Introduction
Malaria is an infectious disease caused by a protozoan parasite belonging to the genus Plasmodium. These parasites are transmitted through mosquito vectors to a diverse range of vertebrate hosts including mammals like primates, bats and rodents, but also to reptiles and birds (Fricke et al., Reference Fricke, Vardo-Zalik and Schall2010; Schaer et al., Reference Schaer, Perkins, Decher, Leendertz, Fahr, Weber and Matuschewski2013; Templeton et al., Reference Templeton, Asada, Jiratanh, Ishikawa, Tiawsirisup, Sivakumar and Kaneko2016). Plasmodium species differ in the vector species they are transmitted by, the range of hosts they can infect, their pathogenicity and in their distribution across the world (Levine, Reference Levine1988; Escalante and Ayala, Reference Escalante and Ayala1994). In this sense, over 200 morphological species of Plasmodium have been formally described based on morphology where 5 of them can infect humans: Plasmodium falciparum, Plasmodium vivax, Plasmodium malariae, Plasmodium ovale and Plasmodium knowlesi (Sato, Reference Sato2021). Despite their broad range of infection, some Plasmodium species are extremely host specialists such as P. falciparum, which infects humans but not African apes that are phylogenetically very close to humans (Liu et al., Reference Liu, Li, Learn, Rudicell, Robertson, Keele and Hahn2010). P. falciparum is transmitted by several anopheline species, where Anopheles gambiae is one of the most well-known vectors of human malaria (Gouagna et al., Reference Gouagna, Ferguson, Okech, Killeen, Kabiru, Beier, Githure and Yan2004). This mosquito–parasite association has been widely studied in the last century to bring information to design new strategies to reduce malaria transmission.
Another well-studied group of Plasmodium species are those affecting wild birds, i.e. avian malaria parasites (LaPointe et al., Reference LaPointe, Atkinson and Samuel2012). Avian malaria encompasses more than 40 morphologically described Plasmodium species (Atkinson, Reference Atkinson1991) but over 500 different lineages have been described using sequence divergence in the mitochondrial cytochrome b gene (Bensch et al., Reference Bensch, Pérez-Tris, Waldenströum and Hellgren2004, Reference Bensch, Hellgren and Pérez-Tris2009). These parasites are mainly transmitted by Culex mosquitoes (Fonseca et al., Reference Fonseca, Keyghobadi, Malcolm, Mehmet, Schaffner, Mogi, Fleischer and Wilkerson2004). Within this mosquito genera, Culex pipiens species complex may act as vector for Plasmodium species such as Plasmodium relictum (Lapointe et al., Reference LaPointe, Goff and Atkinson2010) and Plasmodium gallinaceum (Pruck-Ngern et al., Reference Pruck-Ngern, Pattaradilokrat, Chumpolbanchorn, Pimnon, Narkpinit, Harnyuttanakorn, Buddhirakkul and Saiwichai2015). The importance of studying P. relictum and its association with both its vertebrate and invertebrate hosts relies on the fact that it is one of the most widespread avian malaria parasites in the world (Kazlauskienė et al., Reference Kazlauskienė, Bernotienė, Palinauskas, Iezhova and Valkiūnas2013; Valkiūnas et al., Reference Valkiūnas, Ilgūnas, Bukauskaitė, Fragner, Weissenböck, Atkinson and Iezhova2018). Moreover, this malaria species is responsible for several bird species extinctions (Atkinson and Samuel, Reference Atkinson and Samuel2010) and is currently listed as one of the 100 most dangerous invasive species in the world (Boudjelas et al., Reference Boudjelas, Browne, De Poorter and Lowe2020).
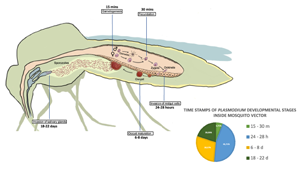
All Plasmodium species share a similar but complex life cycle (Votýpka et al., Reference Votýpka, Modrý, Oborník, Šlapeta and Lukeš2016) that involves 2 separate hosts: a vertebrate host and a mosquito vector (invertebrate host). Inside each host, the parasite undergoes multiple developmental stages. The life cycle of Plasmodium begins in the vertebrate host, when the sporozoites are expelled with the saliva of the female mosquito while is taking a blood meal. Inside the vertebrate host, the parasite undergoes different developmental stages that conclude with the production of gametocytes. The next step is the transmission of gametocytes to an invertebrate host (Fig. 1), which is achieved when a female mosquito feeds on infected blood. The ingested gametocytes of Plasmodium develop into male and female gametes in the midgut lumen. Inside the mosquito vector, the parasite reproduces sexually (Bennink et al., Reference Bennink, Kiesow and Pradel2016), and the fertilized gametes produce zygotes, the only diploid stage of the parasite, which develop into motile ookinetes that invade the epithelium of the midgut cells in the mosquito to reach its basal side. Ookinetes then develop into oocysts that produce several sporozoites, which mature in a period that varies depending on the Plasmodium species. Once the maturation period is over, the midgut sporozoites are released into the haemolymph and migrate to the salivary glands where they are ejected along the saliva into a new vertebrate to start a new life cycle (Vaughan, Reference Vaughan2007).
Developmental stages of Plasmodium during sexual reproduction inside its invertebrate host. Time points in the figure correspond to time post blood feeding (PBF). Gametogenesis occurs 15 min PBF when ingested gametocytes of Plasmodium develop into male and female gametes, followed by fecundation 30 min PBF leading to the production of zygotes, that develop into motile ookinetes. Invasion of the midgut cells by ookinetes takes place between 24 and 28 h PBF. Oocyst maturation takes place between 6 and 8 days PBF followed by release of sporozoites and migration to salivary glands that conclude with their ejection along the saliva into a new vertebrate 18–22 days PBF.

The family Culicidae comprises several genera, including Culex mosquitoes that diverged from Anopheles during the early Jurassic period (~160–200 million years ago, da Silva et al., Reference da Silva, Machado, de Paula, da Silva Pessoa Vieira, de Morais Bronzoni, de Melo Santos and Wallau2020; Lorenz et al., Reference Lorenz, Alves, Foster, Suesdek and Sallum2021). They are species of medical and veterinary importance that act as vectors for shared pathogen groups, such as Plasmodium spp. Most genomic studies are currently focused on Anopheles species since An. gambiae genome was completely sequenced more than 20 years ago (Holt, et al., Reference Holt, Subramanian, Halpern, Sutton, Charlab, Nusskern and Hoffman2002), and since then it has been widely used to investigate mosquito DNA expression patterns to Plasmodium infection. The genome of Culex quinquefasciatus was reported more recently (Arensburger et al., Reference Arensburger, Megy, Waterhouse, Abrudan, Amedeo, Antelo and Atkinson2010), showing great differences in genome size and in the total number of genes between the 2 mosquito species. In this sense, the genome of An. gambiae is smaller (278 Mb) than Cx. quinquefasciatus genome (579 Mb) and, therefore, the number of annotated genes is slightly bigger in Cx. quinquefasciatus (Severson and Behura, Reference Severson and Behura2012). However, although the information of the genomes of both mosquito species is available since long time ago, there is an important knowledge gap concerning gene expression in response to Plasmodium infection in the C. pipiens complex.
Here, we extensively review the current knowledge on the regulation of key genes of the avian mosquito vector Cx. quinquefasciatus, relevant during P. relictum infection. We also aim to compare the activation of genes expressing important immune and metabolic pathways during Plasmodium infections between the human and avian malaria mosquito vectors during Plasmodium infection, Anopheles and Culex, respectively, to highlight the limited number of genomic studies focusing on Culex. We further describe problems that may limit genomic research in Plasmodium-infected vectors, such as the time elapsing since the mosquito takes a blood meal to sampling point, the proportion of malaria-infected mosquito cells (parasitaemia), the variability of vector gene expression among collected tissues and specific parasite–vector associations.
Materials and methods
Our literature search was conducted in September 2022. Initial title, keyword, and abstract screening was performed using the items for systematic review and meta-analysis established in PRISMA (Moher et al., Reference Moher, Liberati, Tetzlaff and Altman2009) modified for Ecology and Evolution, PRISMA-Eco Evo (O'Dea et al., Reference O'Dea, Lagisz, Jennions, Koricheva, Noble, Parker, Gurevitch, Page, Stewart, Moher and Nakagawa2021). A systematic search on the available literature on genomic analysis of Anopheles and Culex mosquitoes infected with Plasmodium was performed on the Web of Science (WoS) database. The search was conducted in English. We included articles published between 2002 and 2023 years (see Supplementary Table 1) searching with specific Booleans (see below). The search string comprised 2 substrings. The first substring targeted mosquito genomic changes during infection using the following Boolean search keywords [Genom* AND transcriptom* Vector AND mosquito AND Anopheles OR Culex AND malaria AND association AND infection]. We retrieved 358 articles on WoS. The second substring aimed at the evolution of Anopheles and Culex linked to genomic and transcriptomic analysis during infection using the following Boolean search keywords [Genom* AND transcriptom* Vector AND mosquito AND Anopheles OR Culex AND malaria AND evolut*]. We retrieved 531 articles on WoS. After the removal of duplicates between the 2 substrings, we obtain a subtotal of 635 articles that were screened at title, keyword and abstract stage. Five hundred and sixty-eight articles were excluded for further analyses because they were not related to our aim of study. In consequence, full-text of the remaining 67 articles showing genomic and transcriptomic analysis of mosquitoes infected with Plasmodium were screened in a decision tree containing our inclusion/exclusion criteria, following the guidelines for systematic search and study screening for literature reviews in Ecology and Evolution proposed by Foo et al. (Reference Foo, O'Dea, Koricheva, Nakagawa and Lagisz2021) (Fig. 2). Twenty-three studies met all our inclusion criteria. All these studies examined genes involved in important immune and metabolic pathways in human or avian malaria mosquito vectors Anopheles and Culex during natural and non-natural Plasmodium infection.
Decision tree based on PRISMA framework. Inclusion/exclusion criteria were used to filter studies focused on genes involved in important immune and metabolic pathways in Anopheles and Culex during Plasmodium infection.

Current knowledge on the genomics of mosquito during malaria infections
An. gambiae has been used in many research studies to gain genetic insights that might help prevent and eradicate malaria. In this sense, transcriptomic analyses are a useful tool to understand the role of the mosquito in Plasmodium transmission, as they bring information about the regulation of RNA expression during an exact moment of a specific event during the parasite infection (Domingos et al., Reference Domingos, Pinheiro-Silva, Couto, do Rosário and de la Fuente2017). There is a huge number of studies providing information about fundamental aspects of An. gambiae gene expression during both non-natural (i.e. when infection experiments uses a combination of parasite and vector species that have not been observed in the wild) and natural malaria infection (Dong et al., Reference Dong, Aguilar, Xi, Warr, Mongin and Dimopoulos2006; Baton et al., Reference Baton, Robertson, Warr, Strand and Dimopoulos2009; Mead et al., Reference Mead, Li, Tu and Zhu2012; Biryukova et al., Reference Biryukova, Ye and Levashina2014; Ruiz et al., Reference Ruiz, Yerbanga, Lefèvre, Ouedraogo, Corces and Gómez-Díaz2019). Thanks to these studies the information regarding immune and physiological response linked to various parasite developmental stage is quite broad. However, transcriptomic studies focused on the Culex complex are still scarce, even during P. relictum infection.
Genes involved in immune responses
When a mosquito bites a non-infected vertebrate host, the expression of genes related with several biological processes important for reproduction and survival such as egg production or cell homoeostasis is affected (Bryant et al., Reference Bryant, Macdonald and Raikhel2010). Infected blood that contains malaria sporozoites activates different mechanisms linked to immune response (Luckhart et al., Reference Luckhart, Vodovotz, Cui and Rosenberg1998) or cell apoptosis (Ahmed and Hurd, Reference Ahmed and Hurd2006). In dipterans, there are 3 genes' categories involved in the innate immune response against Plasmodium which regulation is well described in An. gambiae: (i) recognition proteins of pathogen's components (Dong et al., Reference Dong, Aguilar, Xi, Warr, Mongin and Dimopoulos2006; Gendrin et al., Reference Gendrin, Yerbanga, Ouedraogo, Lefèvre, Cohuet and Christophides2016), (ii) components of signalling pathways related to the modulation, amplification, and transduction of cell signals (Chen et al., Reference Chen, Dong, Sandiford and Dimopoulos2012) and (iii) antimicrobial peptides (AMPs), complement factors and enzymes (Dixit et al., Reference Dixit, Sharma, Patole and Shouche2008; Clayton et al., Reference Clayton, Dong and Dimopoulos2014).
Mosquitoes, like the rest of invertebrates, rely on innate immunity as their only defence system (Christophides et al., Reference Christophides, Vlachou and Kafatos2004). Immune response occurs in several tissues of the mosquito: the midgut epithelium, lumen, haemolymph and within the salivary glands (Osta et al., Reference Osta, Christophides, Vlachou and Kafatos2004). When Plasmodium (or other infectious microorganisms) infect the mosquito, 2 main responses might be activated against the parasites: the humoral and cellular responses. Humoral response is formed by 3 main immune pathways: Toll, Imd and JAK/STAT (Dimopoulos et al., Reference Dimopoulos, Richman, Müller and Kafatos1997; Tikhe and Dimopoulos, Reference Tikhe and Dimopoulos2021). These immune pathways include different immune cascades that conclude with the transcriptional regulation of mechanisms that aims to clear the parasite from the vector (Dong et al., Reference Dong, Fu, Dong, Simões, Zhu and Dimopoulos2020). In cellular response, different immune components like enzymes (Dong et al., Reference Dong, Aguilar, Xi, Warr, Mongin and Dimopoulos2006) or specific cells like haemocytes physically isolate and destroy the parasite (Clayton et al., Reference Clayton, Dong and Dimopoulos2014). Interestingly, these immune responses reducing Plasmodium parasitaemia take place in 3 events of Plamodium life-cycle inside the mosquito vector: (i) the ookinete maturation, also limited by molecules from vertebrate host and digestive molecules from mosquito secreted into the bloodmeal (Sinden et al., Reference Sinden, Alavi and Raine2004), (ii) the invasion of the midgut by ookinetes and (iii) the sporozoite migration through the haemocoel to the salivary glands (Ghosh et al., Reference Ghosh, Ribolla and Jacobs-Lorena2001; Shahabuddin and Costero, Reference Shahabuddin and Costero2001; Sinden et al., Reference Sinden, Alavi and Raine2004). These mechanisms, are attributed to haemocyte-mediated immune responses (Frolet et al., Reference Frolet, Thoma, Blandin, Hoffmann and Levashina2006) that activate genes in the midgut of the mosquito (Dong et al., Reference Dong, Aguilar, Xi, Warr, Mongin and Dimopoulos2006).
Genomic studies in An. gambiae show a variety of results depending on the vector–parasite association. Most genomic studies of Anopheles mosquitoes have explored non-natural parasite–vector associations, such as, An. gambiae infected with a rodent malaria parasite (Plasmodium berghei). In genomic studies of immune response, rodent malaria parasite P. berghei is commonly used to experimentally infect An. gambiae for the identification of key genes for the innate immune system (Baton et al., Reference Baton, Robertson, Warr, Strand and Dimopoulos2009; Raddi et al., Reference Raddi, Barletta, Efremova, Ramirez, Cantera, Teichmann, Barillas-Mury and Billker2020). Nonetheless, to get a more realistic figure of how Plasmodium parasites trigger immune responses in the mosquito, such associations should be better assessed in natural parasite–vector associations, as many responses both in the vector and in the parasite might have co-evolved over long periods of time. For example, the infection of An. gambiae with the non-natural parasite P. berghei and An. gambiae led to a greater activation of the Toll pathway during the mosquito immune response (Clayton et al., Reference Clayton, Dong and Dimopoulos2014). On the contrary, the infection of An. gambiae with its natural parasite P. falciparum induced a greater activation of genes involved in the Imd pathway (Garver et al., Reference Garver, Dong and Dimopoulos2009; Dong et al., Reference Dong, Das, Cirimotich, Souza-Neto, McLean and Dimopoulos2011, Reference Dong, Aguilar, Xi, Warr, Mongin and Dimopoulos2006). Thus, studies focused on natural parasite-mosquito associations are essential for a better understanding of how malaria impacts on their vector gene expression. Up to date, transcriptomic studies analysing Cx. quinquefasciatus gene expression using natural parasite association provide valuable information about immune response in other parasite–vector associations apart from Anopheles. For example, Garcia-Longoria et al. (Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022) have recently presented the first transcriptomic study analysing the effect of P. relictum infection on Cx. quinquefasciatus immune response. In this natural parasite–vector association, infected mosquitoes showed a greater activation in the Toll pathway, followed by Imd pathway, through the up regulation of several receptors, translation factors and effectors. They also found up regulation of genes related to cellular response in melanization cascade elements, CLIP-domain serine proteases and serpins genes, indicating that these processes may play an important role in the defence against P. relictum. However, other natural avian malaria parasite–vector associations, such as the one between Culex species and P. relictum lineages, have received less attention, and further studies are required to need to verify the up or down regulation of immune pathways and cascades over the avian malaria infection.
Genes involved in metabolic pathways
During malaria infection, mosquitoes exhibit several changes in the expression of its genes involved in metabolism that are key to Plasmodium development (Vlachou et al., Reference Vlachou, Schlegelmilch, Christophides and Kafatos2005). For instance, malaria parasite gametogenesis is triggered by a molecule called xanthurenic acid (XA) (Garcia et al., Reference Garcia, Wirtz, Barr, Woolfitt and Rosenberg1998; McRobert et al., Reference McRobert, Taylor C, Deng, Fivelman, Cummings, Polley, Billker and Baker2008) which is an intermediate metabolite of tryptophan in the mosquito (Billker et al., Reference Billker, Lindo, Panico, Etienne, Paxton, Dell, Rogers, Sinden and Morris1998). This molecule induces intracellular rise in Ca2+ concentration to activate a protein kinase within the parasite, that regulate gametogenesis (gametocyte differentiation into male and female gametes) and Plasmodium transmission (Billker et al., Reference Billker, Dechamps, Tewari, Wenig, Franke-Fayard and Brinkmann2004; Brochet and Billker, Reference Brochet and Billker2016). Specifically, in P. berghei (Billker et al., Reference Billker, Dechamps, Tewari, Wenig, Franke-Fayard and Brinkmann2004) and in P. falciparum (McRobert et al., Reference McRobert, Taylor C, Deng, Fivelman, Cummings, Polley, Billker and Baker2008) this intracellular Ca2+ is essential for the exflagellation process.
Likewise, Guttery et al. (Reference Guttery, Pittman, Frénal, Poulin, Mcfarlane, Slavic, Wheatley, Soldati-Favre, Krishna, Tewar and Staines2013) demonstrated in laboratory conditions that environmental Ca2+ has an impact on P. berghei sexual development. They genetically modified PbCAX gene, a P. berghei Ca2+/H+ exchanger, which is important to maintain Ca2+ homoeostasis. As a result, parasites with experimentally disrupted genes failed to produce zygotes. Moreover, this process could be reversed in vitro by removing environmental Ca2+. They concluded that PbCAX is essential to tolerate Ca2+ within the ionic environment of the mosquito midgut, and ultimately, for ookinete development and differentiation within the mosquito. Interestingly, Ferreira et al. (Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022) experimentally infected wild-caught mosquitoes from the Hawaiian Islands and simulated natural conditions to reflect more reliable effects of the P. relictum–Cx. quinquefasciatus association. They showed important differences between infected and uninfected mosquitoes in the expression of genes related to calcium transportation 24 h and 5 days PFB. More specifically, they found that infected mosquitoes had higher expression levels of genes involved in calcium transportation or binding at 24 h PBF. Also, biological process related with endoplasmic reticulum calcium ion homoeostasis was significantly higher at 5 days PFB in infected mosquitoes (Ferreira et al., Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022).
Glucose is the main source of energy not only for the mosquito but also for the malaria parasite (Liu et al., Reference Liu, Dong, Huang, Rasgon and Agre2013). Blood stages of malaria parasites are dependent on glucose catabolism components such as adenosine triphosphate to obtain their main source of energy (Kirk et al., Reference Kirk, Horner and Kirk1996). Meireles et al. (Reference Meireles, Sales-Dias, Andrade, Mello-Vieira, Mancio-Silva, Simas, Staines and Prudêncio2017) demonstrated the role of glucose levels in the successful hepatic infection of P. berghei, where glucose levels below a standard medium concentration led to failed infection. In addition, they showed that there is an increase in glucose uptake via the GLUT1 transporter (class I facilitative glucose transporter expressed in liver cells) in P. berghei-infected hepatic cells.
Following this idea, Wang and Wang (Reference Wang and Wang2020) examined the function of the glucose transporter Asterglut1 in the non-natural association Anopheles stephensi–P. berghei. They found that the knockdown of the glucose transporter genes significantly increased the glucose level in the midgut of the mosquito prior to blood feeding and increased P. berghei infection, hence suggesting that Asteglut 1 participate in the defence against malaria infection. Ferreira et al. (Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022) evaluated the gene-expression response of Culex mosquitoes exposed to Plasmodium infection in a natural parasite–vector association, reporting a lower expression level in infected mosquitoes compared to control in a gene involved in glucose metabolism (6-phosphogluconate dehydrogenase) during ookinete invasion (24 h post feeding), and in an unknown sugar transporter gene (CPIJ008946) 10 days post feeding.
The same level of metabolic importance is attributed to solute carriers transporting ions helping to maintain ionic homoeostasis (Hirata et al., Reference Hirata, Czapar, Brin, Haritonova, Bondeson, Linser, Cabrero, Thompson, Dow and Romero2012). Recent studies have shown the importance of these solute carriers in both natural and non-natural parasite- vector associations. In An. gambiae experimentally infected with rodent malaria (P. berghei), infected mosquitoes showed an upregulation of solute carrier genes involved in cell homoeostasis in the salivary glands 18 days post blood feeding (PBF) (Couto et al., Reference Couto, Antunes, Pinheiro-Silva, do Rosário, de la Fuente and Domingos2017). Moreover, the infection of An. gambiae mosquitoes with P. berghei with knocked down solute carrier genes induced a reduction of the number of sporozoites in the salivary glands and increased mosquito death rate (Couto et al., Reference Couto, Antunes, Pinheiro-Silva, do Rosário, de la Fuente and Domingos2017). Accordingly, in a natural parasite–vector association, Cx. quinquefasciatus infected by P. relictum showed significantly higher expression of several anion and ion transporter genes 5 days PBF (Ferreira et al., Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022). These results might indicate that Plasmodium may exploit An. gambiae (Couto et al., Reference Couto, Antunes, Pinheiro-Silva, do Rosário, de la Fuente and Domingos2020) and Cx. quinquefasciatus cellular mechanisms to obtain resources to maximize its reproduction and transmission.
Problems related to work with mosquito–parasite association through transcriptomics
Parasitaemia levels (the proportion of malaria infected cells inside an organism) might affect host transcriptome response (Videvall et al., Reference Videvall, Palinauskas, Valkiūnas and Hellgren2020) as differences in the parasite load may harm and affect mosquitoes at a different scale. Accordingly, organisms with higher parasitaemia usually show higher amounts of differentially expressed genes compared to those with lower parasitaemia. In this sense, a previous study has suggested that a strong response to infection is accompanied by high parasitaemia rates (Videvall et al., Reference Videvall, Palinauskas, Valkiūnas and Hellgren2020). For example, in the case of Culex mosquitoes, 2 recent studies have shown different transcriptomic responses probably due to mosquitoes harbouring different levels of parasitaemia caused by 2 different avian malaria strains (P. relictum pGRW04 and pSGS1) (Ferreira et al., Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022; Garcia-Longoria et al., Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022). Garcia-Longoria et al. (Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022) experimentally infected Culex mosquitoes with P. relictum pSGS1, achieving fairly high levels of parasitaemia and, a significant amount of up-regulated immune genes. However, Ferreira et al. (Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022), analysed the gene expression of Culex mosquitoes naturally infected with P. relictum pGRW04, showing a low parasitaemia in these mosquitoes and a low number of significant genes responding to the infection.
Another weakness when analysing transcriptomic in insects during mosquito–parasite associations is limitation of tissue sampling, which can lead to tissue-biased expression (Baker et al., Reference Baker, Nolan, Fischer, Pinder, Crisanti and Russell2011). Plasmodium infection in mosquitoes is quite restricted to specific tissues where sporozoites are mainly detected in salivary glands, ookinetes in the gut walls and gametes inside the gut (Valkiūnas, Reference Valkiūnas2005). This differentiation potentially might complicate the detection of mosquito readings since the detection of transcriptomic signals can be masked by mosquito tissues that are more abundant in the sample. This is an important issue to deal with because it can lead to false negatives and, therefore, to lose information about differentially gene expression.
An important caveat in the study of mosquito–parasite association is the lack of a well assembled and annotated genome. In the case of Culex mosquitoes, previous studies have detected around 20% of uncharacterized genes in their analyses (see Ferreira et al., Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022, but also see Garcia-Longoria et al., Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022), thus highlighting the need to improve the annotation and gene prediction of the Culex assembled genomes. Nevertheless, this is not the case for genomes related to Anopheles family. Since the publication of An. gambiae genome by Holt et al. (Reference Holt, Subramanian, Halpern, Sutton, Charlab, Nusskern and Hoffman2002) several updates from this genome have been made resulting in high-quality reference genome where a high number of annotated genes can be detected when analysing transcriptome response (Sharakhova et al., Reference Sharakhova, Hammond, Lobo, Krzywinski, Unger, Hillenmeyer, Bruggner, Birney and Collins2007; George et al., Reference George, Sharakhova and Sharakhov2010; Kingan et al., Reference Kingan, Heaton, Cudini, Lambert, Baybayan, Galvin, Durbin, Korlach and Lawniczak2019).
Time points of sampling PBF is another crucial aspect in transcriptomic research, and it is directly related to malaria parasite development inside the mosquito (Fig. 3). The duration and timing of the different stages of the malaria life cycle differ between Plasmodium species, and it is determined by factors such as internal mosquito temperature and pH (Beier, Reference Beier1998). The arrangement of time points in these genomic analyses include a few minutes and hours after the blood meal was taken to several days after blood meal ingestion. There is a reduced number of studies analysing vector gene expression on a single time point in comparison to those focused on a range of different time points. In this sense, only 5 studies analysed the gene expression on a single time point PBF, whereas 18 studies were done using and an arrangement of different time points PBF (Fig. 3; See Supplementary Table 1).
Number of studies analysing gene expression at different times post blood feeding (PBF). X axis shows the number of articles focusing a specific sampling time interval and Y axis shows the different times of sampling. Single time points are shown in dark blue and arrangement of time points are shown in light blue.

Early time points PBF (from 30 min PFB to 12 h PFB) are used for detecting the initial effect of Plasmodium. For example, in Cx. quinquefasciatus it has been shown that only 2 receptors of Toll pathway (CPIJ019764, CPIJ018343) were significantly up-regulated at 30 min PBF, but there were not Toll transcription factors expressed at this time point (Garcia-Longoria et al., Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022). Most studies use a range from 18 to 28 h PBF (Dimopoulos et al., Reference Dimopoulos, Christophides, Meister, Schultz, White, Barillas-Mury and Kafatos2002; Vlachou et al., Reference Vlachou, Schlegelmilch, Christophides and Kafatos2005; Xu et al., Reference Xu, Dong, Abraham, Kocan, Srinivasan, Ghosh, Sinden, Ribeiro, Jacobs-Lorena, Kafatos and Dimopoulos2005; Baton et al., Reference Baton, Robertson, Warr, Strand and Dimopoulos2009; Mead et al., Reference Mead, Li, Tu and Zhu2012; Alout et al., Reference Alout, Ndam, Sandeu, Djégbe, Chandre, Dabiré, Djogbénou, Corbel and Cohuet2013; Nsango et al., Reference Nsango, Pompon, Xie, Rademacher, Fraiture, Thoma, Awono-Ambene, Moyou, Morlais and Levashina2013). Specially, 24 h PBF is a fairly used time point in genomic studies, because it is the time where Plasmodium ookinetes invade the epithelium of midgut cells and reach its basal side (Osta et al., Reference Osta, Christophides, Vlachou and Kafatos2004). Accordingly, Garrigós et al. (Reference Garrigós, Ylla, Martínez-de la Puente, Figuerola and Ruiz-López M2023) found that at 24 h PBF, P. relictum induced the expression of spätzle gene (CPIJ006792), a ligand of Toll receptors. This initial stage is then followed by the development of oocyst into sporozoites for 6–10. A large amount of transcription factors and its inhibitors are expected to be regulated during this period. In line with this idea, a striking up regulation of Toll receptors like Dorsal transcription factor within the Toll pathway (CPIJ002469) has been shown at 8 days PBF in Cx. quinquefasciatus, whereas its inhibitor the cactus protein CPIJ004774), is down regulated at this stage (Garcia-Longoria et al., Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022). Finally, a reduced number of studies use the range between 18–22 days (Xu et al., Reference Xu, Dong, Abraham, Kocan, Srinivasan, Ghosh, Sinden, Ribeiro, Jacobs-Lorena, Kafatos and Dimopoulos2005; Couto et al., Reference Couto, Antunes, Pinheiro-Silva, do Rosário, de la Fuente and Domingos2017; Zhang et al., Reference Zhang, Huang, Zhu and Zhang2017; Carr et al., Reference Carr, Rinker, Dong, Dimopoulos and Zwiebel2021; Garrigós et al., Reference Garrigós, Ylla, Martínez-de la Puente, Figuerola and Ruiz-López M2023), which is the stage related to sporozoite migration to the salivary glands (Amino et al., Reference Amino, Giovannini, Thiberge, Gueirard, Boisson, Dubremetz, Prévost, Ishino, Yuda and Ménard2008). A smaller number of transcription factors and receptors are expected to be expressed 22 days PBF. According to this, Garcia-Longoria et al. (Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022) reported no differences in the expression of both the toll transcription factors and its inhibitor proteins between Plasmodium-infected and uninfected Culex mosquitoes at 22 days PBF.
Discussion
Metabolic and immune response of mosquitoes during Plasmodium infection affects parasite fitness, by limiting its capacity to survive within the host, to reproduce and to be transmitted into new hosts. Since Plasmodium reproduction and transmission is linked to mosquito derived molecules such as XA, the expression of genes related to tryptophan metabolism could be a targeted by the parasite to increase its fitness. Following this idea, Ferreira et al. (Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022) suggested that since calcium is essential for Plasmodium ookinete motility and gametogenesis (Luckhart et al., Reference Luckhart, Vodovotz, Cui and Rosenberg1998), an enhanced expression of mosquito calcium transporters could supply malaria parasites with Ca2+ to facilitate midgut invasion. Alternatively, an enhanced expression of calcium transporters (as shown in Ferreira et al., Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022) could be a mosquito response to malaria infection, because high levels of environmental Ca2+ can be a threat to parasite homoeostasis and limit Plasmodium development (Guttery et al., Reference Guttery, Pittman, Frénal, Poulin, Mcfarlane, Slavic, Wheatley, Soldati-Favre, Krishna, Tewar and Staines2013). Nevertheless, there is not an agreement about the real effect of mosquito-derived calcium and the implication of mosquito transporters during Plasmodium development within the mosquito.
Also, while most of the studies focused on metabolic components relevant in malaria transmission are primarily focused on Anopheles (Adedeji et al., Reference Adedeji, Ogunlana, Fatumo, Beder, Ajamma, Koenig and Adebiyi2020), there is little information about the role of metabolic gene regulation during avian malaria infection in Culex. Whether these components of metabolism might play a role in the success of avian malaria infection in invertebrate vector requires further investigation.
Natural and non-natural Plasmodium-vector associations have significantly different profiles for immune activation (Sreenivasamurthy et al., Reference Sreenivasamurthy, Dey, Ramu, Kumar, Gupta, Mohanty, Harsha, Sharma, Kumar, Pandey, Kumar and Prasad2013). As showed in previous sections, non-natural associations in Anopheles–Plasmodium activate the Toll pathway while in natural associations in Anopheles–Plasmodium takes place to the Imd pathway activation (Garver et al., Reference Garver, Dong and Dimopoulos2009; Dong et al., Reference Dong, Das, Cirimotich, Souza-Neto, McLean and Dimopoulos2011; Reference Dong, Aguilar, Xi, Warr, Mongin and Dimopoulos2006). Additionally, natural Culex–Plasmodium associations take place to the activation of both immune pathways (Garcia-Longoria et al., Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022). Interestingly, both immune pathways are supposed to be established early in the evolution of metazoa (Hoffmann et al., Reference Hoffmann, Kafatos, Janeway and Ezekowitz1999); Imd pathway is supposed to be more effective towards Plasmodium (Meister et al., Reference Meister, Kanzok, Zheng, Luna, Li, Hoa, Clayton, White, Kafatos, Christophides and Zheng2005), whereas Toll pathway is more specific towards bacteria and fungi in mosquitoes (Tikhe and Dimopoulos, Reference Tikhe and Dimopoulos2021). However, differences in immune gene expression depending on the mosquito–parasite association are largely unknown among vector species. It could be hypothesized that these differences between pathway activation in natural and non-natural associations between Anopheles and Culex may be linked to differences in parasitaemia, or disparities in co-evolutionary history between hosts and parasites, among others. Nonetheless, more studies comparing side-by-side natural and non-natural mosquito vector–malaria parasite associations need to be explored.
Regarding the problem of parasitaemia and transcriptome response, the different outcomes in gene expression showed by Garcia-Longoria et al. (Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022) and Ferreira et al. (Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022) highlight that parasitaemia is an important limitation when analysing transcriptomic response during a host–parasite association. The different levels of parasitaemia achieved during the infection of the mosquito Cx. quinquefasciatus by 2 closely related P. relictum parasites (pSGS1 and pGRW04) might explain differences in transcriptomic responses between these 2 studies. Future researchers should take these restrictions into account during the experimental design and try to go deeper in the effects that parasitaemia caused on transcriptomic results depending on the avian malaria strain.
The limitation imposed by the potential bias in gene expression determined by the origin of the collected tissue should be addressed in further studies. In this sense, it becomes essential to differentiate the RNA expression analyses depending on the tissue through a prior dissection of the mosquitoes and sequencing specific tissues as a whole. However, including a dissection step into the analysis might also affect the observed gene regulation due to the possibility that the RNA will degenerate during the process. In avian malaria–vector associations, tissue-specific research is still a pending task, but it has been explored in several studies in human malaria (Dixit et al., Reference Dixit, Sharma, Mourya, Kamaraju, Patole and Shouche2009; Sreenivasamurthy et al., Reference Sreenivasamurthy, Madugundu, Patil, Dey, Mohanty, Kumar, Patel, Wang, Kumar, Pandey and Prasad2017). Previous studies in Anopheles mosquitoes focused on specific tissues have been able to detect not only parasite strategies for avoiding mosquito immune system (Xu et al., Reference Xu, Dong, Abraham, Kocan, Srinivasan, Ghosh, Sinden, Ribeiro, Jacobs-Lorena, Kafatos and Dimopoulos2005) but also to differentiate gene expression of individual sporozoites through single-cell RNA sequencing (Ruberto et al., Reference Ruberto, Bourke, Merienne, Obadia, Amino and Mueller2021).
Perhaps, the key stone to understand mosquito gene regulation during malaria infection would be to increase the number of annotated genes. Future improvements in Culex genomes would help to further understand how these mosquito families respond to malaria infection and the degree to which they have evolved immunity along different or similar evolutionary routes.
Finally, sampling time PBF should be considered a crucial issue in genomic studies, since different time points PBF are linked to different outcomes in expression patterns. Studies using a wide arrangement of time points PBF are useful to understand the changes of mosquito gene expression along the parasite development, and could be important for designing new tools for malaria control. Future studies should analyse vector gene expression over a wider range of time points PBF to extend the knowledge about malaria in early and long-term effects.
Although early time points are the less studied, a recent study has reported that a great number of genes are differentially expressed in Culex at 24 h PBF (Garrigós et al., Reference Garrigós, Ylla, Martínez-de la Puente, Figuerola and Ruiz-López M2023). After this time point, there is a substantial reduction in the number of genes expressed by the mosquito over time. Studies focusing on initial stages of infection could bring important information about gene expression during these early steps of infection, and even clues to examine potential adaptive parasite manipulations on the invertebrate host.
Concluding remarks
Genomic studies are an essential tool to understand the dynamic of vector borne diseases like malaria. It is crucial to reveal the regulatory genomic changes in vectors during parasite development, as it might lead to key information that can be used to prevent the spreading of the disease. Many genomic studies have been centred on natural and to a large degree non-natural associations between Plasmodium species and the mosquito vector An. gambiae. Regarding the human system, more studies based on natural parasite–vector associations P. falciparum and An. gambiae are needed to clearly identify the coevolution between these 2 organisms. The information focusing on the natural association between Plasmodium and other genera of mosquito vectors, such as Culex, is even scarcer. A very representative example is the C. pipiens complex transmitting avian malaria parasites, where only 3 natural Culex–Plasmodium associations have been explored. Specifically, between 1 mosquito species, Cx. quinquefasciatus and 2 P. relictum lineages (pGRW04, pSGS1) (Ferreira et al., Reference Ferreira, Videvall, Seidl, Wagner, Kilpatrick, Fleischer and Fonseca2022; Garcia-Longoria et al., Reference Garcia-Longoria, Ahrén, Berthomieu, Kalbskopf, Rivero and Hellgren2022) and 1 P. cathemerium lineage (PADOM02) (Garrigós et al., Reference Garrigós, Ylla, Martínez-de la Puente, Figuerola and Ruiz-López M2023), which have shown differences in transcriptomic responses. Future research should use up-to-date RNA-sequencing techniques and optimized sampling protocols to efficiently explore the effects of natural associations in Culex gene expression during different stages of parasite development, and compare their results with those obtained from natural and non-natural vector–parasite associations in human malaria. By doing this, genomic differences and similarities between Anopheles and Culex mosquitoes infected with Plasmodium parasites would help for a better understanding on how these 2 distantly related vector species respond over the infection, and also on how the parasite might manipulate vector gene expression for its own benefit. This understanding would be useful for the development of new molecular techniques for malaria control.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182023000756
Authors’ contribution
Conceptualization and methodology I.H.-C., L.G.-L. and O.H; literature search I.H.-C.; writing – original draft, I.H.-C. and L.G.-L.; writing – review and editing, I.H.-C., L.G.-L. and O.H.; funding acquisition, L.G.-L. and O.H. All authors have read and agreed to the published version of the manuscript.
Financial support
Funding was provided by the Consejería de Economía e Infraestructura of the Junta de Extremadura and the European Regional Development Fund, a Way to Make Europe (research project IB20089) to L.G.-L. and I.H.-C.; Swedish Research Council (grants 2016-03419 and 2021-03663) and Nilsson-Ehle foundation to O.H.
Competing interest
None.
Ethical standards
Not applicable.