Introduction
Trypanosomatidae is a family of protozoan parasites possessing a single large mitochondrion, which encompasses a network of catenated circular DNA molecules, the so-called kinetoplast or kDNA (Maslov et al., Reference Maslov, Opperdoes, Kostygov, Hashimi, Lukeš and Yurchenko2019). These species have been attracting research attention because of numerous unique or rare biochemical and molecular traits, such as trans-splicing and polycistronic transcription (Clayton, Reference Clayton2019; Michaeli, Reference Michaeli2011), mitochondrial RNA editing (Aphasizheva et al., Reference Aphasizheva, Alfonzo, Carnes, Cestari, Cruz-Reyes, Goringer, Hajduk, Lukeš, Madison-Antenucci, Maslov, McDermott, Ochsenreiter, Read, Salavati, Schnaufer, Schneider, Simpson, Stuart, Yurchenko, Zhou, Zíková, Zhang, Zimmer and Aphasizhev2020), presence of modified nucleotides (van Luenen et al., Reference van Luenen, Farris, Jan, Genest, Tripathi, Velds, Kerkhoven, Nieuwland, Haydock, Ramasamy, Vainio, Heidebrecht, Perrakis, Pagie, van Steensel, Myler and Borst2012) and unusual organelles (Szöör et al., Reference Szöör, Haanstra, Gualdrón-López and Michels2014; Docampo, Reference Docampo2016), or a bizarre variation of the nuclear genetic code (Záhonová et al., Reference Záhonová, Kostygov, Ševčíková, Yurchenko and Eliáš2016). Most of these flagellates are monoxenous (with one host in their life cycle) parasites restricted to invertebrates (Maslov et al., Reference Maslov, Votýpka, Yurchenko and Lukeš2013), while members of the genera Endotrypanum, Leishmania, Phytomonas, Porcisia and Trypanosoma have switched to dixeny (two-host life cycle) and infect vertebrates or plants in addition to invertebrates (Lukeš et al., Reference Lukeš, Butenko, Hashimi, Maslov, Votýpka and Yurchenko2018). It is established beyond a reasonable doubt that the dixenous species have evolved from the monoxenous ancestor(s) independently several times (Lukeš et al., Reference Lukeš, Skalický, Týč, Votýpka and Yurchenko2014). Notably, several Leishmania and Trypanosoma spp. are of medical importance, as they cause severe human diseases, and are fairly well-studied (Stuart et al., Reference Stuart, Brun, Croft, Fairlamb, Gurtler, McKerrow, Reed and Tarleton2008; Nussbaum et al., Reference Nussbaum, Honek, Cadmus and Efferth2010).
Telomeres typically represent repetitive physical ends of linear eukaryotic chromosomes, variable in length and sequence in different groups of organisms (Fulnečková et al., Reference Fulnečková, Ševčíková, Fajkus, Lukešová, Lukeš, Vlček, Lang, Kim, Eliáš and Sýkorová2013). Their main role is to protect chromosome ends from being recognized and processed as DNA double-strand breaks by the cellular repair machinery in order to prevent chromosomal end-to-end fusions (Pfeiffer and Lingner, Reference Pfeiffer and Lingner2013). Such shielding is provided by the telomere-associated protein complexes (Lewis and Wuttke, Reference Lewis and Wuttke2012) or specific complementary DNA structures, such as telomere loops (t-loops) facilitating the protection of chromosome ends (Tomáška et al., Reference Tomáška, Nosek, Kar, Willcox and Griffith2019). It is generally assumed that telomeres undergo gradual shortening with each round of cell division because of incomplete lagging strand synthesis of linear DNA templates by DNA polymerases, known as the ‘end replication problem’ (Olovnikov, Reference Olovnikov1973; Greider, Reference Greider1990; Hackett and Greider, Reference Hackett and Greider2002). In order to overcome this problem and, thus, prevent telomere shortening, cells engage a dedicated enzyme called telomerase (Greider and Blackburn, Reference Greider and Blackburn1985).
Telomeres of kinetoplastids share many traits with those of other eukaryotes. They have the canonical sequence (5′-ttaggg-3′) found in vertebrates, end with a t-loop, are associated with capping protein complexes and maintained by telomerases (Muñoz-Jordán et al., Reference Muñoz-Jordán, Cross, de Lange and Griffith2001; Conte and Cano, Reference Conte and Cano2005; Fulnečková et al., Reference Fulnečková, Ševčíková, Fajkus, Lukešová, Lukeš, Vlček, Lang, Kim, Eliáš and Sýkorová2013). Similar to the situation in other eukaryotic pathogens, genes encoding trypanosomatid virulence factors are often located in the sub-telomeric regions and their expression may be co-regulated with telomeres (Chiurillo et al., Reference Chiurillo, Cano, Da Silveira and Ramirez1999; Dobson et al., Reference Dobson, Scholtes, Myler, Turco and Beverley2006; Hovel-Miner et al., Reference Hovel-Miner, Boothroyd, Mugnier, Dreesen, Cross and Papavasiliou2012). Moreover, transposable elements are often found in association with telomeres (Pardue et al., Reference Pardue, Danilevskaya, Traverse and Lowenhaupt1997; Rahnama et al., Reference Rahnama, Novikova, Starnes, Zhang, Chen and Farman2020). In agreement with this, a sub-telomeric region of Leptomonas pyrrhocoris chromosome contains an integrated copy of an RNA-dependent RNA polymerase putatively originating from an RNA virus of the family Tombusviridae infecting this flagellate (Grybchuk et al., Reference Grybchuk, Akopyants, Kostygov, Konovalovas, Lye, Dobson, Zangger, Fasel, Butenko, Frolov, Votýpka, d'Avila-Levy, Kulich, Moravcová, Plevka, Rogozin, Serva, Lukeš, Beverley and Yurchenko2018), and possibly contributing to the retrotransposon translocation within the trypanosomatid genome. Telomeric regions of kinetoplastid chromosomes also possess several features distinguishing them from their counterpart in most of the other eukaryotes. For example, the telomeres of Trypanosoma brucei increase in length (by approximately 10 bp per generation) until they reach an equilibrium (Bernards et al., Reference Bernards, Michels, Lincke and Borst1983; Pays et al., Reference Pays, Laurent, Delinte, Van Meirvenne and Steinert1983; Horn et al., Reference Horn, Spence and Ingram2000). In trypanosomatids, a modified nucleobase, base J (β-D-gluco-pyranosyl-oxy-methyl-uracil) is involved in RNA polymerase II transcription termination and is preferentially localized to telomeres (Borst and van Leeuwen, Reference Borst and van Leeuwen1997; Genest et al., Reference Genest, Ter Riet, Cijsouw, van Luenen and Borst2007; van Luenen et al., Reference van Luenen, Farris, Jan, Genest, Tripathi, Velds, Kerkhoven, Nieuwland, Haydock, Ramasamy, Vainio, Heidebrecht, Perrakis, Pagie, van Steensel, Myler and Borst2012).
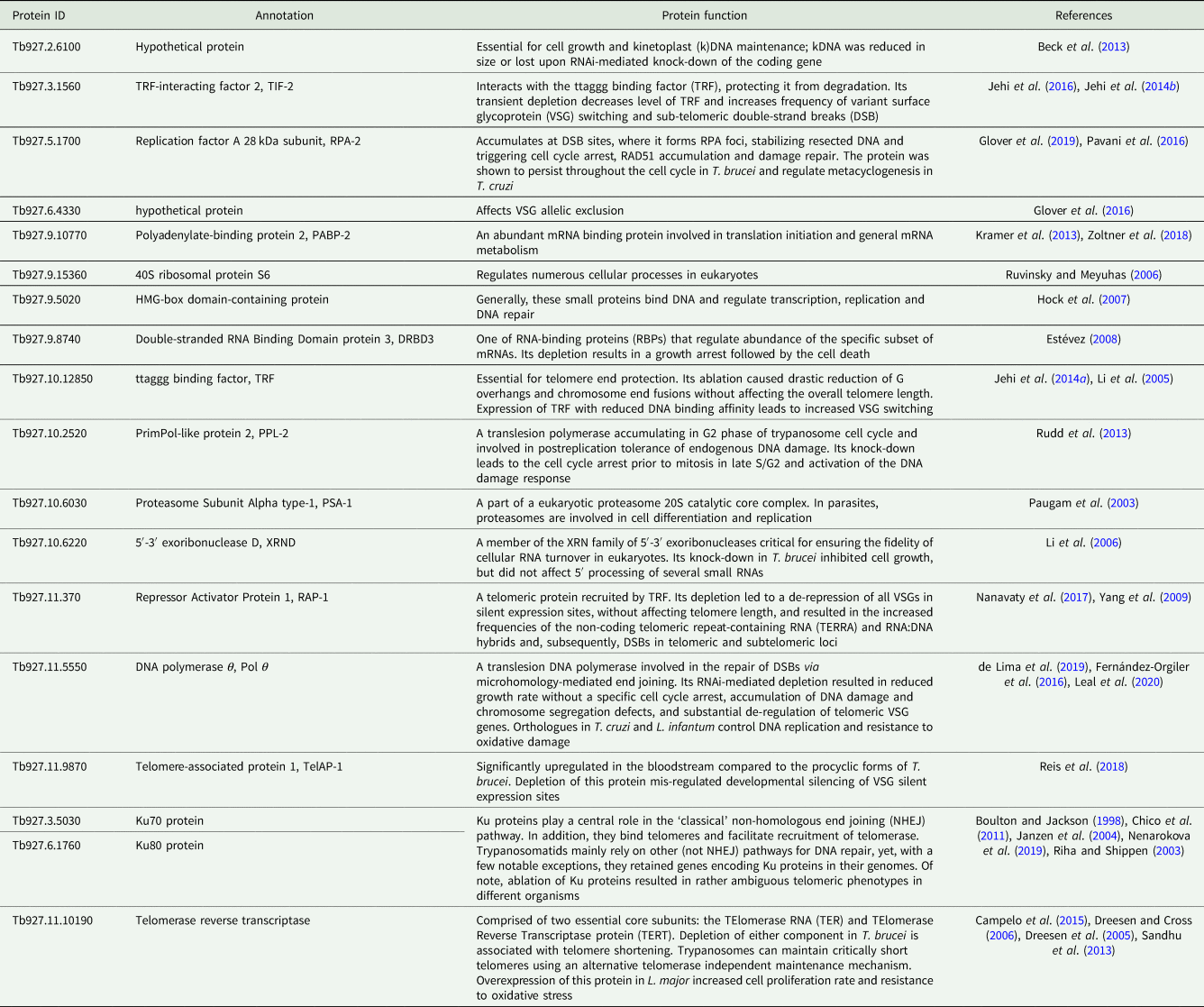
To the best of our knowledge, there has been very little systematic effort to analyse telomeres in trypanosomatids outside the medically relevant Trypanosoma and Leishmania spp. (Fu et al., Reference Fu, Perona-Wright and Barker1998; Fu and Barker, Reference Fu and Barker1998a, Reference Fu and Barker1998b; Chiurillo et al., Reference Chiurillo, Cano, Da Silveira and Ramirez1999, Reference Chiurillo, Peralta and Ramirez2002; Muñoz-Jordán et al., Reference Muñoz-Jordán, Cross, de Lange and Griffith2001; Janzen et al., Reference Janzen, Lander, Dreesen and Cross2004; Conte and Cano, Reference Conte and Cano2005; Genest and Borst, Reference Genest and Borst2007). Therefore, we decided to do that for a wide range of trypanosomatids with a special emphasis on largely neglected parasites of insects, which are not pathogenic to humans. We selected more than 20 proteins from a set of recently defined putative trypanosomatid telomere-associated proteins (Reis et al., Reference Reis, Schwebs, Dietz, Janzen and Butter2018) for more detailed in silico analyses. For most of these proteins, some functional information is available [Table 1; the TriTrypDB (Aslett et al., Reference Aslett, Aurrecoechea, Berriman, Brestelli, Brunk, Carrington, Depledge, Fischer, Gajria, Gao, Gardner, Gingle, Grant, Harb, Heiges, Hertz-Fowler, Houston, Innamorato, Iodice, Kissinger, Kraemer, Li, Logan, Miller, Mitra, Myler, Nayak, Pennington, Phan, Pinney, Ramasamy, Rogers, Roos, Ross, Sivam, Smith, Srinivasamoorthy, Stoeckert, Subramanian, Thibodeau, Tivey, Treatman, Velarde and Wang2010) gene IDs are used throughout the text]. The predicted telomere-associated complex appears to be a cohort of proteins with widely variable functions, from ribosome and proteasome subunits to telomerase and even DNA repair proteins (Boulton and Jackson, Reference Boulton and Jackson1998; Paugam et al., Reference Paugam, Bulteau, Dupouy-Camet, Creuzet and Friguet2003; Riha and Shippen, Reference Riha and Shippen2003; Janzen et al., Reference Janzen, Lander, Dreesen and Cross2004; Dreesen et al., Reference Dreesen, Li and Cross2005; Ruvinsky and Meyuhas, Reference Ruvinsky and Meyuhas2006; Chico et al., Reference Chico, Ciudad, Hsu, Lue and Larriba2011; Sandhu et al., Reference Sandhu, Sanford, Basu, Park, Pandya, Li and Chakrabarti2013; Nenarokova et al., Reference Nenarokova, Záhonová, Krasilnikova, Gahura, McCulloch, Ziková, Yurchenko and Lukeš2019). In this work, we analysed the evolutionary history of telomere-associated proteins in Kinetoplastea, performed a systematic analysis of telomere length variation among trypanosomatids on the dataset, which incorporates a wide range of understudied monoxenous members of the family Trypanosomatidae, and established a correlation between the level of transcription for several analysed telomere-associated proteins and the telomere length.
Predicted telomere-associated protein complex composition in T. brucei
Materials and methods
In silico analyses
A putative set of telomere-associated proteins of T. brucei brucei TREU927 (Reis et al., Reference Reis, Schwebs, Dietz, Janzen and Butter2018) were used as queries for BLAST searches (Altschul et al., Reference Altschul, Gish, Miller, Myers and Lipman1990) against a dataset of annotated proteins of 64 trypanosomatids and the eubodonid Bodo saltans. First, BLASTp searches were performed with an E-value set to 1 and all the hits with an E-value not exceeding 10−15 were retained. If the respective sequence was not identified among annotated proteins, the searches were repeated with the tBLASTn algorithm against a database of genome sequences. In case no protein was identified in the genome, HMMER v.3.3 (Eddy, Reference Eddy2009), a more sensitive method for the identification of divergent homologues based on hidden Markov models was employed. Annotated proteins and assembled genome sequences were downloaded from the NCBI Genome (Sayers et al., Reference Sayers, Agarwala, Bolton, Brister, Canese, Clark, Connor, Fiorini, Funk, Hefferon, Holmes, Kim, Kimchi, Kitts, Lathrop, Lu, Madden, Marchler-Bauer, Phan, Schneider, Schoch, Pruitt and Ostell2019) and TriTrypDB v. 45/46 (Aslett et al., Reference Aslett, Aurrecoechea, Berriman, Brestelli, Brunk, Carrington, Depledge, Fischer, Gajria, Gao, Gardner, Gingle, Grant, Harb, Heiges, Hertz-Fowler, Houston, Innamorato, Iodice, Kissinger, Kraemer, Li, Logan, Miller, Mitra, Myler, Nayak, Pennington, Phan, Pinney, Ramasamy, Rogers, Roos, Ross, Sivam, Smith, Srinivasamoorthy, Stoeckert, Subramanian, Thibodeau, Tivey, Treatman, Velarde and Wang2010) databases. The validity of the hits was confirmed using reciprocal BLAST searches against T. brucei proteins and alignments including the query and all identified proteins, if necessary. The resulting gene presence/absence table and a cladogram manually written in a Newick format based on recent publications (Butenko et al., Reference Butenko, Kostygov, Sádlová, Kleschenko, Bečvář, Podešvová, Macedo, Žihala, Lukeš, Bates, Volf, Opperdoes and Yurchenko2019; Kostygov et al., Reference Kostygov, Grybchuk-Ieremenko, Malysheva, Frolov and Yurchenko2014, Reference Kostygov, Frolov, Malysheva, Ganyukova, Chistyakova, Tashyreva, Tesařová, Spodareva, Režnarová, Macedo, Butenko, d'Avila-Levy, Lukeš and Yurchenko2020; Kostygov and Yurchenko, Reference Kostygov and Yurchenko2017; Lukeš et al., Reference Lukeš, Butenko, Hashimi, Maslov, Votýpka and Yurchenko2018; Frolov et al., Reference Frolov, Malysheva, Ganyukova, Spodareva, Yurchenko and Kostygov2019; Kato et al., Reference Kato, Caceres, Seki, Silupu Garcia, Holguin Mauricci, Castro Martinez, Moreno Paico, Castro Muniz, Troyes Rivera, Villegas Briones, Guerrero Quincho, Sulca Jayo, Tineo Villafuerte, Manrique de Lara Estrada, Arias, Passara, Ruelas Llerena, Kubo, Tabbabi, Yamamoto and Hashiguchi2019) were used for Dollo parsimony analysis in the Count software (Csűrös, Reference Csűrös2010) and results were visualized in a graphical editor.
Trypanosomatid isolates and cultivation
Cultures of Crithidia expoeki (BJ08.175), C. fasciculata (Cf-Cl), C. fasciculata (ATCC20358), C. termophilla (ATCC30817), L. pyrrhocoris (H10), L. seymouri (ATCC30220), Novymonas esmeraldas (E262AT), Strigomonas oncopelti (TCC290E) and Zelonia costaricensis (15EC) were grown in BHI medium (Oxoid/Thermo Fisher Scientific, Basingstoke, UK) supplemented with 2 μg mL−1 Hemin (Sigma-Aldrich, St. Louis, USA) and 50 units mL−1 of Penicillin/Streptomycin (BioSera, Nuaillé, France) at 23°C. Cultures of Endotrypanum monterogeii (ATCC30507), Herpetomonas samuelpessoai, Leishmania (Leishmania) major (LV39), L. (L.) amazonensis (Josefa, LV78, LV79 and PH8), L. (L.) mexicana (M379), L. (Mundinia) martiniquensis (LEM2494), L. (M.) orientalis (PCM2), L. (Sauroleishmania) tarentolae (ParrotTarII), L. (Viannia) braziliensis (human2017), L. (V.) guyanensis (M4147), Phytomonas sp. (EM1), Porcisia deanei (TCC258) and P. hertigi (TCC260) were grown in M199 medium supplemented with 2 μg mL−1 Biopterin, 2 μg mL−1 Hemin (all Sigma-Aldrich), 25 mm HEPES (Lonza, Basel, Switzerland), 50 units mL−1 of Penicillin/Streptomycin (BioSera) and 10% fetal bovine serum (BioWest, Nuaillé, France) at 23°C. Cultures of Angomonas deanei (CT-IOC 044), A. desouzai (CT-IOC 109), Blastocrithidia sp. (p57), Blastocrithidia triatomae, Blechomonas ayalai (B08-376), Blechomonas pulexsimulantis (ATCC50186), Herpetomonas muscarum (MMO-01), Jaenimonas sp. (Finn-01.02), Kentomonas sorsogonicus (MF-08.02), Lafontella sp. (GMO-01), Vickermania ingenoplastis (CP21), Wallacemonas collosoma (ATCC30261) and W. rigidus (PL11) were maintained in SDM medium (BioWest) supplemented with 10% fetal bovine serum (BioWest) and 50 units mL−1 of Penicillin/Streptomycin (BioSera) at 23°C. In the cases of Lafontella and Endotrypanum, cultures were grown in a bi-phasic medium, overlaying blood agar. All species were validated by amplifying and sequencing the 18S rRNA gene as described previously (Kostygov et al., Reference Kostygov, Grybchuk-Ieremenko, Malysheva, Frolov and Yurchenko2014).
Quantification of transcription level of genes encoding telomeric proteins using RT-qPCR
RNA was isolated and transcript levels of the telomeric proteins were assessed by RT-qPCR as described previously (Záhonová et al., Reference Záhonová, Hadariová, Vacula, Yurchenko, Eliáš, Krajčovič and Vesteg2014; Kraeva et al., Reference Kraeva, Leštinová, Ishemgulova, Majerová, Butenko, Vaselek, Bespyatykh, Charyyeva, Spitzová, Kostygov, Lukeš, Volf, Votýpka and Yurchenko2019). Sequences of the specific primers for L. mexicana/amazonensis orthologues of T. brucei genes are listed in Supplementary Table 1. Expression values were normalized to those of 18S rRNA.
Southern blotting
The previously established terminal restriction fragment analysis of telomere lengths protocol was followed (Janzen et al., Reference Janzen, Lander, Dreesen and Cross2004). In brief, total genomic DNA from the log-phase grown cells was isolated and digested with AluI, HinfI and RsaI overnight. Restriction fragments were separated in 0.75% agarose gel, transferred to a ZetaProbe blotting membrane (Bio-Rad, Hercules, USA), probed with the DIG-labelled telomeric probe [CCCTAA]x25 in the DIG Easy Hyb buffer (Roche Diagnostics, Indianapolis, USA), and visualized with the DIG Luminescent Detection Kit (Roche Diagnostics). The probe was labelled by the Dioxigenin NT Labeling Kit (Jena Bioscience GmbH, Jena, Germany). Statistics of the telomere lengths were obtained with an online tool WALTER (Web-based Analyser of the Length of TElomeRes) (Lyčka et al., Reference Lyčka, Peška, Demko, Spyroglou, Kilar, Fajkus and Fojtová2021). For the loading and integrity control in the L. mexicana complex analysis, DNAs were processed as above, and the membrane was probed against a fragment of a gene encoding telomerase (LmxM.36.3930) (Supplementary Table 1).
Results and discussion
The core set of proteins putatively involved in telomere maintenance in kinetoplastids is conserved
To study the phylogenetic distribution of proteins predicted to be involved in telomere maintenance (Reis et al., Reference Reis, Schwebs, Dietz, Janzen and Butter2018), we analysed the presence/absence of the corresponding 24 genes in the available genomes of trypanosomatids and their close eubodonid relative, B. saltans (Table 2). Most of the studied proteins (20 of 24) are well conserved and we consider them as a core set putatively involved in telomere maintenance in kinetoplastids. It is worth noting that the telomere association and function in telomere maintenance has already been confirmed for some of these proteins, while some others have not been functionally characterized yet. Thus, the composition of the core set of proteins involved in telomere maintenance, as defined previously (Reis et al., Reference Reis, Schwebs, Dietz, Janzen and Butter2018) and discussed herein, should be taken with caution. Despite the fact that most of the respective genes are conserved across Kinetoplastea (Fig. 1, Table 2) and, thus appear to be present in the kinetoplastid common ancestor, we came across several interesting exceptions that are discussed in detail below.
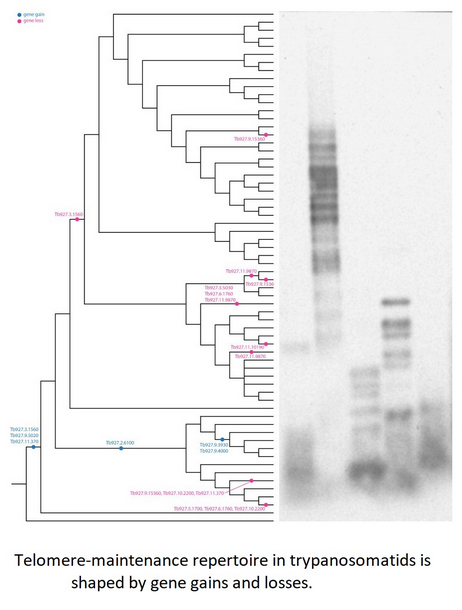
Gains and losses of genes encoding putative telomere-associated proteins in kinetoplastids.

Presence of genes putatively involved in telomere maintenance in kinetoplastids
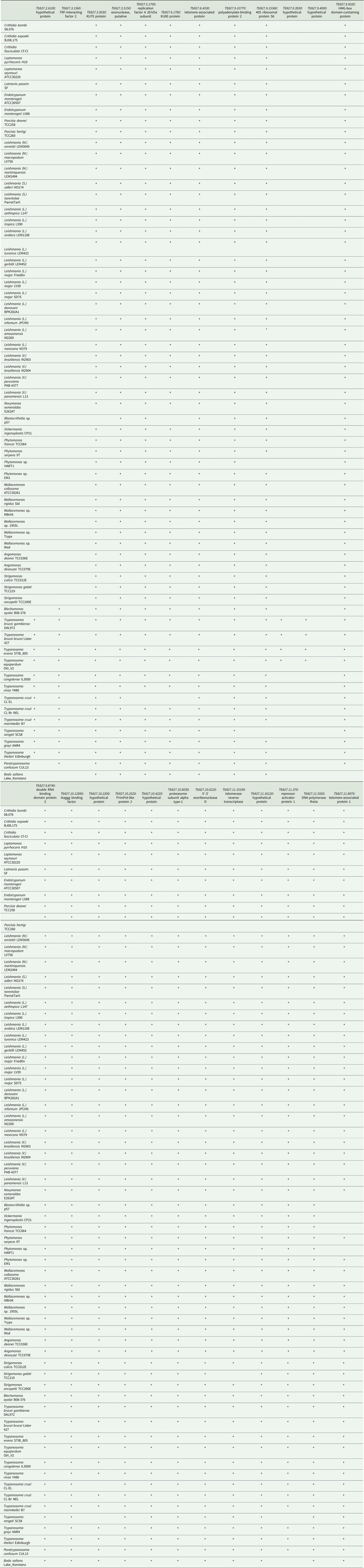
Species analysed by Southern blotting are shaded.
+, identified; empty, not identified; *, identified in strain UA301.
A set of three proteins (orthologues of T. brucei Tb927.3.1560, Tb927.9.5020 and Tb927.11.370) was acquired by the common ancestor of trypanosomatids upon the separation from bodonids (Fig. 1). One of them, Tb927.3.1560 [TIF-2, an orthologue of mammalian TINF2 (Jehi et al., Reference Jehi, Li, Sandhu, Ye, Benmerzouga, Zhang, Zhao and Li2014a; Reference Jehi, Wu and Li2014b)] was suggested to be essential, as it is involved in shelterin (a protein complex implicated in telomere protection) assembly and telomerase-mediated telomere length maintenance in other organisms (Walne et al., Reference Walne, Vulliamy, Beswick, Kirwan and Dokal2008; Frank et al., Reference Frank, Tran, Qu, Stohr, Segal and Xu2015). Yet, it is not present in bodonids and was secondarily lost in all other trypanosomatids outside of the genera Paratrypanosoma, Trypanosoma and Blechomonas (Fig. 1), raising a question of how do they cope with its absence or whether they replaced it with a functional analogue? Tb927.2.6100 is Trypanosoma-specific, confirming previous report (Beck et al., Reference Beck, Acestor, Schulfer, Anupama, Carnes, Panigrahi and Stuart2013). Surprisingly, this protein was shown to be specifically associated with kDNA, so its role in telomere maintenance, if any, remains to be elucidated by functional genetics approaches. Two proteins (orthologues of T. brucei Tb927.9.3930 and Tb927.9.4000) are present only in four species of the T. brucei group and may determine specific traits of these parasites.
An orthologue of Tb927.11.9870 (TelAP-1) is present in most species, but it is conspicuously absent from the representatives of two monoxenous groups (Blastocrithidia and Vickermania spp.) and most Phytomonas spp., plant pathogens with streamlined genomes (Porcel et al., Reference Porcel, Denoeud, Opperdoes, Noel, Madoui, Hammarton, Field, Da Silva, Couloux, Poulain, Katinka, Jabbari, Aury, Campbell, Cintron, Dickens, Docampo, Sturm, Koumandou, Fabre, Flegontov, Lukeš, Michaeli, Mottram, Szoor, Zilberstein, Bringaud, W and Dollet2014). While we cannot rule out a possibility that the protein is divergent beyond recognition by available bioinformatics tools, there may exist another component fulfilling the role of TelAP-1 in these species.
Of special attention is the absence of Tb927.3.5030 (Ku70) and Tb927.3.5030 (Ku80) orthologues in Blastocrithidia sp., which has a non-canonical nuclear genetic code with all three stop codons reassigned to encode amino acids (Záhonová et al., Reference Záhonová, Kostygov, Ševčíková, Yurchenko and Eliáš2016). It has been recently proposed that such an absence may lead to the accumulation of numerous insertions in many protein-coding genes of these organisms (Nenarokova et al., Reference Nenarokova, Záhonová, Krasilnikova, Gahura, McCulloch, Ziková, Yurchenko and Lukeš2019).
Trypanosomatid telomeres are variable in length
We performed a systematic screen of the telomere length across Trypanosomatidae by Southern blotting (Fig. 2, Table 3). Our analysis revealed that monoxenous Leishmaniinae (Kostygov and Yurchenko, Reference Kostygov and Yurchenko2017) of the genera Leptomonas, Novymonas and Zelonia have fairly short telomeres (weighted medians 900–1200 bp; hereafter only rounded weighted median data are compared in the text, see Table 3 for minimum and maximum values), while telomeres in analysed Crithidia spp. ranged from 400 bp in C. fasciculata Cf-C1 to 4300 bp in C. expoeki. These numbers correlate well with previous reports on telomere length in the selected representatives of the genera Crithidia, Leishmania and Trypanosoma (Genest et al., Reference Genest, Ter Riet, Cijsouw, van Luenen and Borst2007). Of note, the repertoire of genes implicated in telomere maintenance is identical in these flagellates (Table 2), so these differences can be explained by either the presence of other proteins involved in this process, or (more likely) differences in gene expression. Telomeres of Blechomonas, Herpetomonas, Jaenimonas and Wallacemonas spp. are 1300–2100 bp long. The endosymbiont-containing Strigomonadinae [Angomonas, Kentomonas and Strigomonas spp. (Votýpka et al., Reference Votýpka, Kostygov, Kraeva, Grybchuk-Ieremenko, Tesařová, Grybchuk, Lukeš and Yurchenko2014)] differ in telomere length, with S. oncopelti bearing the shortest chromosome ends of ~400 bp.
Southern blotting analysis of telomere repeats in selected species of Trypanosomatidae. Marker sizes are indicated on the left. The vertical lines denote a composite image from the same blot. DNA integrity controls are presented in Supplementary Fig. 1 (left and middle panels).

Telomere lengths (weighted median, minimum–maximum) in selected species of Trypanosomatidae

Representatives of three genera (Blastocrithidia, Leishmania and Trypanosoma) deserved special attention. Uniquely among trypanosomatids, Blastocrithidia spp. lack Ku proteins (Nenarokova et al., Reference Nenarokova, Záhonová, Krasilnikova, Gahura, McCulloch, Ziková, Yurchenko and Lukeš2019), yet their telomeres are of similar length to telomeres of other trypanosomatids (600 and 1500 bp in B. triatomae and Blastocrithidia sp., respectively), arguing that either Ku proteins are dispensable for the telomere length maintenance in these species, or their loss can be compensated by other factors. Telomere sizes vary in different Trypanosoma spp. represented by short telomeres in T. mega (400 bp) and substantially longer telomeres in T. brucei Lister 427 (3100–3400 bp). In contrast to the previous report (Dreesen and Cross, Reference Dreesen and Cross2008), we did not document differences in telomeres' length between the procyclic and bloodstream stages of T. brucei. However, both strains in our analysis have the same origin (Lister 427), while the abovementioned study compared Lister 427 and TREU927 strains. Similar to the cases discussed above, despite possessing the same repertoire of telomere-bound proteins, the distribution of telomere sizes in the Leishmania–Porcisia–Endotrypanum clade (Espinosa et al., Reference Espinosa, Serrano, Camargo, Teixeira and Shaw2018) is wide, exemplified by two extreme cases of P. hertigi (5000 bp) and L. mexicana (400 bp, Fig. 2). Variable telomere length in Leishmania spp. (and possibly other Leishmaniinae) may be explained by the presence of a stress-sensitive telomere-proximal replication activity outside S phase of the cell cycle in these species (Damasceno et al., Reference Damasceno, Marques, Beraldi, Crouch, Lapsley, Obonaga, Tosi and McCulloch2020, Reference Damasceno, Marques, Black, Briggs and McCulloch2021).
RNA level of telomerase and several telomere-associated proteins correlates with telomere length in the species of L. mexicana complex
We analysed telomere length and expression of the core set of proteins putatively involved in telomere maintenance in closely related species forming the L. mexicana complex (Eresh et al., Reference Eresh, McCallum and Barker1994). Similar to the cases discussed above, telomeres in L. mexicana and four isolates of L. amazonensis greatly differed in length from ~400 bp in L. mexicana M379 to ~3400 bp in L. amazonensis LV79 (Fig. 3, Table 3, Supplementary Fig. 1). Such a wide range of telomere lengths correlated well with the expression of the Leishmania spp. telomerase (orthologue of Tb927.11.10190) and a telomere-associated hypothetical protein (orthologue of Tb927.6.4330). The higher expression of these proteins correlated with longer telomeres. The specific roles of these and other proteins remain to be further elucidated by functional studies.
Transcript levels of telomere-associated proteins and telomere lengths in the species of L. mexicana complex. (A) Quantitative RT-PCR analysis of the core set of proteins implicated in telomere maintenance. Gene expression is presented as normalized means and standard deviations of three replicates. Data are presented in two graphs to account for differences in expression values. (B, C) Southern blotting analysis of telomere repeats (B) and telomerase-encoding gene (C, used as an additional DNA integrity control) in L. amazonensis LV78, LV79, PH8, Josefa and L. mexicana M379. Marker sizes are indicated on the left. DNA integrity controls are presented in Supplementary Fig. 1 (right panel).

Conclusions
The genome analysis has allowed us to identify a core set of 20 conserved proteins predicted to be responsible for telomere maintenance in trypanosomatids. Several proteins, previously identified in T. brucei pull-downs, are trypanosome-specific. Out of 20 proteins conserved in Trypanosomatidae, two (Ku70 and Ku80) are conspicuously missing in Blastocrithidia spp., yet telomeres in these species do not appear to be affected by their loss. We documented that telomeres of trypanosomatids are diverse in length, even within groups of closely related species. One such group is a complex of species, related to L. mexicana. Our analysis demonstrated that the expression of several telomere-associated proteins correlates with the documented differences in telomere length within species of the L. mexicana complex, which is indicative of a potential role these proteins may play in the telomere length maintenance.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182021000378.
Acknowledgements
We thank Dr A Kostygov (University of Ostrava) for his advice on trypanosomatid cultivation, Dr P. Volf and Dr J. Sádlová (both Charles University) for providing Leishmania spp. isolates, Dr C. Janzen (University of Würzburg) for sharing the plasmid pSP73 bearing the telomeric probe, and Dr C. Janzen and Dr R. McCulloch (University of Glasgow) for specific advice on methodology.
Financial support
Computational resources were funded by the project ‘e-Infrastruktura CZ’ (e-INFRA LM2018140) provided within the program ‘Projects of Large Research, Development and Innovations Infrastructures’. This work was supported by the Grant Agency of Czech Republic (18-15962S and 20-07186S) and the European Regional Funds (project ‘Centre for Research of Pathogenicity and Virulence of Parasites’ CZ.02.1.01/16_019/0000759) to V.Y. and J.L., and the grant SGS/PrF/2021 from the University of Ostrava (to E.P. and A.T.S.A.). A.T.S.A. and E.P. were also supported by the Moravskoslezský kraji research initiative (RRC/02/2020) and City of Ostrava (0493/ZM1822/8), rescpectively. A part of this work on telomere diversity in Leishmania spp. was funded by the Russian Science Foundation (grant 19-15-00054) to V.Y. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Conflict of interest
None.
Ethical standards
Not applicable.