Introduction
Giardia intestinalis continues to be the most encountered parasitic disease worldwide. It is recognized as a significant public health concern and is included in the World Health Organization’s Neglected Diseases Initiative (Savioli et al. Reference Savioli, Smith and Thompson2006). Although more frequently detected in developing countries with limited access to clean water, sanitation and hygiene facilities, ongoing disease surveillance in developed nations has observed a sharp increase in the number of Giardia sp.-associated waterborne outbreaks of gastrointestinal illness (Efstratiou et al. Reference Efstratiou, Ongerth and Karanis2017; Ryan et al. Reference Ryan, Lawler and Reid2017). Sporadic cases are also on the rise in countries like Australia (Zajaczkowski et al. Reference Zajaczkowski, Mazumdar, Conaty, Ellis and Fletcher-Lartey2018). In New South Wales (NSW), Australia, giardiasis remains the most common notifiable parasitic infection in NSW with an average (before the COVID-19 pandemic) of around 3000 cases notified by laboratories each year (Communicable Diseases Branch 2017; Zajaczkowski et al. Reference Zajaczkowski, Mazumdar, Conaty, Ellis and Fletcher-Lartey2018). It may be that these case numbers are reflective of better diagnostic methodologies used in pathology laboratories and the implementation of sensitive assays such as multiplex-PCR. Although it should be noted that culture-independent DNA-based testing methods were only implemented in Australian diagnostic laboratories in late 2013 and onwards, the complete impact of these testing methods on Giardia case notifications remains unquantified (Zajaczkowski et al. Reference Zajaczkowski, Mazumdar, Conaty, Ellis and Fletcher-Lartey2018). The increase of children attending child-care centres across NSW may also play a part in larger case numbers. By state, NSW has the largest share of children attending child-care centres, and use of these centres has increased by 27·8% just within the past 10 years (Department of Education, Employment and Workplace Relations 2022). However, only a few Australian studies have examined the risk factors that drive local transmission of giardiasis (Waldron et al. Reference Waldron, Ferrari, Cheung-Kwok-Sang, Beggs, Stephens and Power2011; Fletcher et al. Reference Fletcher, Caprarelli, Merif, Andresen, Van Hal, Stark and Ellis2014; Zajaczkowski et al. Reference Zajaczkowski, Mazumdar, Conaty, Ellis and Fletcher-Lartey2018).
Annual notifications of G. intestinalis peak between January and April each year and are highest among children aged 0 to 4 years, and adults aged 30 to 39 years (Communicable Diseases Branch 2017). Incidence rates (per 100 000 population) also vary across communities, ranging from 11·2 to 63·2; and high rates (∼39·0) are reported from both rural and urban health districts (Communicable Diseases Branch 2018). In NSW, it remains unclear if associations exist between high-risk age groups. A recent epidemiological study found that among hospitalized patients in NSW, giardiasis was the second most identified enteric protozoa (after Blastocystis spp.) affecting mainly children at school age and younger (Fletcher et al. Reference Fletcher, Sibbritt, Stark, Harkness, Rawlinson, Andresen, Van Hal, Merif and Ellis2015). Additionally, a more recent study based in South-Western Sydney confirmed that children aged 5-years of age and under were 7 times at greater risk of contracting giardiasis (Zajaczkowski et al. Reference Zajaczkowski, Mazumdar, Conaty, Ellis and Fletcher-Lartey2018). Despite this, it remains unclear whether there are other factors at play, or even whether fluctuations across NSW communities are reflecting different disease dynamics.
Recent molecular studies have determined that G. intestinalis can be split into 8 morphologically identical genetic assemblages (assemblages A to H) which can only be distinguished by molecular typing methods (Feng and Xiao Reference Feng and Xiao2011; Adam et al. Reference Adam, Dahlstrom, Martens, Bruno, Barbian, Ricklefs, Hernandez, Narla, Patel, Porcella and Nash2013). Both assemblages A and B are potentially zoonotic; they are responsible for most human infections; however, they have also been successfully isolated from other mammals. The remaining assemblages (C to H) are host-specific, although in some rare cases have been detected in humans (Lasek-Nesselquist et al. Reference Lasek-Nesselquist, Welch, Thompson, Steuart and Sogin2009; Feng and Xiao Reference Feng and Xiao2011). Previous work has shown conflicting results regarding the relationship between G. intestinalis assemblages A and B and their clinical presentation. Some studies have suggested that assemblage A is associated with more severe clinical symptoms in Peru, Bangladesh and Spain (Haque et al. Reference Haque, Roy, Kabir, Stroup, Mondal and Houpt2005; Peréz Cordón et al. Reference Peréz Cordón, Cordova Paz Soldan, Vargas Vásquez, Velasco Soto, Sempere Bordes, Sánchez Moreno and Rosales2008; Minetti et al. Reference Minetti, Lamden, Durband, Cheesbrough, Fox and Wastling2015); however, the opposite has also been suggested by others (Homan and Mank Reference Homan and Mank2001). Currently, studies from Australia have yet to find a clear correlation between assemblages and symptoms in humans despite several genotyping studies being reported (Nolan et al. Reference Nolan, Jex, Koehler, Haydon, Stevens and Gasser2013; Asher et al. Reference Asher, Holt, Andrews and Power2014, Reference Asher, Hose and Power2016; Ebner et al. Reference Ebner, Koehler, Robertson, Bradbury, Jex, Haydon, Stevens, Norton, Joachim and Gasser2015; Zahedi et al. Reference Zahedi, Field and Ryan2017). Two studies, however, did investigate a link between clinical symptoms and assemblage type (Read et al. Reference Read, Walters, Robertson and Thompson2002; Yang et al. Reference Yang, Lee, Ng and Ryan2010). Both studies were based in Western Australia. Read et al. (Reference Read, Walters, Robertson and Thompson2002) observed a strong association between assemblage A infection and diarrhoea, while Yang et al. (Reference Yang, Lee, Ng and Ryan2010) did not find similar correlations. It is difficult to ascertain whether these conflicting results were the result of differences in study methodology; however, it is an issue that needs further investigation.
Clinical symptoms of G. intestinalis infection can differ according to each individual, and some cases can even remain asymptomatic. Symptoms can include acute and/or chronic diarrhoea, stomach cramps, nausea, vomiting, dehydration and weight loss (Muhsen and Levine Reference Muhsen and Levine2012). Although it is still unclear why certain cases remain asymptomatic, it has been suggested that host-parasite factors and the genotypic differences within a parasite can influence the subsequent clinical presentation of an infected individual (Rafiei et al. Reference Rafiei, Roointan, Samarbafzadeh, Shayesteh, Shamsizadeh and Borujeni2013; Coelho and Singer Reference Coelho and Singer2018).
The aim of this study was to identify G. intestinalis assemblages contributing to human infections in NSW, Australia, and to detect any significant associations between assemblages and the demographic, clinical and geographical factors. This study provides information on the impact of giardiasis on human health in NSW, and a better understanding of the continuing rise in infection. This will increase the capacity of NSW to apply advanced analyses to disease surveillance and will inform the application of similar methodologies to other intestinal protozoan diseases in NSW.
Materials and methods
Faecal specimen collection
Faecal specimens were collected between June 2018 and December 2019 from individuals who had tested positive for Giardia species. While the collection process began following ethics approval in 2018, some samples accessed from participating hospitals and private laboratories were retrospectively included, with dates back to 2016. These archived samples were de-identified and their inclusion was permitted under the approved ethics protocol. Samples were collected from 2 hospitals, the Centre for Infectious Diseases and Microbiology at Westmead Hospital, NSW and Sydpath at St. Vincent’s Hospital, NSW. To mitigate geographical bias, samples were also collected from 2 private pathology laboratories, namely Laverty Pathology and Douglass Hanly Moir Pathology (DHM), both situated in NSW, Australia. These private laboratories cover a broader geographical scope within NSW when compared to the hospital laboratories.
Diagnosis of Giardia in the hospital pathology laboratories involved a combination of multiplex-PCR detection and immunoassays (Weitzel et al. Reference Weitzel, Dittrich, Möhl, Adusu and Jelinek2006; Stark et al. Reference Stark, Al-Qassab, Barratt, Stanley, Roberts, Marriott, Harkness and Ellis2011; Couturier et al. Reference Couturier, Jensen, Arias, Heffron, Gubler, Case, Gowans and Couturier2015). In both private pathology laboratories, Giardia diagnosis was made by visualizing Giardia cysts and/or trophozoites in faecal smears of prepared concentrates using microscopy. Stool samples from DHM were initially prepared using the Mini-Parasep solvent-free (Apacor, England, UK) faecal parasite concentrator with a formalin and Triton X/ethyl acetate solution. Both private laboratories also utilized commercial antigen tests such as the Remel ProSpecT Giardia/Cryptosporidium microplate immunoassay (Thermo Fisher Scientific) to detect positive antigens.
For each positive stool sample collected, efforts were made to obtain the corresponding patient’s gender, age and postcode region of residence was obtained from the electronic medical records (eMR). To protect the sensitive personal information of the patients, no identifiers were collected from the eMR, and ages of the patients were replaced by age groups to further reduce the possibility of re-identification. All patient ages were categorized into 1 of the 6 age groups: ≤ 5, 6–15, 16–29, 30–49, 50–69 and 70 + years. The history of the patient’s symptoms and potential risk factors were collected from the clinical notes recorded on laboratory requests or medical records. All faecal samples were transported to University of Technology Sydney, provided a unique identification number and stored unpreserved at 4°C before DNA extraction. Note that for individuals with multiple faecal samples collected at the same time, only 1 sample was included in the analyses.
Multiplex RT-PCR
To confirm the presence of G. intestinalis and to detect any co-infections within the collected samples, the specimens were analysed by a multiplexed real-time PCR (RT-PCR) EasyScreen assay (Genetic Signatures, Newtown, Australia) at Sydpath at St. Vincent’s Hospital, Sydney, Australia (Stark et al. Reference Stark, Al-Qassab, Barratt, Stanley, Roberts, Marriott, Harkness and Ellis2011). The assay includes a DNA extraction step as part of the workflow, using proprietary 3base technology to enhance detection sensitivity. The EasyScreen kit tests for a variety of enteric pathogens including common enteric protozoan parasites: (a) Dientamoeba fragilis, (b) Cryptosporidium spp., (c) Blastocystis hominis, (d) Entamoeba complex, (e) G. intestinalis; bacterial pathogens: (a) Salmonella spp., (b) Campylobacter spp., (c) Shigella spp., (d) Yersinia enterocolitica, (e) toxigenic Clostridium difficile and (f) Listeria monocytogenes; and viruses: (a) Norovirus group I, (b) Norovirus group II, (c) Adenovirus hexon, (d) Adenovirus 40/41, (e) Rotavirus A and B, (f) Astrovirus (group 1–7) and (g) Sapovirus.
Stratified sample selection
During the study period, a total of 410 G. intestinalis-positive faecal samples were collected. Of these, 107 were excluded due to missing or incomplete demographic and clinical data (i.e. more than half the fields unavailable), leaving 303 samples. From this group, 169 (55·8%) were selected for genotyping using a stratified sampling strategy to ensure proportional representation by age, sex and geographic location.
While efforts were made to obtain full demographic data for all 169 selected samples, a small proportion (4·1%) had 1 or 2 missing variables (e.g. gender, age, postcode, seasonality, travel history or risk factors). These samples were still included, as they provided valuable insight into assemblage distribution across NSW. This limitation is acknowledged, and readers are advised that conclusions drawn from the demographic trends should be interpreted with caution due to the small number of incomplete records.
Sample collection occurred continuously over the 42-month study period, and samples underwent DNA extraction and genotyping in sequential batches rather than in a single selection at the end of the study. To account for temporal variation, stratified sampling was applied within defined 3-month intervals, aligning with seasonal time periods. Within each interval, samples were selected to proportionally represent the demographic and geographic characteristics of the total case population during that timeframe.
All faecal specimens were stored unpreserved at 4°C before DNA extraction. The final genotyped cohort of 169 samples was constructed by aggregating each stratified batch, ensuring the resulting dataset reflected the demographic, geographic and temporal composition of the full sample population. This methodology helped minimize potential bias related to sample reduction.
Genomic DNA extraction
To prepare samples for genotyping, a DNA extraction was performed on 150 mg of each of the 169 collected faecal samples using the ISOLATE II Fecal DNA Kit (Bioline, Sydney, Australia) according to the manufacturer’s instructions with minor modifications. Specifically, the wash step was performed 3 times with Fecal DNA Wash Buffer (rather than once as per the manufacturer’s instructions). Elution was accomplished by adding 100 μl elution buffer. The eluted DNA was stored at −20°C until PCR amplification. Samples with sterile water were used as a negative control to monitor contamination during nucleic acid extraction. Samples spiked with Cryptosporidium spp. DNA templates were used as a positive control during the extraction process. DNA concentration and purity were determined via 260/280 and 260/230 ratios measured on the NanoDrop One microvolume UV-Vis Spectrophotometer (Thermo Fisher Scientific, United States).
Nested PCR amplification of the G. intestinalis SSU rRNA
Primers originally designed by Hopkins et al. (Reference Hopkins, Meloni, Groth, Wetherall, Reynoldson and Thompson1997) amplify the small subunit ribosomal RNA (SSU-rRNA) gene of G. intestinalis producing a 292 bp product from the primary PCR reaction, and a 130 bp product from the secondary PCR reaction (Hopkins et al. Reference Hopkins, Meloni, Groth, Wetherall, Reynoldson and Thompson1997). Due to the small product size and innate low genetic variation within the SSU-rRNA gene, distinguishing between assemblage A and B sequences is entirely dependent on identifying 4 single nucleotide polymorphisms (SNPs). To increase discriminatory power, new primers were designed to capture an area with a greater distribution of polymorphisms between assemblages A and B. The Clustal W multiple sequence alignment program was used to align SSU-rDNA sequence reads of assemblages A and B, and a total of 7 SNPs were identified between the 2 assemblages (Larkin et al. Reference Larkin, Blackshields, Brown, Chenna, McGettigan, McWilliam, Valentin, Wallace, Wilm, Lopez, Thompson, Gibson and Higgins2007).
A 447 bp fragment of the SSU-rRNA gene was first amplified using the previously described forward primer RH11 (Hopkins et al. Reference Hopkins, Meloni, Groth, Wetherall, Reynoldson and Thompson1997) and the newly designed reverse primer RH4.1: TGGCACCAGACCTTGCCCT. This reaction was followed by a secondary amplification step, which used the internal primer GiarF (Read et al. Reference Read, Walters, Robertson and Thompson2002) and the newly designed reverse primer GiarR.1: ACTCCCCGTCGCTGCCT. The use of GiarR.1 resulted in a nested PCR amplicon of 363 bp long. Both PCR amplifications were prepared in a final volume of 50 μl and carried out using conditions previously described (Hopkins et al. Reference Hopkins, Meloni, Groth, Wetherall, Reynoldson and Thompson1997). Negative controls (no template added) and positive controls (containing DNA from previously sequenced and confirmed G. intestinalis samples) were included in each assay reaction. Reactions were performed on an Eppendorf Mastercycler Nexus (Sigma-Aldrich).
Specificity of the novel primers was tested using a panel of 3 protozoan parasite-positive and 2 bacteria-positive clinical samples previously submitted to St Vincent’s Hospital (including Cryptosporidium parvum, D. fragilis, B. hominis, Campylobacter spp. and Clostridium spp.). Sensitivity was estimated using a series of 10-fold dilutions of DNA from extracted G. intestinalis DNA samples to assess the lowest detection threshold of each PCR assay. Reaction templates corresponded to decreasing concentrations from 102 to 10−3 ng/µL DNA per PCR tube.
To confirm successful amplification, 4 µL of the PCR product was subjected to electrophoresis on a 2·0% agarose gel containing GelRed Nucleic Acid Gel Stain (Sigma-Aldrich). PCR products of the correct band length (363 bp) were purified by using a PCR purification kit (Qiagen, GmbH. Germany) and sequenced (Macrogen, Seoul, Korea) on both strands using the PCR primers. Sequence data were trimmed and analysed using SeqTrace and for each PCR product, a consensus contig was generated from the sequence data (Stucky Reference Stucky2012). The final sequences were then compared to sequences (>99% similarity) contained in GenBank using the nucleotide-BLAST tool. The identification of homologous sequences allowed the determination of the G. intestinalis assemblage. Mixed-assemblage infections were identified by the presence of clear and consistent double peaks at all 7 SNP positions in the chromatograms, indicating the simultaneous presence of both Assemblage A and B templates within the same sample.
Assemblage-specific nested PCR amplification of the TPI gene
Two G. intestinalis assemblage-specific nested PCR assays were used to amplify the triose phosphate isomerase (TPI) gene. A 605 bp fragment of the gene was first amplified using previously described primers AL3543 and AL3546 (Sulaiman et al. Reference Sulaiman, Fayer, Bern, Gilman, Trout, Schantz, Das, Lal and Xiao2003). The secondary PCR reaction involved 2 separate assays using assemblage-specific primers; the assemblage A-specific primers Af and Ar amplifying a 332 bp PCR product (Geurden et al. Reference Geurden, Geldhof, Levecke, Martens, Berkvens, Casaert, Vercruysse and Claerebout2008) and the assemblage B-specific primers Bf and Br amplifying a 400 bp product (Levecke et al. Reference Levecke, Geldhof, Claerebout, Dorny, Vercammen, Cacciò, Vercruysse and Geurden2009). Both PCR amplifications were prepared in a final volume of 50 μl and carried out using conditions previously described (Levecke et al. Reference Levecke, Geldhof, Claerebout, Dorny, Vercammen, Cacciò, Vercruysse and Geurden2009). Negative controls (no template added) and positive controls (containing DNA from previously sequenced and confirmed G. intestinalis samples) were included in each assay reaction. Reactions were performed on an Eppendorf Mastercycler Nexus (Sigma-Aldrich) and PCR products were separated by electrophoresis in a 2·0% agarose gel containing GelRed Nucleic Acid Gel Stain (Sigma-Aldrich).
Defining single and mixed assemblage infections
Further genetic characterization of G. intestinalis involved combining the results of both the assemblage-specific PCR (tpi) and the nested PCR (SSU-rRNA). Characterizing a sample as a single assemblage (either assemblage A or B) would require both PCR assays to have an identical result. A mixed assemblage was defined by either an identical A + B result for both PCR assays or discordant results from each assay.
Statistical analyses and mapping of spatial data
Statistical analysis was performed using SPSS Statistics 27 (IBM, USA). Categorical variables are reported in terms of percentages, with corresponding confidence intervals (CI) at 95%. The existence of association between categorical variables was evaluated using Pearson’s Chi-Square test (or Fisher’s Exact test for sparse data). Statistical significance was set as a P-value < 0·05.
Positive human cases of giardiasis in NSW were also geographically mapped using ArcGIS. Case postcode data was initially geocoded using ArcGIS, then spatially joined to 2 polygon layers: NSW Local Government Area (LGA) boundaries (2019) and Local Health District (LHD) boundaries (2014). NSW is divided into 8 metropolitan LHDs and 7 rural/regional LHDs34. The LHDs are further split into 128 LGAs. Cases were then aggregated according to the postcode-matched LGA and LHD, to calculate the total number of G. intestinalis cases and G. intestinalis assemblages for each region in NSW.
Results
Detection and identification of G. intestinalis genotypes
Of the 169 samples selected through stratified sampling and subjected to genotyping, the majority (87·0%) were sourced from private pathology laboratories, while a smaller proportion (13·0%) originated from hospital laboratories. The tpi assemblage A/B-specific PCR amplified in 87·0% (147/169) of these samples; 18·4% (n = 27) were only assemblage A, 54·4% (n = 80) were only assemblage B and 27·2% (n = 40) were classified as mixed assemblages. The SSU-rRNA PCR amplified 80·5% (136/169) of the specimens; of which 18·4% (n = 25) were only assemblage A, 73·5% (n = 100) were only assemblage B and 8·1% (n = 11) were mixed assemblages A + B. An overview of these sample numbers and distributions is provided in Supplementary Table 1.
Among the subset of G. intestinalis-positive samples that were collected, 162 (95·9%) were successfully amplified at 1 or more loci; of which 9·3% (n = 15) were only assemblage A, 46·9% (n = 76) were only assemblage B and 43·8% (n = 71) were mixed infections of assemblage A + B (see Supplementary Table 1).
Co-infecting pathogens were detected in 49·1% (n = 83) of all G. intestinalis-positive faecal samples. Of these co-infections, 56·6% (n = 47) were parasitic, 12·0% (n = 10) were bacterial and 8·4% (n = 7) were viral. Joint parasitic/viral co-infections were also identified in 14·5% (n = 12), followed by parasitic/bacterial co-infections (4·8%, n = 4), bacterial/viral co-infections (2·4%, n = 2) and parasitic/bacterial/viral co-infections (1·2%, n = 1). Overall, the most common pathogens detected were the enteric protozoa B. hominis (31·3%, n = 25) and D. fragilis (15·0%, n = 12), followed by the pathogens Campylobacter spp. (5·0%, n = 4) and Enterovirus (5·0%, n = 4). Other co-infections are reported in Supplementary Table 2.
Socio-demographics of G. intestinalis assemblage cases
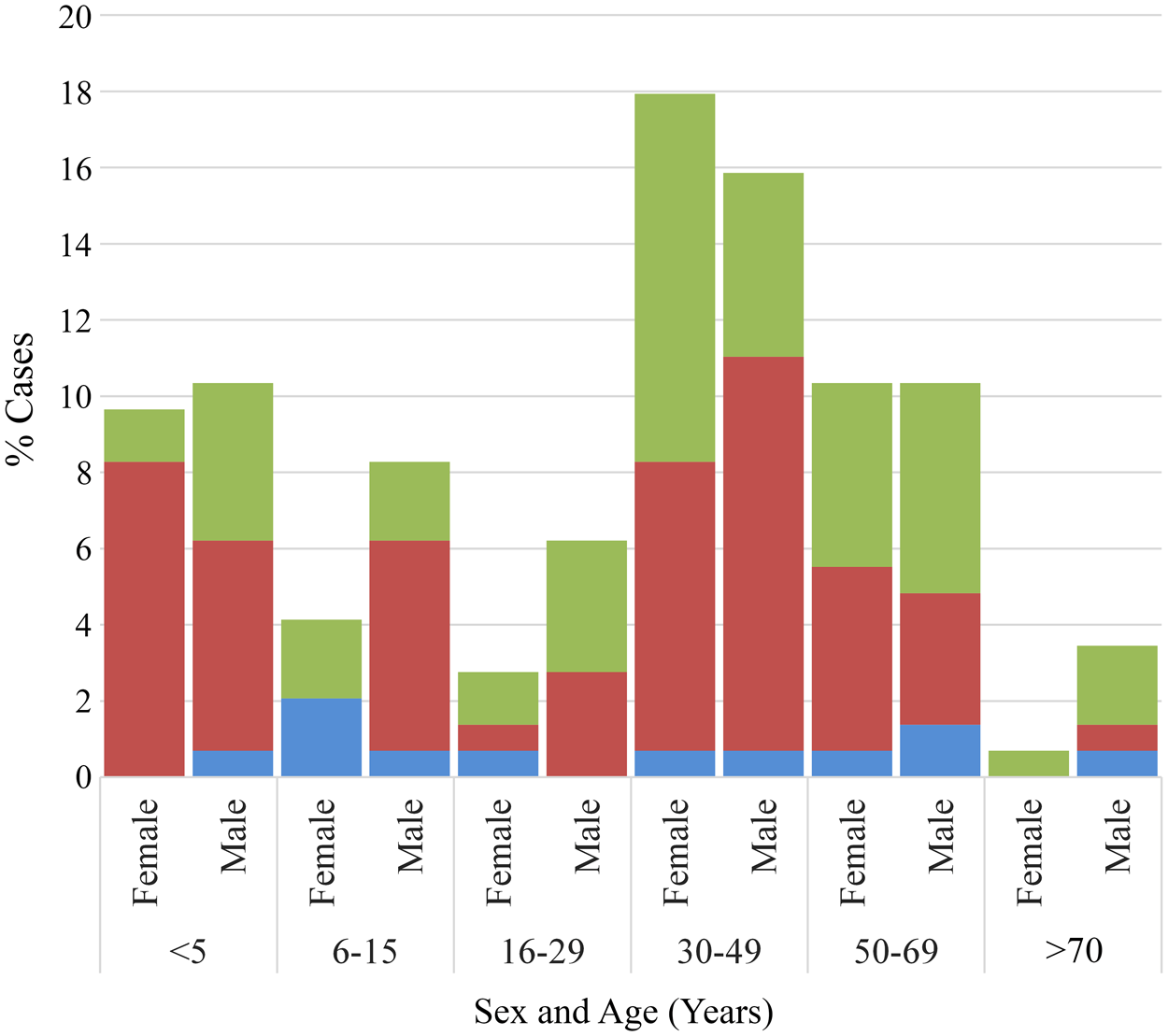
Of the 169 G. intestinalis positive cases selected to represent the broader population, 85·8% (145/169) had complete age metadata and 91·1% (154/169) had valid gender metadata; these cases were included in subsequent analyses (see Supplementary Table 1). Combined age and gender distributions are presented in Figure 1. Individuals infected with G. intestinalis ranged in age from infancy to over 70 years. A bimodal age distribution was observed, with peaks among children aged ≤ 5 years and adults in their 30s. Cases aged 30–49 (33·8%, n = 49) and 50–69 years (20·7%, n = 30) made up the largest age groups, followed closely by those ≤ 5 years (20·0%, n = 29).
Distribution of G. intestinalis assemblages A, B and A + B by age and sex (%). Assemblage A, blue; assemblage B, red; mixed-assemblage A + B, green.

Overall, giardiasis was observed in more males (55·2%, n = 85) than females (44·8%, n = 69), although there was no significant difference between these 2 groups (Table 1). Within male and female groups, the assemblage distribution appeared uniform. For males, single infections with assemblages A or B were identified in 53·8 and 57·5% cases, respectively. Among females, 46·2% cases were assemblage A only infections, whilst 42·5% cases were only B. Mixed assemblage A + B cases were identified in 52·9% males and 47·1% females. Assemblages were distributed across all age groups; however, single assemblage A infections were mostly seen in children aged 6–15 years, and adults aged 50 years and greater. In comparison, single assemblage B infections were more common among middle-aged individuals aged 30–49 years old, and children aged 5 years and under. When categorizing the cases into 2 age categories (≤5 and > 5 years), it was found that children ≤ 5 years old were more commonly infected by assemblage B only (OR = 2·74; 95% CI: 1·15–6·51; P = 0·020) than assemblage A only (Table 1). Additionally, females aged ≤ 5 years old had a greater risk of assemblage B-only infection than their male counterparts (OR = 2·61; 95% CI: 1·12–6·07; P = 0.001).
Distribution of G. intestinalis assemblages based on age (n = 145) and sex (n = 154)

* P-value < 0·05 is significant.
Clinical and travel history of G. intestinalis cases
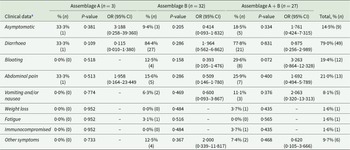
Overall, no significant association was found between symptoms and infection with specific G. intestinalis assemblages. Of the 9 asymptomatic cases, 5 had mixed assemblages A + B, 3 had assemblage B and 1 had assemblage A. Common clinical symptoms identified from the G. intestinalis cases included diarrhoea (79 0%, n = 49), abdominal pain (21 0%, n = 13) and bloating (19 4%, n = 12) (Table 2). Vomiting and/or nausea, weight loss and fatigue were also reported, albeit rarely (8 1, 1 6 and 1 6%, respectively). A few individuals positive for G. intestinalis also reported being asymptomatic (14 5% (n = 9)) at the time of sampling. These included cases who had recently travelled overseas and/or arrived in the country as a refugee (n = 3), as well as family members, friends and household members of confirmed giardiasis cases (n = 5). The remaining asymptomatic case (1 6%) had reported having a lowered immunity. A total of 62 cases whose isolates were successfully genotyped (and who did not have co-infections with other enteropathogens) had clinical data available (Table 2). More cases from whom assemblage B and mixed assemblages A + B were identified reported symptoms (n = 32 and n = 27, respectively); however, comparisons could not be made with assemblage A as clinical data was only available for 3 individuals. Regarding the travel history of cases, 7 8% (n = 10/129) of cases reported travelling overseas prior to illness onset. Out of the 10 cases, none were infected with assemblage A only, 20 0% had assemblage B and the remaining 80 0% were found to have mixed A + B infection. Additionally, those who had travelled overseas were 6 times more likely to be infected with mixed assemblages A + B (OR = 5 917; 95% CI: 1 20–29.08, P = 0.02) as opposed to single infections.
G. intestinalis assemblages and recorded clinical symptoms

a Excluding clinical data for samples with co-infections with other enteropathogens.
Spatial and seasonal distribution of G. intestinalis assemblages across NSW LHDs
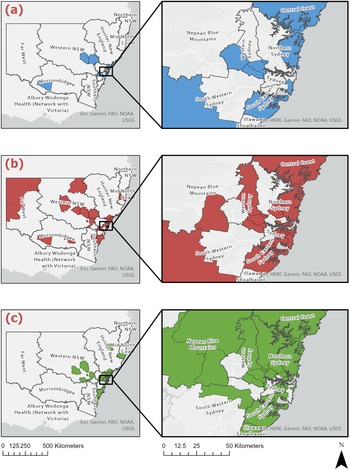
Of the 169 giardiasis cases selected to reflect the broader population, only 141 had valid geographic information available and were therefore included in the spatial mapping across NSW (Figure 2), aggregated into assemblage groups. Cases appeared in metropolitan areas of Sydney, the Blue Mountains (west of Sydney) and regional inland and coastal centres of NSW. Most cases, however, were from metropolitan areas, in particular the Northern Sydney district. The surrounding areas of Inner-West Sydney also showed a high frequency of giardiasis infection (Figure 2). In the regional/rural areas, cases were often seen in the Newcastle and lower Hunter region as well as mid-western regional locations including Dubbo, Orange and Bathurst. Allocation of postcode data to LHDs supported this: a total of 67 4% (n = 95) samples were from metropolitan LHDs (see Table 3). The Northern Sydney LHD accounted for over a quarter (29 5%, n = 28) of all metropolitan samples, and of these 35 7% (n = 10) were from the Northern Beaches LGA. A total of 32 6% (n = 46) of cases were from rural/regional LHDs, including Western NSW (41 3%, n = 19), Hunter New England (26 1%, n = 12) and the Far West (10 9%, n = 5). A smaller number of regional cases were from Southern NSW (n = 4), Murrumbidgee (n = 3) and Mid North Coast (n = 3).
Geospatial distribution of G. intestinalis assemblages A, B and A + B across NSW local health districts (n = 141). This figure shows the geospatial distribution of (a) G. intestinalis assemblage A (blue), (b) G. intestinalis assemblage B (red) and (c) G. intestinalis mixed-assemblage A + B (green) across NSW local health districts.

Distribution of G. intestinalis assemblages based on region of residence in NSW (n = 141)

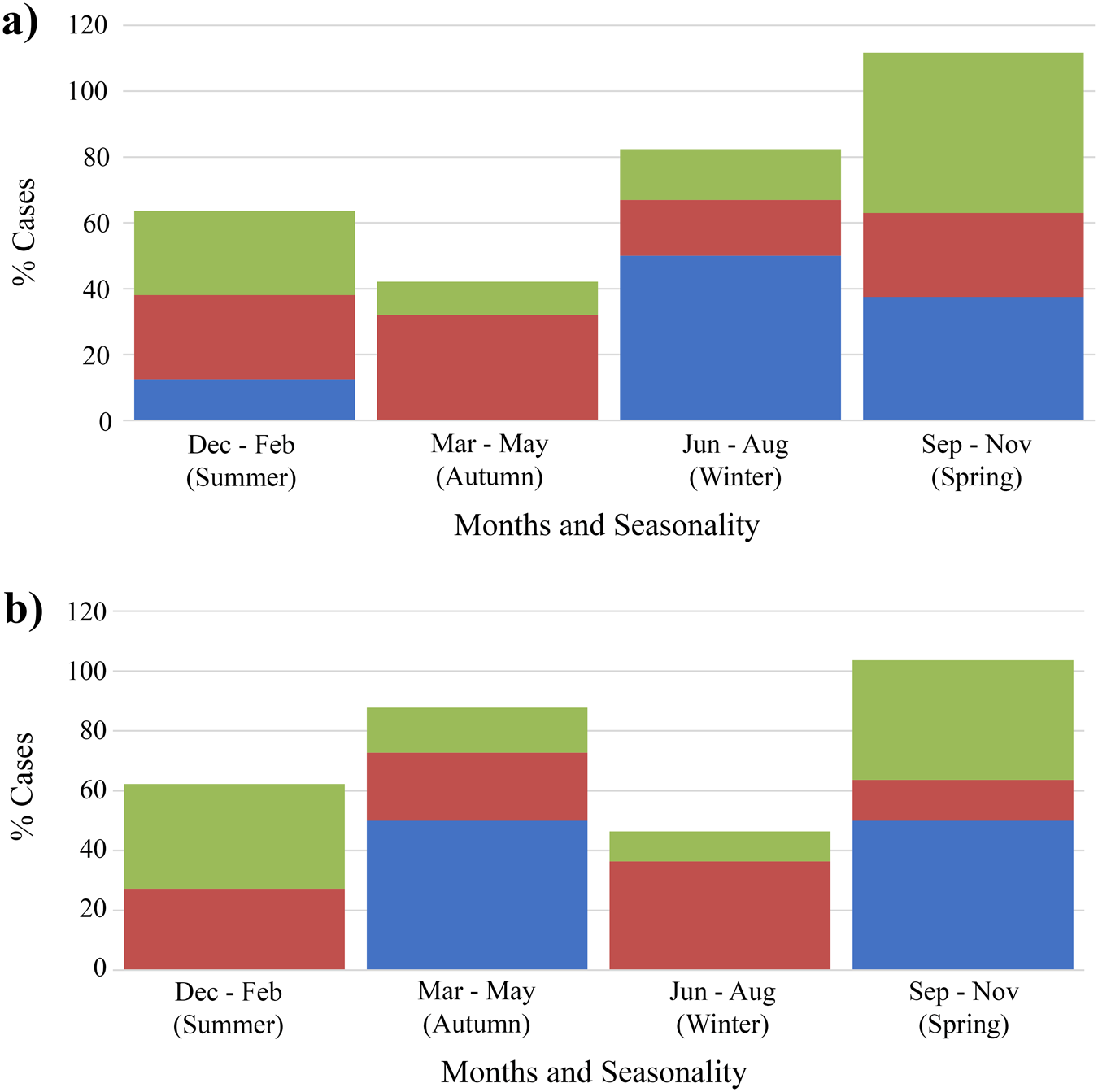
Using ArcGIS, the G. intestinalis assemblages (n = 141) were mapped to LGA boundaries. The maps (Figure 2) showed that the assemblages were distributed across the entirety of NSW and did not show obvious geographic clustering. There were no correlations found between region of residence and specific assemblage type, and most infections were commonly reported in metropolitan regions of Sydney and the eastern coast of NSW. There were, however, significant associations found between seasons and assemblage B only infections (P = 0·004) as well as mixed assemblage infections (P = 0·005). In metropolitan NSW, single assemblage A infections were not observed in autumn, and were only detected in summer, spring and winter (P = 0·048) (Figure 3). Additionally, mixed assemblage infections were most often found in spring (P = 0·004), whilst single assemblage B infections made up most cases in summer, autumn and winter. In rural/regional areas of NSW (Figure 3), single assemblage B infections made up 31·0% of all cases, and peaked in autumn and winter.
Distributions of G. intestinalis assemblages and seasonal dispersal across metropolitan and rural/ regional local health districts. This figure shows the seasonal distribution of genotyped cases occurring in (a) metropolitan local health districts (n = 94) and (b) regional/ rural local health districts (n = 46) in NSW between 2016 and 2019. Assemblage A, blue; assemblage B, red; mixed-assemblage A + B, green.

Discussion
The present study aimed to identify the genetic diversity of G. intestinalis in humans across metropolitan and rural/regional areas of NSW, Australia during 2016–2019. Faecal samples that were positive for G. intestinalis were collected (n = 410) from participating hospital and private pathology laboratories participating in the study. From this pool, a representative subset of samples was chosen for genotyping (n = 169) and 95 9% (n = 162) were successfully amplified using PCR. The genotyping results indicate the presence of both assemblages A and B in NSW. Assemblage B infections were predominant, accounting for 46 9%, followed by mixed infections of assemblages A and B at 43 8% and single assemblage A infections at 9 3%. There is still a lot of controversy surrounding the distribution of assemblages around the world mainly because the results are difficult to compare. Studies in Australia, Austria and Sweden determined that assemblage B was the most dominant (Read et al. Reference Read, Walters, Robertson and Thompson2002; Lee et al. Reference Lee, Lee, Park, Yong and Hwang2006; Lebbad et al. Reference Lebbad, Petersson, Karlsson, Botero-Kleiven, Andersson, Svenungsson and Svärd2011). Assemblage B infections are not only more virulent than assemblage A, but have been reported more commonly in giardiasis outbreaks (Xiao and Feng Reference Xiao and Feng2017). The higher virulence of assemblage B infections suggests that those infected are more likely to seek medical assistance, which in turn could account for the higher detection rates. High re-infection rates have also been attributed to assemblage B infections, wherein poor hygiene and environmental contamination allow this assemblage to recirculate within a community (Thompson Reference Thompson2008).
Of note is the high number of mixed-assemblage infections identified (43 8%). In other studies, mixed-assemblage infections are rarely identified accurately, and most studies report only a 3–10% prevalence (Kohli et al. Reference Kohli, Bushen, Pinkerton, Houpt, Newman, Sears, Lima and Guerrant2008). PCR-based studies that use assemblage-specific primers targeting the tpi locus are often more likely to observe mixed-assemblage infections in comparison to other PCR methodologies (Huey et al. Reference Huey, Mahdy, Al-Mekhlafi, Nasr, Lim, Mahmud and Surin2013; Elhadad et al. Reference Elhadad, Abdo, Tolba, Salem, Mohamed, El-Abd and El-Taweel2021). In part, this is due to the polymorphic nature of the tpi markers that allow them to reliably distinguish between assemblage A and B isolates. While standard primer sets may overlook mixed assemblage cases due to the variable proportions of assemblage A and B DNA, assemblage-specific primers excel in this regard (Zajaczkowski et al. Reference Zajaczkowski, Lee, Fletcher-Lartey, Alexander, Mahimbo, Stark and Ellis2021). This was confirmed in the present study, where the tpi-PCR successfully genotyped 27 2% mixed assemblage A + B infections, in comparison to the 8 1% obtained by the SSU-rRNA-PCR assay.
In the present study, cases of G. intestinalis infection were found across all age groups ranging from 0 years to over 70 years. Data show that the number of cases peaked at ages ≤ 5 years and 30–49 years regardless of gender (Figure 1). Similar bimodal age distributions have been observed in G. intestinalis surveillance reports from the USA (Yoder et al. Reference Yoder, Gargano, Wallace and Beach2012; Painter et al. Reference Painter, Gargano, Collier and Yoder2015) and England (Breathnach et al. Reference Breathnach, McHugh and Butcher2010). Although the distribution of G. intestinalis assemblages was found across all age groups in NSW, adults aged in their 30s and 40s, and children under 5 years of age maintained a higher prevalence of assemblage B. In fact, children under 5 years old were more commonly infected by assemblage B (OR = 2 74; 95% CI: 1 15–6 51; P = 0 020) than assemblage A. This strong association between assemblage B infection and children under 5 years might be indicative of age-specific risk factors and transmission routes. The exposure to assemblage B organisms in children can occur either through high-risk activities such as day-care attendance or schooling, as well as poor hygiene behaviours (Thompson Reference Thompson2000; Oliveira-Arbex et al. Reference Oliveira-Arbex, David, Oliveira-Sequeira, Bittencourt and Guimaraes2016). Children with assemblage B infections have also demonstrated a higher level of cyst shedding, which would facilitate a faster spread within institutional settings and areas where children frequent (Kohli et al. Reference Kohli, Bushen, Pinkerton, Houpt, Newman, Sears, Lima and Guerrant2008). In Spain, G. intestinalis-positive children were 10 times more likely to be infected with assemblage B in comparison to adults (Wang et al. Reference Wang, Gonzalez-Moreno, Roellig, Oliver, Huguet, Guo, Feng and Xiao2019). Additionally, children may play a critical role in ongoing transmission cycles by facilitating secondary spread to family members, day-care centre staff and other children. Nappy changing and toilet training are particularly high-risk activities for parents and caregivers, due to frequent contact with faecal material and possible environmental contamination. The burden of exposure from infected children likely contributes to the increased rates of assemblage B infection observed in parent-aged adults. Supporting this, a Brazilian study observed a predominance of assemblage B infection in middle-aged adults aged 30 to 39 years old (Faria et al. Reference Faria, Zanini, Dias, da Silva and Sousa2016), a pattern also reported in England (Minetti et al. Reference Minetti, Lamden, Durband, Cheesbrough, Fox and Wastling2015). Co-infecting pathogens were detected in nearly half (49 1%, n = 83) of all-G. intestinalis-positive faecal samples. Infection of G. intestinalis with concomitant infections with a variety of gut bacteria, viruses and parasites are incredibly common in most countries. Most co-infections were parasitic (56 6%, n = 47) and bacterial (12 0%, n = 10). Overall, the most common pathogens detected were the enteric protozoa B. hominis (31 3%, n = 25) and D. fragilis (15 0%, n = 12), followed by Campylobacter spp. (5 0%, n = 4) and Enterovirus (5 0%, n = 4). Infection of G. intestinalis with concomitant infections with a variety of gut bacteria, viruses and parasites are incredibly common in most countries. In India, Giardia infections with Vibrio cholerae and rotavirus were commonly identified in children aged under 10 years old (Mukherjee Reference Mukherjee2014) while in Nicaragua, the majority of G. intestinalis cases (70 4%) were co-infected with either Norovirus, Sapovirus or enteropathogenic Escherichia coli (EPEC). (Becker-Dreps et al. Reference Becker-Dreps, Bucardo, Vilchez, Zambrana, Liu, Weber, Peña, Barclay, Vinjé, Hudgens, Nordgren, Svensson, Morgan, Espinoza and Paniagua2014) In the USA, multiple parasites including G. intestinalis, Cryptosporidium spp., and Entamoeba spp. were responsible for drinking water outbreaks (Craun et al. Reference Craun, Brunkard, Yoder, Roberts, Carpenter, Wade, Calderon, Roberts, Beach and Roy2010). Interestingly, a study in Uganda observed a link between Giardia assemblage B and Helicobacter pylori infection (Ankarklev et al. Reference Ankarklev, Hestvik, Lebbad, Lindh, Kaddu-Mulindwa, Andersson, Tylleskär, Tumwine and Svärd2012). There are limited studies on the associations between G. intestinalis assemblages and co-infecting pathogens, so comparisons remain difficult to make. In the present study, no associations were found between assemblage type and co-infecting pathogen. The high level of co-infections can be explained as most enteric pathogens are transmitted via the same route of infection: the faecal-oral route. This underscores the value of continued examination of faecal specimen from symptomatic persons for multiple pathogens in developed settings such as Australia, where the practice appears to be diminishing in clinical settings.
In the present study, it was found that clinical symptoms were not associated with assemblage type. This was consistent with previous studies in Brazil (Kohli et al. Reference Kohli, Bushen, Pinkerton, Houpt, Newman, Sears, Lima and Guerrant2008), Iran (Bahrami et al. Reference Bahrami, Zamini, Haghighi and Khademerfan2017; Kashinahanji et al. Reference Kashinahanji, Haghighi, Bahrami, Fallah, Saidijam, Matini and Maghsood2019; Rafiei et al. Reference Rafiei, Baghlaninezhad, Köster, Bailo, Hernández de Mingo, Carmena and Panabad E, Beiromvand2020), Thailand (Tungtrongchitr et al. Reference Tungtrongchitr, Sookrung, Indrawattana, Kwangsi, Ongrotchanakun and Chaicumpa2010) and China (Liu et al. Reference Liu, Zhang, Zhang, Wang, Li, Shu, Zhang, Shen, Zhang and Ling2012). Despite this, there have been other studies that have reported a close association between assemblages and clinical symptoms (Helmy et al. Reference Helmy, Abdel-Fattah and Rashed2009; Al-Mohammed Reference Al-Mohammed2011; Pestehchian et al. Reference Pestehchian, Rasekh, Babaei, Yousefi, Eskandarian, Kazemi and Akbari2012). Assemblage B has been associated with severe diarrhoea, vomiting, abdominal pain and bloating (Al-Mohammed Reference Al-Mohammed2011; ElBakri et al. Reference ElBakri, Samie, Bessong, Potgieter and Odeh2014; Hussein et al. Reference Hussein, Ismail, Mokhtar, Mohamed and Saad2017; Wang et al. Reference Wang, Gonzalez-Moreno, Roellig, Oliver, Huguet, Guo, Feng and Xiao2019), although it is equally plausible that since younger children and their middle-aged parents appear to be predominantly affected with this assemblage and are more likely to seek care, these assemblages become overrepresented among notified cases. In other studies, assemblage A has been affiliated with more serious clinical symptoms (Read et al. Reference Read, Walters, Robertson and Thompson2002; Haque et al. Reference Haque, Roy, Kabir, Stroup, Mondal and Houpt2005; Breathnach et al. Reference Breathnach, McHugh and Butcher2010; Sarkari et al. Reference Sarkari, Ashrafmansori, Hatam, Motazedian, Asgari and Mohammadpour2012; El Basha et al. Reference El Basha, Zaki, Hassanin, Rehan and Omran2016). It remains difficult to determine a true correlation between assemblages and symptoms. It may be that the virulence of assemblages A and B in humans relies on a variety of factors, including human host age and gender, parasite growth rates, metabolic products or toxins and even drug resistance.
Another interesting finding was that only 8 0% of G. intestinalis positive cases reported travelling overseas prior to illness onset, suggesting that most giardiasis cases in NSW are a result of endemic transmission. There is a misconception that G. intestinalis infection in industrialized countries is mainly associated with international travel to developing nations. Several studies have observed that most giardiasis cases in industrialized countries are in fact a result of endemic transmission and local risk factors (Espelage et al. Reference Espelage, an der Heiden, Stark and Alpers2010; Plutzer et al. Reference Plutzer, Ongerth and Karanis2010; Woschke et al. Reference Woschke, Faber, Stark, Holtfreter, Mockenhaupt, Richter, Regnath, Sobottka, Reiter-Owona, Diefenbach and Gosten-Heinrich2021). In the present study, individuals with a history of overseas travel were 6 times more likely to be infected with mixed assemblages A and B (OR = 5 917; 95% CI: 1 20–29 08, P = 0 02) as opposed to being infected with a single assemblage infection. This is a novel finding which has not been observed elsewhere, and it remains important to investigate this further. However, a recent study (Samie et al. Reference Samie, Tanih, Seisa, Seheri, Mphahlele, ElBakri and Mbati2020) noted that the occurrence of mixed-assemblage infections is higher in developing countries as opposed to developed regions of the world. Exposure to conditions where environments are contaminated with human faeces, have poorer access to or less well-maintained hygiene facilities may be more common in developing settings, which increases the risk of both assemblages co-circulating in communities and being picked up by travellers.
Analyses of 141 cases of sporadic human giardiasis showed that infections were widely dispersed across eastern regions of Sydney and NSW, where the majority of the NSW population resides (Australian Bureau of Statistics 2017) (Figure 2). Among the NSW LHDs, most (67 4%) sporadic cases occurred in metropolitan LHDs. These findings are consistent with historical state-wide surveillance trends that identified significant positive associations between area-level advantage and an increased likelihood of giardiasis notifications (Mazumdar et al. Reference Mazumdar, Fletcher-Lartey, Zajaczkowski and Jalaludin2020). However, it cannot be ignored that the high incidence rates of giardiasis detected in urban Sydney may also be artefactual, particularly as these locations often have highly transient populations. It must also be considered that individuals residing in metropolitan areas have better access to primary healthcare facilities and greater access or inclination to submitting stool samples for testing when compared with those living in rural areas. In addition to this, densely populated cities such as Sydney have a higher risk of exposure to an infected individual, whether that be through contaminated environment, wastewater, sewage or recreational waters or transmission through day-care centres, schools, and other institutional settings. The plausibility of this was confirmed by a study in the USA that found a positive correlation between giardiasis prevalence and population density and population size (Dreelin et al. Reference Dreelin, Ives, Molloy and Rose2014).
Seasonal trends in the dispersal patterns of assemblages A and B were also observed (Figure 3.a and Figure 3.b). Single assemblage A cases were not detected in metropolitan LHDs during autumn (P = 0 048). Alternatively, single assemblage A cases were also missing in regional areas across summer and winter. This finding may be artefactual and is likely due to the lower numbers of assemblage A cases identified throughout the study. The low number of single Assemblage A infections may also reflect methodological limitations, particularly the reduced sensitivity of some assays to detect minor assemblage components in mixed infections. Rather than indicating a true scarcity of Assemblage A, it is possible that these infections are under-detected when co-occurring with Assemblage B. Additionally, Assemblage A may cause milder or more asymptomatic infections, meaning individuals are less likely to seek diagnosis. This could help explain why our study observed very few single A infections but a relatively high number of A and B mixed infections. Overall, the giardiasis infection rates peaked in spring and dropped in early autumn and winter. This is consistent with other reports of seasonality (Hoque et al. Reference Hoque, Hope and Scragg2002, Reference Hoque, Hope, Scragg, Baker and Shrestha2004). A peak incidence of giardiasis in NSW during October through to December coincides with high prevalence of outdoor and higher risk activities in these warmer months.
Conclusions
In summary, this study provides new insights into the molecular diversity of G. intestinalis in NSW, Australia, and helps to inform enhanced surveillance and prevention strategies in developed metropolitan areas. During the study period, a higher prevalence of assemblage B was observed among human cases in NSW. Factors which possibly influence this higher incidence in NSW may be behavioural, climatic, environmental or related to the virulence or assemblage of the parasite. Higher numbers of mixed assemblage infections were also identified, which is a novel finding for a developed country like Australia. The distribution of assemblages A and B remained relatively uniform across genders and no clear differences were observed in clinical presentation between assemblages; however, assemblage B was more commonly observed among children. Further high-powered studies are needed to investigate the prevalence and clinical manifestations of assemblage B in children. While most giardiasis cases were transmitted locally, those individuals who had reported travelling overseas prior to illness onset were 6 times more likely to be infected with mixed assemblages A and B as opposed to single assemblages. This novel discovery underscores the importance of additional investigation into ‘travel’ as a risk factor for Australians, particularly delving into the differences observed in giardiasis transmission dynamics between endemic and international cases. Among metropolitan LHDs, G. intestinalis cases were consistently identified in the Nepean Blue Mountains, Northern Sydney, Western Sydney, South-eastern Sydney, Sydney CBD and Central Coast regions, which persisted throughout all seasons, and have highlighted these locations as potential disease hotspots in NSW. It remains essential to improve our knowledge of giardiasis and its molecular epidemiology among host populations; to help better inform surveillance strategies and response actions aimed at preventing further spread of infection. Further studies involving the geospatial and spatiotemporal distribution of G. intestinalis assemblages are recommended, and in particular targeting the metropolitan and urban areas of NSW.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182025100991.
Data availability statement
The data that support the findings of this study are available from the corresponding author, [author initials], upon reasonable request.
Acknowledgements
The authors wish to acknowledge the assistance of the Centre for Infectious Diseases and Microbiology (CIDM) at Westmead Hospital, Sydpath at St. Vincent’s Hospital as well as Laverty Pathology and Douglass Hanly Moir Pathology (DHM) for providing the samples and patient data required for this study.
Author contributions
J.E., D.S. R.L. and S. F-L. conceived and designed the study. D. S., R. L. and M. W. organised sample and data collection. P. Z. conducted all experiments and with A. M., K. A. and S. F-L. completed data analyses. P. Z. wrote the manuscript with revisions from J. E., R.L. D. S. and S. F-L. All authors approved the final draft.
Financial support
Funding to support this research was provided by (a) the NSW Ministry of Health under the NSW Health PhD Scholarship Program, (b) Department of Microbiology, St Vincent’s Hospital Sydney.
Competing interests
None.
Ethical approval
Ethics approval for the conduct of this study was received from the South-Western Sydney Local Health District Human Research Ethics Committee (HREC) which is accredited by the NSW Ministry of Health (HREC approval number: HE18/059 LNR), and the University of Technology Sydney (UTS approval number: ETH21-5951).