Introduction
In rice (Oryza sativa L.) agriculture of the southern United States, the largest biotic constraint on yield is competition from agricultural weeds. The United States is the fifth-largest global exporter of rice (USDA-FAS 2023), and thus it is of the utmost importance for worldwide food security to maximize rice crop yield while simultaneously working to minimize agricultural input (e.g., fertilizer, fuel, land, herbicides, and labor). Among the multitude of weedy species that infest southern U.S. rice fields, the weed consistently ranked most harmful is barnyardgrass (Echinochloa spp.) (Butts et al. Reference Butts, Kouame, Norsworthy and Barber2022). Belowground competition from Echinochloa can divert up to 80% of available soil nitrogen to the weed, reducing rice yields by up to 70% in heavily infested fields (Holm et al. Reference Holm, Pluknett, Pancho and Herberger1977; Smith Reference Smith1988). Echinochloa species are highly adapted to rice agroecosystems due to their high seed production (Tahir and Roma-Burgos Reference Tahir and Roma-Burgos2021), propensity to rapidly evolve multi-herbicide resistance (Rouse et al. Reference Rouse, Roma-Burgos, Norsworthy, Tseng, Starkey and Scott2018), adaptability to the various rice-cropping systems (Butts et al. Reference Butts, Kouame, Norsworthy and Barber2022), and cryptic seedling vegetative forms that hinder early detection among rice plants (Ye et al. Reference Ye, Tang, Wu, Jia, Qiu, Chen, Mao, Lin, Xu, Yu, Lu, Wang, Olsen, Timko and Fan2019).
Current weed control efforts collectively treat all Echinochloa species as a single entity. However, infestations are in reality a complex of several Echinochloa species. While infestations of barnyardgrass [Echinochloa crus-galli (L.) P. Beauv.] (E. crus-galli sensu stricto; 2n = 6x = 54; AABBCC subgenomes) were noted by the early 1900s, it became the most prevalent weed in rice fields of the southern United States only after deployment of the herbicide 2,4-D in the 1940s began to effectively control broadleaf weeds (Godar and Norsworthy Reference Godar and Norsworthy2023). Overreliance on the herbicide propanil, to which E. crus-galli evolved resistance after almost 30 yr (Talbert and Roma-Burgos Reference Talbert and Burgos2007), further promoted proliferation of populations into the late 1980s. Echinochloa crus-galli is particularly devastating to rice fields because the seedling and tillering stages of this species are nearly indistinguishable from those of rice, leading to season-long competition where escapes from weed control operations are undetected. The only obvious phenotypic difference between the two species at the early vegetative stage is the lack of a ligule in E. crus-galli, a general characteristic of the genus. After panicle exsertion, inflorescences of the weed are easily differentiated from rice, but are highly variable in length, (5 to 25 cm) with primary racemes of 2 to 8 cm in length (Tahir Reference Tahir2016).
Echinochloa crus-galli in the United States includes two recognized subspecies varietal groups. Echinochloa crus-galli (L.) Beauv. var. crus-galli is primarily associated with wetland areas such as flooded rice paddies and is phenotypically considered a rice mimic (Wu et al. Reference Wu, Shen, Jiang, Feng, Tang, Lao, Jia, Lin, Xie, Weng, Dong, Qian, Lin, Xu and Lu2022). Conversely, Echinochloa crus-galli var. praticola Ohwi is a weedy form found in drier lands outside rice paddies, with a significantly larger tiller angle on average. Anecdotal reports (NR-B, personal observation) have suggested that furrow-irrigated rice fields (fields not permanently flooded) have seen invasion by E. crus-galli var. praticola.
By the early 1990s, agricultural shifts to mechanized direct drill seeding into dry fields, in place of traditional broadcast seeding into pre-flooded fields, led to the appearance of a second Echinochloa species, junglerice [Echinochloa colona (L.) Link] (E. colona; 2n = 6x = 54; DDEEFF subgenomes). In addition, a wider selection of herbicides and herbicide-resistant rice technologies were used to control E. crus-galli, further promoting E. colona (Godar and Norsworthy Reference Godar and Norsworthy2023). Echinochloa crus-galli and E. colona are highly morphologically similar, making them exceedingly difficult to distinguish, even by trained taxonomists and weed scientists. As with E. crus-galli, E. colona seedlings are often confused for rice, with a nearly identical vegetative morphology. Inflorescences of E. colona at maturity are, on average, shorter than those of E. crus-galli (6 to 12 cm), but significantly overlap the potential phenotypic space. Primary branches are short and non-overlapping at 1 to 2 cm in length. Unlike some congeners, both E. crus-galli and E. colona are glabrous and produce small, mostly awnless seeds (Tahir Reference Tahir2016).
While E. crus-galli and E. colona are the major members of the weedy Echinochloa complex of southern U.S. rice fields, two additional minor species may also be rising to frequencies of agricultural concern. Rough barnyardgrass [Echinochloa muricata (P. Beauv.) Fernald] is native to the United States (unlike E. crus-galli and E. colona, which are of Eurasian origin) and grows much taller than rice (∼1.5 m tall), with prominent stem nodes and large, awned seeds (Tahir Reference Tahir2016). Coast cockspurgrass [Echinochloa walteri (Pursh) A. Heller], another native North American species, is the newest Echinochloa weed of concern, with rare sightings in Arkansas (NR-B, personal observation). Echinochloa walteri can be readily distinguished from its congeners in the region by dense pubescence on the culm.
While E. crus-galli historically has been the dominant weedy species of rice fields in the southern United States, recent studies have suggested a pronounced shift toward E. colona. However, these studies relied on morphological characteristics and have been dismissed as “anomalous reports” in a recent review (Godar and Norsworthy Reference Godar and Norsworthy2023). One such study in Texas identified 52 of 54 collected ecotypes as E. colona (Liu et al. Reference Liu, Singh, Abugho, Lin, Zhou and Bagavathiannan2022), while another study in Arkansas found 73 of 94 accessions to be E. colona (Tahir Reference Tahir2016). Both studies used phenotypic characterization of taxonomic traits to distinguish between species and relied heavily on access to voucher specimens from local herbaria for comparisons. Such morphological analyses for species determination are time-, space-, and resource-consuming and require trained observer expertise to avoid errors. The recent publication of E. crus-galli and E. colona genomes (Wu et al. Reference Wu, Shen, Jiang, Feng, Tang, Lao, Jia, Lin, Xie, Weng, Dong, Qian, Lin, Xu and Lu2022; Ye et al. Reference Ye, Wu, Mao, Jia, Qiu, Lao, Chen, Jiang, Tang, Peng, Pan, Wang, Feng, Guo and Zhang2020) could be used to distinguish the species through comparative genomics approaches; however, these methods would be expensive and slow and require bioinformatics expertise. Comparative genomics is not useful for quick species ID. The current lack of clear data on the relative prevalence of E. crus-galli and E. colona in southern U.S. rice production systems highlights the need for easy and cheap PCR-based methods of identification to distinguish between species so that precision weed control techniques can be developed and implemented.
Another open question in the biology of Echinochloa weeds of rice fields is the degree to which they form interspecific hybrids. One recent survey of worldwide Echinochloa samples identified three samples in China with evidence of introgression between hexaploid E. crus-galli and tetraploid late watergrass (Echinochloa oryzicola var. oryzicola Ohwi), which is considered the tetraploid progenitor of E. crus-galli (2n = 4x = 36; AABB subgenome) (Wu et al. Reference Wu, Shen, Jiang, Feng, Tang, Lao, Jia, Lin, Xie, Weng, Dong, Qian, Lin, Xu and Lu2022). In the United States, E. oryzicola var. oryzicola is a problem weed in Californian rice agriculture but is not present in the southern rice-growing region (Godar and Norsworthy Reference Godar and Norsworthy2023). As such, this particular hybrid pair is unlikely to be observed in Arkansas rice fields, but its occurrence in China establishes that interspecies, and even interploidy, hybrids can form in this species complex. Separately, anecdotal reports of potential hybrids have also been received from southern U.S. rice farmers and extension agents based on morphological observations (TR Butts, Purdue University, personal communication), but no genomic work has confirmed or refuted their hybrid status.
In this study, we use genotyping-by-sequencing (GBS) and chloroplast genome sequencing approaches to develop a test that is simple, cheap, and effective at distinguishing between E. colona and other Echinochloa species. We then apply this test to assess the dominance of E. colona in eastern Arkansas, the primary region of southern U.S. rice production. We additionally use the GBS data to investigate whether putative hybrids, identified based on morphology, are in fact the product of interspecific hybridization.
Materials and Methods
Plant Materials for GBS and Chloroplast Genome Sequencing
Plant materials for sequencing were obtained as seed samples from three sources (Supplementary Table S1). Provisional species identifications were designated by the donating party based on morphology (hereafter referred to as “provisional species identifications”). Eight samples were provided by NR-B, nominally encompassing E. colona (n = 3), E. crus-galli (n = 3), E. muricata (n = 1), and E. walteri (n = 1). Twenty-eight samples representing 5 nominal taxa were obtained from the USDA-GRIN collection: E. colona (n = 7), E. crus-galli (n = 14), E. muricata (n = 1), E. walteri (n = 5), and E. colona var. frumentacea (n = 1). An additional 12, putatively hybrid, samples of unknown species identity were acquired from rice fields in five Arkansas counties during August of 2022 with the help of collaborator Thomas R. Butts (then Arkansas Weed Extension Specialist; now at Purdue University). All samples were grown from seed (one plant per seed sample) in the Washington University in St Louis greenhouse under standard midsummer ambient conditions (21 to 27 C, 16:8 h light:dark) until panicle maturity for DNA extraction using the Qiagen DNeasy Plant extraction Kits (Qiagen, Germantown, MD, USA) and visual inspection. The combination of the three seed sources resulted in a dataset of 48 samples with varying confidence of correct taxonomic identification.
GBS
To definitively identify samples to species, we utilized a GBS approach. Extracted DNA was sent to the University of Minnesota Genomics Center (UMGC) where GBS libraries were prepared by UMGC. Follow-up quality control and one lane of 1 × 151 bp short-read sequencing using the AVITI platform was then performed. Libraries were demultiplexed by UMGC and raw reads were returned to Washington University for in-house bioinformatic processing and analysis, as described in “Data Analysis.”
Chloroplast Genome Sequencing
As most Echinochloa weeds in the southern United States are hexaploids, which can pose challenges for PCR, we also performed chloroplast genome sequencing as a complementary approach for species determination. Chloroplast genome libraries were prepared at Washington University using the NEBNext Ultra II FS DNA library prep kit with NEBNext universal i5/i7 index primers for Illumina for the same 48 samples. After library preparation, samples were sent to UMGC for quality control and 2 × 150 short-read sequencing on the AVITI Freestyle platform (New England Biolabs, Ipswich, MA, USA). Libraries were demultiplexed by UMGC, and raw reads were returned to Washington University for in-house analysis.
Data Analysis
Raw reads were trimmed, aligned, and converted to.bam files using cutadapt v1.18, bwa v0.7.18, and samtools v1.15 packages, respectively, as part of the fastgbs_V2 pipeline v1.0 (Torkamaneh et al. Reference Torkamaneh, Laroche and Belzile2020). The GBS reads were aligned to the STB_08 E. crus-galli genome (Ye et al. Reference Ye, Wu, Mao, Jia, Qiu, Lao, Chen, Jiang, Tang, Peng, Pan, Wang, Feng, Guo and Zhang2020), while the chloroplast genome reads were aligned to an E. crus-galli chloroplast genome sequence found in GenBank accession number KJ000047 (Ye et al. Reference Ye, Lin, Li, Wang, Qiu, Fu, Zhang, Chen, Ye, Song, Jin, Zhu, Lu, Guo and Fan2014). In both datasets, variants were identified using the mpileup program in the bcftools software (Danecek et al. Reference Danecek, Bonfield, Liddle, Marshall, Ohan, Pollard, Whitwham, Keane, McCarthy, Davies and Li2021). Resultant .vcf files were further filtered using vcftools (Danecek et al. Reference Danecek, Auton, Abecasis, Albers, Banks, DePristo, Handsaker, Lunter, Marth, Sherry, McVean and Durbin2011) to remove indels, variants with a minor allele frequency <0.05, and variants with more than 20% missing data. For the GBS dataset, sites significantly out of Hardy-Weinberg equilibrium (P < 1 × 10−10) were also filtered using vcftools.
Population Structure Analysis
The pca flag in Plink V1.9 (Purcell et al. Reference Purcell, Neale, Todd-Brown, Thomas, Ferreira, Bender, Maller, Sklar, De Bakker, Daly and Sham2007) was used with the GBS dataset to visualize genetic differentiation among samples by principal component analysis (PCA). Samples were re-classified to species using the results of this PCA, and updated species designations were used for further analysis. To identify cpDNA single-nucleotide polymorphisms (SNPs) that were diagnostic of E. crus-galli versus E. colona, the --weir-fst-pop flag was used in vcftools, and only sites with an F ST = 1 were retained.
All contigs and scaffolds (>4,500) of the E. crus-galli nuclear genome were converted to chromosome “0” using the --allow-extra-chr 0 flag in plink2.0 (Galinsky et al. Reference Galinsky, Bhatia, Loh, Georgiev, Mukherjee, Patterson and Price2016) to allow for creation of the.bed,.bim, and.fam files required for further analysis, as the hard-coded limit of allowed chromosomes is 99. The ADMIXTURE program (Alexander et al. Reference Alexander, Novembre and Lange2009) was then used to identify population structure within the GBS dataset with the --cv flag enabled for cross-validation. Results from ADMIXTURE were visualized using Distruct v2.3 (Chhatre 2018) at K = 2 to 10.
Species Identification using Restriction Enzyme Assay
Using the list of cpDNA SNPs identified by F ST analysis (E. crus-galli vs. E. colona), we interrogated each SNP for potential restriction enzyme cut site polymorphisms using the online NEBcutter v3.0 tool (https://nc3.neb.com/NEBcutter/). The results of this analysis were used to identify a single potential SNP/enzyme combination, and PCR primers ech_ID1_F1 (5′-TCGAACCGTAGACCTTCTCG-3′) and ech_ID1_R1 (5′-GAGCACAGGGAGCCATCTTA-3′) (Supplementary Table S2) were designed to surround it. Thermocycler conditions for 20 µl PCR were as follows: initial denaturation at 94 C for 2 min, followed by 35 cycles of denaturation at 94 C for 30 s, annealing at 51 C for 30 s, and extension at 72 C for 30 s. PCR finished with a final extension at 72 C for 7 min. After PCR, 10 µl of PCR product were removed and set aside for later visualization as a control for restriction enzyme digestion. One microliter (5 units) of the restriction enzyme AciI, the enzyme associated with the restriction enzyme cut site polymorphism induced by the chosen SNP, was added directly to the remaining 10 µl of PCR product for 1 h of digestion in the thermocycler at 37 C (per manufacturer’s protocol). Pre- and post-digestion products were visualized side by side on a 1% agarose gel. One known E. crus-galli (ECR-34) and one known E. colona (ECO-22) were included once on each new day the restriction enzyme was used to validate enzyme function.
Plant Materials for Validation of cpDNA-based Species Identification
A second dataset, built from a long-standing collection of Echinochloa samples in the Roma-Burgos lab that have been identified via taxonomic characteristics by experts, was used to validate the chloroplast-based species identification test (Supplementary Table S3). This dataset consisted of E. colona (n = 20), E. crus-galli (n = 6), and E. muricata (n = 3). Approximately 20 seeds of a single genotype were scarified and grown in a growth chamber at the University of Arkansas with 16:8 h light:dark, 450 µmol m−2 s−1 light intensity, and 50% humidity until DNA extraction. Extractions were performed with the hexadecyltrimethylammonium bromide (CTAB) method by pooling tissue from 10 to 15 individual seedlings per genotype.
Plant Materials for Assessment of Echinochloa Species Prevalence in Eastern Arkansas Rice Fields
A third dataset was obtained from field-collected samples in the summer of 2024 (Supplementary Table S2). Twenty-three rice fields from nine counties were sampled opportunistically by driving state highways between Stuttgart and Jonesboro, AR. Field selection criteria were: visual confirmation of Echinochloa infestation from the road, a distance of at least 10 miles between sampled fields, and no more than four fields sampled per county. Panicles from approximately four Echinochloa plants were collected per field; these were selected to maximize collected strain phenotypic diversity and were not necessarily proportional to phenotypic frequencies within a field (hereafter referred to as a “diversity-first” collection). In total, 84 Echinochloa panicle samples with unknown species designations were collected. One seed per sampled plant was grown in the Washington University greenhouse under standard midsummer conditions (21 to 27 F, 16:8 h light:dark) until enough tissue was available for high-quality DNA extraction (typically 21 d). DNA extraction was performed using Qiagen DNeasy Plant extraction Kits (QIAGEN. Germantown, MD, USA).
Results and Discussion
GBS
GBS yielded approximately 2 million SNPs after filtering, which were distributed roughly evenly across the E. crus-galli reference genome. Using all available SNPs, PCA of all samples revealed three genetically distinct clusters (Figure 1). PC1 (accounting for 37.8% of total variation) separates E. colona (left) from the rest of the samples (right), while PC2 (accounting for 12.7% of total variation) separates E. crus-galli (top) from the remaining samples (bottom). The samples from the third (lower right) genetic cluster represent E. muricata and E. walteri, which were easily distinguished by phenotype (dense pubescence on E. walteri culms; large, awned seeds on E. muricata). Of note, several samples in this whole-dataset PCA are likely morphological misidentifications of species (e.g., green squares and purple circle in the E. crus-galli cluster). These misidentifications underscore the need for easy and reliable identification tools. Using the results of this PCA, we reclassified samples from their provisional species identification to their genomic species identification.
Principal component analysis (PCA) of genotyping-by-sequencing (GBS) single-nucleotide polymorphism (SNP) data for 48 Echinochloa samples using provisional species identification (i.e., before species reassignment). In the main figure, shape denotes origin of the sample, while color indicates the nominal species identification before genetic analysis. Inset figure is a reanalysis of the E. crus-galli samples alone to identify within-species varietal differences.

Closer investigation of the E. crus-galli samples alone further subdivided that group into three additional genetic clusters (Figure 1, inset). Metadata stored in the USDA-GRIN database for samples in the lower-left portion of the inset cluster suggest that all of these samples were collected from outside rice fields (parking lots, riverbanks, sidewalks, etc.), which is consistent with the varietal group E. crus-galli var. praticola (Godar and Norsworthy Reference Godar and Norsworthy2023). Indeed, the complete nuclear genome of a single sample from this group (PI 649368, from Iowa) was previously sequenced and given the same varietal designation (Wu et al. Reference Wu, Shen, Jiang, Feng, Tang, Lao, Jia, Lin, Xie, Weng, Dong, Qian, Lin, Xu and Lu2022). The lower-right genetic cluster of the inset contained three samples, two of which were part of the unknown collection, which were collected from inside southern U.S. rice fields. This makes it likely that this cluster represents E. crus-galli var. crus-galli samples; however, phylogenetic analysis with additional samples of known species identity would be necessary to make this determination definitively. The final genetic cluster of the inset (top) is a geographically diverse group (originating from Afghanistan, Florida, California, India, and other locations) with no metadata available in the USDA-GRIN database, making varietal designations impossible without further analysis.
Cross-validation error analysis of ADMIXTURE results suggested an optimal subpopulation number of K = 4 or K = 6 (Supplementary Figure S1). At face value, this agrees with the biological interpretation of four species (E. crus-galli, E. colona, E. muricata, and E. walteri). Interestingly, however, E. muricata and E. walteri are grouped as one population at K = 4, with E. crus-galli split into two distinct subpopulations (Figure 2). This within-species resolution may be an artifact of using E. crus-galli as the reference genome, which would bias SNP calling toward this species. At K = 6, a third subpopulation of E. crus-galli appears, and E. walteri splits from E. muricata. There is some discordance between subpopulation clustering in E. crus-galli between the analysis types. At K = 4, ADMIXTURE identifies only two clusters. Additionally, in the PCA, the lower right cluster of the inset contains only three samples, while the smallest genetic cluster in ADMIXTURE contains four samples (purple) at K = 6. With regard to the unknown samples taken from Arkansas rice fields in the summer of 2022, ADMIXTURE clearly identifies five as E. colona, five as E. muricata, and two as E. crus-galli. At K = 4, both of these E. crus-galli samples have admixed population assignments consistent with intervarietal admixture; however, this pattern disappears at K = 6, a common result of increasing K in ADMIXTURE as the program is given more degrees of freedom to maximize within-group Hardy-Weinburg equilibrium.
Population structure of 48 Echinochloa samples based on genome-wide genotyping-by-sequencing (GBS) single-nucleotide polymorphism (SNP) data using ADMIXTURE at K = 3, 4, and 6. Known samples are grouped by genomic species identification obtained by PCA. Unknown samples represent the 12 putative E. colona × E. crus-galli hybrid samples collected in Arkansas rice fields in the summer of 2022. Optimal number of populations (K = 4) is shown in addition to a secondary peak in CV error (K = 6) as well as a low K (K = 3) to identify any potential between-species hybridization. “Unknown” samples are those collected as putative E. colona × E. crus-galli hybrids in 2022 from inside rice fields. No interspecific hybridization is identified in Unknowns; however, there appears to be a within-species admixture of several E. crus-galli samples.

Published reports of interspecific hybridization from China and anecdotal reports from U.S. rice farmers led to the open question of the degree to which Echinochloa forms interspecific hybrids in southern U.S. rice fields (Wu et al. Reference Wu, Shen, Jiang, Feng, Tang, Lao, Jia, Lin, Xie, Weng, Dong, Qian, Lin, Xu and Lu2022). Hybridization was hypothesized as one potential avenue of rapid herbicide-resistance evolution, as the sharing of herbicide-resistance alleles has been shown to be the primary evolutionary driver in other weedy species of rice fields (Wedger et al. Reference Wedger, Roma-Burgos and Olsen2022, 2024). However, our ADMIXTURE results reveal no evidence of interspecific hybridization between E. colona and E. crus-galli. Despite the small sample size (n = 12), the deliberate sampling of weeds that farmers pointed to as examples of hybrids lends strength to this conclusion.
Chloroplast Genome Sequencing
To complement GBS analyses of nuclear genome sequence data, we used a chloroplast genome sequencing approach to mine for species-diagnostic SNPs in the same 48 samples analyzed by GBS. Alignment to an E. crus-galli chloroplast genome identified 493 SNPs. Analysis of interspecific differentiation identified 19 candidate SNPs with complete differentiation between E. colona and E. crus-galli (F ST = 1). After interrogation for restriction site polymorphisms, a single SNP—a C-to-T polymorphism at nucleotide 52654 of the KJ000047.1 E. crus-galli chloroplast genome—was determined to be diagnostic by altering the AciI recognition site from 5′-CCGC-3′ in E. crus-galli (recognized) to 5′-CTGC-3′ in E. colona (unrecognized) (Figure 3).
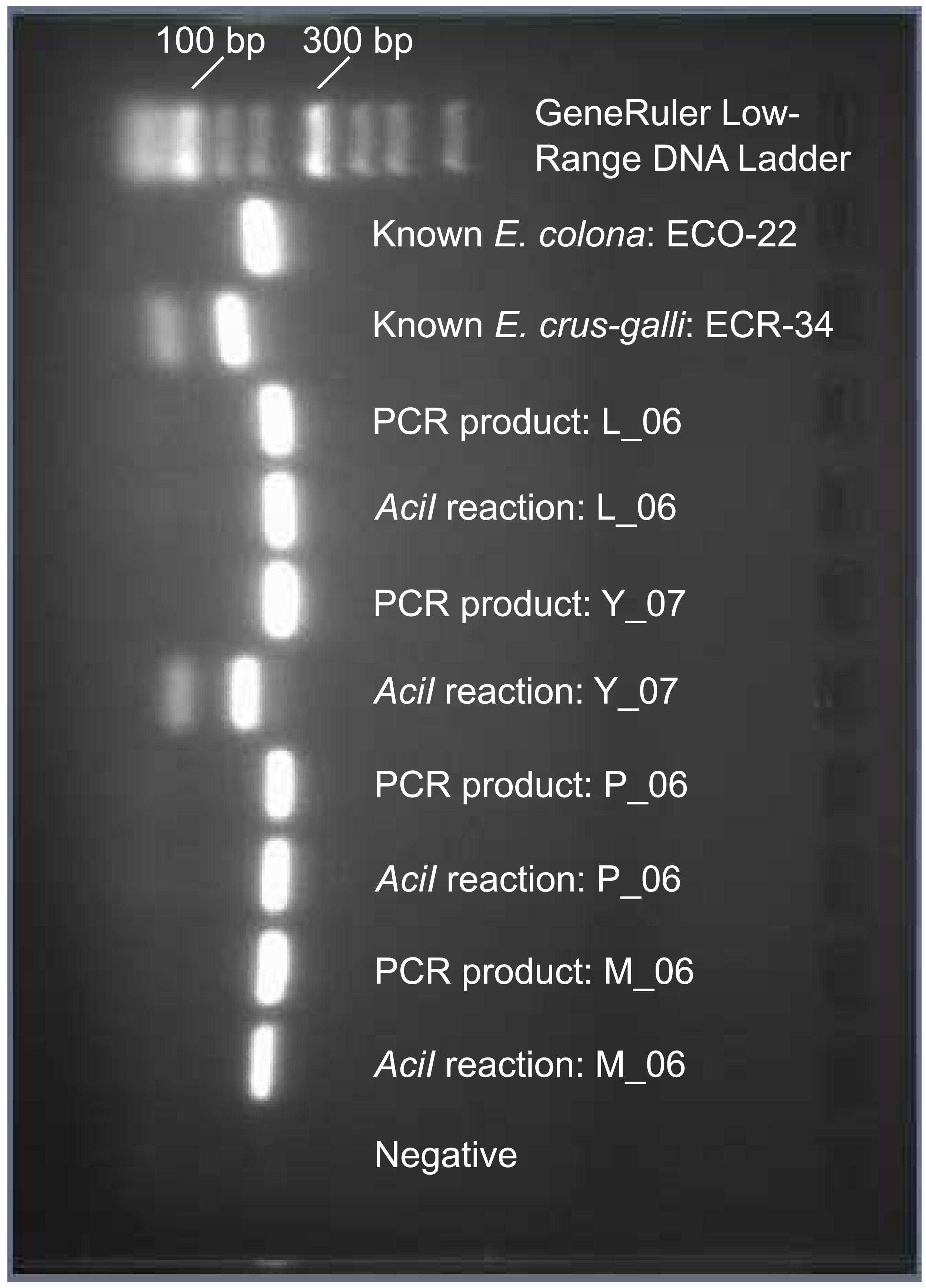
Gel image of typical results from PCR-based restriction enzyme test. PCR product (187 bp) remains uncut in Echinochloa colona samples (L_06, M_06, and P_06 in image), while product is cut by restriction enzyme AciI (139 and 48 product sizes) in Echinochloa crus-galli, Echinochloa muricata (Y_07 in image), and Echinochloa walteri samples.

Further validation of this test was performed by the Roma-Burgos lab using 29 known samples of Echinochloa (Supplementary Table S3). As designed, every PCR product from E. colona remained uncut (187 bp), while all other samples were cut by AciI (139 and 48 product sizes) (Supplementary Table S3). Of note, E. muricata was also cut by the AciI restriction enzyme in this assay. Several E. walteri samples were evaluated in the Olsen lab and were, again, cut by the enzyme (Supplementary Table S1). All three species that contain the restriction site sequence (are cut by AciI) are readily distinguished based on morphological characteristics. No cpDNA SNPs with species-diagnostic potential could be identified in the chloroplast genome between E. crus-galli and E. muricata or between E. crus-galli and E. walteri.
Assessment of Echinochloa Species Prevalence in Eastern Arkansas Rice Fields
Using the restriction enzyme test designed for this study, we found that PCR products from 75/84 (89.3% of our diversity-first collection) samples remained uncut by AciI, confirming an E. colona species designation (Supplementary Table S4). Of the nine PCR products cut by our restriction enzyme assay, four were visually identified as E. muricata (Supplementary Figure S2). Thus, only 5/84 (5.9% of our diversity-first collection) samples in our collections are likely E. crus-galli (Table 1). No E. walteri samples were identified in our field collections. Given the diversity-first nature of our 2024 field collections, it is likely that we oversampled rare genotypes. This implication would suggest the true field frequency of E. colona genotypes to be higher, and 89.3% could be a “minimum prevalence.” Echinochloa crus-galli samples were distributed throughout the sampling locations (i.e., not clustered in a single field) with no clear geographic bias (Table 1).
Echinochloa collection counties and identifications in eastern Arkansas rice fields using restriction enzyme assay with follow-up phenotypic analysis when necessary (n = 84) in a diversity-first collection.a

a The diversity-first nature of these collections suggests that rare phenotypes were likely oversampled and, as such, these values might not represent the true occurrence frequencies of these species.
Despite the century-long dominance of E. crus-galli as the major agricultural weed in southern U.S. rice fields, surveys from the last decade have hinted at a pronounced shift in population sizes, with E. colona increasing to be the more dominant weed, with only small populations of E. crus-galli remaining (Liu et al. Reference Liu, Singh, Abugho, Lin, Zhou and Bagavathiannan2022; Tahir Reference Tahir2016). Here, our cpDNA-based results definitively document this increase, with the overwhelming majority of our 2024 collections from eastern Arkansas positively identified as E. colona. This restriction enzyme–based test, which can be performed easily by any laboratory equipped with pipettes, a thermocycler, and a gel electrophoresis system, is economical, at less than US$7 per sample, and it does not require a skilled bioinformatician or taxonomist for data interpretation. This test can thus be readily employed to document infestations throughout the southern U.S. rice production region. While “cut” results will require further phenotypic analysis, and hypothetical future hybrids will be missed, the ability to confidently identify weedy E. colona samples will accelerate our understanding of the species complex and aid in the constant fight against agricultural weeds. As a final consideration, while no E. crus-galli × E. colona hybrid samples were identified in this study, and no hybrids of this ancestry have ever been reported, this assay would be unable to identify such a sample, given its chloroplast genome anchoring. Additional nuclear assessments of Echinochloa populations should be undertaken periodically to monitor this evolutionary direction.
Future Control Efforts for Echinochloa
As rice agriculture continues to adapt to 21st-century pressures for ever-increasing yield with diminishing resources, one of the biggest problems to solve is biotic constraints from agricultural weeds. To date, the primary approach to controlling agricultural weeds has been the broad application of herbicides. Unfortunately, the Echinochloa complex of species has evolved multi-resistance to a wide variety of herbicide modes of action, including inhibition of: fatty acid biosynthesis (molinate and thiobencarb), acetyl-CoA carboxylase (cyhalofop-butyl, fenoxaprop-ethyl, and quizalofop-P-ethyl), acetolactate synthase (bispyribac-sodium, penoxsulam, and imazethapyr), 1-Deoxy-D-xylulose 5-phosphate (DOXP) synthase (clomazone), auxin (quinclorac), and photosynthesis (propanil) (Godar and Norsworthy Reference Godar and Norsworthy2023; Liu et al. Reference Liu, Singh, Zhou and Bagavathiannan2021; Rouse et al. Reference Rouse, Roma-Burgos, Norsworthy, Tseng, Starkey and Scott2018). As farmers utilize more elaborate crop rotations, alternate dry- and wet-seeding seasons, and transition to furrow-irrigation practices, an understanding of population structure, intraspecific admixture, and continuous monitoring of Echinochloa species dynamics is foundational to the development of informed precision chemical weed management strategies.
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/wsc.2026.10091
Data availability
Genotyping-by-sequencing data are available through NCBI under BioProject accession number PRJNA1427233. Chloroplast genome sequence data are available under Bioproject accession number PRJNA1428746.
Acknowledgments
The authors of this article would like to thank Thomas R. Butts (Purdue University) for providing samples used in hybridization analysis. We would also like to thank members of the Olsen and Roma-Burgos lab groups for helpful comments on drafts of this article and assistance in the laboratory.
Funding statement
Funding for Echinochloa work in the Olsen lab was provided by a Donald Danforth Plant Science grant (Washington University in Saint Louis). Funding for Echinochloa work in the Roma-Burgos lab was provided by BASF Corporation (Research Triangle Park, NC, USA) grant no. GR015959.
Competing interests
The authors declare no conflicts of interest.