bloodstream (Mantel et al., Reference Mantel, Hjelmqvist, Walch, Kharoubi-Hess, Nilsson, Ravel, Ribeiro, Grüring, Ma, Padmanabhan, Trachtenberg, Ankarklev, Brancucci, Huttenhower, Duraisingh, Ghiran, Kuo, Filgueira, Martinelli and Marti2016). Notably, EV-derived miRNAs directly reflect the pathological characteristics of their cellular origin (Xu et al., Reference Xu, Di, Fan, Wu, Gu, Sun, Khan, Li and Li2022). EVs derived from P. falciparum cultures with infected red blood cells (iRBCs) showed high expression levels of specific small RNAs, while miR-451a, miR-486-5p, miR-92a-3p, miR-103a-3p, let-7b-5p, miR-181a and miR-106b-5p were particularly prominent in the small RNA profile of these EVs (Mantel et al., Reference Mantel, Hjelmqvist, Walch, Kharoubi-Hess, Nilsson, Ravel, Ribeiro, Grüring, Ma, Padmanabhan, Trachtenberg, Ankarklev, Brancucci, Huttenhower, Duraisingh, Ghiran, Kuo, Filgueira, Martinelli and Marti2016; Wang et al., Reference Wang, Xi, Hao, Deng, Liu, Wei, Gao, Zhang and Wang2017; Babatunde et al., Reference Babatunde, Mbagwu, Hernández-Castañeda, Adapa, Walch, Filgueira, Falquet, Jiang, Ghiran and Mantel2018). P. falciparum EVs carrying miRNA-451 downregulate target genes (CAV-1 and ATF-2) in endothelial cells, affecting vascular function (Mantel et al., Reference Mantel, Hjelmqvist, Walch, Kharoubi-Hess, Nilsson, Ravel, Ribeiro, Grüring, Ma, Padmanabhan, Trachtenberg, Ankarklev, Brancucci, Huttenhower, Duraisingh, Ghiran, Kuo, Filgueira, Martinelli and Marti2016). Wang et al. demonstrated host-miRNA regulation, specifically showing that miR-451 and miR-140 from large EVs downregulate VAR genes encoding P. falciparum erythrocyte membrane protein (PfEMP1) (Wang et al., Reference Wang, Xi, Hao, Deng, Liu, Wei, Gao, Zhang and Wang2017). The expression of miRNA varies in different physiological conditions associated with the development of pathological diseases and organ damage. However, the expression of EV-derived miRNA from patient plasma in Plasmodium knowlesi malaria is not well studied.
P. knowlesi primarily resides in monkeys but can infect humans as a zoonotic disease transmitted through mosquito vectors. Infections caused by P. knowlesi range from asymptomatic cases to severe malaria, presenting with anaemia, acute respiratory distress, renal failure and thrombocytopenia, and can potentially be fatal. Its 24-h intraerythrocytic cycle accelerates parasitaemia and disease progression (Anstey et al., Reference Anstey, Grigg, Rajahram, Cooper, William, Kho and Barber2021), leading to severe outcomes or death if not promptly diagnosed and treated (Cox-Singh et al., Reference Cox-Singh, Hiu, Lucas, Divis, Zulkarnaen, Chandran, Wong, Adem, Zaki, Singh and Krishna2010; Rajahram et al., Reference Rajahram, Barber, William, Menon, Anstey and Yeo2012; Chantaramongkol and Buathong, Reference Chantaramongkol and Buathong2016). In Thailand, P. knowlesi cases, once rare, have been steadily increasing. According to the Thai National Malaria Control Program, reported cases rose from 6 out of 11 595 in 2017 to 259 out of 16 680 in 2023 (ThaiMOPH, 2024).
The identification of EVs and miRNAs can help elucidate the molecular processes that contribute to these complications, potentially leading to better diagnostic tools and targeted interventions.
Materials and methods
Blood sample collection and processing
Peripheral blood samples were obtained from individuals infected with the P. knowlesi parasite and from healthy donors. The blood specimens were obtained using leftover EDTA-containing tubes from the hospital in Southern Thailand. This study was approved by the Ethical Committee of the Faculty of Medical Technology, Prince of Songkla University (EC66-03). The characteristics of 13 P. knowlesi patients and 10 healthy donors are shown in Table 1. All P. knowlesi patients were confirmed by nested polymerase chain reaction (PCR) (Putaporntip et al., Reference Putaporntip, Buppan and Jongwutiwes2011). The blood samples were centrifuged at 1500 g for 10 min. The resulting plasma was then subjected to another round of centrifugation at 2500 g for 15 min at 4°C, twice, to obtain platelet-free plasma. A 0.5–1-mL aliquot of the plasma supernatant was preserved at −80°C until further isolation of EVs.
Characteristics of the study subjects
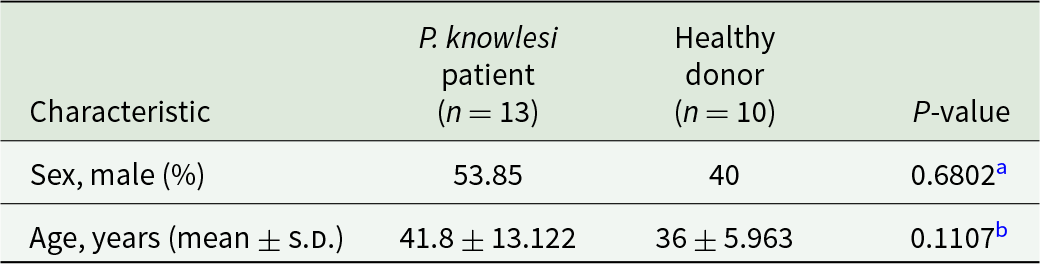
a Fisher’s exact test.
b Mann–Whitney U test.
EVs enrichment using size exclusion chromatography and ultracentrifugation
A commercial size exclusion chromatography (SEC) qEV1/35 nm column from iZON Sciences (Christchurch, New Zealand) was used to isolate EVs from the plasma samples following the manufacturer’s instructions (Fig 1A). The column was equipped with an automatic fraction collector. Before sample separation, the column was pre-equilibrated with phosphate-buffered saline (PBS), which was previously filtered in a sterile manner through a 0.22-μm filter. For each isolation, 700 µL of plasma was loaded onto the top of the qEV column. Subsequently, 4 fractions (700 μL per fraction) were immediately collected into separate 1.5-mL microcentrifuge tubes. To further concentrate the EVs, the pooled fractions were subjected to ultracentrifugation (UC) at 120 000 g for 90 min. After centrifugation, the EV pellet was resuspended in PBS, resulting in a final volume of 100 μL.
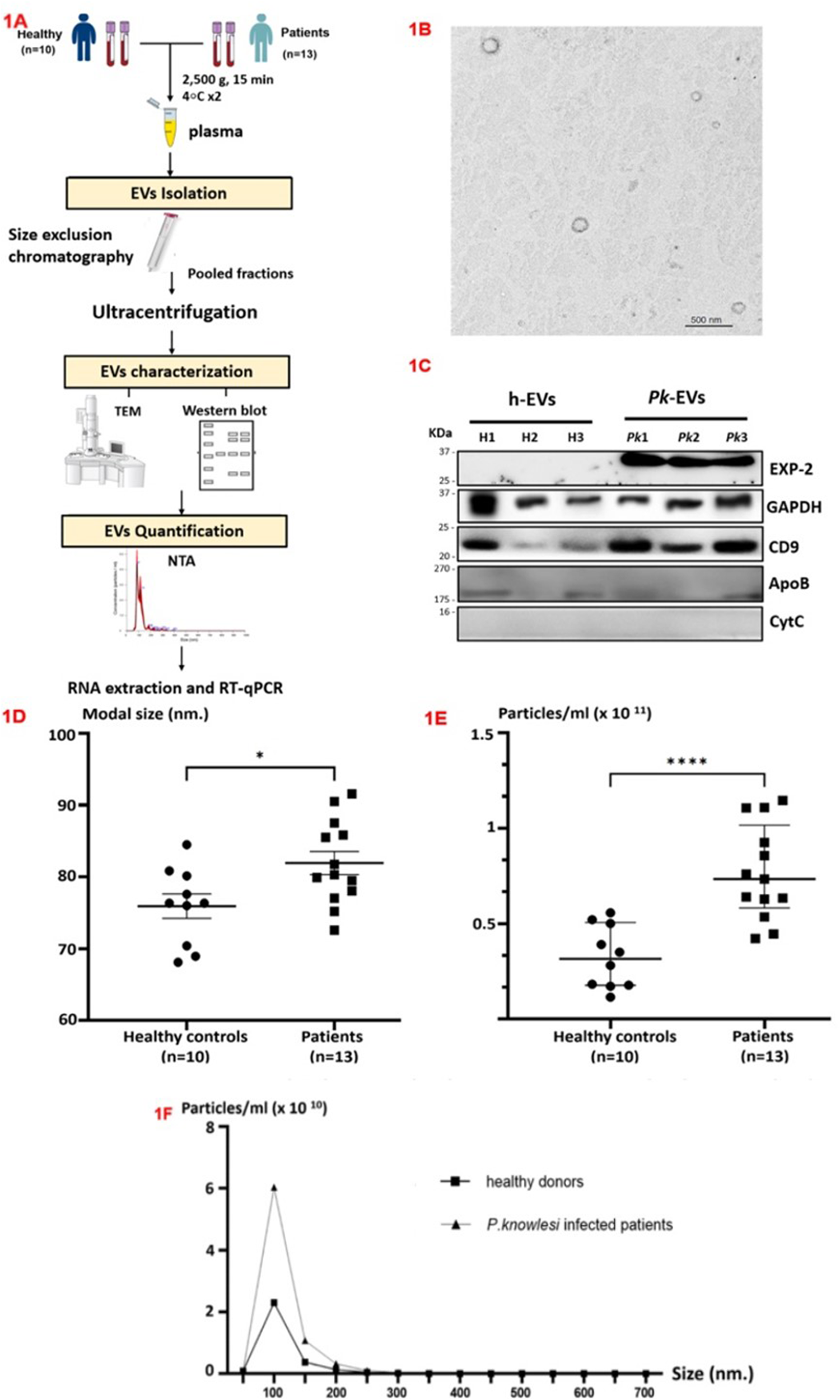
Isolation and characterization of plasma EVs. (A) Schematic overview of experimental methods. (B) Transmission electron microscopy. (C) Western blot detection of EXP-2, GAPDH, CD9, ApoB and Cytochrome C1 (CYC1). (D) Mode size and (E) concentration between patients and uninfected individuals, compared using the unpaired 2-tailed t-test. Data are presented as mean values with SEM. (F) Size distribution of extracellular particles (nm) versus concentration (particles/mL) (*P < 0.05, ****P < 0.0001).
Transmission electron microscopy
The EVs were subjected to a fixing process using a 2.5% glutaraldehyde solution. Subsequently, these fixed EVs were carefully placed onto a Formvar carbon grid. To enhance visualization, the grids were treated with a 2% uranyl acetate stain and then allowed to dry at ambient temperature. Finally, the dried grids were observed and analysed using an advanced Talos™ F200i transmission electron microscope manufactured by Thermo Scientific (MA, USA).
Western blot analysis
For immunoblotting analysis, 10 μg of isolated EVs were mixed with 5× loading buffer and then heated at 100°C for 5 min. Subsequently, the EV proteins were separated on a 12% polyacrylamide gel. Proteins were then transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were blocked using 5% non-fat milk in tris-buffered saline (TBS) for 1 h. After blocking, the membrane was washed thrice with TBS-T (TBS with 0.1% Tween 20) for 10 min. The primary antibodies, including anti-rabbit CD9 (Abcam ab223056) (Abcam, Cambridge, UK) at a dilution of 1:2000 and anti-mouse GAPDH (Abcam ab8245) at a dilution of 1:20 000, were diluted in 1% non-fat milk in TBS-T and used as positive markers for EVs. The anti-cytochrome C1 (BioLegend, CA, USA) and anti-rabbit APOB (Abcam ab139401) were diluted at 1:1000 as non-EV markers and indicators of lipoprotein contamination, respectively. The anti-mouse EXP-2 (The European Malaria Reagent Repository, Cat# 7.7) was diluted at 1:600 to detect against the parasitophorous vacuole membrane (PVM). After incubating with the primary antibodies, the PVDF membrane was washed thrice with TBS-T buffer for 10 min. For secondary detection, secondary antibodies were applied at a dilution of 1:5000 and incubated at room temperature for 1 h. An ultra-sensitive enhanced chemiluminescent (ECL) horseradish peroxidase substrate was employed to visualize the specific proteins.
Quantification of EVs
Nanoparticle tracking analysis (NTA) was employed to determine the size and concentration of the EVs. The NTAs were carried out using a NanoSight NS300 NTA 3.4 system equipped with a 488-nm laser. To conduct the analyses, aliquots of the isolated EVs were diluted 500–1000-fold with 0.22-μm filtered deionized water, ensuring that the number of particles per frame fell within the range of 20–100. The camera level for each sample was set at 13, and the detection threshold was set at 5. The NTA software analysed 5 videos for 60 s each in duplicate, while the temperature of the laser unit was maintained at 25°C. The obtained NTA data were compared for size distribution, modal size, and concentration of particles between the P. knowlesi infection and healthy donor groups.
RNA extraction and reverse transcriptase quantitative PCR
The instructions provided by the manufacturer of an RNA extraction kit were followed to isolate total RNA. Initially, 350 µL of lysis buffer was added to the EV sample. The mixture was briefly vortexed and incubated for 5 min at room temperature. Next, 100% ethanol was added to the sample, and the resulting mixture was transferred to a spin column and centrifuged at 13 000 rpm for 1 min. After washing, the isolated RNA was eluted from the spin column using 50 µL of elution buffer. The quantification of RNA was performed using a NanoDrop spectrophotometer. The complementary DNA (cDNA) was generated using a QuantiMir reverse transcription kit from System Biosciences. Subsequently, amplification was performed with specific miRNA primers, as detailed in Table 2. The amplification was carried out in a total volume of 20 µL PCR mixture which contained 10 µL of 2X qPCRBIO SyGreen Mix, 0.4 µM of each primer, and 1 µL of cDNA. The amplification thermal condition consisted of 95°C for 1 min followed by 40 cycles of 95°C for 15 s and 60°C for 1 min. Each sample was analysed in triplicate, with the negative control as nuclease-free water. Technical replicates with a standard deviation exceeding 0.5 were excluded and retested.
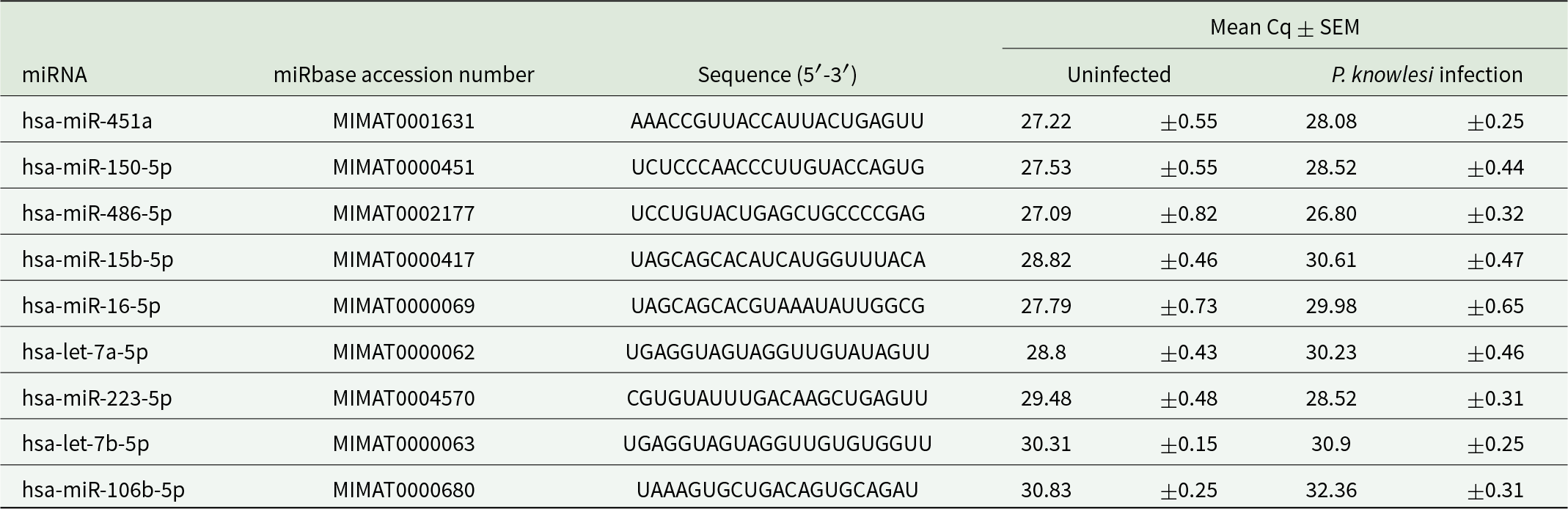
MiRNA sequences and Cq values
miRNA expression analysis
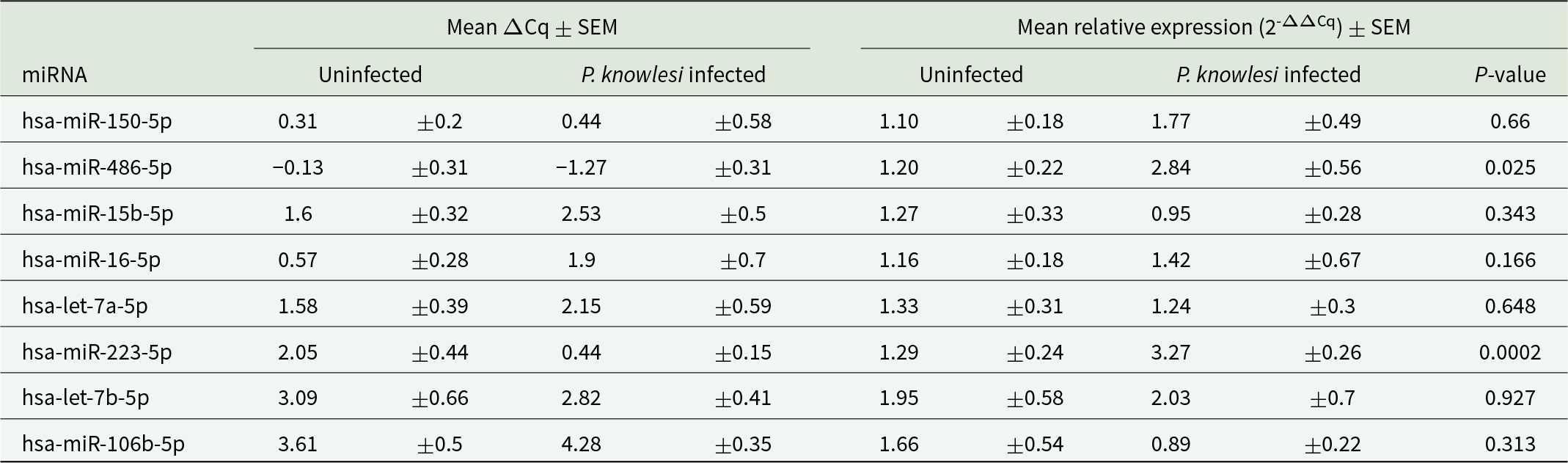
Quantification cycle (Cq) values were used to compare the relative quantities of miRNAs between the malarial and healthy samples. The Delta Cq (ΔCq) of the target miRNA was calculated by averaging triplicate values and subtracting them from the average Cq values of miR-451a in the corresponding samples. The hsa-mir451a-5p was used as a normalization control. ΔΔCq was then normalized by comparing the ΔCq of each sample to the mean ΔCq of the healthy control group. The expression of EV miRNA was assessed using the comparative Cq method (−2ΔΔCq).
Target gene prediction and enrichment analysis
The target genes of upregulated miRNAs were predicted using TargetScan Release 8.0 (https://www.targetscan.org/vert_80/), miRDB (https://mirdb.org/), miRDIP (https://ophid.utoronto.ca/mirDIP/), and miRTarBase (https://mirtarbase.cuhk.edu.cn/). The human genes related to malaria infection were retrieved from the Kyoto Encyclopedia of Genes and Genomes (KEGG) malaria pathway. The candidate targets that overlapped between the miRNA target database and the malaria pathway in the KEGG database were selected using a Venn diagram. The Gene Ontology (GO) and KEGG enrichment analyses of individual miRNAs were conducted using the DIANA-miRPath v.4.0 (https://diana-lab.e-ce.uth.gr/app/miRPathv4) and DAVID gene annotation tool (https://david.ncifcrf.gov/). The prediction in DIANA-miRPath was based on experimentally supported targets from TarBase 8.0. The pathway union option was used, with false discovery rate (FDR) correction. The bar and bubble plots were generated using SRplot (https://www.bioinformatics.com.cn/srplot). The P-values were calculated by Fisher’s exact test and the FDR was obtained by the Benjamini–Hochberg method. FDR values less than 0.05 were considered statistically significant for enrichment analysis. The DAVID online tool was conducted for upregulated miRNA, and P-value <0.05 was set as a significant threshold.
The transcriptome data of P. knowlesi were retrieved from the PlasmoDB 8.0 database to determine the target binding of human miRNAs with P. knowlesi transcripts. Subsequently, an analysis of the selected miRNA sequences was conducted against the P. knowlesi strain H transcriptome using the psRNATarget tool (http://plantgrn.noble.org/psRNATarget/) with default recommended value of 5 (Dai et al., Reference Dai, Zhuang and Zhao2018).
Statistical analysis
All data were presented as the mean and standard error of the mean (SEM), with normality assessed using the D’Agostino–Pearson test. Student’s t-test was used to assess differences in mode size and concentration of EVs between groups. The non-parametric Mann–Whitney U test was used to compare the expression levels of miRNAs between the plasma-derived EVs of healthy individuals and patients. This statistical analysis was conducted using GraphPad Prism software (version 9.0; GraphPad Software, San Diego, CA, USA). Receiver operating characteristic (ROC) curve analysis was performed to evaluate the diagnostic potential of miRNA expression levels for P. knowlesi infection. ROC curve analysis was carried out using delta Ct values (Canatan et al., Reference Canatan, Yılmaz, Sonmez, Cim, Baykara, Savas, Coskun, Goksu and Aktekin2022). The accuracy of the test was assessed by measuring the area under the ROC curve (AUC), which identified optimal sensitivity and specificity levels for distinguishing normal individuals from patients.
Results
Characterization of plasma-derived EVs
Transmission electron microscopy (TEM) and Western blot analysis were used for EV characterization to evaluate the presence of the EVs isolated through SEC- and UC-based methods (Fig 1A). TEM revealed spherical phospholipid bilayer structures with an diameter of approximately 100 nm (Fig 1B). As shown in Fig 1C, EVs demonstrated enrichment of exosomal protein markers including CD9 and glyceraldehyde-3-phosphate dehydrogenase (GAPDH), while Cytochrome C1 (CYC1) (non-EV marker) was not detected. The presence of parasite proteins EXP-2 was determined in EVs from P. knowlesi malaria patients. A signal for ApoB lipoprotein found in chylomicrons very-low-density lipoprotein, intermediate-density lipoprotein and low-density lipoprotein (LDL) particles was observed.
Quantification of EVs’ size and concentration
The size and concentration distribution for each biological replicate were determined through NanoSight analysis. The particles displayed diameters ranging from under 50 to 600 nm, with most falling within the 50–100 nm size category. The 250–600 nm size range accounted for less than 1% of the total, indicating the presence of aggregated vesicles. The average modal size of isolated EVs from P. knowlesi-infected individuals (Pk-EVs) was 81.9 ± 1.75 nm, which was significantly larger than those from healthy donors (h-EVs) at 75.9 ± 1.25 nm (P-value = 0.019) (Fig 1D). The size was consistent with the results of TEM. Larger EVs observed by the nanoparticle tracking system were possibly the result of aggregate formation. A statistically significant 2.4-fold increase in particle concentration was recorded for P. knowlesi-infected patients, measuring 7.64 × 1010 particles/mL (range: 4.21 × 1010–1.15 × 1011 particles/mL) compared to healthy donors, with concentration 3.24 × 1010 particles/mL (range: 1.15 × 1010–5.56 × 1010 particles/mL), P-value < 0.0001 (Fig 1E). The size versus concentration of the 2 groups is illustrated in Fig 1F. The mean concentration of extracellular particles within the 1–50 nm size range was higher in healthy donors compared to those with P. knowlesi infection (8.32 × 108 vs. 7.2 × 108). In the peak graph showing the 50–100 nm size range, the P. knowlesi infection group exhibited a significantly higher concentration at 6.03 × 1010 particles/mL, in contrast to 2.35 × 1010 particles/mL in healthy donors.
Relative expression of miRNAs
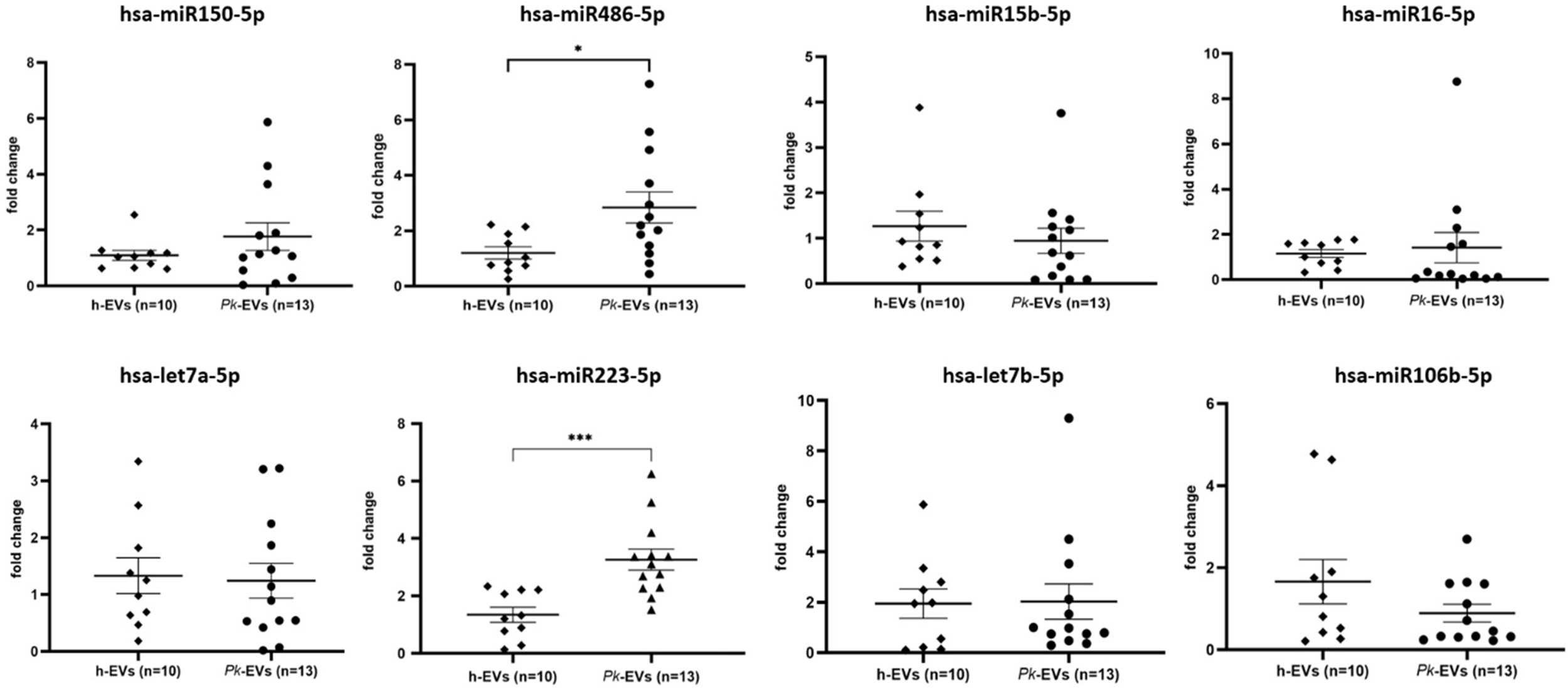
The SYBR Green quantitative PCR (qPCR)-based detection methods were used to evaluate the expression of 8 specific human miRNAs. The average Cq, ΔCq and 2-ΔΔCq values for all miRNAs were calculated (Tables 2 and 3), with hsa-miR-451a-5p serving as the endogenous control for data normalization. The relative expression of EV-miRNAs in malaria compared to healthy donors is shown in Fig 2. The reverse transcriptase qPCR (RT-qPCR) results revealed a significant increase in levels of hsa-miR-223-5p (P-value = 0.0002) and hsa-miR-486-5p (P-value = 0.008) in Pk-EVs group. The other 6 miRNAs showed no significant differences in expression levels between P. knowlesi infection and healthy individual groups.
Dot plots of miRNA relative expression in EVs derived from healthy controls and P. knowlesi-infected individuals. Graphs display mean levels with SEM. Differences between the 2 groups were assessed by using the non‐parametric Mann–Whitney U test (*P < 0.05, ***P < 0.001).
Evaluation of miRNA expression between healthy controls and patients
miRNA target prediction, GO and KEGG enrichment analysis
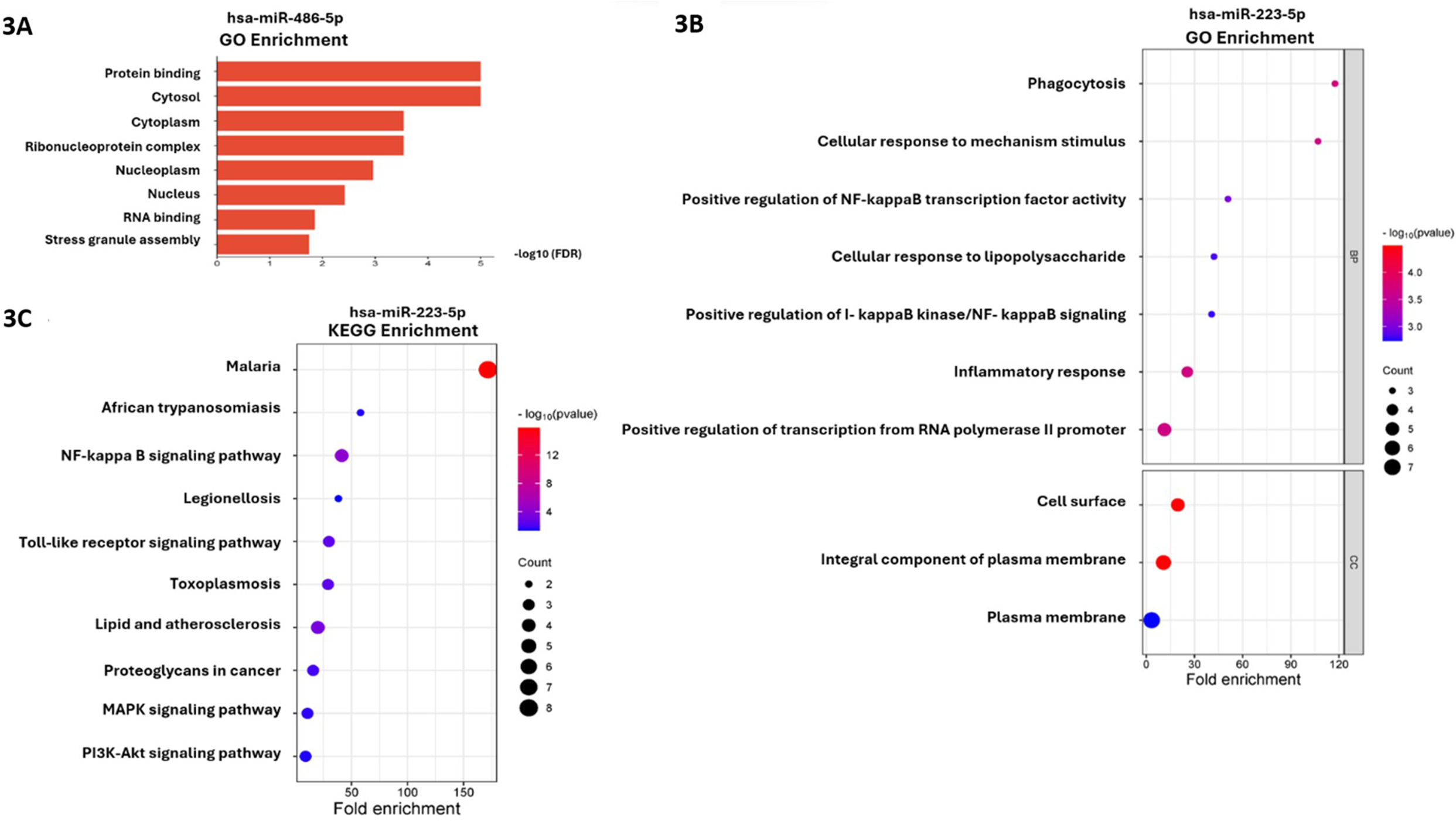
Prediction of the human-host gene targets for hsa-miRNA-223-5p and hsa-miRNA-486-5p utilized information from 4 databases. Results showed that 8 and 1 computationally predicted targets were identified for hsa-miR-223-5p and hsa-miR-486-5p, respectively (Table 4). Intriguingly, CD40 was a shared target gene for both miRNAs. A pathway analysis of individual miRNAs was conducted using DIANA miRPath. Notably, no enriched pathways associated with hsa-miRNA-223-5p were observed in this analysis. By contrast, hsa-miRNA-486-5p showed enrichment in GO terms related to protein binding for molecular function; cytosol, cytoplasm and ribonucleoprotein complex for cellular component (CC) (Fig 3A), demonstrating significant enrichment in the KEGG pathways of the EGFR (epidermal growth factor receptor) tyrosine kinase inhibitor resistance pathway. DAVID was employed to determine overrepresented GO terms and KEGG pathways associated with hsa-miRNA-223-5p. This tool conducted functional and pathway enrichment analysis based on the predicted target genes of hsa-miRNA-223-5p. The top 10 ranked GO terms and KEGG pathways are displayed in Fig 3B and 3C, respectively. For GO BP, the predicted target genes were significantly enriched in phagocytosis, cellular response to mechanism stimulus, and positive regulation of NF-κB transcription factor activity, while for GO CC, the enriched GO terms were cell surface, an integral component of plasma membrane and plasma membrane. In the KEGG pathway analysis, several predicted target genes were enriched in malaria, African trypanosomiasis and NF-κB signaling pathway.
Gene Ontology (GO) and KEGG enrichment analysis of hsa-miR-223-5p and hsa-miR-486-5p. (A) Enriched GO terms of hsa-miR-486-5p performed using DIANA-miRPath. (B) Top 10 enriched GO terms associated with the predicted target genes of hsa-miR-223-5p and (C) top 10 enriched KEGG pathway associated with the predicted target genes of hsa-miR-223-5p. BP, biological process; CC, cellular component; DAVID, Database for Annotation, Visualization, and Integrated Discovery.
Potential human target genes of hsa-miR-223-5p- and hsa-miR-486-5p-related malaria pathway
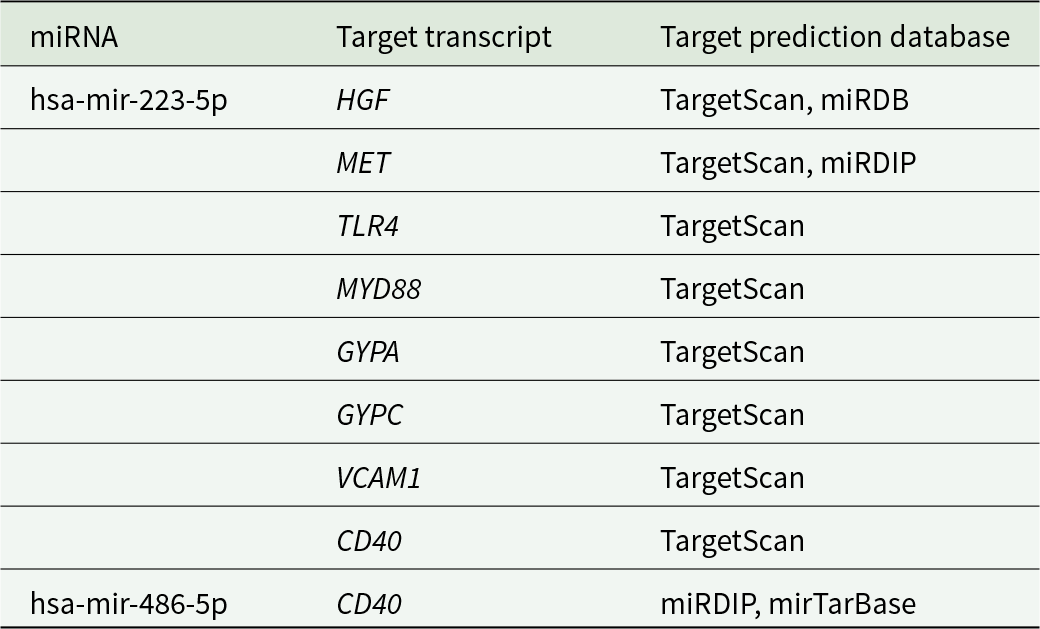
GYPC: glycophorin C, HGF: hepatocyte growth factor, CD40: cluster of differentiation 40, TLR4: toll-like receptor 4, GYPA: glycophorin A, MYD88: myeloid differentiation primary response protein 88, MET: MET Proto-Oncogene, Receptor Tyrosine Kinase, VCAM1: vascular cell adhesion molecule 1.
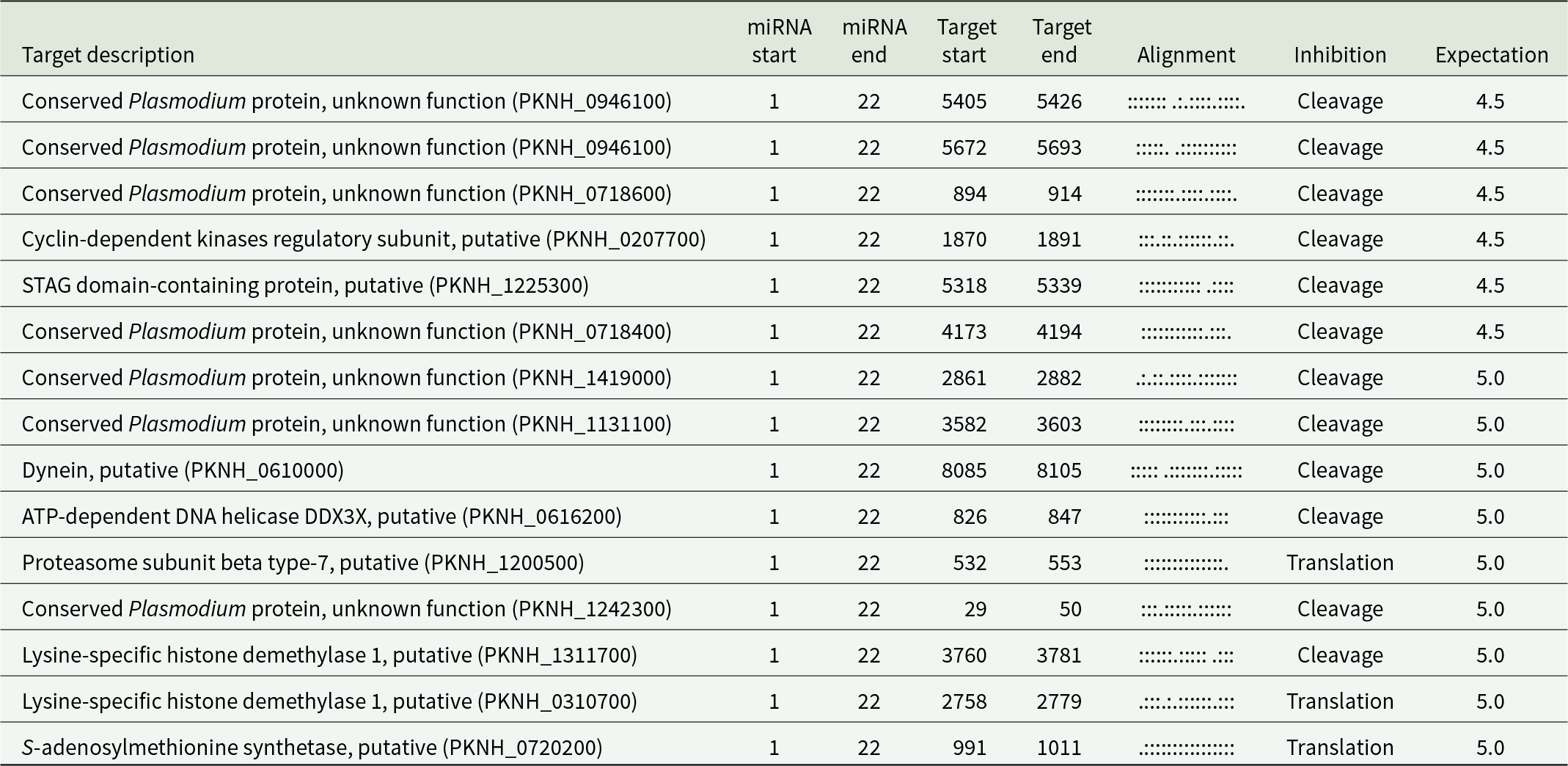
To explore potential interactions between miRNAs and P. knowlesi mRNAs and identify the potential gene targets of miRNAs in P. knowlesi mRNAs, the computational prediction software for miRNA–mRNA interactions, psRNATarget, was used. A total of 35 and 15 target candidates, with expectation values higher than 4.5, were identified as targets for hsa-miRNA-223-5p and hsa-miRNA-486-5p, respectively, as presented in Tables 5 and 6. Both miRNAs were shown to downregulate various genes in P. knowlesi by cleaving mRNA or translation inhibition.
Predicted P. knowlesi mRNA targets by miRNA-223-5p

Predicted P. knowlesi mRNA targets by miRNA-486-5p
Potential of EVs-derived miRNAs as diagnostic markers
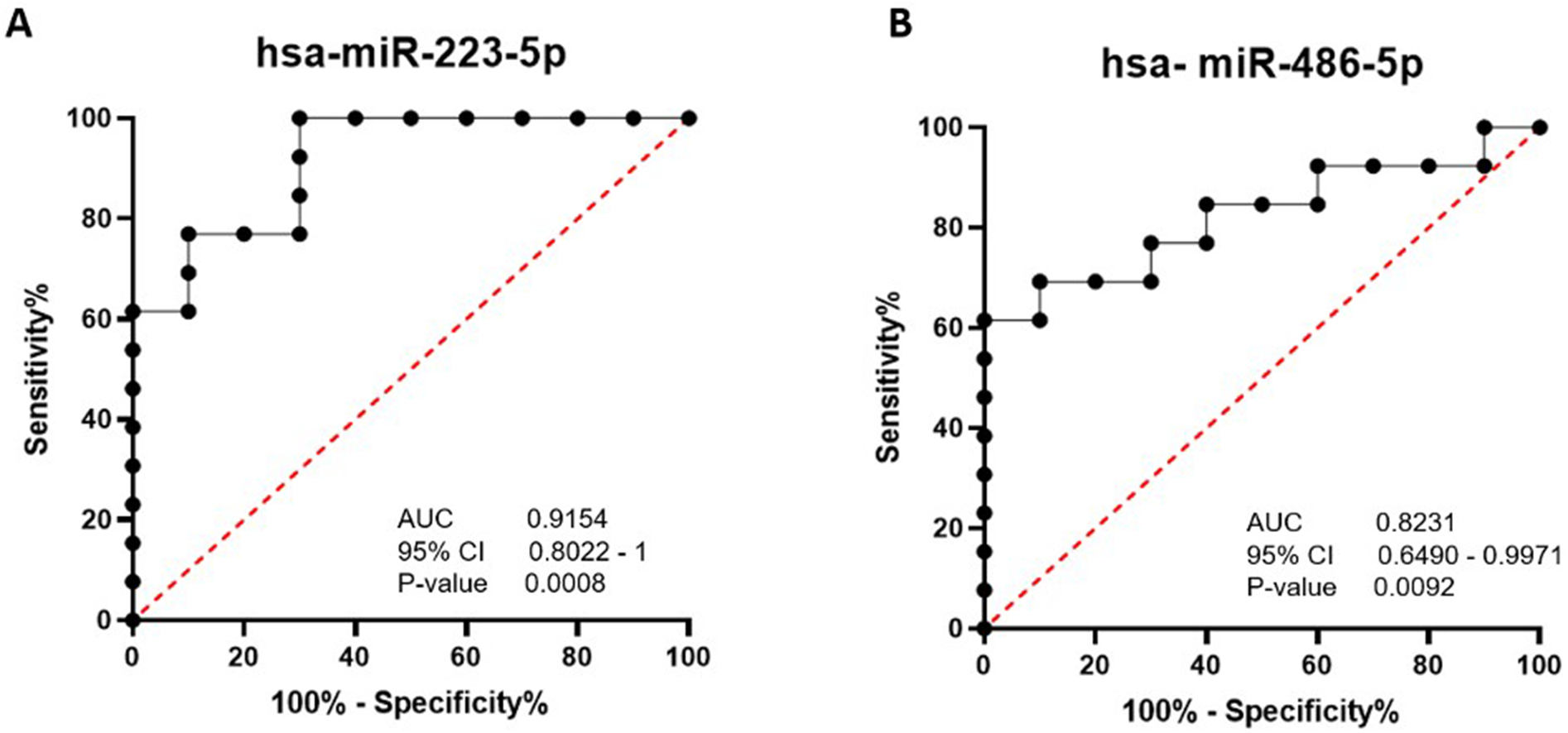
The AUC analysis was performed to investigate the possibility of miRNA-223-5p and miRNA-486-5p as potential diagnostic biomarkers. ROC curve illustrates the plot between true-positive rate (sensitivity) and false-positive rate (100-specificity). The optimal cut-off was identified as the maximum likelihood ratio, computed as sensitivity divided by (100 – specificity). The AUC was used to distinguish between the disease and healthy groups, where a value of 1 represents optimal discrimination. As shown in Fig 4, the AUC for miRNA-223-5p was 0.9154 (P-value = 0.0008). At the relative expression level of miRNA-223-5p at the optimal cut-off value of 7.692, the sensitivity was 76.92% (95% CI: 49.74–91.82%), and the specificity was 90% (95% CI: 59.58–99.49%). For hsa-miRNA-486-5p, the AUC was determined to be 0.8231 (P-value = 0.0092), with sensitivity and specificity 61.5% and 100%, respectively, when the optimal cut-off value was 6.923.
Area under the ROC curve for miRNAs based on the RT-qPCR data. (A) hsa-miRNA-223-5p, and (B) hsa-miRNA-486-5p.
Discussion
The investigation of exosomal miRNAs in samples from malaria patients is an expanding area of research that holds the potential to improve our understanding of malarial pathophysiology. The identification of miRNAs involved in the biology and pathology of P. knowlesi malaria has not been extensively studied. This study characterized plasma-derived EVs and presented quantitative data on miRNAs derived from EVs. Significant differences in plasma-derived EVs were noted between malaria patients and healthy individuals, with those from patients being both larger in size and more abundant. This increase in size may be associated with pathological changes in host cells due to cellular stress, such as membrane blebbing, alterations in cell morphology and the formation of apoptotic bodies. These pathological changes can lead to the biogenesis and release of EVs of varying sizes, including larger subpopulations (Avalos-Padilla et al., Reference Avalos-Padilla, Georgiev, Lantero, Pujals, Verhoef, N Borgheti-Cardoso, Albertazzi, Dimova and Fernàndez-Busquets2021; Minwuyelet and Abiye, Reference Minwuyelet and Abiye2022). Additionally, the immune response to Plasmodium infection may enhance EV production as part of the inflammatory response, resulting to an overall increase in both the quantity and size of EVs. The molecular mechanisms underlying the biogenesis of these different EV subpopulations may involve intracellular pathways such as endosomal sorting complexes required for transport (Opadokun and Rohrbach, Reference Opadokun and Rohrbach2021; Minwuyelet and Abiye, Reference Minwuyelet and Abiye2022). The increased levels of plasma EVs observed during malaria infections result from the activation of circulating cells, particularly in severe cases (Babatunde et al., Reference Babatunde, Yesodha Subramanian, Ahouidi, Martinez Murillo, Walch and Mantel2020). However, our study had limited data regarding clinical symptoms, haematological information or parasitaemia, which hindered the understanding of their relationships with the level of EVs. In this study, the isolated EVs included those originating from P. knowlesi parasites expressing the EXP-2 protein, which are situated on the PVM and are crucial for transporting proteins from the vacuole to the cytoplasm of red blood cells. These EVs may be internally generated and subsequently released externally, similar to the production of exosomes.
EV-miRNA quantification was performed using RT-qPCR targeting hsa-miR451-5p, hsa-miR150-5p, hsa-miR486-5p, hsa-miR15b-5p, hsa-miR16-5p, hsa-miR106b-5p, hsa-miR223-5p, let7a-5p and let7b-5p. The hsa-miR451-5p was used as an endogenous control in our study due to its stable expression and relevance to the erythroid system. The miRNA-451a, which regulates erythroid differentiation and maturation, was found to be abundant in EVs under parasite culture conditions (Rathjen et al., Reference Rathjen, Nicol, McConkey and Dalmay2006; Mantel et al., Reference Mantel, Hjelmqvist, Walch, Kharoubi-Hess, Nilsson, Ravel, Ribeiro, Grüring, Ma, Padmanabhan, Trachtenberg, Ankarklev, Brancucci, Huttenhower, Duraisingh, Ghiran, Kuo, Filgueira, Martinelli and Marti2016; Wang et al., Reference Wang, Xi, Hao, Deng, Liu, Wei, Gao, Zhang and Wang2017). EVs from plasma of patients infected with P. knowlesi exhibited significantly higher levels of hsa-miR-223-5p and hsa-miR-486-5p compared to those from healthy individuals, while the others showed no significant differences between the 2 groups. Exosomal miRNA expression levels changes in response to varying conditions. Different isolation techniques can capture distinct EV subpopulations based on physical properties such as size and density. Consequently, the isolated EVs may represent different subpopulations, which can influence the miRNA and biomolecule profiles observed (Llorens-Revull et al., Reference Llorens-Revull, Martínez-González, Quer, Esteban, Núñez-Moreno, Mínguez, Burgui, Ramos-Ruíz, Soria, Rico, Riveiro-Barciela, Sauleda, Piron, Corrales, Borràs, Rodríguez-Frías, Rando, Ramírez-Serra, Camós, Domingo and Costafreda2023). In a recent study that isolated EVs via UC, EV-derived miR-150-5p and miR-15b-5p were identified in patients infected with P. vivax. Furthermore, upregulation of let-7a-5p was observed in both P. vivax and P. falciparum infections (Ketprasit et al., Reference Ketprasit, Cheng, Deutsch, Tran, Imwong, Combes and Palasuwan2020). The miR-223 is involved in the proliferation and function of granulocytes and platelets (Fazi et al., Reference Fazi, Rosa, Fatica, Gelmetti, De Marchis, Nervi and Bozzoni2005; Shi et al., Reference Shi, Zhou, Ji, Zhang, Ma, Zhang and Li2015), implying that platelets and granulocytes might serve as potential sources of EVs during P. knowlesi infection. The upregulation of miR-223 in platelets is associated with increased platelet activation and aggregation (Gatsiou et al., Reference Gatsiou, Boeckel, Randriamboavonjy and Stellos2012). During malaria infection, iRBCs could interact with platelets, leading to the release of chemokines and inflammatory cytokines (Srivastava and Srivastava, Reference Srivastava and Srivastava2015). Hsa-miR-223 also forms complexes with lipoproteins circulating in the blood (Vickers et al., Reference Vickers, Palmisano, Shoucri, Shamburek and Remaley2011). LDL has been observed interacting with the surface of EVs, making it interesting to determine whether this interaction is the result of contamination from the circulation. Isolating EVs from lipoproteins in the blood is still challenging due to the overwhelming abundance of lipoproteins, exceeding EVs by at least 105-fold (Zhang et al., Reference Zhang, Borg, Liaci, Vos and Stoorvogel2020). SEC is effective in largely eliminating high-density lipoprotein, although LDL of comparable density might be co-isolated with EVs. Employing subsequent UC as a washing step can aid in minimizing contamination (Koster et al., Reference Koster, Rojalin, Powell, Pham, Mizenko, Birkeland and Carney2021). miR-223 has been documented as being upregulated in sickle cell erythrocytes infected with P. falciparum (LaMonte et al., Reference LaMonte, Philip, Reardon, Lacsina, Majoros, Chapman, Thornburg, Telen, Ohler, Nicchitta, Haystead and Chi2012) and in the plasma of P. vivax patients (Hadighi et al., Reference Hadighi, Heidari, Fallah, Keshavarz, Tavakoli, Mansouri and Sadrkhanloo2022). Moreover, in mice with cerebral malaria, miR-223-3p, miR-19b-3p and miR-142-3p exhibited significant upregulation compared to non-infected mice (Martin-Alonso et al., Reference Martin-Alonso, Cohen, Quispe-Ricalde, Foronda, Benito, Berzosa, Valladares and Grau2018).
Computational target predictions were conducted for hsa-miR-223-5p and hsa-miR-486-5p on both human and malaria transcripts, along with an enrichment analysis in GO and KEGG pathways. The prediction indicated that hsa-miR223-5p targets several human transcripts including hepatocyte growth factor (HGF), MET proto-oncogene (MET), vascular cell adhesion molecule 1 (VCAM-1), toll-like receptor 4 (TLR4), myeloid differention primary response 88 (MyD88), glycophorin A (GYPA), glycophorin C (GYPC) and CD40 molecule (CD40). The HGF/MET signalling pathway plays a critical role in diverse cellular processes such as cell growth, survival, motility and morphogenesis (Organ and Tsao, Reference Organ and Tsao2011). Moreover, it is essential for the initial development of parasites within the host liver by preventing cell apoptosis and involving host-cell actin cytoskeleton reorganization (Carrolo et al., Reference Carrolo, Giordano, Cabrita-Santos, Corso, Vigário, Silva, Leirião, Carapau, Armas-Portela, Comoglio, Rodriguez and Mota2003). VCAM-1 contains 6 or 7 immunoglobulin domains, is expressed on endothelial cells in blood vessels, and acts as a cell adhesion molecule (Cook-Mills et al., Reference Cook-Mills, Marchese and Abdala-Valencia2011). The elevated VCAM-1 expression is associated with the sequestration of P. falciparum in vessels, particularly in severe cases (Armah et al., Reference Armah, Wired, Dodoo, Adjei, Tettey and Gyasi2005). TLR4 is expressed in immune cells including monocytes, macrophages, B cells, dendritic cells and epithelial cells. Monocytes expressing TLR4 are activated by parasite-derived GPI anchors and hemozoin, leading to the production of pro-inflammatory cytokines. MyD88 acts as a key signalling protein, connecting the activated receptors with downstream signalling molecules. MyD88 activation leads to the activation of transcription factors such as NF-κB and MAPKs, which results in the production of pro-inflammatory cytokines like IL-1, TNF-α and IL-6 (Dobbs et al., Reference Dobbs, Crabtree and Dent2020). GYPA and GYPC are cell surface receptors found on red blood cells that engage with parasite antigens such as EBA-175 and EBA-140, facilitating the invasion of parasites (Jaskiewicz et al., Reference Jaskiewicz, Jodłowska, Kaczmarek and Zerka2019). In this study, all predicted human host targets of hsa-miR-223-5p were found to be associated with specific biological processes and CCs based on GO analysis. These targets are enriched in processes like phagocytosis, cellular response to mechanical stimuli, and NF-κB activation, which are critical for immune cell function and host defence. The enriched CCs include the cell surface, integral components of the plasma membrane and the plasma membrane, emphasizing the role of miR-223-5p in membrane-related immune interactions. Based on these findings, we hypothesized that miR-223-5p may target HGF, which acts as an anti-inflammatory mediator. If miR-223-5p reduces HGF levels, immune responses could shift toward a more pro-inflammatory state, thereby activating pathways such as NF-κB, TLR4 and MyD88 (Zhou et al., Reference Zhou, Pal, Hsu, Gurol, Zhu, Wirbisky-Hershberger, Freeman, Kasinski and Deng2018). A previous study suggested a role for miRNAs in modulating inflammatory signalling pathways (Lee et al., Reference Lee, Howitt, Cheng, King, Stawiski, Fendler, Chowdhury, Matulonis and Konstantinopoulos2021); however, direct evidence linking miR-223-5p to HGF suppression and subsequent activation of pro-inflammatory processes remains limited. Further experimental validation is needed to clarify whether miR-223-5p directly influences HGF-mediated immune modulation during P. knowlesi infection. Additionally, while miR-223-5p overexpression may help regulate inflammation and prevent tissue damage, it could also compromise the immune system’s ability to clear the Plasmodium parasite, which utilizes various immune evasion strategies (Su et al., Reference Su, Xu, Stadler, Teklemichael and Wu2025). Moreover, it is important to note that hsa-miR-223-5p can exert varying effects on its target genes, depending on factors such as its subcellular location, expression level, the degree of complementarity with the target and other regulatory elements in the cellular environment (O’Brien et al., Reference O’Brien, Hayder, Zayed and Peng2018). Further investigation is needed to clarify its specific role in P. knowlesi infection.
Our results indicated an increase in hsa-miR-486-5p expression in plasma-derived EVs from P. knowlesi-infected patients. This finding concurred with a previous study, wherein miRNA profiling of EVs isolated from P. falciparum culture revealed that hsa-miR-486-5p exhibited one of the highest expression levels (Babatunde et al., Reference Babatunde, Mbagwu, Hernández-Castañeda, Adapa, Walch, Filgueira, Falquet, Jiang, Ghiran and Mantel2018). MiR-486-5p is expressed abundantly in RBCs and is involved in RBC maturation. CD40 was predicted as the target of miR-486-5p. CD40, a membrane glycoprotein, is prominently expressed in B lymphocytes, dendritic cells, monocytes and platelets. CD40 expression is induced by pro-inflammatory factors and regulated by transcriptional factors including NF-κB, responsible for the inflammatory response (Antoniades et al., Reference Antoniades, Bakogiannis, Tousoulis, Antonopoulos and Stefanadis2009). The results of this study demonstrated that miR-486-5p was enriched in protein binding and cytosol. Moreover, the parasite mRNA has been predicted as a target of human miR-223-5p, with the SICAvar gene being the most frequently identified target in the database. The P. knowlesi schizont-infected cell agglutination (SICA) var gene family, expressed on the surface of infected erythrocytes, codes for antigenic variation, analogous to the virulence-associated PfEMP1 gene (Lapp et al., Reference Lapp, Korir-Morrison, Jiang, Bai, Corredor and Galinski2013). The SICAvar gene is associated with cytoadhesion of P. knowlesi iRBCs with endothelial cells in the umbilical vein and the gastrointestinal tract (Chuang et al., Reference Chuang, Sakaguchi, Lucky, Yamagishi, Katakai, Kawai and Kaneko2022; Peterson et al., Reference Peterson, Joyner, Lapp, Brady, Wood, Cabrera-Mora, Saney, Fonseca, Cheng, Jiang, Soderberg, Nural, Hankus, Machiah, Karpuzoglu, DeBarry, Tirouvanziam, Kissinger, Moreno, Gumber and Galinski2022). EVs are found to harbour promising miRNA candidates, serving as robust biomarkers with an AUC close to 1. This characteristic makes them valuable for diagnostic purposes. The resistance of miRNAs to degradation, facilitated by their protection within vesicles during circulation, adds to the appeal of exosome miRNAs as stable and reliable disease biomarkers. However, the sorting process of miRNAs in EVs is not well understood and may exhibit either selective or non-selective characteristics, potentially influenced by the subtypes of EVs (Temoche-Diaz et al., Reference Temoche-Diaz, Shurtleff, Nottingham, Yao, Fadadu, Lambowitz and Schekman2019). Recognition by RNA-binding proteins such as hnRNPA2B1 and Argonaute-2, specific structural features, post-transcriptional modifications of miRNA and cellular content can contribute to the loading of miRNAs into EVs (Qiu et al., Reference Qiu, Li, Zhang and Wu2021). Our findings shed light on miRNAs associated with P. knowlesi infection, providing valuable insights into their molecular mechanisms. Further research is required to explore the role of miRNAs in malaria pathogenesis. In-depth in vitro and in vivo studies, along with investigations in human populations are required to confirm these findings and assess the potential of miRNAs as a therapeutic target or diagnostic marker.
Conclusions
This study established a foundation for understanding the pathophysiology of P. knowlesi, highlighting the increased expression of EV-miRNAs, specifically hsa-miR-223-5p and hsa-miR-486-5p, which could reflect the underlying processes of P. knowlesi infection. Further studies should validate the functionality of EV-miRNAs, as they have the potential to provide crucial insights to develop new diagnostic or patient monitoring biomarkers.
Author contributions
Conceptualization: ST and RN, Software: TK and NN, Investigation: TK and PM. Resources: CW, JS, SJ, CR, and RN, Formal Analysis: HB, KS, NT, RP, CS, and NS, Writing – Original Draft Preparation: TK and ST, Writing – Revised Draft and Editing: ST, RN, HB, JS, and CW. All authors have read and approved the final version of the revised manuscript. RN and ST contributed equally as co-corresponding authors.
Financial support
This research was supported by Prince of Songkla University (Grant No. MET6602031S and MET6402038S), National Science, Research and Innovation Fund (NSRF) and Prince of Songkla University (Grant No. MET6601206S), PSU Ph.D. Scholarship contract No. PSU_PHD2562-004, the NSRF via the Program Management Unit for Human Resources & Institutional Development, Research and Innovation (Grant No. B13F660074) and Thailand Science Research and Innovation (TSRI), Chulabhorn Research Institute (grant number 49890/4759798).
Competing interests
The authors declare no competing interests.
Ethical standards
All samples were obtained as leftover specimens from routine laboratory procedures. This study received ethical approval from the Ethics Committee of the Faculty of Medical Technology, Prince of Songkla University, under approval number EC66-03.