Introduction
β-Carotene is a tetra-terpenoid consisting of a C40 structure including two β-ionone rings. Together with lycopene, it is among the most frequently consumed dietary carotenoids in human subjects(Reference Biehler, Alkerwi and Hoffmann1–Reference Wawrzyniak, Hamulka and Friberg3), also ranking among the highest in blood / plasma concentrations(Reference Wawrzyniak, Hamulka and Friberg3, Reference Fraser, Jaceldo-Siegl and Henning4). β-Carotene is the most important source of non-preformed vitamin A, as this molecule can, following absorption, be cleaved to form vitamin A (retinal). The maintenance of normal vision, enhancement of growth, tissue differentiation and reproduction was associated with β-carotene's dietary intake via fruits and vegetables as well as its blood concentrations, and have been further also associated with reduced incidence or disease biomarkers of several chronic complications such as type-2 diabetes(Reference Sluijs, Cadier and Beulens5, Reference Sugiura, Nakamura and Ogawa6) and other CVD(Reference Wang, Chung and McCullough7, Reference Karppi, Kurl and Ronkainen8).
However, highly dosed supplemental use of β-carotene has been correlated with smokers with negative outcomes, increasing total mortality(Reference Bjelakovic, Nikolova and Gluud9, Reference Bjelakovic, Nikolova and Gluud10) and enhancing lung cancer rate(Reference Omenn, Goodman and Thornquist11, 12). While nutritional relevant levels of β-carotene have been in the range of 0·1–8·8 (median 3·9) mg/d as reviewed recently(Reference Böhm, Borel and Corte-Real13), isolated administered β-carotene at rather high doses (>30 mg/d) may initiate previously mentioned negative effects. The reasons for these negative effects are not entirely clear but indicate that such high levels of β-carotene, which are usually non-harmful, may affect precancerous lesions, which are a hallmark of cancer development, and result from smoking(Reference van Helden, Godschalk and Swarts14). Studies in ferrets indicated altered non-beneficial retinoid signalling in the lung(Reference Wang, Liu and Bronson15). These studies suggest prudency with regards to form and dosing of β-carotene, while smoking appears as the major trigger of disease formation.
While earlier studies have emphasised direct antioxidant effects of β-carotene, including quenching singlet oxygen and lipid peroxides(Reference van Helden, Godschalk and Swarts14, Reference Krinsky and Johnson16, Reference Krinsky and Yeum17), more recent studies have especially highlighted that besides carotenoids, mainly carotenoid-metabolites obtain a role in altering gene expression(Reference Piga, van Dartel and Bunschoten18) reviewed in Kaulmann and Bohn(Reference Kaulmann and Bohn19), mainly via the interaction of the β-carotene- / retinol-metabolite all-trans-retinoic acid (ATRA)(Reference Rühl, Bub and Watzl20) with nuclear receptors, including retinoic acid receptor (RAR) and retinoid X receptor (RXR)(Reference Prakash, Liu and Hu21, Reference Ben-Dor, Nahum and Danilenko22). Nuclear hormone-mediated signalling is not directly but indirectly related to β-carotene intake. Initially, further cleavage and cleavage metabolites, especially the active vitamin A compounds (ATRA) are formed by β-carotene oxygenase 1 (BCO1) centric cleave or following eccentric cleavage by BCO2(Reference Amengual, Widjaja-Adhi and Rodriguez-Santiago23), which underlie a complex regulation in the organism, as explained further in the present paper. In addition and with an unclear nutritional and physiological relevance, transcription factors such as NF-κB and nuclear factor erythroid 2-related factor 2 (Nrf2) may also be involved in β-carotene-mediated signalling.
Recent studies, including gene-association studies(Reference Borel, Desmarchelier and Nowicki24–Reference Borel, Desmarchelier and Nowicki26), have suggested that the bioavailability and bioactivity of β-carotene are related to key steps determining β-carotene absorption, distribution, metabolism and excretion (ADME), and are summarised as following:
(a) the dietary release of β-carotene from the food matrix and its micellisation;
(b) its cellular uptake into the enterocytes;
(c) transport and metabolism of β-carotene, i.e.
(c1) the intracellular metabolism of β-carotene in the enterocyte;
(c2) or alternatively, the further transport of β-carotene within the organism and metabolism in target tissues;
(d) further transport and bio-distribution of carotenoids or carotenoid-metabolites;
(e) transmission and regulation of biological-mediated functions;
(f) excretion.
Consequently, there is a large variability of β-carotene bioavailability in human subjects, due to both nutritional- and host-related factors, including genetic variations(Reference Desmarchelier and Borel27, Reference Bohn, Desmarchelier and Dragsted28).
In the present review and position paper, we aim to pinpoint and highlight these critical steps in the metabolism of β-carotene, which likely constitute the strongest levers regarding bioavailability and bioactivity. Focusing on gaps of knowledge, especially the further metabolism following cleavage by BCO1 and transmission of biological-mediated signalling and regulation/autoregulation of these pathways, are in the focus of this review.
β-Carotene during digestion
The majority of β-carotene in native plant matrices is present in the all-trans form(Reference Lessin and Schwartz29). Following food processing and especially heat treatment, various cis-isomers are formed(Reference Marx, Stuparic and Schieber30), such as the 9-cis, 13-cis and 15-cis isomers(Reference Lessin and Schwartz29), while other isomers are much less abundant. Processes such as novel emerging non-thermal food processing technologies, such as high-pressure processing, high-intensity pulsed electric fields and ultrasonication(Reference Cilla, Bosch and Barberá31) can cause some structural changes in carotenoids such as trans–cis isomerisation, potentially altering their solubility and their micellisation efficacy.
Following ingestion and mastication of the food matrix in the oral phase of digestion, the bolus is passed on to the stomach where the food matrix is further mixed and plant cells further macerated. Using in vitro studies, human mastication was determined to enhance the release of β-carotene from the plant matrix in one study by approximately 35 % during in vitro gastric and intestinal digestion. The particle size and the type of chewing had more impact on carotenoid bioaccessibility than cell wall presence, with smaller particle size and more fine chewing significantly enhancing bioaccessibility(Reference Low, D'Arcy and Gidley32), presumably due to enhanced access of digestion enzymes. In line with these results, emulsions with small droplet diameters (0·2 v. 23 µm) improved β-carotene transfer from lipid droplets to mixed micelles and bioaccessibility from approximately 35–60 %(Reference Salvia-Trujillo, Verkempinck and Sun33).
To a low extent, the acidic pH (3–5) of the stomach may lead to small losses of β-carotene, resulting in the formation of carotenoid-cations (CarH+) first(Reference Konovalov and Kispert34), which may then result in trans–cis isomerisation. However, most studies investigating digestion do not suggest that significant isomerisation takes place under physiological conditions(Reference Ferruzzi, Lumpkin and Schwartz35–Reference Tyssandier, Reboul and Dumas37). Losses of β-carotene during digestion were reported from in vitro studies to range in the area of 30–70 %(Reference Blanquet-Diot, Soufi and Rambeau38, Reference Courraud, Berger and Cristol39), especially in the presence of oxidising compounds such as iron(Reference Kopec, Gleize and Borel40), likely resulting in the formation of β-apo-carotenals and epoxides(Reference Sy, Dangles and Borel41, Reference Sy, Dangles and Borel42). However, a recent clinical study using C13 β-carotene supports the conclusion of the only published clinical study dedicated to this topic, i.e. that β-carotene is fairly robust to human digestive conditions, with total losses <2 %(Reference Kopec, Caris-Veyrat and Nowicki43). Regarding host-produced enzymes, gastric lipase has been reported to be responsible for the digestion of approximately 5–40 % of ingested lipids(Reference Alminger, Aura and Bohn44), breaking down TAG, which are present together with carotenoids in lipid droplets. To which extent variations in gastric lipase concentrations translate into altered β-carotene bioaccessibility is not known. A practical limitation to study such effects is the non-availability of recombinant gastric-lipase or a suitable replacement which would allow studying effects more realistically in vitro (Reference Alminger, Aura and Bohn44).
In the small intestine, under the influence of bile acids, free fatty acids (FFAs), mono- and di-acylglycerols and phospholipids, originating mostly from the diet following digestion and under the influence of pancreatic lipase, lipid droplets are further processed to allow for mixed micelles formation of about 3–8 nm diameter(Reference Sy, Gleize and Dangles45, Reference Parker46). Proteins may aid in emulsification(Reference Soukoulis and Bohn47), but may also act as inhibitors thereof, preventing their transfer to mixed micelles(Reference Saini, Nile and Park48). A higher content of lipase and bile likely foster micellisation(Reference Bohn, Desmarchelier and Dragsted28, Reference Corte-Real, Desmarchelier and Borel49). Likewise, the presence of lipids in a meal rather foster micellisation(Reference Bohn50, Reference Borel51), while fibre(Reference Palafox-Carlos, Ayala-Zavala and Gonzalez-Aguilar52) or high minerals may hamper micelle formation(Reference Biehler, Hoffmann and Krause53, Reference Borel, Desmarchelier and Dumont54). The more apolar carotenoids, including β-carotene, are hypothesised to accumulate in core of the mixed micelle. The mixed micelles then diffuse through the mucus layer to the unstirred water layer of the enterocytes in the small intestine, where β-carotene may be taken up (Fig. 1).
(Colour online) Processing of β-carotene during digestion. All factors that impinge on matrix-release, transfer from lipid droplets to mixed micelles, and their diffusion to the enterocyte surface can alter bioaccessibility and thus bioavailability of β-carotene. By contrast, the influence of the colon and its microbiota remains unclear.
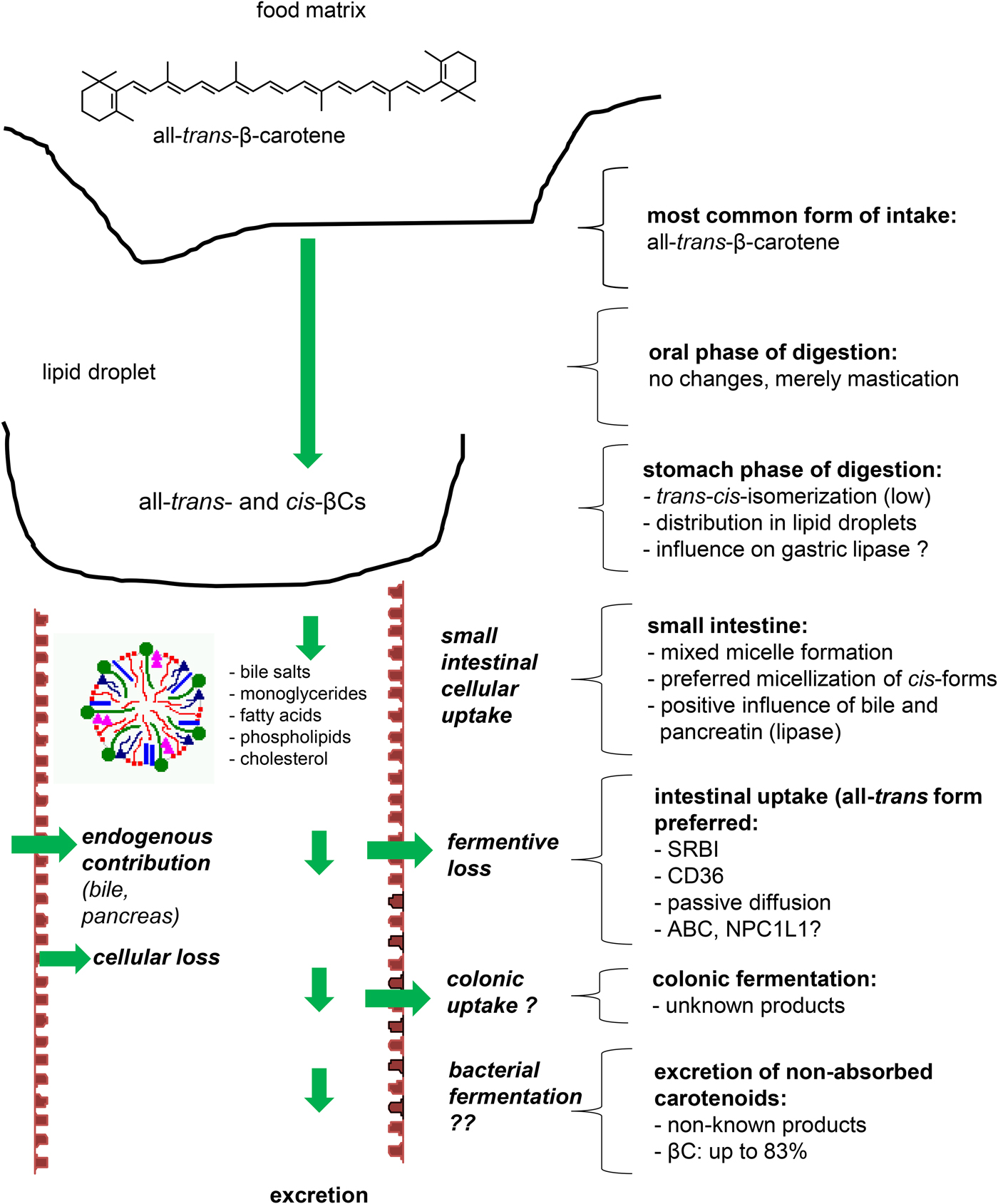
Though micellisation of cis-isomers is higher than their all-trans forms(Reference Ferruzzi, Lumpkin and Schwartz35), cellular uptake appears higher for the all-trans-form(Reference During, Hussain and Morel55). Whether and to what extent cellular uptake depends on the mucus layer is still unclear, but the mucus layer does not appear to constitute a barrier to absorption(Reference Kaulmann, Andre and Schneider56). While β-carotene absorption was earlier thought to occur mostly by passive diffusion(Reference Hollander and Ruble57), the involvement of several apical membrane proteins (transporters), including scavenger-receptor class B-type I (SR-B1(Reference Sun58)), also known as SR-BI(Reference During, Dawson and Harrison59) as well as CD36(Reference During, Dawson and Harrison59–Reference Borel, Lietz and Goncalves62) has meanwhile strongly been suggested, following studies with compounds inhibiting these proteins or in models over-expressing them. Genetic association studies have indeed found that SNP in SCARB1 (Reference Borel, Lietz and Goncalves62) and ABCA1 (Reference Borel, Desmarchelier and Nowicki25) were correlated with β-carotene bioavailability as determined by plasma and plasma-triglyceride-rich lipoproteins appearance, respectively (summarised in Supplementary table 1). However, it is not known whether these proteins have a direct or an indirect effect on β-carotene absorption. Indeed, recent results suggest that CD36 may indirectly modulate the apical to intracellular flux of β-carotene by modulating the synthesis rate of chylomicrons(Reference Buttet, Traynard and Tran63) in which β-carotene is incorporated. A recent candidate gene association study has suggested that ABCG5, and thus the heterodimer ABCG5/G8, could be involved in the efflux of a fraction of absorbed β-carotene back to the intestinal lumen(Reference Borel, Desmarchelier and Nowicki25). It is not clear if and to which extent Niemann-Pick disease, type C1, gene-like 1 or other membrane proteins participate in β-carotene absorption(Reference During, Dawson and Harrison59).
Finally, following gastro-intestinal digestion, a large proportion of non-absorbed β-carotene reaches the colon (as much as 50–95 %). It is unclear what happens under the influence of the gut microbiota(Reference Bohn64), but it has been shown that a large proportion of β-carotene is degraded into unknown compounds(Reference Kaulmann, Andre and Schneider56, Reference Goni, Serrano and Saura-Calixto65), as much as 98 % for pure β-carotene(Reference Serrano, Goni and Saura-Calixto66). However, a report by Mosele et al.(Reference Mosele, Macia and Romero67) points out to a high stability of β-carotene following in vitro colonic fermentation. The remainder, according to a report up to 83 %(Reference Shiau, Mobarhan and Stacewicz-Sapuntzakis68), is thus excreted in the faeces. It is likely that the matrix and microbiota differences add significantly to this variation. However, at present, there is no evidence that the colon plays a significant part in the absorption of β-carotene or its metabolites.
In summary, all factors that influence micellisation are likely to influence further β-carotene bioavailability and ADME aspects. The crucial part that micellisation plays for bioaccessibility is also well reflected by the high correlation between in vitro derived bioaccessibility and measures of bioavailability in vivo (Reference Tyssandier, Reboul and Dumas37, Reference Reboul, Richelle and Perrot69).
Intracellular metabolism and basolateral secretion of β-carotene by enterocytes
After having crossed the apical membrane, β-carotene must cross the polarised intestinal cell to be secreted at its basolateral side (Fig. 2A). Little is known about the intracellular transport and metabolism of β-carotene in the enterocyte. Nevertheless, since β-carotene is insoluble in water, intracellular binding protein(s) is/are likely to be involved(Reference Reboul and Borel70). This protein could be BCO1, which is mainly localised in the cytosol of mature enterocytes from the jejunum(Reference Duszka, Grolier and Azim71), as it is the main enzyme cleaving β-carotene(Reference Amengual, Widjaja-Adhi and Rodriguez-Santiago23, Reference Grolier, Duszka and Borel72–Reference dela Sena, Narayanasamy and Riedl74), and because it has a great affinity for β-carotene. An intracellular transport protein could also be BCO2, although we assume that its mitochondrial localisation(Reference Amengual, Lobo and Golczak75) is not compatible with its involvement as an intracellular transporter of β-carotene. An intracellular β-carotene transporter could also be a fatty acid binding protein (FABP), more likely liver FABP (L-FABP / FABP1), which is also present in the intestine and displays high-affinity binding for various hydrophobic ligands(Reference Gajda and Storch76).
(Colour online) (A) Candidate proteins for β-carotene metabolism within the enterocyte. When genetic variants have been associated with β-carotene bioavailability(Reference Borel, Desmarchelier and Nowicki25), the encoded proteins are coloured in grey. Dotted lines indicate regulations, i.e. regulation of BCO1 and SR-BI expression by ISX and regulation of chylomicron synthesis by SR-BI and CD36. (B) Candidate proteins that can modulate postprandial blood chylomicron β-carotene concentrations. When genetic variants have been associated with postprandial chylomicron β-carotene response to dietary β-carotene(Reference Borel, Desmarchelier and Nowicki25), the encoded proteins are coloured in grey. The dotted line indicates that this pathway is assumed but not demonstrated. (C) Proteins involved in the liver metabolism of β-carotene. Note that, to focus on β-carotene and for improved clarity, the fate of chylomicron retinyl esters in the liver is not shown, as well as the liver metabolism of retinol that involves numerous proteins(Reference Borel and Desmarchelier235). The liver is the hub of β-carotene metabolism: it is the main organ that stores β-carotene and distributes it to the peripheral tissues. β-Carotene reaches the liver mainly as β-carotene and retinyl esters, mainly RP, originating from β-carotene cleavage in the enterocyte and incorporated in chylomicrons. β-Carotene is then mostly stored in hepatic stellate cells. When genetic variants have been associated with blood β-carotene concentrations(Reference Hendrickson, Hazra and Chen80, Reference Yabuta, Urata and Wai Kun81, Reference Ferrucci, Perry and Matteini236), the encoded proteins are coloured in grey. βC: β-carotene, ABCA1: ATP binding cassette subfamily A member 1, ABCB1: ATP-binding cassette, sub-family B (MDR/TAP), member 1, ABCG5/G8: ATP-binding cassette, sub-family G member 5 and 8, ATRA: all-trans-retinoic acid, BCO1: β-carotene oxygenase 1, BCO2: β-carotene oxygenase 2, BCO2: β-carotene oxygenase 2, CD36: CD36 molecule, CXCL8: C-X-C motif chemokine ligand 8, ELOVL2: elongation of very long chain fatty acids protein 2, FABP: fatty acid binding protein, GPIHBP1: glycosylphosphatidylinositol-anchored high density lipoprotein binding protein 1, HL: hepatic lipase (encoded by LIPC), HSPGs: heparan sulphate proteoglycans, ISX: intestine specific homoeobox (transcription factor under the control of retinoic acid, regulating expression of SR-BI and BCO1), LDLR: LDL-receptor, LPL: lipoprotein lipase, LRP1: LDL-receptor-related protein 1, MTP: microsomal TAG transfer protein, NPC1L1: Niemann Pick C1-like 1, PKD1L2: polycystin 1-like 2 (gene/pseudogene), RBP4: serum retinol-binding protein, ROL: retinol, RP: retinyl palmitate and other retinyl esters coming from βC cleavage in the enterocyte, RPE65: retinal pigment epithelium-specific 65 kDa protein, SAR1B: secretion associated Ras-related GTPase 1B, SOD2: superoxide dismutase 2, SR-BI: scavenger receptor class B type I, TCF7L2: transcription factor 7-like 2, TTR: transthyretin.
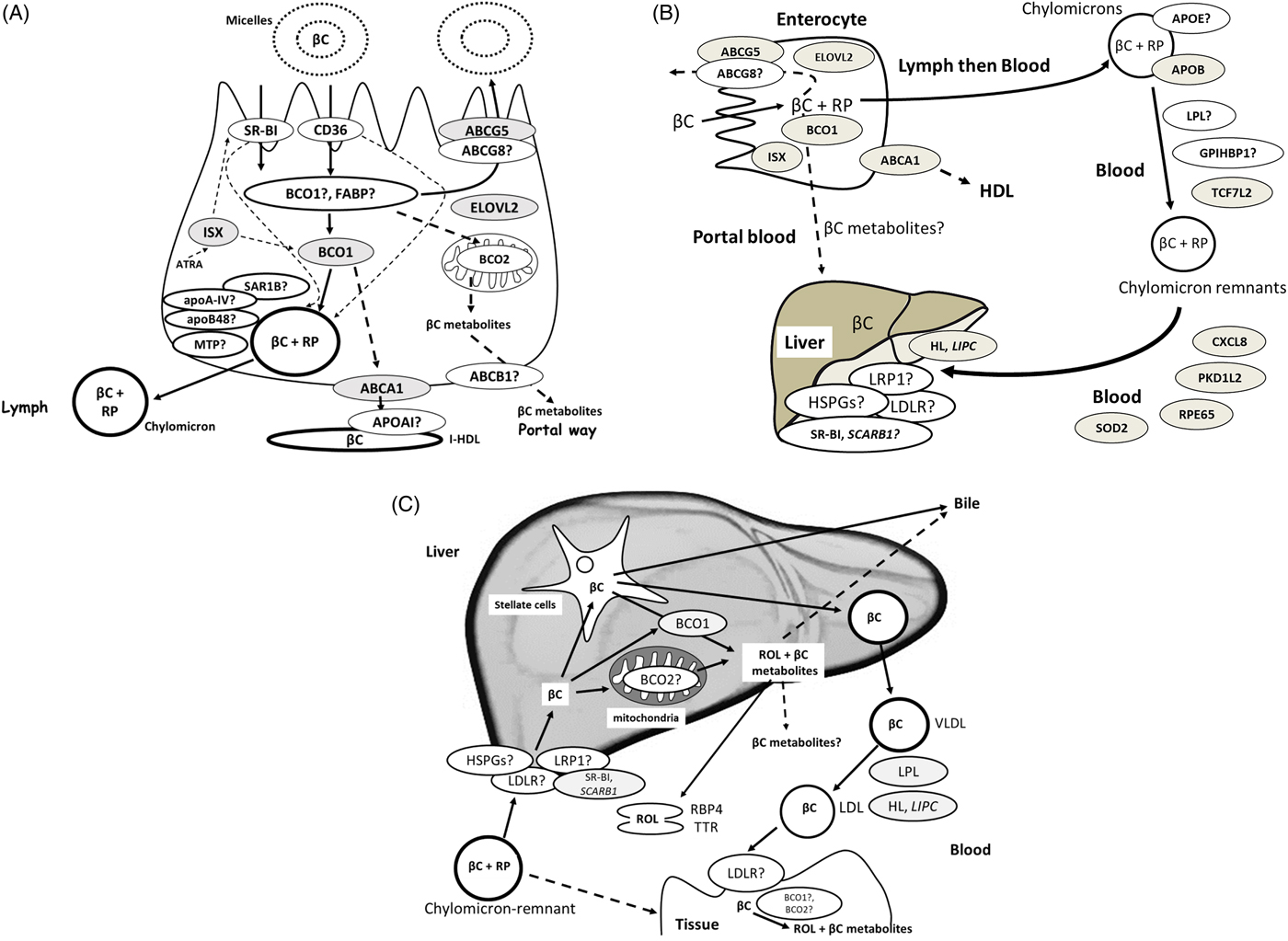
At this step, it is important to emphasise that only a fraction of absorbed β-carotene is metabolised in the enterocyte. The importance of this fraction, which was estimated at about 70 % by using stable isotope methods(Reference Tang, Qin and Dolnikowski77), likely depends on the vitamin A status of the body (see the next section). The secretion mechanism of β-carotene at the basolateral side of the enterocyte likely depends on its centric cleavage by BCO1, producing retinal, which is then, following conversion to retinol, mainly re-esterified by lecithin-retinol acyl-transferase (LRAT). It is assumed that the parent molecule is incorporated in nascent chylomicrons(Reference Borel, Grolier and Mekki78), while the less apolar β-carotene metabolites, which are produced by eccentric cleavage by BCO2, are secreted in the portal blood. Indeed, one genome-wide association study(79) and two candidate gene association studies(Reference Hendrickson, Hazra and Chen80, Reference Yabuta, Urata and Wai Kun81) have shown that SNP in BCO1, the main β-carotene metabolising enzyme, were associated with blood plasma β-carotene concentration. Other gene-association studies involving SNP in BCO1 and postprandial β-carotene and retinyl palmitate responses(Reference Lietz, Oxley and Leung82, Reference Leung, Hessel and Meplan83) confirmed that this gene and its variants are key regulators of blood concentrations of these vitamin A forms.
The mechanisms responsible for the incorporation of β-carotene in chylomicrons are poorly understood. It is hypothesised that they involve enzymes/Apo responsible for the assembly of chylomicrons, e.g. microsomal TAG transfer protein (MTP), apoA-IV, secretion associated Ras-related GTPase 1B (SAR1B) and apoB48. A recent candidate gene association study(Reference Borel, Desmarchelier and Nowicki25) has also suggested that the protein involved in intestinal HDL secretion(Reference Brunham, Kruit and Iqbal84), i.e. ATP binding cassette subfamily A member 1 (ABCA1), may also be involved in β-carotene secretion in intestinal HDL.
Finally, we suggest that the secretion of β-carotene metabolites in the portal blood might involve basolateral membrane proteins that can aid in the efflux of these metabolites, e.g. ATP binding cassette subfamily B member 1 (ABCB1), which encodes for P-glycoprotein(Reference Harrison85). Furthermore, several candidate gene association studies have shown that genetic variants in elongation of very long chain fatty acids protein 2 (ELOVL2, also termed ELOVL fatty acid elongase 2) play a significant role in carotenoid absorption(Reference Borel, Desmarchelier and Nowicki24–Reference Borel, Desmarchelier and Nowicki26). This is possibly due to the inhibitory effect of EPA, which is further elongated to docosapentaenoic acid and DHA by ELOVL2, on β-carotene absorption(Reference Mashurabad, Kondaiah and Palika86).
It is now acknowledged that vitamin A status can regulate β-carotene absorption and cleavage efficiency via a negative feedback loop: the higher the vitamin A status, the lower β-carotene absorption efficiency and cleavage, and inversely. The mechanism involves an intestinal transcription factor termed intestine specific homoeobox (ISX), which acts as a repressor of SCARB1 and BCO1 upon ATRA activation(Reference Lobo, Amengual and Baus87, Reference Lobo, Hessel and Eichinger88). Following vitamin A uptake, the intracellular concentrations of ATRA increase, inducing ISX expression. Consequently, less β-carotene is taken up by the enterocyte, and less β-carotene can be converted to retinal. When the intracellular concentration of ATRA drops, which is assumed to be the case when dietary vitamin A intake is low, ISX exerts less repressor activity towards SCARBI and BCO1 and consequently β-carotene uptake and conversion increase. A study in Zambian children with hypervitaminosis A supports this regulation. Indeed, these children had high serum carotenoid concentrations(Reference Mondloch, Gannon and Davis89) and many of them experienced hypercarotenodermia during mango season, a period of high provitamin A carotenoid intake. This might indicate as a possible explanation that conversion of provitamin A carotenoids to retinal by BCO1 was more inhibited by the hypervitaminosis A than their absorption via SR-BI, which is encoded by SCARB1. This is not surprising, as provitamin A carotenoid absorption involves not only SR-B1 but also CD36(Reference Borel, Lietz and Goncalves62), which is not assumed to be regulated by ISX. In a recent candidate genes association study a SNP in ISX together with SNP in other genes, was associated with the variability in β-carotene bioavailability(Reference Borel, Desmarchelier and Nowicki25). It was also reported in another study that a SNP in the ISX binding site in the BCO1 promoter (rs6564851) was associated with decreased conversion rates of β-carotene by 50 % and increased fasting blood concentrations of β-carotene(Reference Lobo, Amengual and Baus87). These associations support that genetic variations in this gene are key determinants of blood β-carotene concentrations.
In summary, it is assumed that the interplay between the β-carotene metabolising enzymes BCO1/2 and potential intracellular transporters possibly in conjunction with yet unidentified efflux transport proteins and those involved in chylomicron synthesis are among the most crucial actors influencing β-carotene bioavailability. Furthermore, the host vitamin A status which is detected by ISX and likely further mediated via retinoic acid signalling pathways also constitutes a paramount ‘critical control point’ in the metabolism of β-carotene.
Postprandial blood transport of newly absorbed β-carotene from the intestine to the liver
The enterocytes are assumed to secrete most of the newly absorbed β-carotene into chylomicrons, though it has been suggested that water-soluble β-carotene metabolites, e.g. apo-carotenals, could be secreted in the portal circulation and therefore directly reach the liver (Fig. 2B(Reference Harrison85)). In general, it is assumed that for compounds with a log P below approximately 5, portal absorption would predominate(Reference Charman and Stella90), which would be the case for some of the β-apo-carotenoids. Chylomicrons also contain retinyl esters, mainly retinyl palmitate(Reference Sauvant, Mekki and Charbonnier91), which originate either from esterification of retinol produced by the BCO1-mediated cleavage of β-carotene, or from re-esterification of preformed vitamin A present in the diet. It has been shown that most retinyl palmitate and β-carotene are not exchanged between lipoproteins and remain in chylomicrons and their remnants during their intravascular metabolism(Reference Blomhoff, Helgerud and Dueland92, Reference Tyssandier, Choubert and Grolier93). Thus, most β-carotene incorporated into chylomicron remnants, which are produced during vascular lipolysis of chylomicron TAG by both lipoprotein lipase (LPL) and glycosylphosphatidylinositol anchored high density lipoprotein binding protein 1 (GPIHBP1)(Reference Dallinga-Thie, Franssen and Mooij94), is taken up by hepatocytes during the postprandial period(Reference Blomhoff, Helgerud and Rasmussen95). This uptake involves several proteins, e.g. the LDL-receptor (LDLR), the LDL-receptor-related protein 1 (LRP1), SR-B1 and heparan sulphate proteoglycans (HSPGs)(Reference Dallinga-Thie, Franssen and Mooij94). Candidate gene association studies have also found that SNP in LPL (Reference Herbeth, Gueguen and Leroy96) were associated with blood β-carotene concentration.
The fact that β-carotene is carried in the blood by chylomicrons during the postprandial period implies that its metabolism is closely related to that of these TAG-rich lipoproteins. This is supported by a recent study that has shown that genetic variants in genes involved in chylomicron metabolism, i.e. transcription factor 7-like 2 (TCF7L2), ApoB, LIPC (which encodes hepatic lipase) and ABCA1 modulate the postprandial chylomicron β-carotene response to a meal that contained dietary β-carotene(Reference Borel, Desmarchelier and Nowicki25). Note also that a SNP in LIPC (Reference Borel, Moussa and Reboul97) was also associated with fasting blood β-carotene concentration.
In short, the transport from the intestine to the liver is mostly governed by proteins involved in chylomicron metabolism in the blood (e.g. ApoB and hepatic lipase) and likely also by proteins involved in chylomicron uptake by the liver (e.g. LRP1 and LDLR).
Liver metabolism and blood transport of β-carotene and its metabolites from the liver to extra-hepatic tissues
The liver is the main storage organ for vitamin A mainly in the form of retinyl esters. It has been estimated that for healthy, well-nourished individuals, approximately 70 % of vitamin A present in the body is stored in the liver(Reference O'Byrne and Blaner98). Following chylomicron-remnant uptake by the liver, which involves cell surface receptors (see the previous section), it is assumed that chylomicron remnant retinyl palmitate and β-carotene are released in hepatocytes during chylomicron remnant metabolism (Fig. 2C). They are then assumed to follow different metabolic pathways. Retinyl palmitate is assumed to be hydrolysed by a retinyl ester hydrolase to retinol. Retinol is then assumed to bind to cellular retinol-binding protein, type I (CRBPI / RBP1)(Reference Ong99) and to be transported to either the site where it is transferred to retinol-binding protein 4 (RBP4) or to hepatic stellate cells (also known as fat-storing cells, lipocytes or Ito cells) where it is re-esterified by LRAT(Reference Ong, MacDonald and Gubitosi100, Reference Rose101). Interestingly, hepatic LRAT expression is regulated by vitamin A status(Reference Blomhoff, Helgerud and Rasmussen95). This regulation likely involves ATRA and its respective response elements activated by the liganded nuclear hormone receptors RAR and RXR and further interaction with DNA. This regulation is proposed to give rise to a positive feedback loop when cellular ATRA concentrations are high, turning on hepatic stellate cell LRAT expression(Reference Nagatsuma, Hayashi and Hano102) and increasing the synthesis of retinyl esters(Reference O'Byrne and Blaner98) in these cells(Reference Wake103, Reference Wake104). These cells store approximately 70–90 % of liver vitamin A(Reference O'Byrne and Blaner98).
Contrarily to chylomicron retinyl palmitate, the fate of chylomicron β-carotene in the liver is barely known. How β-carotene is released from chylomicrons and how it is transported into hepatocytes remains unanswered. Concerning its cleavage, it is assumed that it is either cleaved to retinal by BCO1, which is highly expressed in hepatic stellate cells(Reference Shmarakov, Fleshman and D'Ambrosio105), or by BCO2, which is apparently more expressed in hepatocytes(Reference Shmarakov, Fleshman and D'Ambrosio105). The fraction of β-carotene that does not undergo this cleavage is either incorporated into VLDL and secreted into the blood, or stored in lipid droplets in parenchymal and hepatic stellate cells(Reference Shmarakov, Fleshman and D'Ambrosio105, Reference Lakshman, Asher and Attlesey106). The mechanism involved in the mobilisation of β-carotene stores is not known, but we hypothesise that it requires the hydrolysis of lipid droplet TAG.
The liver secretes vitamin A in the form of retinol either into the bile partly also as oxidised and/or further conjugated, i.e. glucuronidated metabolites(Reference Zachman and Olson107, Reference Zachman, Singer and Olson108), or directly into the blood. The liver secretes retinol into the blood arising partly from β-carotene metabolism but also as pro-vitamin A carotenoids, mainly β-carotene. Retinol is bound to serum retinol binding protein (RBP4), which in turn binds to transthyretin (TTR), stabilising the complex(Reference Peterson109). β-Carotene is incorporated in VLDL. Retinol associated with RBP4/TTR is taken up by two structurally related membrane receptors: stimulated by retinoic acid 6 (STRA6)(Reference Kawaguchi, Yu and Honda110) and the recently discovered STRA6-like receptor, also known as RBP4 receptor-2 (RBPR2)(Reference Alapatt, Guo and Komanetsky111). Retinol uptake via STRA6 depends on a functional coupling with intracellular LRAT(Reference Amengual, Golczak and Palczewski112). STRA6 and RBPR2 exhibit different tissue expression patterns: STRA6 is expressed in numerous tissues but not in the liver and intestine, where RBPR2 is mostly expressed(Reference Alapatt, Guo and Komanetsky111). VLDL-β-carotene and LDL-β-carotene, which originate from VLDL metabolism, are most likely taken up by tissues via LDL-receptor dependent mechanisms(Reference Thomas and Harrison113), requiring the tissue to express the LDL-receptor. However, candidate gene association studies have also found SNP associated with blood β-carotene concentration for SR-BI(Reference Borel, Moussa and Reboul114), also participating in the cellular uptake of HDL(Reference Hoekstra115), in addition to cellular uptake of β-carotene by enterocytes(Reference van Bennekum, Werder and Thuahnai60, Reference Borel, Lietz and Goncalves62, Reference Harrison85).
In summary, β-carotene can be stored mainly in the liver or alternative organs within the organism (and further cleaved by BCO1/2), secreted into the blood or into the bile. Bile excretion occurs via single or multiple oxidations at various locations of the derivative and further glucuronidation. Secretion into the bloodstream may follow targeted β-carotene cleavage at various locations or further incorporation and transport by VLDL.
β-Carotene metabolism, biodistribution, bioactivation and excretion
β-Carotene intake depends on the individual food intake in addition with an important influence of the individual food matrix and well as the food quality as outlined earlier. Further β-carotene metabolism includes isomerisation of the conjugated double bond system to various geometric isomers such as 9-cis- and 13-cis-β-carotene (Fig. 3). These cis-isomers are present in minor amounts in the raw food matrix(Reference Ben-Amotz and Fishier116–Reference Vasquez-Caicedo, Sruamsiri and Fau-Carle118) and isomer concentration may increase also due to food processing including simple heat based cooking(Reference Lessin and Schwartz29). It may further be induced in the human organism by various processes(Reference Hieber, King and Fau-Morioka119–Reference Relevy, Rühl and Harari121), where cis-isomers seem to be preferred v. all-trans-isomers. Whether this accumulation is due to favourable biophysical properties or by targeted and preferred uptake of cis-isomers is currently not known. For retinoids with a cis-isomeric structure, some enzymes were found which mediate targeted and non-targeted isomerisation, such as retinal pigment epithelium-specific 65 kDa protein(Reference Shyam, Gorusupudi and Nelson122, Reference Redmond, Poliakov and Kuo123) conversion to 11-cis-isomers, and Sphingolipid delta(4)-desaturase / desaturase 1 (DES1) (Reference Kaylor, Yuan and Cook124) to 9-cis- / 13-cis-isomers. In addition, binding proteins such as RBP1 and RLBP1 and retinol dehydrogenases RDH4/5 have specific cis-isomer preferred binding/transporter and synthesis mechanisms(Reference Parker and Crouch125–Reference Huang, Possin and Saari127). If these targeted isomerisation and isomer-binding properties also exist for carotenoids and apo-carotenoids seems likely via the same or alternative enzymes and binding proteins but has not been studied so far.
(Colour online) Metabolism of β-carotene with major metabolites formed in vivo. Involved enzymes, binding proteins, receptors and target genes involved in β-carotene metabolism towards bioactive retinoids. Derivatives marked with ‘*’ have been conclusively identified to be endogenously present. At – all-trans; SCARB1 – scavenger receptor class B type I; CD36 – cluster of density 36; ABCG5 / 8 – ATP binding cassette member 5 / 8; BCO1 – β-carotene oxygenase 1; BCO2 – β-carotene oxygenase 2; LRAT1 / 2 – lecithin retinol acyltransferase, DGAT1 / 2 – diacylglycerol O-acyltransferase 1 / 2; ISX – intestinal transcription factor; STRA6 – stimulated by retinoic acid 6; RBPR2 – retinol-binding protein receptor 2; RDH 5 / 10 – retinol dehydrogenase 5 / 10; DHRS3 / 9 – short-chain dehydrogenase/reductase 3 / 9; RBP1 / 2 / 4 – retinyl-binding protein 1 / 2 / 4; REH – retinyl-esterase; RETSAT – all-trans-retinol 13,14-reductase; ALDH1A1 / 2 / 3 – aldehyde dehydrogenase 1 family, member A1 / 2 / 3; CRABP1 / 2 – cellular-retinoic acid binding protein 1 / 2; RPE65 – retinal pigment epithelium-specific 65 kDa protein; DES1 – sphingolipid delta(4)-desaturase; RLBP1 – retinal-binding protein 1; RAR – retinoic acid receptor; RXR – retinoid-X receptor; TG2 – transglutaminase 2; SCD1 – stearoyl-CoA desaturase / (Δ−9-)desaturase-1; ELOVL6 – elongation of very long chain fatty acids protein 6; HOXB6 / 8 – homoeobox protein 6 / 8, HBEGF – heparin-binding-epidermal growth factor; RARRES2 – retinoic acid receptor responder protein 2 / chemerin; ADIPOQ – adiponectin; UCP1 – uncoupling protein 1, UGT2B7 – UDP-glucuronyltransferase-glucuronosyltransferase-2B7.
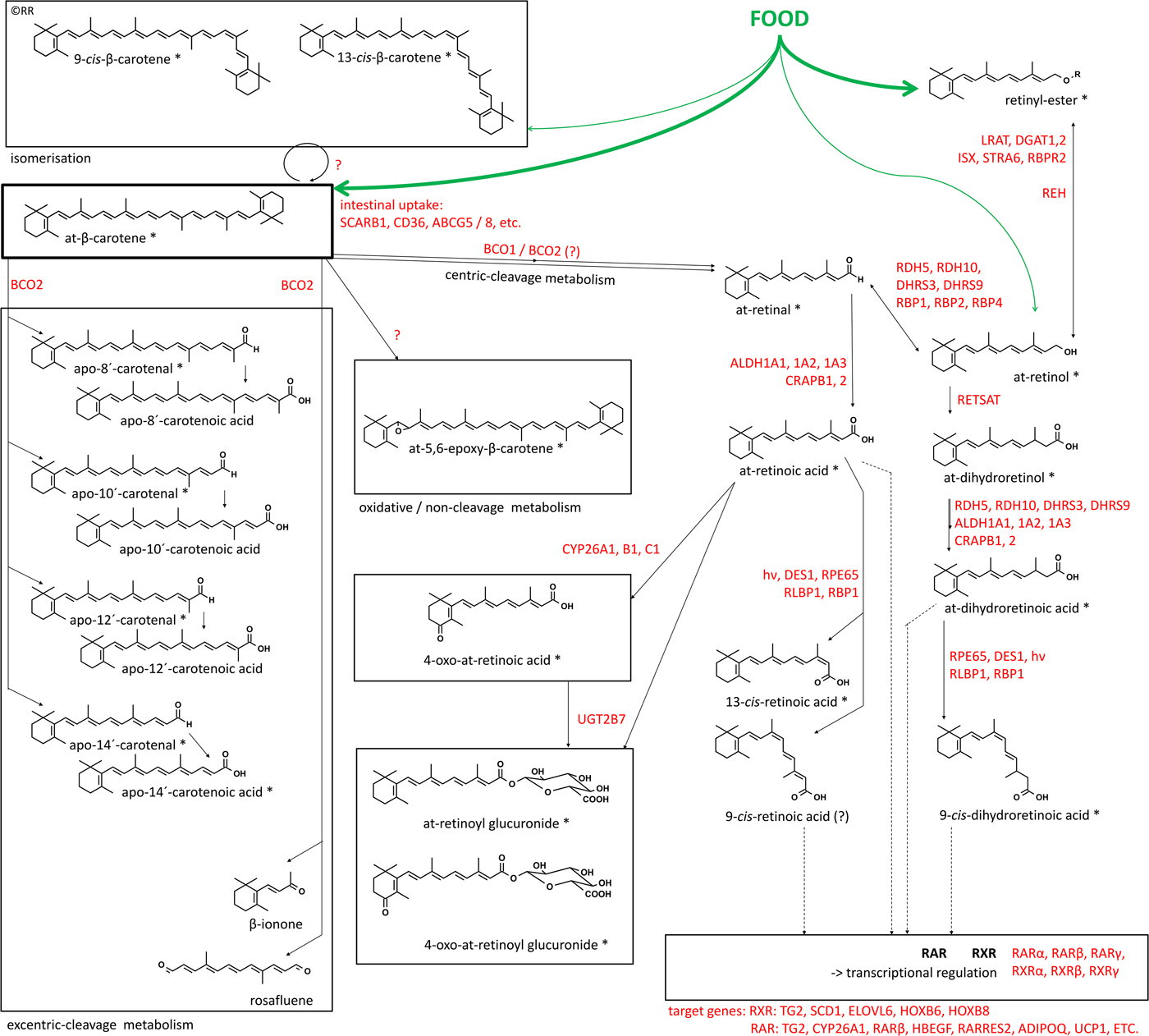
Following absorption, a large proportion of carotenoid and later retinoid metabolism is under control of nuclear hormone receptor signalling, as a partly auto-regulatory homoeostatic regulated process. Many steps, involving receptors such as RARβ, to anabolic enzymes including BCO1(Reference Bachmann, Desbarats and Pattison128), BCO2 and aldehyde dehydrogenase 1 family, member A3 (ALDH1A3), catabolic enzymes including CYP26A1 and LRAT and binding proteins, including RBP1, RBP4 and cellular-retinoic acid binding protein 2 (CRABP2) are under control of RAR-RXR- and PPAR-RXR-mediated signalling(Reference Gericke, Ittensohn and Mihaly129, Reference Balmer and Blomhoff130). RAR-RXR- and PPAR-RXR-mediated signalling also controls various other important lipid metabolic processes and places carotenoids as precursors of important regulators of general lipid metabolism as reviewed earlier(Reference Evans and Mangelsdorf131). This auto-regulation of general lipid metabolism and nuclear hormone-mediated signalling includes various target genes and several of these specific target genes are key for eliciting beneficial health effects of β-carotene, especially regarding cancer, allergic inflammatory disorders such as asthma and various CVD. As a consequence, carotenoid metabolism and usage towards bioactive retinoids for further bioactive signalling seems to be likely dependent on: (a) sufficient levels of available carotenoids in the human organism and (b) a targeted metabolic bioactivation pathway to elicit beneficial activities of carotenoids.
A long list of enzymes and binding proteins (summarised in Fig. 3) is responsible for the metabolism of carotenoids to bioactive retinoids in a temporal and spatial highly controlled manner. The initial steps are the cleavage of carotenoids by the two known human carotenoid-oxygenases BCO1 and 2(Reference Lindqvist, Sharvill and Sharvill132, Reference Wu, Guo and Wang133). The resulting apo-carotenals (named retinal in the case of apo-15′-carotenal), can then further be reduced to alcohols and esterified to store retinoids as retinyl esters, a reaction mediated by LRAT and diacylglycerol O-acyltransferase 1/2 (DGAT1/2)(Reference Orland, Anwar and Cromley134–Reference Ross and Zolfaghari137). Retinyl-esters can further be de-esterified by esterases (REHs) to alcohols and especially retinol to serve as precursors for later bioactivation(Reference Schreiber, Taschler and Preiss-Landl138, Reference Schreiber, Taschler and Wolinski139). Retinol can later be transported by various retinol binding proteins and further be oxidised in target tissues by retinol dehydrogenases, mainly RDH4/5 and 10 as well as short-chain dehydrogenase / reductase 3 and 9 (DHRS 3 and 9) (Reference Napoli140, Reference Kumar, Sandell and Trainor141). Inter- and intra-cellular transport of various forms of retinals and retinols is mediated by specific binding proteins including RBP1, 2 and 4 and RLBP1(Reference Napoli142). The physiological and nutritional relevance of additional apo-carotenals and apo-carotenoic acids remains unclear, and they were described as low affinity activators / competitive antagonists of nuclear hormone receptors, as reviewed previously(Reference Eroglu and Harrison143). Unfortunately, a clear link between the low physiological and nutritional relevant levels in human subjects or high level and further dependent biological-mediated signalling were not described yet, and thus no nutritional or physiological relevance can currently clearly be associated with these derivatives.
The major intermediate bioactive precursor is retinal, the visual pigment, which can be obtained possibly from multiple sources: (a) direct cleavage of carotenoids via BCO1(Reference Lindqvist and Andersson144) or (b) via a physiological unclear and indirect cascade via BCO2(Reference Amengual, Widjaja-Adhi and Rodriguez-Santiago23) or (c) via oxidation from retinol(Reference Napoli145). Retinol is present at low levels in the food matrix and in highly homoeostatically regulated levels in blood. However, retinal becomes also significantly available via cleavage of retinyl-esters, the major dietary relevant and major storage form, which can be cleaved to retinol and oxidised to retinal in target tissues(Reference Napoli142, Reference D'Ambrosio, Clugston and Blaner146). The oxidation of retinal to ATRA is the key step to yield the lipid-hormone ATRA. Retinoic acid dehydrogenases (aldehyde dehydrogenase 1 family, member A1, A2 and A3), in association with cellular retinoic acid binding proteins 1 and 2 are strictly controlling this bio-activation(Reference D'Ambrosio, Clugston and Blaner146). All-trans retinoic acid (ATRA) can further initiate, via ligand activation of the RARα, β, γ, transcriptional signalling of a large array of target genes(Reference Balmer and Blomhoff130) (Fig. 5) including: TG2 – transglutaminase 2; HBEGF – heparin-binding-epidermal growth factor; RARRES2 – RAR responder protein 2 / chemerin; ADIPOQ – adiponectin; UCP1 – uncoupling protein 1, as well as the afore-mentioned auto-regulated targets in the retinoid metabolism cascade.
While ATRA as the endogenous RAR-ligand is well accepted(Reference Petkovich147), the existence of an endogenous RXR-ligand has largely been mysterious(Reference de Lera, Krezel and Rühl148, Reference Calleja, Messaddeq and Chapellier149). The ATRA isomer 9-cis-retinoic acid (9CRA) has been described to be ‘the’ endogenous ligand for this crucial heterodimer nuclear hormone receptor(Reference Allenby, Bocquel and Saunders150–Reference Heyman, Mangelsdorf and Dyck152). Unfortunately, the endogenous, physiological and nutritional relevance of 9CRA has been questioned and remained unclear(Reference Rühl, Krzyzosiak and Niewiadomska-Cimicka153–Reference Schmidt, Brouwer and Nau159). Recently, the lipid hormone 9-cis-13,14-dihydroretinoic acid (9CDHRA) has been identified as the endogenous and physiological relevant RXR ligand(Reference de Lera, Krezel and Rühl148, Reference Rühl, Krzyzosiak and Niewiadomska-Cimicka153). Further examinations about the nutritional relevance are currently under investigation(Reference Rühl, Krezel and de Lera160). 9CDHRA can activate RXR and via this route also initiate non-permissive heterodimers, such as RXR-PPAR, RXR-liver X receptors (LXRs), RXR-farnesoid receptors (FXR) and RXR-nuclear hormone receptor 4A (NR4A) protein, involved in the expression of a wide array of genes involved in inflammation and lipid metabolism, as reviewed in Desvergne(Reference Desvergne161). A large overlap was found between beneficial anti-inflammatory effects of carotenoids in general and lipid metabolism for many RXR-heterodimer-mediated signalling pathway targets(Reference Evans and Mangelsdorf131, Reference Desvergne161–Reference Mangelsdorf, Ong and Dyck166).
The precise metabolic pathway leading to 9CDHRA is not known yet and likely involves retinol-saturase (RETSAT(Reference Rühl, Krzyzosiak and Niewiadomska-Cimicka153, Reference Moise, Kuksa and Imanishi167), and / or DES-1) and the binding protein RBP1(Reference Kaylor, Yuan and Cook124, Reference Rühl, Krzyzosiak and Niewiadomska-Cimicka153), as well as retinaldehyde binding protein 1 (RLBP1 / CRALBP(Reference Saari, Huang and Possin168, Reference Huang, Jarjour and Oumesmar169)). Additionally, novel still non-identified carotenoids are speculated to be more direct precursors for 9CDHRA (postulated and also outlined in Fig. 5).
Other retinoids such as 13-cis- or 9,13-dicis-RA(Reference Allenby, Bocquel and Saunders150, Reference Horst, Reinhardt and Goff170, Reference Chen, Sass and Seltmann171), all-trans-13,14-dihydroretinoic acid(Reference Vahlquist172, Reference Torma, Asselineau and Andersson173) and phase 1 metabolites including 4-hydroxy- or 4-oxo-retinoic acid, or other hydroxyl- / oxo-metabolites are likely of minor importance for RAR- or RXR-mediated signalling processes(Reference Schmidt, Brouwer and Nau159, Reference Pijnappel, Hendriks and Folkers174–Reference Baron, Heise and Blaner177). Alternative phase 1 metabolism may also occur for β-carotene and would result in epoxidation and hydroxylation to produce a large array of epoxy-, hydroxyl- and oxo-carotenoids (examples shown in Fig. 3, reviewed earlier(Reference Bohm and Bitsch178)). These hydroxy- / oxo-retinoids can further be conjugated via phase 2 metabolism by UDP-glucuronyltransferase-glucosyltransferase to yield water-soluble excretion metabolites such as retinyl-, retinoic acid (retinoyl) and oxo-retinoic acid glucuronides, which were found in serum, faeces and urine(Reference Sass, Masgrau and Saurat179–Reference Samokyszyn, Gall and Zawada181).
BCO1 as a critical bottleneck for the cleavage of β-carotene and enabling RAR- / RXR-mediated signalling
Human subjects centrally cleave β-carotene to retinal and following oxidation and reduction a larger array of multi-functional retinoids are created which have been detected endogenously (Fig. 3(Reference Böhm, Borel and Corte-Real13)). This BCO1-mediated conversion of β-carotene to retinal is therefore an important bottleneck, which is highly controlled and mediated by various factors: (A) availability of the substrate and saturation of the enzymatic conversion potential, (B) presence and relative levels of food derived inhibitors, (C) spatial and temporal regulation and localisation of the enzyme, (D) sex-specific regulations, (E) feedback regulations by bioactive products transcriptionally controlling BCO1-expression and (F) the previously reported polymorphisms of BCO1 as well as assisting proteins.
Starting with the overall conversion in the human organism, β-carotene was reported to vary largely in absorption efficiency (30–70 %) upon intestinal uptake, as explained earlier. This variation is in part due to the variation in BCO1 cleavage potency, partly explainable by frequently occurring polymorphisms of BCO1(Reference Lietz, Oxley and Leung82, Reference Leung, Hessel and Meplan83). Alternatively, many factors which are summarised here describe the inter- and auto-regulatory pathways in BCO1-mediated cleavage to retinoids, enabling RAR- or / and RXR-mediated signalling.
The main question is how BCO1 and its mediated cleavage to centric cleavage metabolites are regulated in the human organism. As described earlier, six major steps (A–F) have been reported and identified. First, the availability of the substrate is usually a major factor for increased conversion and resulting product levels (Fig. 4A). This conversion was presented as a saturating curve, plateauing at β-carotene levels in the range of 15 000–40 000 and 80 000–240 000 nm for β-cryptoxanthin(Reference Lindqvist and Andersson144). For comparison, the endogenous levels for β-carotene were in average range of 360 nm in serum and up to 31 830 nm in organs, while being highest in the adrenals and β-cryptoxanthin in the average range of 230 nm in serum and with highest tissue levels of 2900 nm in adrenals, as reviewed recently(Reference Böhm, Borel and Corte-Real13). We can thus conclude that these active ranges were not reached in serum, while tissue levels approach the saturation of enzyme conversion(Reference Böhm, Borel and Corte-Real13, Reference Lindqvist and Andersson144). It should be noted that serum and tissue levels do not represent freely available carotenoids, but mainly carotenoids attached to binding proteins and carotenoids associated in lipid vesicles in the membranes and lipid accumulating vesicles such as in the adipose tissue(Reference Böhm, Borel and Corte-Real13). In one study examining children(Reference Rühl, Taner and Schweigert182–Reference Rühl184), a direct correlation between all-trans-β-carotene (ATβC) and ATRA serum levels was observed, resulting in a linear correlation with r 0·68, while in adults(Reference Mihaly, Marosvolgyi and Szegedi185–Reference Lucas, Mihály and Lowe187) no such correlation was observed (r −0·14), likely due to auto-regulative processes. The highest relevant level of ATβC in serum of children was 756 nm (Fig. 4B) and 1628 ng/ml (3031 nm) in adults (Fig. 4C), displaying no plateau for conversion to ATRA, based on endogenous relevant β-carotene levels (Fig. 4B and C).
(Colour online) BCO1 localisation and metabolic properties. (A) In vitro kinetic analysis of purified recombinant human BCO1 with β-carotene and β-cryptoxanthin, as published earlier from Lindqvist and Andersson(Reference Lindqvist and Andersson144). (B) Direct correlation newly calculated based on of serum ATβC to ATRA in children in Germany with different ethical backgrounds(Reference Gruber, Taner and Mihaly183, Reference Rühl184). (C) Direct correlation based on serum ATβC to ATRA levels in Hungarian adults (n 40, Lucas et al.(Reference Lucas, Mihály and Lowe187)) This figure is just present in the original study in ng/ml, while 1 ng/ml ATRA corresponds to 3.3 nM and 1 ng/ml ATβC to 1.86 nM. (D) Distribution of BCO1 mRNA expression in human tissues, as published previously in Lindqvist and Andersson(Reference Lindqvist and Andersson144) (PBL – peripheral blood lymphocytes). (E) Differentially expressed genes and pathways by β-carotene v. control diet. Gene expression analysis of different tissues on a control diet supplemented with βC v. control diet (containing adequate vitamin A)-fed mice. A description of the mouse study can be found in van Helden et al.(Reference van Helden, Godschalk and Swarts14). The global transcriptome data were extracted from Gene Expression Omnibus (GEO, Superseries GSE98847), containing lung (GSE98845), liver (GSE98846) and inguinal white adipose tissue (iWAT; GSE27271) and were normalised per tissue and genotype with both sexes included for comparison between sexes. Sex-specific number of differential expressed genes (P < 0·05) are given in number and fold change (FC) of males v. females. (F) ATRA levels in serum (nm) and lung ((pmol/ml / 10−2 m) of control treated (CTRL), low-β-carotene (βC)-diet supplemented (LBC) and high-βC supplemented ferrets (HBC) adapted from Liu et al.(Reference Liu, Wang and Bronson237). Panels A, B, D and F are adapted from van Helden et al.(Reference van Helden, Godschalk and Swarts14) and Lindqvist and Andersson(Reference Lindqvist and Andersson144) and were permitted to reproduction under copyright.
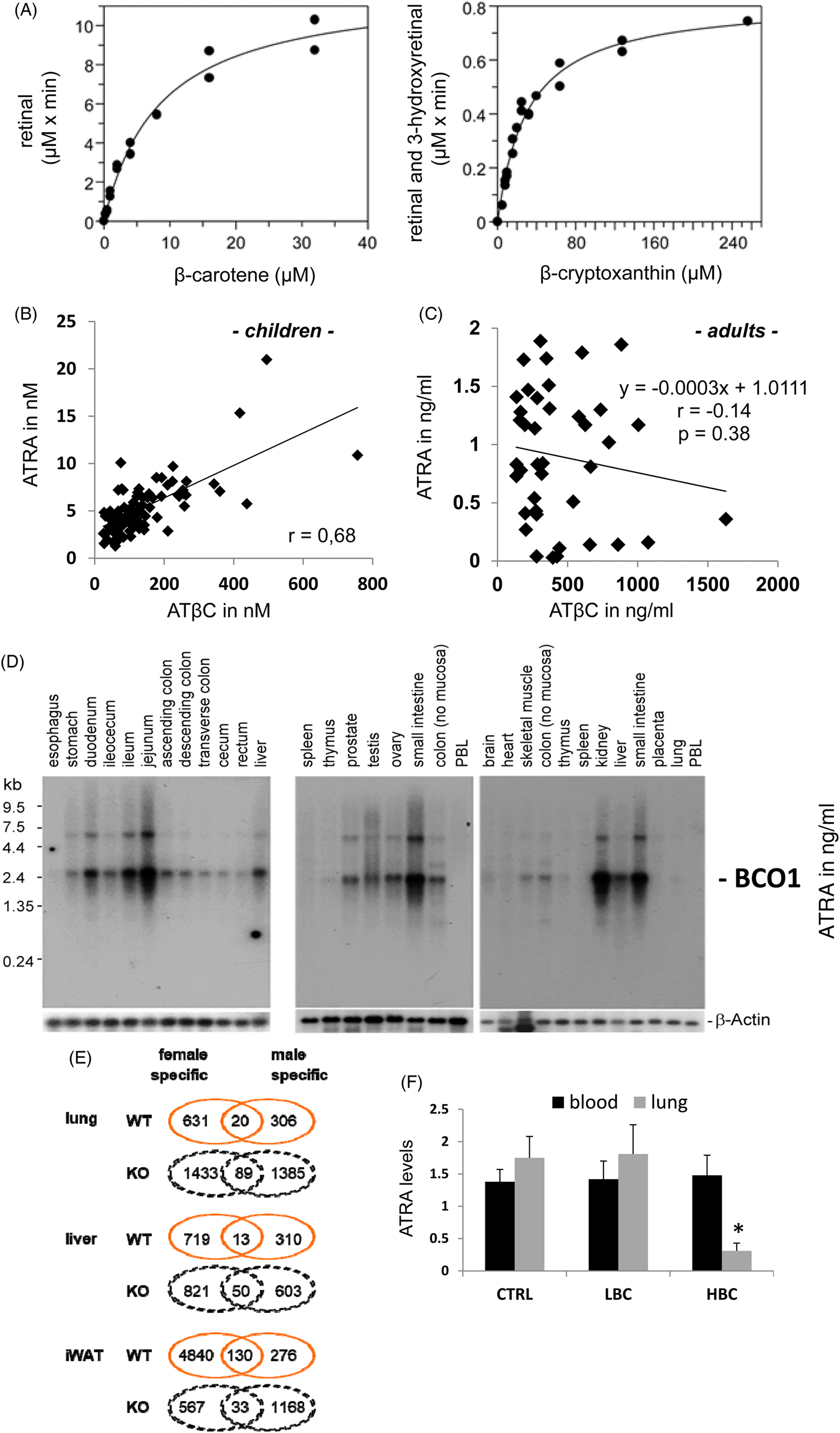
The second modification factor are other carotenoids such as canthaxanthin, lutein and zeaxanthin(Reference Grolier, Duszka and Borel72, Reference van Vliet, van Schaik and Schreurs188), which can inhibit BCO1-mediated conversion partly in a competitive manner. The specific mechanisms of these phenomena were not investigated deeper, and neither have nutritional relevant ranges and relevant ratios been examined. It is likely that these three carotenoids can attach and bind to the active site of the BCO1, thereby inhibiting the binding and enzymatic conversion of known pro-vitamin A carotenoid BCO1 substrates (β-carotene, α-carotene, β-cryptoxanthin, apo-8′-carotenal and lycopene). In in vitro studies, it was reported that three times higher levels of lutein (compared with β-carotene) interfered with β-carotene conversion. These ratios reflect relevant physiological conditions(Reference van Vliet, van Schaik and Schreurs188), which can be obtained after targeted lutein/zeaxanthin supplementation or dietary intake of fruits and vegetables high in these carotenoids, as reviewed recently(Reference Böhm, Borel and Corte-Real13). Additionally assisting proteins such as RBP1 and RBP2, acting as intracellular sensors of endogenous retinoid status are thereby important contributors for ATβC conversion to ATRA(Reference Lietz, Lange and Rimbach189–Reference Nagao191).
The third modifying factor is the specific spatial and temporal regulation of BCO1 expression in the human organism(Reference Lindqvist and Andersson144). Highest BCO1 expression was found in different parts of the intestine, with highest expression levels in the jejunum (Fig. 4D). Other relevant tissues are reproduction organs testis and prostate in males as well as ovaries in females, comparable with levels found in kidney, liver, skeletal muscle and stomach (Fig. 4D). Not displayed in this figure are the relatively high expression levels observed in the eye(192).
As a fourth modification, sex specific regulations were observed. In male mice and rats, a connection between testosterone and carotenoid as well as BCO1-expression was found(Reference Ford, Moran and Smith193–Reference Campbell, Stroud and Nakamura195), while oestrogen / testosterone correlated in older woman with carotenoid levels(Reference Maggio, de Vita and Lauretani196). If non-reproduction-related organs also display this regulation, depending on sexual steroid hormones, was not further investigated. One indicator are higher ATRA and lower retinol serum levels in women v. men(Reference Soderlund, Sjoberg and Svard197), likely as a consequence of higher levels of β-carotene in their serum / plasma, mainly due to the less healthy nutritional status(Reference Böhm, Borel and Corte-Real13, Reference El-Sohemy, Baylin and Kabagambe198, Reference Tucker, Chen and Vogel199), or higher BCO1 presence and activity.
Recently, also gene expression microarray studies were conducted, using wild type (WT) and BCO1-knockout male and female mice, on a low but sufficient vitamin A diet with or without additional β-carotene supplementation(Reference van Helden, Godschalk and Swarts14, Reference van Helden, Godschalk and von Lintig200–Reference van Helden, Heil and van Schooten202), which were used to provide further insights into the differential effects of dietary β-carotene supplementation. It was observed that β-carotene supplementation alters only a small number of overlapping genes in the lung of wild-type male and female mice (n 20, Fig. 4E), while a larger number of the genes altered by β-carotene supplementation were sex specific, n 631 in female and n 306 in male mice(Reference van Helden, Godschalk and Swarts14) (Fig. 4E). This difference was even more striking in the BCO1 knockout mice, where 1433 genes were specifically affected in females and 1385 in males, with only 89 being affected in both sexes and, strikingly, for 85 of these, the direction of expression was oppositely regulated between the sexes (Fig. 4E). The number of genes affected by β-carotene in the liver of BCO1 knockout mice was less than that of the lung (Fig. 4E).
In inguinal white adipose tissue (iWAT) a different pattern emerged with a large number of genes being specifically regulated upon β-carotene exposure in WT female mice (4840) (Fig. 4E). The number of genes specifically regulated by β-carotene exposure in males of WT (276) and BCO1 knockout (1168) genotype was comparable between iWAT v. lung, while BCO1 knockout females showed a reduced number (567 v. 1433) in iWAT. This was strikingly also the case for the much larger number (33 WT v. 130 KO) of common genes in WT mice, and is likely explained by the important role of WAT in steroid hormone metabolism, especially for oestrogens and progestogens in females(Reference DiSilvestro, Petrosino and Aldoori203–Reference Bonet, Canas and Ribot205).
Detailed analysis of the effects of dietary β-carotene supplementation identified a strong down-regulation of RXRα, as well as the pro-adipogenesis trigger PPARγ and its target genes in iWAT of WT female mice(Reference Amengual, Gouranton and van Helden206). This effect was likely dependent on BCO1-mediated retinoid production, since it was not observed in BCO1 knockout female mice and this effect was associated with a reduction in WAT mass, resulting in a reduced adiposity index. This adiposity lowering effect is in line with observations that show that oral ATRA administration induces energy expenditure and fat mass lowering in mice, with WAT being one of the contributing tissues(Reference Mercader, Ribot and Murano207).
The fifth modifying effect is the regulation of BCO1 on the transcriptional level. A feedback mechanism was identified, partly already before a clear identification, characterisation and expression of BCO1 in mice, rats and chicken and human subjects(Reference Lietz, Lange and Rimbach189, Reference Fierce, de Morais Vieira and Piantedosi190, Reference Parvin and Sivakumar208–Reference van Vliet, van Vlissingen and van Schaik210). The conversion and ratio of ATβC to retinoids, especially all-trans-retinal, was used to identify BCO1 activity(Reference Bachmann, Desbarats and Pattison128, Reference Wang, Tang and Fox211). Feedback mechanisms were claimed as a direct involvement of ATRA-RAR-interaction and transcriptional modification of BCO1 expression was shown. ATRA–RAR-mediated signalling is suggested to regulate BCO1 expression identified, either indirectly by retinal conversion per homogenate ratio or directly by mRNA quantification, as a negative feedback mechanism(Reference Bachmann, Desbarats and Pattison128). Treatments of rats with ATRA, retinyl acetate, β-carotene or a synthetic RAR-agonist (Ro41-5253) significantly reduced BCO1 activity identified by retinal conversion(Reference Bachmann, Desbarats and Pattison128). Focusing on ATRA, retinyl acetate and β-carotene treatments to rats, it was found that also serum retinoic acid levels increased and partly negatively correlated with reduced intestinal BCO1 activity(Reference Bachmann, Desbarats and Pattison128). This mechanism was thereby identified as an important negative feedback regulation for retinoid and mainly RAR-mediated signalling. It is noteworthy that nutritional supplementation with high β-carotene can result even in decreased local levels of ATRA with potential negative effects and increased vulnerability towards carcinogenesis as shown in β-carotene-supplemented ferrets (Fig. 4F(Reference Wang, Liu and Bronson15)). This highlights the limits of β-carotene-signalling mediated autoregulation using non-nutritional relevant to high β-carotene stimuli, with even previously reported negative side effects in human subjects as found in the ATBC and Carotene and Retinol Efficacy Trial (CARET) studies(Reference Omenn, Goodman and Thornquist11, 12). That high retinoid stimuli can induce negative side effects was recently described in mice(Reference Rubin, Ross and Stephensen212–Reference Garcia, Rühl and Herz215), and seems also relevant for high-nutritional relevant β-carotene supplementation in ferrets and human subjects(Reference Rühl, Taner and Schweigert182–Reference Rühl184, Reference Melhus, Michaelsson and Kindmark216–Reference Boelsma, van de Vijver and Goldbohm219). In consequence, low / moderate β-carotene supplementation seems to be a tolerable nutritional stimulus to which the mammalian / human organism can respond. Recently it was reported that glucocorticoid regulated pathways and hepatocyte nuclear factor (HNF)1α and HNF4α pathways are important regulators of BCO1 expression(Reference Yamaguchi, Sunto and Goda220).
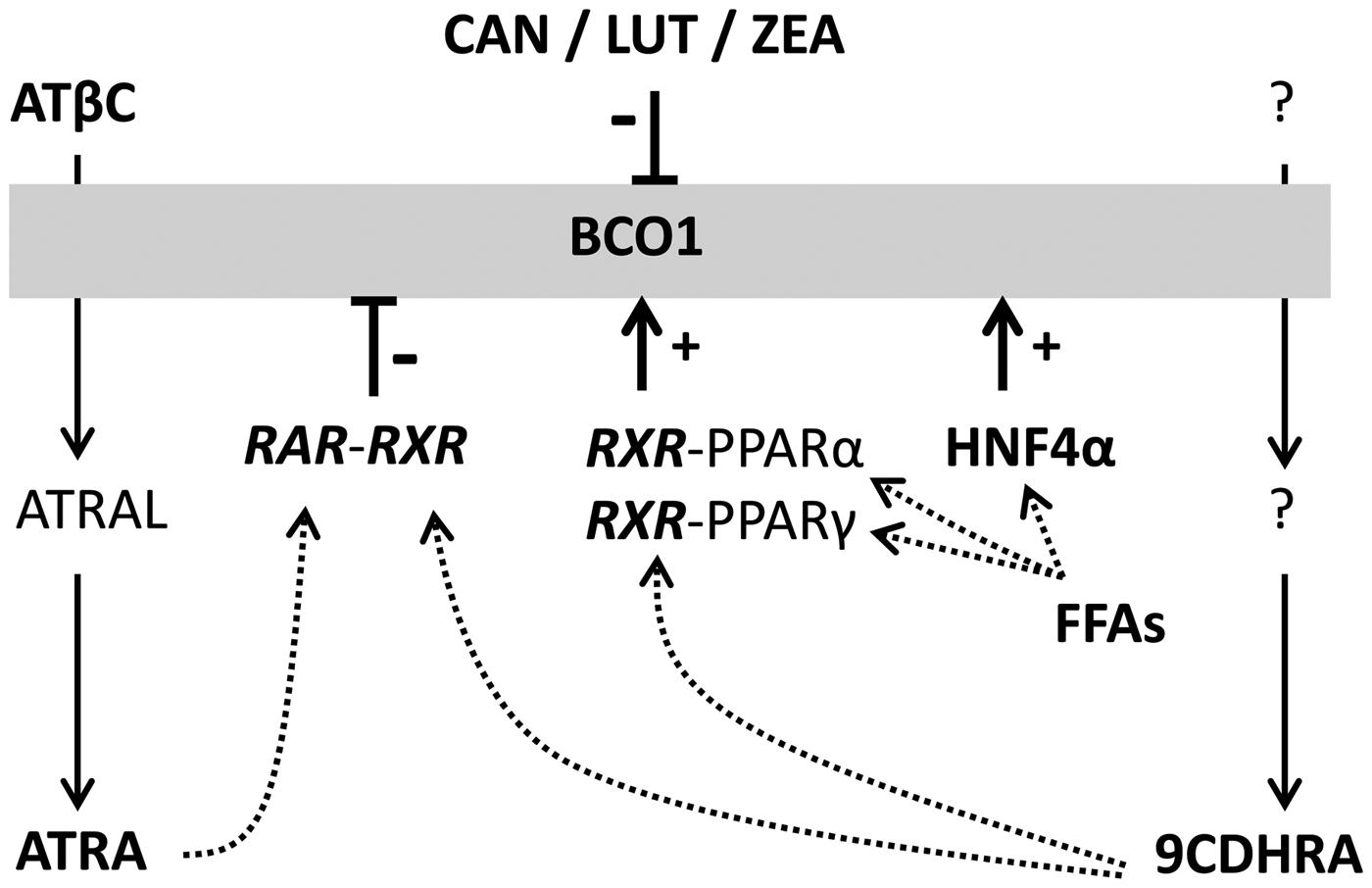
In addition, RXR-PPARα and -PPARγ-mediated signalling was identified as an alternative mechanism, providing positive feedback(Reference Gong, Tsai and Yan221, Reference Gong, Marisiddaiah and Rubin222). The PPARα and PPARγ nuclear hormone receptor heterodimers can be either activated by an RXR-ligand or alternatively by the respective PPAR ligand. For PPARs and HNF4α FFAs and fatty acid metabolites have been identified as natural ligands (Fig. 5(Reference Schupp and Lazar223, Reference Dhe-Paganon, Duda and Iwamoto224)). After a high-dietary intake of fat this important regulatory pathway is initiated by increased levels of FFAs as a direct result of the diet rich in fat and results further in increased BCO1-expression as a direct feedback to this high-fat diet. These two nuclear hormone receptor heterodimers need either an RXR-ligand as well as / or a PPAR-ligand. Dietary transglutaminases can provide PPAR ligands, thus synchronising fat, and concomitantly carotenoid, uptake / availability with BCO1 up-regulation. It was described that the main BCO1-metabolite ATRA is regulating via ATRA–RAR-mediated signalling various important steps in lipid metabolism(Reference Amengual, Gouranton and van Helden206, Reference Kim, Zuccaro and Costabile225, Reference Landrier, Kasiri and Karkeni226), with a focus also on counteracting fat accumulation via energy dissipation in adipose tissue(Reference Landrier, Kasiri and Karkeni226–Reference Bonet, Ribot and Palou229) and regulation of insulin secretion(Reference Takeda, Sriram and Chan230, Reference Brun, Grijalva and Rausch231). The second possibility for negative feedback is the potential synthesis of the endogenous RXR-ligand 9CDHRA, starting from still non-identified carotenoid precursors (Rühl et al., unpublished results(Reference Rühl, Krezel and de Lera160)). This means that the endogenous RAR ligand ATRA and the endogenous RXR-ligand 9CDHRA obtain potential opposite regulation on their own synthesis via positive or negative feedback control mechanism of BCO1 expression and further activity (Fig. 5). As a consequence, levels and dietary intake of specific carotenoid precursors may influence or even control BCO1-mediated synthesis of endogenous RAR- or RXR-ligands and further controlling metabolic processes associated with lipid metabolism with relevance for obesity and diabetes.
Transcriptional regulation of BCO1 metabolism and affected biological processes. Schematic summary of metabolism of the endogenous RAR-activator ATRA starting from ATβC, via all-trans-retinal (ATRAL) to ATRA, which can further activate RAR-RXR-mediated transcriptional signalling. In parallel the newly identified endogenous RXR-ligand 9-cis-13,14-dihydroretinoic acid (9CDHRA) can be created starting from putative carotenoid via putative retinal-analogues to 9CDHRA, which can further activate RXR-hepatocyte nuclear factor (HNF)4α, -PPAR α or -PPARγ-mediated transcriptional signalling. These three receptors (HNF4α, PPARα and PPARγ) can be activated by their ligands, free fatty acids (FFAs) and other metabolites originating from fatty acids. The RAR- or RXR-mediated signalling can positively or negatively alter transcriptional regulated BCO1-expression. LUT, lutein; CAN, canthaxanthin; ZEA, zeaxanthin.
In summary, BCO1 represents a bottleneck for β-carotene-conversion to bioactive retinoids and further RAR-mediated transcriptional activation and signalling. A food matrix high in different natural occurring carotenoids leads to metabolism from pro-vitamin A carotenoids (β-carotene and β-cryptoxanthin) to retinoids, while additional carotenoids such as lutein, zeaxanthin and canthaxanthin may inhibit this conversion. This balanced carotenoid mixture, which is mainly present in fruits and vegetables as a balanced diet, is likely resulting in a much lower conversion to retinoic acids, than an equimolar supplementation with β-carotene as a nutritional supplement. In addition, a diet high in fat induces a strong activation of PPARα-, PPARγ-RXR / HNF4α-mediated signalling, following increased BCO1-expression and increased ATRA levels. Furthermore, this ATRA can induce increased RAR-RXR-mediated signalling as a natural feedback mechanism to stimulate lipid catabolism and blocking fat accumulation. In conclusion, a balanced diet rich in carotenoids originating from fruits and vegetables or alternatively a balanced carotenoid supplementation as present in fruit and vegetable extracts or to be developed ‘smart’ carotenoid supplements and not artificial single high carotenoid supplementation should result in moderate retinoid synthesis with the potential of balancing fat accumulation and stimulating fat usage in the human organism.
Conclusions
β-Carotene availability from the diet, as well as BCO1-mediated cleavage towards bioactive ATRA under consideration of tissue and sex dependent regulation are the two main bottlenecks for enabling retinoid-mediated signalling, as the most well-known processes of β-carotene's metabolic action. Serum β-carotene levels are affected by various aspects concerning our diet and human polymorphisms. Further bioactive signalling starting from serum and tissue ATRA levels is highly homoeostatically auto-regulated by various mechanisms, including nutritional stimuli and sex hormonal regulatory pathways. A strong deficiency of β-carotene in the diet or strong nutritional / supplemental β-carotene stimuli can result in altered retinoic acid levels but without any reported significantly altered further biological-mediated signalling (reviewed in Böhm et al.(Reference Böhm, Borel and Corte-Real13) and Watzl et al.(Reference Watzl, Bub and Brandstetter232–Reference Watzl, Bub and Briviba234)).
How short-term or long-term β-carotene or alternative stimuli affecting BCO1-cleavage and further can alter RAR- / RXR-mediated signalling must be further evaluated in human supplementation trials additionally examining known or postulated β-carotene-dependent health-biomarkers especially including novel omics-based disease marker.
As a final conclusion, the human organism seems to have a high flexibility balancing high and low dietary β-carotene availability based on a complex homoeostatic regulation for maintaining physiological crucial RAR-mediated signalling. The optimal dietary range of β-carotene in concert with other nutrients is highlighted in an additional review(Reference Böhm, Borel and Corte-Real13).
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0029665118002641.
Acknowledgements
The insights from EU-COST action POSITIVE (FA 1403) and EUROCAROTEN (CA 15136) are much appreciated.
Financial support
None.
Conflict of interest
The authors declare no conflict of interest.
Authorship
T. B., C. D., S. N. E., J. K., E. v. S., R. R. and P. B. contributed in setting up and writing the manuscript. RR was the invited speaker at the Glasgow symposium.






