Introduction
Anoplocephala perfoliata is recognized as the most common cestode to colonize the gastrointestinal tract of horses worldwide and is responsible for significant clinical disease (Gasser et al., Reference Gasser, Williamson and Beveridge2005). At present, a dearth of molecular research on A. perfoliata restricts our understanding of the epidemiology and pathogenicity of infections and limits the potential to develop control tools such as vaccines and anthelmintics. For example, molecular diagnostic methods have demonstrated that these parasites are considerably more prevalent than previous coprological methods have suggested (Gasser et al., Reference Gasser, Williamson and Beveridge2005; Lightbody et al., Reference Lightbody, Matthews, Kemp-Symonds, Lambert and Austin2018). Furthermore, necropsy studies in areas of Europe and the US demonstrate prevalence has changed very little over the last 2 decades (Owen et al., Reference Owen, Jagger and Quan-Taylor1988; Fogarty et al., Reference Fogarty, del Piero, Purnell and Mosurski1994; Meana et al., Reference Meana, Luzon, Corchero and Gómez-Bautista1998, Reference Meana, Pato, Martín, Mateos, Pérez-García and Luzón2005; Lyons et al., Reference Lyons, Swerczek, Tolliver, Bair, Drudge and Ennis2000, Reference Lyons, Bolin, Bryant, Cassone, Jackson, Janes, Kennedy, Loynachan, Boll, Burkhardt and Langlois2018; Lightbody et al., Reference Lightbody, Davis and Austin2016). Finally, there are increased concerns for induced disease, such as colic, which can often be fatal (Gasser et al., Reference Gasser, Williamson and Beveridge2005; Back et al., Reference Back, Nyman and Osterman Lind2013). A. perfoliata has long been recognized as a serious equine pathogen (Proudman et al., Reference Proudman, French and Trees1998; Proudman and Trees, Reference Proudman and Trees1999). However, infection has also been regarded as unimportant among surveillance programs (Nielsen et al., Reference Nielsen, Monrad and Olsen2006). Despite this, reports continue to demonstrate that infection poses significant risk to equine health. Indeed, intensifying burdens correlates with an increasing severity of inflammation, intestinal lesions, ulceration, thickening of the mucosa, submucosa, hypertrophy of circular muscle and colic (Williamson et al., Reference Williamson, Gasser, Middleton and Beveridge1997; Proudman et al., Reference Proudman, French and Trees1998; Proudman and Trees, Reference Proudman and Trees1999; Pavone et al., Reference Pavone, Veronesi, Genchi, Fioretti, Brianti and Mandara2011; Lawson et al., Reference Lawson, Pittaway, Sparrow, Balkwill, Coles, Tilley and Wilson2019).
To date, treatment of A. perfoliata relies solely on anthelmintics, namely pyrantel (PRT) or praziquantel (PZQ), to which resistance has not yet been published. Importantly, improved diagnostics have paved the way for reducing the excessive use of anthelmintics (Lightbody et al., Reference Lightbody, Matthews, Kemp-Symonds, Lambert and Austin2018). The discontinuation of the equine tapeworm wormer Equitape® in the U.K. means PZQ will only be available at equine outlets in combination with ivermectin or moxidectin used to treat Cyathostomins, potentially compromising future helminth control. However, PZQ is available as a single active under a ‘special’ consideration from veterinarians. Increasing use of combination anthelmintics as a control option could exacerbate existing resistance occurring within cyathostomins (Nielsen et al., Reference Nielsen, Banahan and Kaplan2020, Reference Nielsen, Littman, Orzech and Ripley2022). These concerns for increased helminth anthelmintic resistance, in addition to the impact of A. perfoliata on equine health, make it imperative to maximize the efficiency of control options.
Given the importance of anthelmintic control of tapeworms, and wider helminth species, there has been increasing interest in using functional genomic approaches, such as proteomics, to understand how parasitic helminths metabolize anthelmintics. Of note has been the proteomic scrutinization of the glutathione transferase protein superfamilies (GST) from a number of helminths (Van Rossum et al., Reference Van Rossum, Brophy, Tait, Barrett and Jefferies2001; Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). GSTs, as phase II detoxification enzymes, have a widespread distribution across parasite tissues, potentially accounting for up to 3% of soluble cytosolic proteins in cestodes (Brophy and Barrett, Reference Brophy and Barrett1990a).
Helminth Sigma, Mu and Omega class GSTs are renowned for their importance in the host–parasite interaction, such as a major detoxification of xenobiotic compounds and endogenously derived toxins (Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Stuart et al., Reference Stuart, Zwaanswijk, Mackintosh, Witikornkul, Brophy and Morphew2021). However, these detoxification mechanisms could also provide protection from oxidative membrane damage via immune assault, internal toxic compounds of helminth metabolism, as well as xenobiotics, such as anthelmintics (van Rossum et al., Reference van Rossum, Jefferies, Rijsewijk, LaCourse, Teesdale-Spittle, Barrett, Tait and Brophy2004; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016). Members of the Omega class of GSTs are also suggested to major roles in the protection of the reproductive system during maturation and as an oxidative response (Board et al., Reference Board, Coggan, Chelvanayagam, Easteal, Jermiin, Schulte, Danley, Hoth, Griffor, Kamath and Rosner2000; Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; Burmeister et al., Reference Burmeister, LÜersen, Heinick, Hussein, Domagalski, Walter and Liebau2008; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Meng et al., Reference Meng, Zhang, Liu, Guo and Xu2014; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016). Furthermore, helminth GSTs have further adapted to the host environment and demonstrate immunomodulatory activities to support establishment and duration (Sato and Kamiya, Reference Sato and Kamiya1995; LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012; Wang et al., Reference Wang, Zhao, Zhang, Zhang, Li, Shang, Ning, Ji, Xia, Cai and Qiao2022). In particular, Sigma class GSTs are involved in prostaglandin synthase activity and a role in the modulation of the host immune response (Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012). Consequently, research has shown evidence that helminth GSTs may have key roles in the establishment of infection and the development of anthelmintic resistance, as demonstrated in the parasitic flatworm Fasciola hepatica (Chemale et al., Reference Chemale, Perally, LaCourse, Prescott, Jones, Ward, Meaney, Hoey, Brennan, Fairweather and Trudgett2010; Scarcella et al., Reference Scarcella, Lamenza, Virkel and Solana2012). To this end, parasitic flatworm GSTs have shown potential as novel a vaccine candidate (Capron et al., Reference Capron, Riveau, Capron and Trottein2005; Dowling et al., Reference Dowling, Hamilton, Donnelly, La Course, Brophy, Dalton and O'Neill2010; LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012).
Global and sub-proteomic studies on the somatic or extracellular proteomes of Anoplocephalidae are limited. However, the recent development of a transcriptome has provided an invaluable source of information for comprehensive molecular analysis of A. perfoliata (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021). This development has led to the proteomic profiling of the A. perfoliata secretome including the free excretory/secretory proteins (ESP) and A. perfoliata derived extracellular vesicles (EVs) (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021); demonstrating proteins released into the host milieu contain a variety of potential immune modulator molecules such as calpains, enolases and importantly GSTs. Thus, the foundation for utilizing in-depth proteomic approaches to better understand A. perfoliata at a molecular level has been provided. This allows the potential identification of the GST superfamily present in this parasitic cestode as has been previously employed for a number of helminths and protein superfamilies (Van Rossum et al., Reference Van Rossum, Brophy, Tait, Barrett and Jefferies2001; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Morphew et al., Reference Morphew, Wilkinson, Mackintosh, Jahndel, Paterson, McVeigh, Abbas Abidi, Saifullah, Raman, Ravikumar and LaCourse2016). Here, global and sub-proteomic approaches were used to investigate the presence of GSTs derived from the equine cestode, A. perfoliata. In doing so, it provides the first evidence of functional expression of 3 A. perfoliata GST (ApGST) classes, namely Mu, Sigma and Omega class GSTs, at the protein level.
Materials and methods
Bioinformatic analysis of the A. perfoliata transcriptome
Potential GST protein sequences were initially identified within the A. perfoliata transcriptome (available at https://sequenceserver.ibers.aber.ac.uk/) according to (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021). Briefly, known confirmed GST protein sequences were used in a tBLASTn (Altschul et al., Reference Altschul, Madden, Schäffer, Zhang, Zhang, Miller and Lipman1997) against the in-house A. perfoliata transcriptome (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021) through BioEdit Sequence Alignment Editor (Version 7.2.6.1; Hall, Reference Hall1999) with the number of expected hits of similar quality (e-value) cut-off set at 1.0 × 10−15. Protein sequences of recognized GST superfamily classes included Alpha, Delta, Epsilon, Zeta, Theta, Mu, Nu, Pi, Sigma and Omega classes from mammals, insects and helminths; all of which were retrieved from Genbank and NCBI Reference Sequence (http://www.ncbi.nlm.nih.gov/) (Supplementary Table S1). Known sequences also included the confirmed Sigma class GSTs identified by Wititkornkul et al. (Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021). The top hit sequences homologous to recognized GST sequences (lowest e-value) were taken as the representative A. perfoliata GST sequences, and subsequently translated into the protein sequences with ExPASy Translate tools (https://web.expasy.org/translate/; Gasteiger et al., Reference Gasteiger, Gattiker, Hoogland, Ivanyi, Appel and Bairoch2003). Selected protein sequences were confirmed as a potential GST sequence by protein super-families, domain prediction and functional site analysis using InterProScan databases (version 77.0; http://www.ebi.ac.uk/interpro/; Jones et al., Reference Jones, Binns, Chang, Fraser, Li, McAnulla, McWilliam, Maslen, Mitchell, Nuka and Pesseat2014; Blum et al., Reference Blum, Chang, Chuguransky, Grego, Kandasaamy, Mitchell, Nuka, Paysan-Lafosse, Qureshi, Raj and Richardson2021) followed by searching against the NCBI (nr) protein database using a protein query (BLASTp; https://blast.ncbi.nlm.nih.gov/Blast.cgi; Altschul et al., Reference Altschul, Madden, Schäffer, Zhang, Zhang, Miller and Lipman1997). The resulting InterPro domains classified as a Glutathione transferase superfamily with either C-terminal or N-terminal domain (IPR036282 and IPR004045 respectively) were kept as a unique GST protein sequence. Non-GST protein sequences, short protein sequences (less than 100 amino acids) and fragmented split sequences were excluded from future analysis. All unique classified GST sequences, or 1 representative if isoforms were present, were taken as a final A. perfoliata GST representative protein sequence for subsequent phylogenetic analysis.
Multiple sequence alignment and phylogenetic analysis of A. perfoliata GSTs
The resulting A. perfoliata GST representative protein sequences were aligned with recognized GST protein sequences from 18 different species covering a range of mammals, nematodes, trematodes, cestodes and insects (Supplementary Table S1.) using ClustalW through BioEdit Multiple Sequence Alignment Editor (Version 7.2.6.1; Hall, Reference Hall1999). Subsequently, an A. perfoliata GST phylogenetic tree was constructed and visualized in MEGA X (version 10.1.7; Hall, Reference Hall2013; Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018). Reliability of the phylogenetic tree was estimated with 1000 bootstrap replicates, using a Maximum Likelihood (ML) method.
Bioinformatic analysis of Omega class GSTs
To investigate and confirm novel A. perfoliata Omega class GST sequences, potential A. perfoliata Omega class GST sequences clustering in previous phylogenetic analyses were aligned separately using ClustalW multiple sequence alignment and used for the generation of an Omega specific phylogenetic tree as described above. Subsequently, the characteristic secondary structure of potential A. perfoliata Omega class GST sequences was predicted using the Predict Secondary Structure (PSIPRED) Protein Analysis Workbench (PSIPRED 4.0; http://bioinf.cs.ucl.ac.uk/psipred/; Jones, Reference Jones1999; Buchan and Jones, Reference Buchan and Jones2019). Omega class GST homologues were further investigated based on the glutathione (GSH)-binding site characteristic of Omega class GSTs in the N-terminal domain including the proline-rich residues in N-terminal extension (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012), the catalytic cysteine residues characteristic of Omega class GSTs (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016) and Omega class GST signature motifs (Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006).
Parasite collection
Live adult cestodes were manually extracted from the caecum at the ileocecal valve and the surrounding caecal walls of naturally infected horses immediately post-slaughter at a local commercial abattoir in Swindon, U.K. All obtained A. perfoliata were extensively washed in phosphate buffered saline to remove host material, flash-frozen, and stored at – 80°C until further use.
Species verification
For DNA extraction, 20 mg of tissue was collected from 6 selected cestodes from the obtained individuals. DNA extraction was carried out using the DNeasy Blood & Tissue Kit (Qiagen Ltd) according to the manufacturer's instructions, as described previously (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012) and the concentration and purity of the extracted DNA quantified using NanoDrop (ThermoFisher Scientific, USA). Species verification PCR amplification was performed using ITS2 forward primer (5’-TGTGTCGATGAAGAGCGCAG) and ITS2 reverse primer (5’-TGGTTAGTTTCTTTTCCTCCGC), as described previously (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). PCR products were subjected to 1% w/v agarose gel electrophoresis. PCR products were extracted from the gel using a Quick Gel Extraction and PCR Purification Combo Kit (Invitrogen, PureLink®) according to manufacturer's instructions and sequenced in-house using the primers described. Sequenced ITS2 regions were then identified by BLASTn against NCBI database.
Cytosolic protein extraction
Replicates of 10 whole A. perfoliata were homogenized in a glass-glass homogenizer on ice in 20 mm potassium phosphate buffer (pH 7.4) containing 50 mm NaCl, 1% v/v Triton X-100 and a cocktail of protease inhibitors (Roche, Complete Mini, EDTA-free). Homogenized samples were delipidated through glass wool at 1000× g for 2 minutes. Samples were then centrifuged at 100 000× g for 45 minutes at 4°C and the soluble protein fraction collected. Protein concentration of the cytosolic protein was quantified using Bradford reagent (Sigma-Aldrich; Bradford, Reference Bradford1976).
GST purification
GSTs were purified as described by Morphew et al. (Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). Briefly, the prepared cytosolic protein was filtered through 0.45 μm syringe filter prior to being applied to a 1 mL Glutathione (GSH)-agarose affinity column (Sigma-Aldrich). The column was equilibrated with equilibration buffer (10 mm KPO4, 150 mm NaCl, pH 7.4) for 20 minutes at a flow rate of 1 mL/minute (20 × bed volumes). The cytosolic protein was then passed over the GSH-agarose 6 times to ensure maximal GST recovery and the flow through was collected. The column was washed with 20 × bed volumes of equilibration buffer to remove any non-specific bound proteins. GST proteins were eluted from the GSH-agarose using 5 mL of elution buffer (50 mm Tris-HCl, 7.5 mm GSH], pH 6.5). Eluted proteins were concentrated through 10 kDa ultra centrifugal filters (Amicon Ultra, Millipore) according to the manufacturer's instructions with a buffer exchange step to ddH2O. Purified GSTs and cytosolic protein flow through were quantified with Bradford Reagent (Bradford, Reference Bradford1976).
GST functional assays
The CDNB (1-Chloro-2,4-dinitrobenzene) glutathione conjugation model substrate assay was used to measure GST activity (Habig et al., Reference Habig, Pabst and Jakoby1974). The assay was completed in a 1 mL volume, with a final concentration of 100 mm potassium phosphate buffer (pH 6.5), 1 mm reduced glutathione, 1 mm CDNB and absorbance change measured at 340 nm for 3 min at 20°C. A second model substrate, ethacrynic acid, was also tested according to the method previously described (LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012) using 1 mm GSH and 0.08 mm ethacrynic acid (EA) at pH 6.5.
Protein precipitation
Whole cytosolic protein and purified GSTs were subjected to precipitation prior to electrophoresis. Briefly, a total of 100 μg of somatic protein and 20 μg of purified GSTs were precipitated with 20% v/v trichloroacetic acid (TCA) in ice cold acetone. After 1 h incubation at −20°C, samples were centrifuged at 21 000 × g to pellet proteins. Each protein pellet was then washed in ice cold 100% v/v acetone 3 times and air dried at −20°C for 15 mins. Precipitated cytosolic and GST samples were then resolubilized in isoelectric focusing buffer containing 8 M Urea, 2% w/v CHAPS, 33 mm DTT and 0.5% v/v ampholytes (pH 3–10).
Gel electrophoresis
GST protein purification fractions or whole A. perfoliata cytosolic samples were run on 12.5% v/v 1D SDS-PAGE gels using the Protean® II xi 2-D Cell (Bio-Rad). Protein samples were mixed 1:1 with SDS loading buffer (0.2 M Tris-HCl, 8% SDS, 40% Glycerol, 0.02% Bronophenol blue, 50 mm DTT, pH 6.8), heated at 95°C for 5 mins and loaded. 1D gels were run at 70 V for 30 minutes followed by 150 V until completion. For 2D SDS-PAGE, GST samples were rehydrated and focused on 7 cm pH 3 − 10NL IPG strips to 10 000 Vh, using a Protean IEF Cell (Bio-Rad). Following focusing, IPG strips were equilibrated for 15 minutes in equilibration buffer (30% v/v glycerol, 6 M urea, 1% w/v DTT) followed by 15 min in alkylating equilibration buffer (30% v/v glycerol, 6 M urea, 4% w/v iodoacetamide). Equilibrated strips were run on 12.5% SDS-PAGE gels at 70 V for 30 minutes and 150 V until completion with Protean® II xi 2-D Cell (Bio-Rad). All gels were fixed for 1 hour in 10% v/v acetic acid and 40% v/v ethanol followed by Colloidal Coomassie blue staining (Phastgel Blue R, Amersham Biosciences) overnight. Gels were de-stained in 1% v/v acetic acid and imaged using a GS-800 calibrated densitometer (BioRad). Individual GST protein spots observed on 2D SDS PAGE were detected and quantified using Progenesis PG220, software version 2006 (Nonlinear Dynamics Ltd.), as described previously (Morphew et al., Reference Morphew, MacKintosh, Hart, Prescott, LaCourse and Brophy2014;, Reference Morphew, Wilkinson, Mackintosh, Jahndel, Paterson, McVeigh, Abbas Abidi, Saifullah, Raman, Ravikumar and LaCourse2016).
GST classification via immunoblotting
A. perfoliata cytosolic protein and purified GSTs were run on 1D or 2D SDS PAGE gels. Proteins were then transferred to a nitrocellulose membrane over 2 hours at 40 V according to the method of Towbin et al. (Reference Towbin, Staehelint and Gordon1979). Post transfer, membranes were stained with amido black (0.1% w/v Amido black, 10% v/v acetic acid and 25% v/v isopropanol) to monitor protein transfer, then de-stained 10% v/v acetic acid and 25% v/v isopropanol. De-stained membranes were washed with Tris buffered saline with Tween-20 (TTBS: 20 mm Tris-HCl,154 mm NaCl, 1% v/v Tween 20, pH 7.5) 3 times for 5 minutes. Membranes were then incubated overnight in blocking buffer (5% w/v skimmed milk powder in TTBS) prior to incubating membranes with primary antibodies. Primary antibodies, at 1:5,000 dilution for anti-Mu class antibody (SjGST26; Pharmacia-Biotech 27-4577) and 1: 30 000 for anti-Sigma class GST antibody (FhGST-S1; LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012), were incubated with the membrane at room temperature for 1.5 hours. Membranes were then washed for 5 minutes in Tris buffered saline (TBS; 20 mm Tris-HCl, 154 mm NaCl, pH 7.5) 3 times. Anti-Mu and anti-Sigma probed membranes were incubated in a secondary antibody diluted at 1: 30 000 in TTBS (anti-Rabbit IgG, conjugated with alkaline phosphatase, produced in Goat, Sigma-Aldrich) and immunogenic proteins visualized using BCIP/NBT (5-bromo-4-chloro-3-indoyl phosphate/nitro blue tetrazolium) liquid substrate system (Sigma-Aldrich). Membranes were images using the GS-800 calibrated densitometer (BioRad).
Trypsin digestion and mass spectrometry
A. perfoliata somatic protein resolved on 1DE gels were subjected to GeLC proteomics. Each lane was cut into a series of 10 ‘bands’. Protein spots resolved from the purified GST were manually excised from the 2DE gels and used for classical in gel digestion. Protein bands and spots excised from the 1DE and 2DE gels were subjected to trypsin digestion prior to tandem mass spectrometry.
Gel pieces were de-stained and digested overnight in 10 ng/μL trypsin, as previously described (Rooney et al., Reference Rooney, Williams, Northcote, Frankl, Price, Nisbet, Morphew and Cantacessi2022). Immediately prior to performing tandem mass spectrometry the dried peptides were re-suspended in 20 μL 0.1% v/v formic acid. For 2D SDS PAGE protein spots, the method of Duncan et al. (Reference Duncan, Cutress, Morphew and Brophy2018) was utilized for LC-MSMS. For whole A. perfoliata Orbitrap Fusion™ Tribrid™ mass spectrometer (Thermo Scientific™), with EASY-Spray™ Source, coupled to an UltiMate™ 3000 RSLCnano System (Thermo Scientific™) as per methods of Rooney et al. (Reference Rooney, Williams, Northcote, Frankl, Price, Nisbet, Morphew and Cantacessi2022).
Protein identification and GST localization
Mass Hunter Qualitative Analysis software (V B.06, Agilent Technologies) was used to generate peak lists. Peak lists were then exported as MASCOT Generic Files. Whole worm and purified GST samples were submitted to an MSMS ion search using MASCOT against the recently produced A. perfoliata transcript database (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021). Parameters for the MASCOT search were set as the following conditions: fixed modification of carbamidomethylation, variable modification set for oxidation of methionines, peptide charge to 2+, 3+ and 4+, instruments used ESI-QUAD-TOF (purified GSTs fractions) and ESI-TRAP (Whole somatic proteome). Identified peptides presenting MASCOT ions scores above 44 for whole cytosolic proteins and 45 for purified GSTs were considered to have a significant identity or extensive homology (p < 0.05).
Results
Characterization of novel A. perfoliata GSTs
The initial identification of A. perfoliata GST sequences within an in-house A. perfoliata transcriptome by tBLASTn produced a total of 330 A. perfoliata putative GST sequences based on homology to recognized GST superfamily sequences from mammals, insects and helminths (cutoff 1 × 10–15). Of these potential A. perfoliata GST sequences, each were classified using InterProScan and BLASTp, leading to 309 sequences predicted as GST superfamily membership (IPR040079), of which 59 sequences were initially predicted as Glutathione transferase, Mu class membership (IPR003081) (Supplementary Table S2).
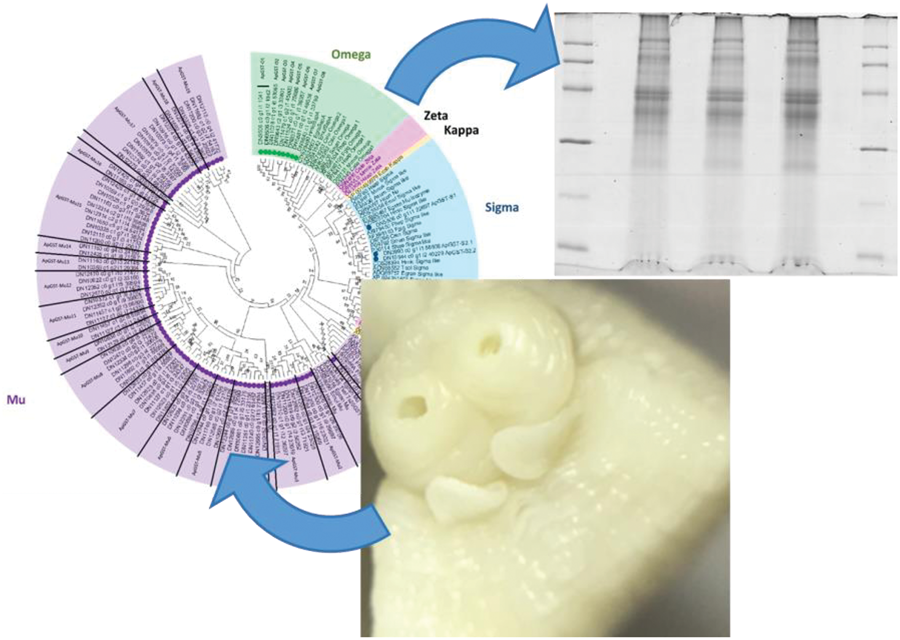
Prior to phylogenetic tree construction, several sequences were filtered out where isoforms were presented (180), short sequences (19) and alternative splice variants or fragmented sequences (15). A maximum likelihood (ML) phylogenetic analysis of the remaining 95 A. perfoliata potential GST representative sequences (Supplementary Table S2) and 58 recognized GST sequences demonstrated that there were no A. perfoliata GST sequences belonging to the Zeta, Delta, Epsilon, Theta, Alpha, Kappa and Pi classes (Fig. 1). Among the group of GST classes, the largest cluster of 83 A. perfoliata GST sequences were clustered in a Mu class GST group, while 3 and 9 A. perfoliata GST sequences were clustered, with strong bootstrap support, in the Sigma (64%) and Omega (98%) GST class groups, respectively (Fig. 1). Of note, only the 3 Sigma class representatives identified previously were identified within the Sigma class.
Phylogenetic analysis of Glutathione transferases (GSTs) identified within the A. perfoliata transcriptome inferred using a Maximum likelihood (ML) tree with JTT matrix-based model. The bootstrap consensus tree is inferred from 1000 replicates. Evolutionary analyses were conducted in MEGA X. Circles denote sequences identified within the bioinformatics analysis of A. perfoliata.

In total the 83 sequences that clustered into 1 large Mu class GST group, alongside known Mu class GSTs, broadly demonstrated 19 clades of A. perfoliata Mu class GSTs. All clades contained more than 1 ApGST-Mu representative, except for ApGST-Mu1 which contained a solitary member. ApGST-Mu1 was the member most closely linked to known platyhelminth Mu class GSTs including members from Fasciolids and Taenia solium, as well as the inclusion of a host, equid, Mu class GST (Fig. 1). Of interest were ApGST-Mu class 16 and 17 which formed a clade with a Mu class representative from Hymenolepis microstoma (Fig. 1).
Characterization of novel A. perfoliata Omega class GSTs
A total of 8 novel A. perfoliata Omega class GSTs represented by 9 sequences used within the phylogenetics (designated ApGST-O1 [1.1 and 1.2], ApGST-O2, ApGST-O3, ApGST-O4, ApGST-O5, ApGST-O6, ApGST-O7 and ApGST-O8) clustered within the Omega class GST branch within the whole GST phylogenetic analysis. All 9 sequences were further subjected to a multiple sequence alignment with 11 recognized Omega class GST sequences from mammals and helminths separately from the remaining GSTs (Fig. 2). The secondary characteristic structure of the multiple aligned A. perfoliata Omega GST homologue sequences predicted 3 β-strands and 5 α-helices demonstrating the consistency of the secondary characteristic structure between ApGST-O1.1 to ApGST-O8 and the 11 recognized Omega class GST sequences (Fig. 2). The investigation of Omega class GST sequences based on the GSH-binding sites in the N-terminal domain demonstrated that all 8 novel A. perfoliata Omega class GST sequences demonstrated a high homology to other GSH-binding sites (Burmeister et al., Reference Burmeister, LÜersen, Heinick, Hussein, Domagalski, Walter and Liebau2008; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016), with the conserved catalytic cysteine residue characteristic of Omega class GSTs particularly at amino acid position 32 (Cys32) (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Meng et al., Reference Meng, Zhang, Liu, Guo and Xu2014; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016). However, each lacked the proline-rich residues in the Omega class characteristic N-terminal extension (PXXP motif) (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). Additionally, all 8 A. perfoliata Omega GSTs were confirmed as Omega class GST according to the presence of amino acid residues related to the Omega class GST signature motifs identified by Chemale et al. (Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006) (Fig. 2). Of note, the sequence length of ApGST-O8 was significantly truncated with much of the C-terminal region absent. Therefore, ApGST-O8 was excluded from further phylogenetic analysis.
Multiple sequence alignment of the 8 novel Omega class GSTs identified in A. perfoliata (ApGST-O1.1 to ApGST-O8). Secondary protein structure prediction using the PSIPRED Protein Analysis Workbench is presented below the alignment is comprised of 3 β-strands, shaded in yellow and 5 α-helices, shaded in pink. The proline-rich residues in the Omega class characteristic N-terminal extension as denoted by Morphew et al. (Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012) are shaded in blue. The catalytic cysteine residues characteristic of Omega class GSTs as denoted by Morphew et al. (Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012), Meng et al. (Reference Meng, Zhang, Liu, Guo and Xu2014) and Kim et al. (Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016) are shaded in green. Amino acids residues directly contact to glutathione are indicated by asterisks (*) (Burmeister et al., Reference Burmeister, LÜersen, Heinick, Hussein, Domagalski, Walter and Liebau2008; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016). Glutathione-binding residues are indicated by closed circles (o) (Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016). The Omega class GST signature motifs as denoted by Chemale et al., (Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006) are marked by arrows (↑). The predicted N- and C-terminal GST domain profiles are indicated by blue- and red-boxes.

A maximum likelihood (ML) phylogenetic tree generated from 7 of the 8 novel A. perfoliata Omega class GST sequences (8 sequences; ApGST-O1.1 to ApGST-O7) and 11 recognized Omega class GST sequences demonstrated that all 7 A. perfoliata Omega class GST sequences clustered strongly together with good support (Bootstrap value 90%). In addition, the ApGST Omega class representatives were clustered, again with strong support (96%), with the homologous Stringent starvation protein A (SspA) sequences from the Cestode clade including representatives from H. microstoma (CDS29490), Echinococcus granulosus (CDS24607) and E. multilocularis (CDS41542) (Supplementary Figure S1).
Species confirmation and GST activity
Live adult cestodes were obtained from the ileocecal valve from naturally infected horses and utilizing DNA extraction and PCR amplification of the ITS2 region were confirmed as A. perfoliata (Supplementary Figure S2). Following confirmation, homogenization of replicates of 10 worms yielded an average of 51.28 mg of cytosolic protein with a total GST activity of 490.34 μmol min−1 and a GST specific activity of 9.56 ± 4.02 μmol min−1 mg−1 prior to GSH based purification (Table 1.). Post GSH affinity chromatography, 0.33% of the total protein purified as GSTs with a specific activity recorded at 32.47 μmol min−1 mg−1 (Table 1). However, given a loss of total activity post purification it is likely that the contribution of GSTs to the total proteome is much higher than 0.33%. In addition, purified ApGST demonstrated significant activity towards EA with a specific activity of 1069.8 ± 194.7 nmol min−1 mg−1.
A. perfoliata GST activity of the cytosolic protein sample pre and post purification against the model substrate CDNB in relation to the total protein used from the sample.

Average taken across the 3 replicates.
Immunological classification
For initial classification of purified ApGST, cytosolic proteins which had been purified through GSH-agarose affinity column and profiled using 1DE were transferred to nitrocellulose membranes and subjected to immunoblotting with parasitic flatworm Anti-Mu and Anti-Sigma class GST antibodies. For Mu class GSTs, membranes probed with the anti-Mu class antibody demonstrated a strong recognition from the 1DE band of purified ApGST indicating a potential abundance of Mu class representatives (Fig. 3). Regarding Sigma class GSTs, transferred 1DE profiles were probed with anti-FhGST-S1 antibodies (Fig. 3) and, in contrast, demonstrated faint reactivity suggesting a lack of purified Sigma class representatives.
Representative 1DE SDS PAGE analysis of the purified A. perfoliata GSTs and western blotting. (A) 1DE of the whole cytosolic protein fraction – (1) Low molecular weight markers (GE Healthcare) – kDa and (2) 10 μg of cytosolic protein fraction extracted from adult A. perfoliata (B) 1DE of the GSH affinity purification fractions demonstrating GST purification – (3) Low molecular weight markers (GE Healthcare) – kDa (4) 15 μl of flow through after GSH affinity, (5) 20 μl of GSH affinity wash and (6) 2 μg of purified A. perfoliata GSTs (boxed). (C) Western blot of the purified A. perfoliata GSTs run on 1DE –and probed with either Anti-Sigma class or Anti-Mu class antibodies (7) Low molecular weight markers (GE Healthcare) – kDa, (8) A. perfoliata GSTs probed with Anti-Mu class antibodies, (9) Anti-Mu class negative control, (10) A. perfoliata GSTs probed with Anti-Sigma class antibodies, anti-FhGST-S1 and (11) Anti-Sigma class negative control. Lanes 8–11 all contained 2 μg of ApGSTs. Very faint reaction to Sigma class GSTs is indicated by an ←. All 1D gels were run on 12.5% acrylamide.

Proteomics of ApGSTs
Three biological replicates of whole A. perfoliata cytosolic proteins were resolved on 1D SDS-PAGE gels (Fig. 3) and identified through MASCOT via MS/MS Ion Search against the A. perfoliata transcriptome. ApGST representatives identified in the A. perfoliata transcriptome were then used to mine the cytosolic protein dataset, along with the A. perfoliata secretome (EV, EV surface and EV free ESP protein data) generated previously by Wititkornkul et al. (Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021), for GSTs. In total, 51 unique ApGST sequences were observed across the 4 datasets (Table 2). The whole worm proteome produced the most abundant number of GST hits with 47 unique ApGST sequences identified within the 797 total proteins identified (Table 2, Supplementary Table S3). Two of which, ApGST-Mu11 (DN11457_c1_g2_i3_56890) and ApGST-Mu9 (DN12300_c0_g1_i9_39697), were present in the top 50 most abundant proteins. Additionally, within the A. perfoliata secretome 1, 2 and 9 GST proteins were observed for EVs, the EV surface and EV-depleted ESP protein datasets, respectively. Only 1 GST protein, ApGST-Mu3 (DN12662_c0_g1_i1_71915), was consistent across all 4 datasets with 6 GST representatives consistent across at least 2 fractions.
ApGST proteins observed from mining of the A. perfoliata proteomic datasets from the whole worm, the EV, the EV surface and the EV-Depleted ESP

EV, EV surface and EV-Depleted ESP datasets obtained from Wititkornkul et al. (Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021)
Dark grey highlights indicate GSTs present in all 4 proteomes.
Light grey highlights are those present across 2 proteomes.
T GST Class predicted by phylogenetic analysis
H Class predicted by homology to isoforms.
Of note, a dominance of Mu class GST members was observed across the complete datasets. In total, 44 out of the 47 unique ApGST sequences identified in the whole worm proteome were classified as, or homologous to, Mu class GST representatives (Table 2). Similarly, all of the ApGST sequences observed in the EVs and EV surface, and 7 of the 9 ApGST sequences in EV-depleted ESP were also classified, or homologous to, Mu class GST representatives. Regarding Sigma class GSTs, ApGST-S1 has previously been highlighted as the only Sigma class GST in the tapeworms secretome (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021). Here, ApGST-S1 was also observed as the only Sigma class GST representative to be localized within the whole worm proteome with a similar relative low abundance. Finally, 2 of the novel A. perfoliata Omega GST representatives were identified in both the whole worm and EV-depleted ESP proteome. Specifically, ApGST-O3 was observed in both the whole worm protein and EV-depleted ESP proteins, whereas ApGST-O4 was only present in the EV-depleted ESP proteins.
Proteomic profiling of affinity purified ApGSTs
GSH agarose affinity purified ApGSTs were resolved on 2D SDS-PAGE gels (Fig. 4). The 2D gels were then subjected to either MASCOT via MS/MS Ion Search against the A. perfoliata transcriptome or transferred to nitrocellulose membranes and subjected to immunoblotting with Mu and Sigma GST antibodies. The 2DE profiling of the purified ApGST fraction using GSH-agarose affinity column presented a total of 11 resolved protein spots ranging from 4.36 to 5.69 in pI and 23.93 to 24.86 kDa in size (Fig. 4., Table 3). Six of these protein spots were consistent across all biological replicates with the remaining 5 consistent across 2 replicates. The visible protein spots (1–11, Fig. 4) present on the replicate 2DE profiles were excised for MSMS analysis. All protein spots contained proteins with significant identity or extensive homology to ApGST representatives (Table 3; Supplementary Table S4). Three spots (Spots 4, 7 and 8) contained 1 additional protein that were not identified as ApGSTs (Importin subunit beta 1 [DN12350_c0_g1_i1_64773], General vesicular transport factor p115 [DN11790_c1_g5_i1_61055] and a Tetratricopeptide repeat protein 35 B [DN8984_c0_g1_i1_28059], respectively). The unique ApGST sequences hit from the 2D spots were solely classified as, or homologous to, Mu class GST representatives (Table 3).
Representative 2DE SDS PAGE analysis of the purified A. perfoliata GSTs and western blotting. (A) 2DE SDS PAGE analysis of the purified A. perfoliata GSTs (20 μg). (B) 2DE SDS PAGE analysis of the purified A. perfoliata GSTs (20 μg) and subsequent western blotting – (1) Resolved spots of 2DE of GSH affinity purification fractions colloidal Coomassie blue stained. Numbered spots correspond to the putative protein identifications in Table 3. (2) Western blot of the purified A. perfoliata GSTs run on 2DE and probed with anti-Mu class antibodies, protein spots with prominent visible reactions are labelled corresponding to 2DE gel. (3) Western blot of the purified A. perfoliata GSTs run on 2DE and probed with anti-Sigma class, anti-FhGST-S1, antibodies. Position of low molecular weight marker (GE Healthcare) indicated – kDa. GSH affinity purification fractions were isoelectric focused on 7 cm pH 3 − 10 nonlinear IPG strips and run on 12.5% polyacrylamide gels. Gels for immunoblotting were transferred onto nitrocellulose membrane membranes for antibody binding and developed using the BCIP/NBT system.

GST proteins identified in the whole worm cytosolic fraction of A. perfoliata following GSH agarose affinity chromatography. The top member of each protein family identified via MASCOT search is reported, remaining family members of significance are shown in Supplementary Table S4.

T GST Class predicted by phylogenetic analysis
H Class predicted by homology to isoforms.
In addition, 8 spots (1, 2 and 6–11), corresponding to the GST 2DE profile, demonstrated strong reactivity to anti-Mu class antibody (Fig. 4). This reactivity was consistent across the 2 biological replicates. The 3 remaining spots (3–5) identified on the 2DE profiles did not demonstrate clear reactivity to the Mu class antibody. However, spots 3–5 were identified as Mu class GSTs via mass spectrometry with all 3 spots containing ApGST-Mu3. The classification of Mu class GST representatives from 2D analysis is again consistent with the dominance of Mu class representatives observed in the transcriptome and localized in the A. perfoliata whole cytosolic proteome.
No visible reaction from 2DE replicates probed with the anti-FhGST-S1 antibodies was observed (Fig. 4). The lack of reaction is consistent with the absence of Sigma class GSTs identified amongst the GSH agarose purified cytosolic proteins via LC-MSMS. Thus, confirming that Sigma GSTs were not purified from the cytosolic protein using GSH affinity.
Discussion
GSTs are multifunctional enzymes and binding proteins that play key roles as the major detoxification system of internally derived toxins and xenobiotics in parasitic helminths (Brophy et al., Reference Brophy, Papadopoulos, Touraki-’, Coles, Körting and Barrett1989a; Brophy and Barrett, Reference Brophy and Barrett1990b), likely as a consequence of limited Phase 1 oxygen dependent cytochrome P450 activities (Nelson, Reference Nelson2009; Pakharukova et al., Reference Pakharukova, Ershov, Vorontsova, Katokhin, Merkulova and Mordvinov2012; Tsai et al., Reference Tsai, Zarowiecki, Holroyd, Garciarrubio, Sanchez-Flores, Brooks, Tracey, Bobes, Fragoso, Sciutto and Aslett2013). Until recently, our knowledge of the GST complement within A. perfoliata has been limited with GSTs only demonstrated in the ESP fractions (Wititkornkul et al., Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021; Hautala et al., Reference Hautala, Pursiainen, Näreaho, Nyman, Varmanen, Sukura, Nielsen and Savijoki2022). Therefore, high-resolution sub-proteomics, coupled with bioinformatics, were utilized to further expand our knowledge, and understanding of the GST soluble superfamily within this equine helminth. Bioinformatic analysis has additionally identified 3 GST classes within the recent A. perfoliata transcriptome including those homologous with recognized Sigma, Omega and Mu class GSTs whilst members representing the Zeta, Delta, Epsilon, Theta, Alpha, Kappa and Pi GST classes were not identified. Mining the high-resolution proteome data of Wititkornkul et al. (Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021) provided additional evidence for Mu and Omega class GSTs in the A. perfoliata secretome with, for the first time, 3 characterized GST classes identified within the A. perfoliata cytosolic proteome.
An assessment of the A. perfoliata transcriptome revealed 83 putative GSTs were classified as belonging to the Mu class GSTs making them by far the most abundant class present; with all representatives clustering well in the Cestode Mu class GST group. In support of the current analysis, a strong relationship of cestode GSTs classified as Mu class has been previously noted amongst alternative cestode species (Brophy et al., Reference Brophy, Papadopoulos, Touraki-’, Coles, Körting and Barrett1989a; Brophy and Barrett, Reference Brophy and Barrett1990b) and an expansion of this GST class has been identified in the genomes of the cestodes E. multilocularis, E. granulosus, T. solium and H. microstoma (Tsai et al., Reference Tsai, Zarowiecki, Holroyd, Garciarrubio, Sanchez-Flores, Brooks, Tracey, Bobes, Fragoso, Sciutto and Aslett2013). Similarly, in the liver fluke F. gigantica Mu class GSTs were also identified as the most abundant (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). The large proportion of A. perfoliata Mu class GSTs in comparison to the other classes accounts for their dominance localized across the whole worm proteome, GSH-affinity purified fractions, and the mined secretome. However, it can be noted that differences were observed between the unique Mu class ApGST sequences identified in the whole cytosolic proteome and the GSH affinity purified GSTs. These differences may likely be explained by the different enrichment in GSTs between the biological samples used. In addition, some Mu class GST, such as FhGST-Mu5 in F. hepatica, have been shown to be a ‘low-affinity’ GST isoform which presents a failure to purify though affinity columns (Stuart et al., Reference Stuart, Zwaanswijk, Mackintosh, Witikornkul, Brophy and Morphew2021). Thus, the A. perfoliata ApGST-ome likely contains ‘low-affinity’ GST isoforms such as ApGST-Mu 5.
The 3 Sigma class GSTs within the transcriptome were previously identified by Wititkornkul et al. (Reference Wititkornkul, Hulme, Tomes, Allen, Davis, Davey, Cookson, Phillips, Hegarty, Swain and Brophy2021) with Wititkornkul and colleagues defining ApGST-S1 as a likely Sigma class GST. ApGST-S1 was also identified in the cytosolic proteome. However, a lack of Sigma class GSTs was observed in the GSH-affinity purified fraction through both MS/MS and immunoblotting with Anti-FhGST-S1. This is likely due to a combination of a single Sigma identified within the cytosol in relatively low abundance and the dominance of Mu class GST present. In addition, only a single purification method was utilized, namely GSH-agarose affinity chromatography. Studies have demonstrated that F. hepatica Sigma GSTs have reduced binding to GSH-agarose affinity if large quantities of Mu class GSTs are present (Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012) and the combination use of GSH-agarose and S-hexyl GSH-agarose affinity chromatography can provide a more complete profile of helminth GSTs (Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). Utilizing this combination method, a novel Sigma GST was identified in F. gigantica (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). Thus, the use of GSH-affinity purifications may partly account for the lack of Sigma class GSTs identified post purification and further investigations may benefit from the incorporation of S-hexyl GSH-agarose affinity purification.
To date, limited research has focused on how A. perfoliata interacts with its host and the surrounding environment with only a recent elucidation toward the ability of A. perfoliata to modulate their host immune system (Lawson et al., Reference Lawson, Pittaway, Sparrow, Balkwill, Coles, Tilley and Wilson2019). ApGST localization demonstrated Mu, Sigma and Omega ApGSTs across the somatic (ApGST-Mu2-9, Mu11-12, Mu15, Mu17, Mu19, ApGST-O3-4 and ApGST-S1) and secreted (EVs: ApGST-Mu3, ESP: ApGST-Mu1, Mu3, Mu11, ApGSTO4 and ApGST-S1) proteomes, with ApGST-Mu3 showing to be the most dominant Mu GST identified across all proteome fractions and 2DE spots. Thus, this localization provides an insight into an important detoxification component for A. perfoliata survival as well as immunomodulators. Interestingly, Hautala et al. (Reference Hautala, Pursiainen, Näreaho, Nyman, Varmanen, Sukura, Nielsen and Savijoki2022) also identified GSTs in A. perfoliata ESP albeit searching against the NCBInr database and thus relying on homology and protein conservation. Utilizing BLAST demonstrates the closest protein match identified showed significant homology with ApGST-15 sequences were identified. Here, ApGST-15 was identified in only the somatic proteome fractions. Differences here again may be explained by difference in GST enrichment in GSTs between samples.
A. perfoliata Omega class GSTs were analysed separately based on the multiple sequence alignment, phylogenetics analysis and secondary characteristic structure prediction. In total, the analysis of the A. perfoliata transcriptome data revealed the presence of 8 novel A. perfoliata GST sequences, corresponding to 7 Omega class members, all of which showed strong homology to recognized platyhelminth Omega GSTs and Stringent starvation protein A (SspA). Of the 7 novel A. perfoliata Omega class GSTs, 2 (ApGST-O3 & ApGST-O4) and 1 (ApGST-O4) were localized within the whole cytosolic and EV-depleted ESP proteomes, respectively. Whilst Omega GSTs have been identified in several helminth species (Girardini et al., Reference Girardini, Amirante, Zemzoumi and Serra2002; Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016), previous studies have suggested they were likely absent from cestodes (Iriarte et al., Reference Iriarte, Arbildi, La-Rocca, Musto and Fernández2012; Lopez-Gonzalez et al., Reference Lopez-Gonzalez, La-Rocca, Arbildi and Fernandez2018). However, proteins labelled SspA, an Omega GST homologue, have also been identified in cestodes species (Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016). Originally identified in bacteria, true SspA's present the characteristic GST fold but lacks GST binding activity due to replacement of the catalytic cystine residue (Hansen et al., Reference Hansen, Gu, Li, Andrykovitch, Waugh, Jin and Ji2005).
The secondary characteristic structure prediction established the consistency and similarity of the β-strand, α-helix and random coils structures within the recognized platyhelminth Omega class GST sequences, which are conserved regions of these proteins. Although all 8 novel A. perfoliata Omega class GSTs lacked the proline-rich residues in the Omega class characteristic N-terminal extension (PXXP motif) (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012). However, each putative Omega representative contained the catalytic cysteine residue characteristic of Omega class GSTs at position 32 (Cys32) (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Meng et al., Reference Meng, Zhang, Liu, Guo and Xu2014; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016) and contained amino acid residues related to the Omega class GST signature motifs identified by Chemale et al. (Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006). In addition, structural alignment analysis in Kim et al. (Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016) and the ApGSTs demonstrates these catalytic residues and motifs were also identifiable in the designated SspA's from the cestodes H. microstoma, E. granulosus and E. multilocularis. Thus, this provides further support that cestode SspA's, in addition to the A. perfoliata sequences, belong to the Omega class of GSTs. Helminth Omega class GSTs have been demonstrated to have critical roles in protection against oxidative stress, the helminth reproduction system, and inhibition of macrophage viability (Meng et al., Reference Meng, Zhang, Liu, Guo and Xu2014; Kim et al., Reference Kim, Ahn, Kim, Bae, Kwon, Kang, Yang, Sohn and Kong2016; Wang et al., Reference Wang, Zhao, Zhang, Zhang, Li, Shang, Ning, Ji, Xia, Cai and Qiao2022). As with Mu and Sigma class GSTs, the presence of Omega class GSTs in the somatic and EV-depleted ESP fractions suggests they could play a vital role in A. perfoliata's reproductive health and potential immune modulation function.
Previously, Zeta class GSTs have been identified in platyhelminthes such as F. hepatica and F. gigantica, the role of which is poorly understood (Morphew et al., Reference Morphew, Eccleston, Wilkinson, McGarry, Perally, Prescott, Ward, Williams, Paterson, Raman and Ravikumar2012; Stuart et al., Reference Stuart, Zwaanswijk, Mackintosh, Witikornkul, Brophy and Morphew2021). Zeta class GSTs do not appear to have been identified in cestode species and in agreement. Bioinformatic analysis of the A. perfoliata transcriptome did not return any GSTs related to the Zeta class. A preliminary homology search using BLAST across the Wormbase Parasite databases of Zeta GSTs against cestode species also retrieved no hits. Thus, it could be suggested that Zeta GSTs are likely absent from cestode species.
The high GSH affinity ApGST fraction purified through GSH agarose accounted for approximately 0.33% of the overall cytosolic proteins. In contrast, cestode and trematode GST have been shown to account for up to 3–4% of the soluble cytosolic protein fractions (Brophy et al., Reference Brophy, Papadopoulos, Touraki-’, Coles, Körting and Barrett1989a, Reference Brophy, Southant and Barrett1989b; Vibanco-Pérez et al., Reference Vibanco-Pérez, Jiménez, Merchant and Landa1999; Chemale et al., Reference Chemale, Morphew, Moxon, Morassuti, LaCourse, Barrett, Johnston and Brophy2006; Duncan et al., Reference Duncan, Cutress, Morphew and Brophy2018). However, a significant proportion of activity was left unpurified. Neither Sigma nor Omega class ApGSTs were purified through the GSH-affinity columns which will have contributed to GST activity. Additionally, previous studies have identified a failure of pure GST from pure cytosol extracts to completely bind to GSH-agarose affinity chromatography (Brophy and Barrett, Reference Brophy and Barrett1990a; Stuart et al., Reference Stuart, Zwaanswijk, Mackintosh, Witikornkul, Brophy and Morphew2021). Future investigation with improved GST purification, such as the addition of alternative chromatography methods, is thus likely warranted. ApGSTs level of specific GST activity in the CDNB assays was comparable to H. diminuta GST activity post affinity chromatography (Brophy and Barrett, Reference Brophy and Barrett1990a). However, the GST activity was much higher than previously demonstrated among other cestodes and trematode species (Brophy et al., Reference Brophy, Papadopoulos, Touraki-’, Coles, Körting and Barrett1989a; Brophy and Barrett, Reference Brophy and Barrett1990a; Duncan et al., Reference Duncan, Cutress, Morphew and Brophy2018). In addition, ApGST demonstrated relatively high activity towards the model substrate EA to that previously shown in Sm28GST and recombinant FhGST (Walker et al., Reference Walker, Crowley, Moreman and Barrett1993; LaCourse et al., Reference LaCourse, Perally, Morphew, Moxon, Prescott, Dowling, O'Neill, Kipar, Hetzel, Hoey and Zafra2012). EA activity is traditionally used as a marker for mammalian Pi class GST, a common GST class expressed in the mammalian intestinal system (Mannervik et al., Reference Mannervik, Guthenberg, Jensson, Kalim Tahir, Warholm and Jornvallt1985). However, no Pi GST representatives were identified in transcriptome yet A. perfoliata GSTs, potentially Mu class, may exhibit similar detoxifying activity, potential evidence of a GST adapting to the mammalian xenobiotic environment.
In conclusion the data obtained from the A. perfoliata transcriptome and bioinformatics analysis demonstrate strong evidence that the A. perfoliata GST superfamily comprises of Mu, Sigma and Omega class representatives. In addition, proteomic analysis has confirmed the expression of ApGSTs across the A. perfoliata cytosolic and secretome proteomes. The discovery of all 3 cytosolic GST classes in the current data provides a greater in-depth characterization of the Equine tapeworm A. perfoliata, specifically the A. perfoliata GST-ome, illuminating potential mechanisms involving the parasites survival and host–parasite relationships. Given the importance of GSTs to helminth survival, the characterization of the A. perfoliata GST-ome provides valuable targets for developing further novel anthelminthics as well as potential usage in vaccine development for this often-neglected Equine tapeworm.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182024000015.
Data availability statement
The A. perfoliata somatic proteome mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://www.ebi.ac.uk/pride/archive/) via the PRIDE partner repository with the data set identifier PXD029998 and 10.6019/PXD029998. The A. perfoliata transcriptome can be provided upon reasonable request or alternatively is available at https://sequenceserver.ibers.aber.ac.uk/ for interrogation.
Authors’ contributions
Conceptualization: ROM, REW; Carried out experimental design: HMN, BW, DJC, NDA; Data analysis: HMN, BW, DJC, NDA; Writing original draft: HMN, BW; Review and Editing: ROM, REW, PMB. ROM and REW contributed equally and share last authorship. All authors have read and agreed to the published version of the manuscript.
Financial support
The authors acknowledge grant funding (Welsh Government: Establishing Cutting Edge Veterinary Research Laboratories for Wales; European Development Regional Fund Sêr Cymru programme Grant – 80761-AU185) for the acquisition of the Ultimate 3000 RSL Nano with ES082 EASY-Spray Source. H.M.N. is the grateful recipient of an Eleanor & David James PhD Scholarship, Aberystwyth University and BBSRC. B.W. was funded by Rajamangala University of Technology Srivijaya, Thailand.
Competing interests
None.
Ethical standards
Not applicable.