INTRODUCTION
Protozoan parasites and the global burden of their diseases
Protozoa (kingdom Protista) are single-cell organisms that can be free-living or parasitic in nature (Baron, Reference Baron and Baron1996). Out of more than 50 000 protozoan species that have been described to-date, relatively few have been identified as major contributors to the global burden of human diseases (Kuris, Reference Kuris2012) and animal agriculture (Dubey, Reference Dubey and Kreier1977). The protozoa represent 19% of all human parasites (83 out of 437 species to-date) and are associated with 30% of parasite-induced human morbidity-mortality (Kuris, Reference Kuris2012).
Of the four groups of infectious protozoa (CDC, 2017), the Mastigophora (flagellates) and Sporozoa contain the Kinetoplastidae and Apicomplexa, respectively. It is to these two phyla that belong many of the causative agents of disease: Mastigophora – the insect vector-borne kinetoplastids Trypanosoma brucei (Human African Trypanosomiasis, HAT), Leishmania spp. (leishmaniasis, cutaneous and visceral) and Trypanosoma cruzi (American trypanosomiasis, Chagas’ disease); Sporozoa – the apicomplexan Toxoplasma gondii (toxoplasmosis), Cryptosporidium spp. (cryptosporidiosis) and Eimeria spp. (coccidiosis in poultry and cattle), Theileria spp. (East Coast Fever in cattle) and Plasmodium spp., including Plasmodium falciparum the causative agent of severe malaria and one of the ‘Big Three’ global infectious diseases alongside HIV and tuberculosis (Torgerson & Macpherson, Reference Torgerson and Macpherson2011).
Historically, the diseases caused by some of these parasites have been classified as Neglected Tropical Diseases (NTDs) or Neglected Zoonotic Diseases (King, Reference King2011) and were associated with the classical model of the ‘poverty trap’ covering tropical and sub-tropical regions in Africa, Latin America and the Indian subcontinent (Kuris, Reference Kuris2012). However, with global changes in climate and human demographics and associated practices, the classical models do not promise safe boundaries that might contain and/or stop the further global spread of many of these parasitic diseases (Colwell et al. Reference Colwell, Dantas-Torres and Otranto2011). The problems associated with these pathogens are further aggravated by the lack of effective vaccines (Dumonteil, Reference Dumonteil2007; Innes et al. Reference Innes, Bartley, Rocchi, Benavidas-Silvan, Burrells, Hotchkiss, Chianini, Canton and Katzer2011; McAllister, Reference McAllister2014; Black & Mansfield, Reference Black and Mansfield2016) and the paucity of reliable drugs (Zofou et al. Reference Zofou, Nyasa, Nsagha, Ntie-Kang, Meriki, Assob and Kuete2014), in addition to the difficulties of vector or reservoir control (Colwell et al. Reference Colwell, Dantas-Torres and Otranto2011). Therefore, there is a recognized need to find new therapeutic targets in these causative agents in order to develop effective treatment regimens to avoid potentially catastrophic outbreaks, both in terms of human health and economic impact.
This review presents sphingolipid (SL) biosynthesis and ceramide (CER) homoeostasis as a potential gold mine of tractable drug targets for these protozoan parasites.
State-of-the-art treatment of apicomplexan and kinetoplastid diseases
In general, available treatments for the diseases caused by the Kinetoplastidae and Apicomplexa are outdated (if not historic), with relatively few examples that were introduced recently, toxic and require a long treatment regimen, and therefore close monitoring of patients.
The kinetoplastid pathogens in focus here all cause NTDs and as such there are significant problems with the available drug regimens:
Leishmania spp
The treatment of leishmaniasis often requires a long course of intravenous pentavalent antimony drugs (e.g. Glucantime and Pentostam), aminosidine (paromomycin) or liposomal amphotericin B (Croft & Coombs, Reference Croft and Coombs2003; Center for Food Security and Public Health, 2004; WHO, 2004; Kedzierski et al. Reference Kedzierski, Sakthianandeswaren, Curtis, Andrews, Junk and Kedzierska2009). The most recent addition was the orally available miltefosine (Sunder et al. Reference Sunder, Jha, Thakur, Engel, Sindermann, Fischer, Jungle, Bryceson and Berman2002; Verma & Dey, Reference Verma and Dey2004), originally developed as anti-neoplastic agent. Despite its teratogenic effects (Sunder et al. Reference Sunder, Jha, Thakur, Engel, Sindermann, Fischer, Jungle, Bryceson and Berman2002), due to the lack of other effective medications, it has been registered and is now used in India, Colombia, Guatemala and Germany (Soto & Berman, Reference Soto and Berman2006). Other regimens of treatment include Pentamidine (Bray et al. Reference Bray, Barrett, Ward and de Koning2003), allopurinol, dapsone, fluconazole, itraconazole and ketoconazole. However, to-date all available chemotherapeutic agents suffer from being toxic (Chappuis et al. Reference Chappuis, Sundar, Hailu, Ghalib, Rijal, Peeling, Alvar and Boelaert2007) or inaccessible, both geographically and financially, in endemic areas where public health is under-resourced, poor and underdeveloped. Additionally, the lack of effective vaccines (de Oliveira et al. Reference de Oliveira, Nascimento, Barral, Soto and Barral-Netto2009) and the alarming emergence of resistance to these drugs (Croft et al. Reference Croft, Sundar and Fairlamb2006), combined with the short-lived prevention resulting from applying measures such as vector and reservoir host control (WHO, 2004; Figueiredo et al. Reference Figueiredo, Rodrigues, Silva, Koeller, Jiang, Jazwinski, Previato, Mendonça-Previato, Ürményi and Heise2012), demand an intensive search for alternative anti-leishmanials to enable effective treatment and control.
Trypanosoma brucei
Another compelling example of the shortcomings of available treatments is HAT (Mina et al. Reference Mina, Pan, Wansadhipathi, Bruce, Shams-Eldin, Schwarz, Steel and Denny2009; Buckner et al. Reference Buckner, Waters and Avery2012), where there is a lack of effective vaccines (Black & Mansfield, Reference Black and Mansfield2016) and treatment depends on the stage of the disease. Whilst in the first stage, the drugs used are less toxic, easier to administer and more effective, treatment in the second stage requires drugs that can cross the blood-brain barrier, specifically the arsenates (Gibaud & Jaouen, Reference Gibaud, Jaouen, Jaouen and Metzler-Nolte2010), making them considerably more toxic and complex to administer (Babokhov et al. Reference Babokhov, Sanyaolu, Oyibo, Fagbenro-Beyioku and Iriemenam2013). Currently, four drugs are registered for HAT treatment and are provided free of charge to endemic countries through a WHO private partnership with Sanofi-Aventis (Pentamidine, melarsoprol and eflornithine) and Bayer AG (suramin) (Schmidt et al. Reference Schmidt, Khalid, Romanha, Alves, Biavatti, Brun, Da Costa, de Castro, Ferreira, de Lacerda, Lago, Leon, Lopes, Amorim, Niehues, Ogungbe, Pohlit, Scotti, Setzer, Soeiro, Steindel and Tempone2012). Unfortunately, all of them exhibit a broad range of adverse effects. Moreover, treatment regimens are usually highly restrictive, particularly in the second stage of the disease, requiring hospital-based I.V. treatment with continuous monitoring.
Trypanosoma cruzi
Despite their toxic side-effects, nifurtimox and benznidazole are the only licenced drugs available for treatment of Chagas’ disease (Carabarin-Lima et al. Reference Carabarin-Lima, González-Vázquez, Rodríguez-Morales, Baylón-Pacheco, Rosales-Encina, Reyes-López and Arce-Fonseca2013; Bermudez et al. Reference Bermudez, Davies, Simonazzi, Pablo Real and Palma2016), with the latter being the first choice due to its lower side effects. Also, benznidazole has been implemented in the treatment of women before pregnancy in order to prevent/reduce vertical transmission (Carabarin-Lima et al. Reference Carabarin-Lima, González-Vázquez, Rodríguez-Morales, Baylón-Pacheco, Rosales-Encina, Reyes-López and Arce-Fonseca2013; Murcia et al. Reference Murcia, Carrilero, Munoz-Davila, Thomas, López and Segovia2013). Due to the lack alternatives, efforts have been directed towards implementing different treatment regimens in order to reduce toxicity, e.g. intermittent administration schedules, combination therapy and re-purposing of commercial drugs (Bermudez et al. Reference Bermudez, Davies, Simonazzi, Pablo Real and Palma2016).
Management of apicomplexan infections is also challenging and faces many of the same shortcomings encountered in the treatment of kinetoplastid infections.
Toxoplasma gondii
Treatment regimens for toxoplasmosis patients have essentially remained the same since the 1950s (Eyles & Coleman, Reference Eyles and Coleman1953). They largely depend on the repurposing of antibacterials (sulfadiazine, spiramycin and clindamycin) and antimalarials (pyrimethamine and atovaquone) (Opremcak et al. Reference Opremcak, Scales and Sharpe1992; Andrews et al. Reference Andrews, Fisher and Skinner-Adams2014; Antczak et al. Reference Antczak, Dzitko and Długońska2016) in combination, therapies that target parasite folic acid synthesis, protein synthesis or oxidative phosphorylation (Greif et al. Reference Greif, Harder and Haberkorn2001; Antczak et al. Reference Antczak, Dzitko and Długońska2016). Most of these chemotherapeutics are not readily bioavailable at the site of infection (e.g. unable to cross the blood-brain barrier); cannot be administered by patients with hypersensitivity to sulphonamides; have suspected teratogenic properties (Montoya & Remington, Reference Montoya and Remington2008; Paquet & Yudin, Reference Paquet and Yudin2013); are threatened by the emergence of resistance (Sims, Reference Sims and Mayers2009); or require adjuvant therapies (folinic acid supplement) to minimize toxic side effects (for a detailed review see Antczak et al. Reference Antczak, Dzitko and Długońska2016).
Toxoplasmosis is a representative of the urgent need for new antiprotozoal targets. In addition to the fact that T. gondii is estimated to infect 2–3 billion people worldwide (Welti et al. Reference Welti, Mui, Sparks, Wernimont, Isaac, Kirisits, Roth, Roberts, Botte, Marechal and McLeod2007), its treatment is complicated due to two main factors: (a) the parasite undergoes a complex life cycle with two predominant forms in the human host, namely, tachyzoites (proliferative form) and bradyzoites (encysted form, chronic toxoplasmosis); (b) bradyzoite burden is widespread but usually asymptomatic, although it has been associated with psychiatric disorders (Webster et al. Reference Webster, Kaushik, Bristow and McConkey2013). However, in immunocompromised individuals encysted T. gondii transform into proliferative tachyzoite forms causing symptomatic disease, toxoplasmic encephalitis. As such T. gondii is an opportunistic parasite. Notably, all the above-mentioned drugs act only against the tachyzoite stage with no notable effect against encysted bradyzoites (Antczak et al. Reference Antczak, Dzitko and Długońska2016). Recent data from our laboratory (Alqaisi et al. Reference Alqaisi, Mbekeani, Llorens, Elhammer and Denny2017) and others (Sonda et al. Reference Sonda, Sala, Ghidoni, Hemphill and Pieters2005) have shown that the Aureobasidin A and analogous depsipeptides, known to target yeast SL biosynthesis (Wuts et al. Reference Wuts, Simons, Metzger, Sterling, Slightom and Elhammer2015), exhibit activity against bradyzoite T. gondii. This class of compounds may offer a potential treatment for chronic toxoplasmosis and, perhaps, some psychiatric disorders; although the mechanism of action is not via inhibition of parasite SL biosynthesis and is yet to be elucidated (Alqaisi et al. Reference Alqaisi, Mbekeani, Llorens, Elhammer and Denny2017).
Plasmodium falciparum
Falciparum malaria remains one of the ‘Big Three’, most prevalent and deadly infectious diseases across tropical and subtropical regions, with an estimated 154–289 million cases in 2010 (212 million cases in 2015), and 660 000 (429 000 in 2015) associated deaths; although the actual numbers might be even higher (Biamonte et al. Reference Biamonte, Wanner and Le Roch2013; WHO, 2016).
Similar to T. gondii, Plasmodium parasite undergoes a complex life cycle with different stages in different organs of the host, rendering treatment challenging: sporozoites and schizonts in the liver, and merozoites, trophozoites and gametocytes in the blood (Dechy-Cabaret & Benoit-Vical, Reference Dechy-Cabaret and Benoit-Vical2012). Artemisinin-based combination therapies (ACTs) are the standard for treating malaria cases with typical partner drugs including lumefantrine and piperaquine, e.g. Coartem™ (Novartis) and Eurartesim™ (Sigma-Tau) (Biamonte et al. Reference Biamonte, Wanner and Le Roch2013). Other regimens include the use of parenteral artesunate (severe malaria) (Dondorp et al. Reference Dondorp, Fanello, Hendriksen, Gomes, Seni, Chhaganlal, Bojang, Olaosebikan, Anunobi, Maitland, Kivaya, Agbenyega, Nguah, Evans, Gesase, Kahabuka, Mtove, Nadjm, Deen, Mwanga-Amumpaire, Nansumba, Karema, Umulisa, Uwimana, Mokuolu, Adedoyin, Johnson, Tshefu, Onyamboko, Sakulthaew, Ngum, Silamut, Stepniewska, Woodrow, Bethell, Wills, Oneko, Peto, von Seidlein, Day and White2010a ), primaquine (liver and transmission, gametocyte, stages) (Dondorp, Reference Dondorp2013), mefloquine and sulfadoxine/pyrimethamine in combination (effective as single dose antimalarial drug) (Biamonte et al. Reference Biamonte, Wanner and Le Roch2013) and atovaquone/proguanil, Malarone™ (GlaxoSmith Kline), as a prophylactic treatment.
However, although combination therapies have now been adopted, resistance against many existing antimalarials has been observed since the 1950s (Bishop, Reference Bishop1951; Hallinan, Reference Hallinan1953; Sandosham et al. Reference Sandosham, Eyles and Montgomery1964) and remains a severe threat (Rieckmann & Cheng, Reference Rieckmann and Cheng2002; Chinappi et al. Reference Chinappi, Via, Marcatili and Tramontano2010; Dondorp et al. Reference Dondorp, Yeung, White, Nguon, Day, Socheat and von Seidlein2010b ; Newton et al. Reference Newton, Caillet and Guerin2016; Parija, Reference Parija2016; Menard & Dondorp, Reference Menard and Dondorp2017; Zhou et al. Reference Zhou, Xia, Wei, Liu and Peng2017). This bleak view of the future of available anti-malarial chemotherapeutics makes it imperative to invest more efforts in identifying new potent chemotypes that will offer both efficacy and safety.
Cryptosporidium spp
Like T. gondii, Cryptosporidium parvum and Cryptosporidium hominis usually cause a self-limiting disease in healthy individuals but represent a manifest problem in immuno-compromised patients, particularly those with AIDS, where infection leads to acute and protracted life-threatening gastroenteritis (Chen et al. Reference Chen, Keithly, Paya and LaRusso2002). More recent data have led to a radical reassessment of the impact of cryptosporidiosis, with the number of Cryptosporidium-attributable diarrhoea episodes estimated at >7·5 million in children aged <24 months in sub-Saharan Africa and South Asia where infection is estimated to contribute to >250 000 infant deaths per year (Sow et al. Reference Sow, Muhsen, Nasrin, Blackwelder, Wu, Farag, Panchalingam, Sur, Zaidi, Faruque, Saha, Adegbola, Alonso, Breiman, Bassat, Tamboura, Sanogo, Onwuchekwa, Manna, Ramamurthy, Kanungo, Ahmed, Qureshi, Quadri, Hossain, Das, Antonio, Hossain, Mandomando, Nhampossa, Acácio, Omore, Oundo, Ochieng, Mintz, O'Reilly, Berkeley, Livio, Tennant, Sommerfelt, Nataro, Ziv-Baran, Robins-Browne, Mishcherkin, Zhang, Liu, Houpt, Kotloff and Levine2016). Current treatment of cryptosporidiosis relies on a single FDA-approved drug, nitazoxanide, which has limited efficacy in those most at risk. More recently, the repurposing of antimalarials, e.g. quinolones and allopurinols, has been proposed (Gamo et al. Reference Gamo, Sanz, Vidal, de Cozar, Alvarez, Lavandera, Vanderwall, Green, Kumar, Hasan, Brown, Peishoff, Cardon and Garcia-Bustos2010; Chellan et al. Reference Chellan, Sadler and Land2017).
The distinctive metabolic features of this parasite from other apicomplexan organisms, e.g. no plastid-derived apicoplast and the absence of the citrate cycle and cytochrome-based respiratory chain (Ryan & Hijjawi, Reference Ryan and Hijjawi2015), confer several limitations for the identification of targets necessary for the development of anticryptosporidial drugs. However, the core metabolic pathways, e.g. energy metabolism and lipid synthesis are still present and exhibit high level of divergence from the mammalian host, thus presenting an opportunity to identify new drug targets that promise effective and selective treatment (Chellan et al. Reference Chellan, Sadler and Land2017).
The biological significance of SLs
SLs are a class of lipids that are ubiquitous in eukaryotic cell membranes, particularly the plasma membrane, as well as in some prokaryotic organisms and viruses (Merrill & Sandhoff, Reference Merrill, Sandhoff, Vance and Vance2002). Since their earliest characterization by Thudichum (Reference Thudichum1884), they have been a subject of controversy. Initially, they had been considered of structural importance only; however, over the last couple of decades, several reports have revealed their indispensability to a plethora of functions including, but not limited to, the formation of structural domains, polarized cellular trafficking, signal transduction, cell growth, differentiation and apoptosis (Huwiler et al. Reference Huwiler, Kolter, Pfeilschifter and Sandhoff2000; Ohanian & Ohanian, Reference Ohanian and Ohanian2001; Cuvillier, Reference Cuvillier2002; Pettus et al. Reference Pettus, Chalfant and Hannun2002; Buccoliero & Futerman, Reference Buccoliero and Futerman2003).
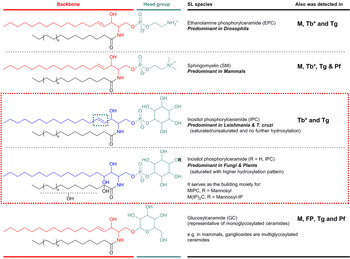
SLs consist structurally of a sphingoid base backbone, e.g. sphingosine (SPH) that can be N-acylated to form CER. To the latter, a variety of head groups: charged, neutral, phosphorylated and/or glycosylated can be attached to form complex SLs, e.g. sphingomyelin (SM), as the primary complex mammalian SL; and inositol phosphorylceramide (IPC) in fungi, plants and numerous protozoa (Fig. 1). These molecules have both polar and non-polar regions giving rise to their amphipathic character, which accounts for their tendency to aggregate into membranous structures, yet retaining the interfacial ability to interact with various partners, e.g. involvement of glycosphingolipids (GSLs) in cellular recognition complexes, cell adhesion and the regulation of cell growth (Gurr et al. Reference Gurr, Harwood and Frayn2002). Furthermore, the diversity of their chemical structures allows for distinctive roles within cellular metabolism, e.g. the signalling functions of SPH and CER vs sphingosine-1-phosphate (S1P) and ceramide-1-phosphate (C1P) (Merrill & Sandhoff, Reference Merrill, Sandhoff, Vance and Vance2002; Metzler, Reference Metzler2003).
Illustration of the predominant species of complex sphingolipid in organisms from different evolutionary clades: EPC in Drosophila; SM in mammals; and IPC in Leishmania and T. cruzi (as representatives of protozoan parasites) and in fungi and plants. IPC is absent from Mammalian cells but essential for many pathogenic organisms (red box). Glycosylated sphingolipids are also ubiquitous across different species. Backbone chain length is commonly C18 derived from palmitoyl-CoA. Mammals M, Fungi and Plants FP, Leishmania spp. L, Trypanosoma cruzi Tc, Trypanosoma brucei Tb, Toxoplasma gondii Tg and Plasmodium falciparum Pf. *Denotes developmental regulation. EPC, ethanolamine phosphorylceramide; IPC, inositol phosphorylceramide; SM, sphingomyelin.
SLs as indispensable structural components
The unique structural features of SLs (the free 3-hydroxy group, the amide functionality and the C4–C5 trans double bond) affect their biophysical properties rendering these molecules different from their glycerolipid counterparts, i.e. SM vs phosphatidylcholine (PC) (Boggs, Reference Boggs1980, Reference Boggs1987; Talbott et al., Reference Talbott, Vorobyov, Borchman, Taylor, DuPré and Yappert2000; Ramstedt & Slotte, Reference Ramstedt and Slotte2002). Such interfacial differences give complex SLs, such as SM, the unique ability to form both intra- and intermolecular hydrogen bonds (Bruzik, Reference Bruzik1988) that are fine-tuned by the trans double bond (Ramstedt & Slotte, Reference Ramstedt and Slotte2002). This ability is reflected in the tendency of SLs to cluster rather than behave like typical ‘fluid’ membrane lipids. Naturally occurring SLs undergo the L β (gel phase) to L α (lamellar phase) transition near the physiological temperature of 37 °C, in contrast, this transition for naturally occurring glycerolipids is near or below 0 °C. Additionally, the long saturated alkyl chains of SLs allow them to pack tightly with sterols, stabilized by hydrogen bonding (Ramstedt & Slotte, Reference Ramstedt and Slotte2002), to form laterally compact hydrophobic micro-domains commonly known as ‘lipid rafts’ (Futerman & Hannun, Reference Futerman and Hannun2004). Similar results have been reported with the fungal/plant counterpart of SM, IPC, where it was shown that IPC was able to form sterol containing ordered domains in model systems (Björkbom et al. Reference Björkbom, Ohvo-Rekilä, Kankaanpää, Nyholm, Westerlund and Slotte2010). These membrane micro-domains can readily segregate from the more disordered and expanded domains of unsaturated acyl chains of glycerolipids (Merrill & Sandhoff, Reference Vance and Vance2002). They have been proposed to function in a diverse array of processes from polarised trafficking of lipid modified proteins (Brown & London, Reference Brown and London1998) and the stabilization of other types of biological structures such as lamellar bodies, to the assembly and activation of signal transduction complexes (Brown & London, Reference Brown and London2000; Magee et al. Reference Magee, Prinen, Alder, Pagakis and Parmryd2002; Pierce, Reference Pierce2002; Vance & Vance, Reference Vance and Vance2002; Hannun & Obeid, Reference Hannun and Obeid2008). They have also been involved in the formation of detergent-insoluble gel-phase domains (Ramstedt & Slotte, Reference Ramstedt and Slotte2002) via the extensive hydrogen-bonding network in the head groups of GSLs that have been implicated during the formation of ‘caveolae’ and surface recognition (Merrill & Sandhoff, Reference Vance and Vance2002).
SLs as indispensable signalling agents
SLs can also function as bioactive signalling molecules due to their biophysical properties, e.g. the low pK a (7–8) of SPH allows it to remain partially uncharged at physiological pH retaining the ability to move across membranes (Merrill & Sandhoff, Reference Vance and Vance2002). Likewise, CER, a neutral species, is able to freely flip flop across membranes (Hannun & Obeid, Reference Hannun and Obeid2008). Many studies have produced evidence of such signalling functions, e.g. SPH exerts pleiotropic effects on protein kinases; CER mediates many cell-stress responses, including the regulation of apoptosis (Georgopapadakou, Reference Georgopapadakou2000); and S1P has crucial roles in cell survival, cell migration and inflammation (Hannun & Obeid, Reference Hannun and Obeid2008)
SL metabolism and the rationale for druggability
The indispensability of SLs for a myriad of cellular processes and functions, ranging from structural integrity to signalling events, makes it is unsurprising that the SL biosynthesis is highly conserved in all eukaryotes where it is, alongside its proposed regulators (Holthuis et al. Reference Holthuis, Tafesse and Ternes2006), an essential pathway (Heung et al. Reference Heung, Luberto and Del Poeta2006; Sutterwala et al. Reference Sutterwala, Creswell, Sanyal, Menon and Bangs2007). This has lead the pathway to be considered vital for protozoan pathogenesis and, therefore, a drug target; e.g. SM synthase activity in Plasmodium (Heung et al. Reference Heung, Luberto and Del Poeta2006). In order to characterise the druggability of protozoan SL biosynthesis, the mammalian pathway, as the most studied system, will be used as the reference model in the following discussions.
SL metabolism constitutes a highly complex network involving critical intersections with various other pathways, particularly glycerolipid biosynthesis (Holthuis & Menon, Reference Holthuis and Menon2014). CER represents the corner stone for both biosynthesis and catabolism, modulating cell fate (Hannun & Obeid, Reference Hannun and Obeid2008). Dysregulation of either SL biosynthesis or catabolism could result in cell death, e.g. of protozoan parasites (Yatsu, Reference Yatsu1971; Brady, Reference Brady1978; Chen et al. Reference Chen, Patterson, Wheatley, O'Brien and Pagano1999; Merrill & Sandhoff, Reference Vance and Vance2002), however here our focus will be on the former pathway.
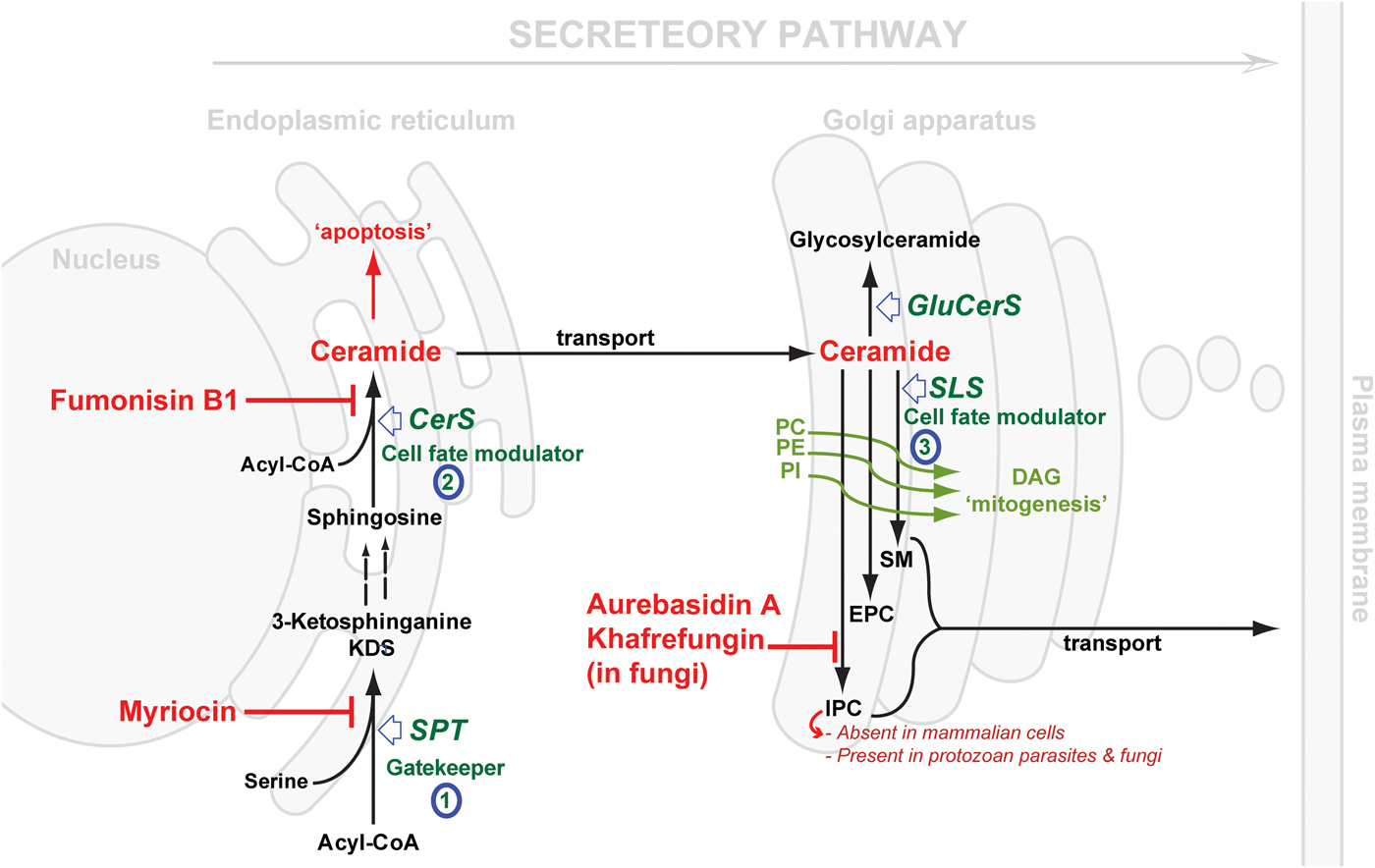
Considering the central position of CER, the druggability of SL metabolism revolves around dysregulation of ‘Ceramide Homeostasis’ (Young et al. Reference Young, Mina, Denny and Smith2012) which in turn leads to ripple effects perturbing the balance between the pro-apoptotic CER and the mitogenic diacylglycerol (DAG), consequently determining cell fate (Fig. 2) – a mechanism that has been associated with resistance to anti-cancer treatments (Ségui et al. Reference Ségui, Andrieu-Abadie, Jaffrézou, Benoist and Levade2006) and has been reported in protozoan parasites, e.g. Plasmodium (Pankova-Kholmyansky et al. Reference Pankova-Kholmyansky, Dagan, Gold, Zaslavsky, Skutelsky, Gatt and Flescher2003; Labaied et al. Reference Labaied, Dagan, Dellinger, Gèze, Egée, Thomas, Wang, Gatt and Grellier2004). The characterisation of several key enzymes involved in SL de novo biosynthesis has revealed divergence between mammalian and protozoan species. Thus, attention has been given to the exploitation of the SL biosynthetic pathway (parasite and/or host) for new drug targets or regimens (Sugimoto et al. Reference Sugimoto, Sakoh and Yamada2004; Zhang et al. Reference Zhang, Hsu, Scott, Docampo, Turk and Beverley2005; Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006; Tanaka et al. Reference Tanaka, Valero, Takahashi and Straus2007; Pruett et al. Reference Pruett, Bushnev, Hagedorn, Adiga, Haynes, Sullards, Liotta and Merrill2008; Mina et al. Reference Mina, Pan, Wansadhipathi, Bruce, Shams-Eldin, Schwarz, Steel and Denny2009; Tatematsu et al. Reference Tatematsu, Tanaka, Sugiyama, Sudoh and Mizokami2011; Young et al. Reference Young, Mina, Denny and Smith2012).
Schematic representation of de novo sphingolipid metabolism. Three key steps are highlighted: (1) SPT, evolutionary divergent in T. gondii; (2) CerS, fewer isoforms in protozoan parasite (c.f. 6 isoforms in mammals); SLS, while predominantly synthesising SM in mammals and to a lesser extent EPC, orthologues in protozoan parasites (Leishmania spp., T. brucei, T. cruzi and T. gondii) can synthesise IPC, an activity that is absent from mammalian cells and the target of the highly specific fungal inhibitors shown. The scheme also illustrates the differential cellular effects of ceramide vs DAG (diacylglycerol). Accumulation of ceramide elicits an apoptotic response while increasing concentrations of DAG promotes cell growth. CerS, ceramide synthase; GluCerS, glucosylceramide synthase; SLS, sphingolipid synthase; SPT, serine palmitoyltransferase; PC, phosphatidylcholine; PE, phosphatidylethanolamine; PI, phosphatidylinositol; SM, sphingomyelin; EPC, ethanolamine phosphorylceramide and IPC, inositol phosphorylceramide.
SL METABOLISM
The key steps in de novo biosynthesis
SL de novo biosynthesis can be simplified into three key steps: a gate-keeper and two cell fate modulator steps. The former comprises the up-stream rate-limiting step of the condensation of acyl-CoA and L-serine, in the endoplasmic reticulum (ER) via serine palmitoyltransferase (SPT), to produce dihydrosphingosine. The latter comprises first the formation of CER in the ER by the action of ceramide synthase (CerS), and then the formation of complex SLs in the Golgi. These products vary depending on the species, and are formed under the catalysis of what could be generically termed SL synthases: SM synthase in mammals and IPC synthase in fungi, plants and protozoa. It is worth mentioning that another Golgi localized metabolic pathway results in the formation of glycosylated CER species, and also contributes to the regulation CER levels (Holthuis & Menon, Reference Holthuis and Menon2014) (Fig. 2).
Protozoan parasites vs host: differences & opportunities
The cross-species differences encountered in the first, SPT-catalysed, step are mostly minor in terms of the chemical structure of the product; mainly due to the chain length of the acyl-CoA utilised in the reaction, e.g. myristoyl-CoA (in Leishmania spp. amongst other odd sphingoid base lengths (Hsu et al. Reference Hsu, Turk, Zhang and Beverley2007)) and palmitoyl-CoA, with the latter more predominant across the Eukaryota (in mammals, Plasmodium and T. brucei) (Richmond et al. Reference Richmond, Gibellini, Young, Major, Denton, Lilley and Smith2010; Botté et al. Reference Botté, Yamaryo-Botté, Rupasinghe, Mullin, MacRae, Spurck, Kalanon, Shears, Coppel, Crellin, Maréchal, McConville and McFadden2013). Further differences may be apparent with respect to the catalysing enzyme, SPT (vide infra). However, clear divergence is observed in the second and the third steps, both of which represent a cell-fate modulator process. CerSs exhibit differential preferences for the chain length of the acyl-CoA substrate (Park et al. Reference Park, Park and Futerman2014) and its hydroxylation pattern (Layre & Moody, Reference Layre and Moody2013), with 6 isoforms present in humans suggesting a different role for each CER species produced (Levy & Futerman, Reference Levy and Futerman2010; Figueiredo et al. Reference Figueiredo, Rodrigues, Silva, Koeller, Jiang, Jazwinski, Previato, Mendonça-Previato, Ürményi and Heise2012). To-date, one or, maximum, two genes encoding CerS function have been identified in protozoan parasite species (Koeller & Heise, Reference Koeller and Heise2011). However, most interesting is the variation in the complex SL formed in the Golgi, reflecting significant differences in the active site of the SL synthases catalysing the transfer reaction. The divergence of the protozoal complex SL synthases, and the synthetic products, with respect to the mammalian host, may provide opportunities to design selective inhibitors. Previously, this step has been validated as a promising drug target in fungi using aureobasidin A (AbA) (Fig. 2) (Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006).
Serine palmitoyl transferase (SPT)
SPTs are members of the pyridoxal 5′-phosphate (PLP)-dependent (Sandmeier et al. Reference Sandmeier, Hale and Christen1994) α-oxoamine synthase family and share a conserved motif (T[FL][GTS]K[SAG][FLV]G) around the PLP-binding lysine (in bold) (Young et al. Reference Young, Mina, Denny and Smith2012). SPT catalyses the first rate-limiting step in the de novo biosynthesis of SLs (Weiss & Stoffel, Reference Weiss and Stoffel1997; Hojjati et al. Reference Hojjati, Li and Jiang2005) (Fig. 2), a reaction involving the decarboxylative Claisen-like condensation of serine and an acyl-CoA (Lowther et al. Reference Lowther, Naismith, Dunn and Campopiano2012), to yield the sphingoid base backbone, 3-ketodihydrosphingosine (3-KDS) (Hanada, Reference Hanada2003; Raman et al. Reference Raman, Johnson, Yard, Lowther, Carter, Naismith and Campopiano2009; Lowther et al. Reference Lowther, Naismith, Dunn and Campopiano2012). Therefore, SPT represents the ‘Gatekeeper’ of the SL biosynthetic pathway.
All eukaryotic SPTs studied to date are ER-resident and membrane bound with a heterodimeric protein core consisting of two subunits sharing ~20% identity: LCB1 and LCB2, ~53 and ~63 kDa respectively (Hanada, Reference Hanada2003; Denny et al. Reference Denny, Goulding, Ferguson and Smith2004; Han et al. Reference Han, Gable, Yan, Natarajan, Krishnamurthy, Gupta, Borovitskaya, Harmon and Dunn2004; Chen et al. Reference Chen, Han, Dietrich, Dunn and Cahoon2006). The latter contains the canonical PLP cofactor binding site while the former has been suggested to be important for complex stability (Lowther et al. Reference Lowther, Naismith, Dunn and Campopiano2012). In contrast, the orthologous SPT from sphingomonad bacteria is a soluble 45 kDa homodimer (Ikushiro et al. Reference Ikushiro, Hayashi and Kagamiyama2001). SPT activity in apicomplexan parasites has been detected and was proposed as a potential drug target (Gerold & Schwarz, Reference Gerold and Schwarz2001; Bisanz et al. Reference Bisanz, Bastien, Grando, Jouhet, Marechal and Cesbron-Delauw2006; Coppens, Reference Coppens2013), however the enzyme(s) responsible have yet to be further characterized (Mina et al. Reference Mina, Thye, Alqaisi, Bird, Dods, Groftehauge, Mosely, Pratt, Shams-Eldin, Schwarz, Pohl and Denny2017). In contrast, kinetoplastid parasites have been shown to possess a heterodimeric SPT similar to the mammalian orthologue (Denny et al. Reference Denny, Goulding, Ferguson and Smith2004). Inhibiting SPT activity (e.g. using myriocin, Fig. 2) results in various effects in different species. Mammalian cells exhibited a loss of viability, with a partial loss of SPT function resulting in a rare SL metabolic disease, Hereditary Sensory Neuropathy type I (HSN1) (Hanada, Reference Hanada2003). In contrast, Saccharomyces cerevisiae were found to be relatively tolerant (Nagiec et al. Reference Nagiec, Baltisberger, Wells, Lester and Dickson1994), and Leishmania major lacking LCB2 were viable but unable to differentiate into infective metacyclic forms (Zhang et al. Reference Zhang, Showalter, Revollo, Hsu, Turk and Beverley2003). However, T. brucei procyclic forms in which SPT expression was reduced were non-viable (Fridberg et al. Reference Fridberg, Olson, Nakayasu, Tyler, Almeida and Engman2008).
The SPT catalysed reaction product, 3KDS, is subsequently reduced by 3-ketosphinganine reductase to form sphinganine (dihydrosphingosine). Subsequent minor metabolic differences are encountered across different species; mainly concerning the order of the hydroxylation (in fungi and higher plants) and acylation to produce CERs (Sugimoto et al. Reference Sugimoto, Sakoh and Yamada2004).
Ceramide synthase
In all eukaryotic systems studied to date, CerSs are ER-resident integral membrane proteins catalysing the N-acetylation of dihydrosphingosine to produce dihydroceramide, which is then oxidized to form CER, the simplest SL species and a key bioactive molecule in numerous cellular pathways (Lahari & Futerman, Reference Lahari and Futerman2007).
Mammalian CerSs are orthologues of longevity-assurance genes, LAG1p and LAC1p identified in yeast (Guillas et al. Reference Guillas, Kirchman, Chuard, Pfefferli, Jiang, Jazwinski and Conzelmann2001). The eukaryotes studied to date have been found to encode at least two CerSs, with humans expressing six – each generating CER with a defined acyl chain length (C18 to C26) (Pewzner-Jung et al. Reference Pewzner-Jung, Ben-Dor and Futerman2006; Levy & Futerman, Reference Levy and Futerman2010). Whilst little is known regarding structure-function relationships or regulation of CerS,, the ubiquitous Lag1 motif has been shown to be important for functionality (Spassieva et al. Reference Spassieva, Seo, Jiang, Bielawski, Alvarez-Vasquez, Jazwinski, Hannun and Obeid2006), likely forming part of the active site.
Experimental evidence (from our laboratory and others) has previously indicated the presence of CerS activity in Leishmania spp (Zhang et al., Reference Zhang, Showalter, Revollo, Hsu, Turk and Beverley2003; Denny et al., Reference Denny, Goulding, Ferguson and Smith2004, Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006) and in T. cruzi (De Lederkremer et al. Reference De Lederkremer, Agusti and Docampo2011). More recently LAG1 orthologues have been identified and functionally and molecularly characterized in the latter parasite (Figueiredo et al. Reference Figueiredo, Rodrigues, Silva, Koeller, Jiang, Jazwinski, Previato, Mendonça-Previato, Ürményi and Heise2012). Other results indirectly suggest the presence of such activity in T. brucei (Patnaik et al. Reference Patnaik, Field, Menon, Cross, Yee and Butikofer1993; Richmond et al. Reference Richmond, Gibellini, Young, Major, Denton, Lilley and Smith2010; Smith & Bütikofer, Reference Smith and Bütikofer2010). Similarly, CerS activity in the Apicomplexa has been inferred (Welti et al. Reference Welti, Mui, Sparks, Wernimont, Isaac, Kirisits, Roth, Roberts, Botte, Marechal and McLeod2007; Zhang et al. Reference Zhang, Bangs and Beverley2010; Pratt et al. Reference Pratt, Wansadhipathi-Kannangara, Bruce, Mina, Shams-Eldin, Casas, Hanada, Schwarz, Sonda and Denny2013), but remains unexplored.
Once formed in the ER, CER is transported, by CER transfer protein CERT in mammals (Kumagai et al. Reference Kumagai, Yasuda, Okemoto, Nishijima, Kobayashi and Hanada2005; Kudo et al. Reference Kudo, Kumagai, Matsubara, Kobayashi, Hanada, Wakatsuki and Kato2010; Rao et al. Reference Rao, Scheffer, Srideshikan, Parthibane, Kosakowska-Cholody, Masood, Nagashima, Gudla, Lockett, Acharya and Acharya2014), to the Golgi apparatus where the synthesis of complex SLs occurs (Ohanian & Ohanian, Reference Ohanian and Ohanian2001; Bromley et al. Reference Bromley, Li, Murphy, Sumner and Lynch2003; Bartke & Hannun, Reference Bartke and Hannun2009; Pata et al. Reference Pata, Hannun and Ng2010). ER CER concentration is kept under tight control as accumulation of CER here has been shown to result in induction of the mitochondrial apoptotic pathway (Vacaru et al. Reference Vacaru, Tafesse, Ternes, Kondylis, Hermansson, Brouwers, Somerharju, Rabouille and Holthuis2009; Tafesse et al. Reference Tafesse, Vacaru, Bosma, Hermansson, Jain, Hilderink, Somerharju and Holthuis2014) via an unknown mechanism (Bockelmann et al. Reference Bockelmann, Mina, Jain, Ehring, Korneev and Holthuis2015).
Sphingolipid synthase
In the Golgi, CER can be phosphorylated by CER kinase (Rovina et al. Reference Rovina, Schanzer, Graf, Mechtcheriakova, Jaritz and Bornancin2009), glycosylated by glucosyl or galactosyl CerS (Raas-Rothschild et al. Reference Raas-Rothschild, Pankova-Kholmyansky, Kacher and Futerman2004), or acquire a variety of neutral or charged head groups under the catalysis of what could be called generically SLSs, to form various complex phosphosphingolipids. Phylogenetic analyses have identified at least 4 clades of SLS (Huitema et al. Reference Huitema, van den Dikkenberg, Brouwers and Holthuis2004; Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006).
In mammals CER is a substrate for the SLS, SM synthase, to produce SM (Huitema et al. Reference Huitema, van den Dikkenberg, Brouwers and Holthuis2004). Whilst in fungi and higher plants phytoceramide is utilized by a different SLS, IPC synthase, to produce IPC as the principal phosphosphingolipid (Nagiec et al. Reference Nagiec, Nagiec, Baltisberger, Wells, Lester and Dickson1997; Wang et al. Reference Wang, Yang, Tangchaiburana, Ndeh, Markham, Tsegaye, Dunn, Wang, Bellizzi, Parsons, Morrissey, Bravo, Lynch and Xiao2008). This landscape is significantly divergent when it comes to protozoa.
In the kinetoplastid Leishmania spp. and T. cruzi, CER acquires a phosphorylinositol head group from phosphatidylinositol (PI) to produce IPC via IPC synthase (Zhang et al. Reference Zhang, Hsu, Scott, Docampo, Turk and Beverley2005; Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006; Mina et al. Reference Mina, Mosely, Ali, Shams-Eldin, Schwarz, Steel and Denny2010), although there are some reports of SM in T. cruzi (Quiñones et al. Reference Quiñones, Urbina, Dubourdieu and Luis Concepción2004) (Fig. 2). Whilst Leishmania encodes a single copy IPC synthase, T. cruzi has two highly related copies (Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006). Further divergence, and possible redundancy, is encountered in T. Brucei, which harbours 4 genes that encode SLSs (Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006; Sutterwala et al. Reference Sutterwala, Hsu, Sevova, Schwartz, Zhang, Key, Turk, Beverley and Bangs2008). This enzyme portfolio results in a diverse profile of the complex SL species (SM, IPC and ethanolamine phosophorylceramide [EPC]) which are developmentally regulated during the life cycle of the parasite (Sutterwala et al. Reference Sutterwala, Hsu, Sevova, Schwartz, Zhang, Key, Turk, Beverley and Bangs2008).
In apicomplexan parasites, previous reports have indicated the presence of glycosyl-ceramide and SM in P. falciparum and T. gondii, as summarized in Zhang et al (Reference Zhang, Bangs and Beverley2010). However, other findings reported the presence of EPC in T. gondii (Welti et al. Reference Welti, Mui, Sparks, Wernimont, Isaac, Kirisits, Roth, Roberts, Botte, Marechal and McLeod2007) and, more recently, IPC (Pratt et al. Reference Pratt, Wansadhipathi-Kannangara, Bruce, Mina, Shams-Eldin, Casas, Hanada, Schwarz, Sonda and Denny2013). The latter study also characterized T. gondii SLS as demonstrating IPC synthase activity in vitro (Pratt et al. Reference Pratt, Wansadhipathi-Kannangara, Bruce, Mina, Shams-Eldin, Casas, Hanada, Schwarz, Sonda and Denny2013).
The divergence of SLS function, with respect to the host, seen in both kinetoplastid and apicomplexan protozoan parasites in intriguing and, perhaps, indicated them as a tractable drug target. In support of this hypothesis, ceramide-analogues with anti-Plasmodium activity have already been identified (Labaied et al. Reference Labaied, Dagan, Dellinger, Gèze, Egée, Thomas, Wang, Gatt and Grellier2004).
In general, SLSs are Golgi-resident transmembrane proteins, presumed to have 6 transmembrane domains with the active site facing the Golgi lumen (Holthuis et al. Reference Holthuis, Tafesse and Ternes2006; Sutterwala et al. Reference Sutterwala, Hsu, Sevova, Schwartz, Zhang, Key, Turk, Beverley and Bangs2008). Those orthologues identified in kinetoplastids demonstrated two conserved regions (CGDX 3SG H T & H YTX D VX 3YX 6FX 2YH) with respect to the animal SM synthases (Huitema et al. Reference Huitema, van den Dikkenberg, Brouwers and Holthuis2004; Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006). These regions contain the so-called the catalytic triad (two Histidines and one Aspartate residues) that mediates a nucleophilic attack on lipid phosphate ester during the transferase/hydrolase activity (Mina et al. Reference Mina, Mosely, Ali, Shams-Eldin, Schwarz, Steel and Denny2010). Apicomplexan orthologues form a separate evolutionary clade, yet retain the catalytic triad (Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006; Pratt et al. Reference Pratt, Wansadhipathi-Kannangara, Bruce, Mina, Shams-Eldin, Casas, Hanada, Schwarz, Sonda and Denny2013), as does the fungal orthologue AUR1p (Heidler & Radding, Reference Heidler and Radding2000). Further evidence for the essentiality of these residues was provided when mutation of the active histidine of the triad was shown to deactivate fungal IPC synthase and mammalian SM synthase-related activity (Levine et al. Reference Levine, Wiggins and Munro2000; Vacaru et al. Reference Vacaru, Tafesse, Ternes, Kondylis, Hermansson, Brouwers, Somerharju, Rabouille and Holthuis2009). Furthermore, recently it has been shown that substrate selectivity, and so the diversity of SLS activity, may depend on key residues close to the transferase active residues or on a luminal loop of the protein (Sevova et al. Reference Sevova, Goren, Schwartz, Hsu, Turk, Fox and Bangs2010; Kol et al. Reference Kol, Panatala, Nordmann, Swart, Van Suijlekom, Cabukusta, Hilderink, Gabrietz, Mina, Somerharju, Korneev, Tafesse and Holthuis2017).
In the Eukaryota SLS's occupy a central position at the intersection of glycerolipids (PI/PC/PE and DAG) and SLs ([phyto]ceramide and IPC/SM/EPC). Accordingly, these enzymes act as regulators of a delicate balance between pro-apoptotic CER and pro-mitogenic DAG (Holthuis et al. Reference Holthuis, Tafesse and Ternes2006).
The most significant previous example of SL biosynthesis inhibition as a drug target was reported in fungi. Aureobasidin A (AbA), a depsipeptide, was first reported by Ikai et al. (Reference Ikai, Takesako, Shiomi, Moriguchi, Umeda, Yamamoto, Kato and Naganawa1991) and soon after its antifungal properties were highlighted (Takesako et al. Reference Takesako, Kuroda, Inoue, Haruna, Yoshikawa, Kato, Uchida, Hiratani and Yamaguchi1993). The target gene was further characterized (Hashidaokado et al. Reference Hashidaokado, Ogawa, Endo, Takesako and Kato1995) revealing its identity to be the IPC synthase (AUR1p). AbA is a specific and potent (low nanomolar) inhibitor of the fungal IPC synthase. This ushered in a new era in the search for anti-fungal chemotherapeutics, positioning IPC synthase as a promising, broad spectrum, anti-fungal drug target (Sugimoto et al. Reference Sugimoto, Sakoh and Yamada2004). Other specific inhibitors were later added to the arsenal of fungal IPC synthase inhibitors: khafrefungin (Mandala et al. Reference Mandala, Thornton, Rosenbach, Milligan, Garcia-Calvo, Bull and Kurtz1997), rustmicin (Harris et al. Reference Harris, Shafiee, Cabello, Curotto, Genilloud, Goklen, Kurtz, Rosenbach, Salmon, Thornton, Zink and Mandala1998; Mandala et al. Reference Mandala, Thornton, Milligan, Rosenbach, Garcia-Calvo, Bull, Harris, Abruzzo, Flattery, Gill, Bartizal, Dreikorn and Kurtz1998) and others (Ohnuki et al. Reference Ohnuki, Yano, Ono, Kozuma, Suzuki, Ogawa and Takatsu2009). Unfortunately, further development of these inhibitors stalled, either due to physical properties, e.g. aureobasidin A is very sparingly soluble in water (Georgopapadakou, Reference Georgopapadakou2000; Sugimoto et al. Reference Sugimoto, Sakoh and Yamada2004), or because their highly complex chemical structures rendered chemical synthesis challenging, with the few synthetic efforts reported resulting in compounds with either reduced or no activity (Sugimoto et al. Reference Sugimoto, Sakoh and Yamada2004; Aeed et al. Reference Aeed, Young, Nagiec and Elhammer2009). However, recent works have highlighted that semi-synthetic strategies may overcome these barriers (Wuts et al. Reference Wuts, Simons, Metzger, Sterling, Slightom and Elhammer2015).
Perhaps reflecting the evolutionary divergence of these enzymes, the protozoan IPC synthase orthologues, from Leishmania major and T. gondii are not susceptible to AbA inhibition (Denny et al. Reference Denny, Shams-Eldin, Price, Smith and Schwarz2006; Pratt et al. Reference Pratt, Wansadhipathi-Kannangara, Bruce, Mina, Shams-Eldin, Casas, Hanada, Schwarz, Sonda and Denny2013). Some studies have reported the inhibitory effects of AbA and analogues against T. gondii in culture (Sonda et al. Reference Sonda, Sala, Ghidoni, Hemphill and Pieters2005; Alqaisi et al. Reference Alqaisi, Mbekeani, Llorens, Elhammer and Denny2017), however this is not associated with inhibition of SL biosynthesis. Despite this, the protozoan SLS's remain tractable drug targets with no functional equivalent in mammalian cells. Surprisingly, at least one SLS isoform from T. brucei was acutely sensitive to AbA treatment (Mina et al. Reference Mina, Pan, Wansadhipathi, Bruce, Shams-Eldin, Schwarz, Steel and Denny2009), although these findings stirred some controversy due, in part, to the redundancy of T. brucei SLSs (4 isoforms) compared with the single copy found, for example, in L. major and T. gondii (Sutterwala et al. Reference Sutterwala, Hsu, Sevova, Schwartz, Zhang, Key, Turk, Beverley and Bangs2008).
THE ENIGMATIC NATURE OF SL DRUGGABILITY
Difficulties in pinpointing SL functionality
Investigation and deciphering of the functions of each specific SL species remains challenging. This is due to the complexity in SL metabolic interconnections, their varied biophysical properties (neutral or charged), chain length variation, the hydrophobic nature of the involved enzymes and the presence of multiple pathways that can operate in parallel (Hannun & Obeid, Reference Hannun and Obeid2008). The interaction with other cellular metabolic pathways (e.g. glycerolipid metabolism) introduces another layer of complexity.
Overall, the signalling effect/role of an individual SL could be defined on spatial-temporal basis with at least five parameters: (a) subcellular localisation, (b) regulation (c) chain length specificity, (d) kinetics of trafficking and (e) mechanism of action. For example, phosphorylation of 1–3% cytosolic SPH may double the levels of S1P that acts on G protein-coupled receptor (GPCR) to elicit a specific response in a particular cellular locality for certain period of time (Hannun & Obeid, Reference Hannun and Obeid2008). Such signalling events can be described as a function of cytosolic S1P that is regulated by S1P Kinase, with the signal caused through the interaction of S1P with a GPCR. The elucidation of such complex systems remains challenging and a comprehensive discussion of the issue is beyond the scope of this review. However, an additional layer of significant complexity in terms of the pathogenic protozoa arises when considering the SL signalling network in the case of obligate intracellular parasites, where host SL biosynthesis, and its interaction with parasite de novo synthesis, must be taken into account.
Parasite–host SL interplay
The intimate parasite–host interaction in terms of SL metabolism has been well documented; L. major pathogenic amastigotes isolated from mammalian hosts showed normal IPC levels (Zhang et al. Reference Zhang, Hsu, Scott, Docampo, Turk and Beverley2005) despite lacking LCB2, a functional SPT and the ability to synthesis CER de novo. Alterations in host, macrophage, cell SL biosynthesis upon infection may compensate for this deficiency (Ghosh et al., Reference Ghosh, Bhattacharyya, Das, Raha, Maulik, Das, Roy and Majumdar2001, Reference Ghosh, Bhattacharyya, Sirkar, Sa, Das, Majumdar, Roy and Majumdar2002). These studies suggest a complex and multifaceted interplay between host and parasite SL metabolism comprising nutritional factors and signalling pathways that could modulate parasite survival and/or host defence (Zhang et al. Reference Zhang, Bangs and Beverley2010). Similar observations have been reported in the apicomplexan parasites (Romano et al. Reference Romano, Sonda, Bergbower, Smith and Coppens2013). This highlights the striking potential of host and parasite SL modulation as an anti-protozoal target, as is similarly proposed for pathogenic fungi (Zhang et al. Reference Zhang, Bangs and Beverley2010; Ramakrishnan et al. Reference Ramakrishnan, Serricchio, Striepen and Bütikofer2013).
PERSPECTIVE
Classically dissecting the role and locale of critical enzymatic steps in SL biosynthesis and assessing the effect on the parasite fitness and virulence could turn into an overwhelmingly challenging task aggravated by: the complexity of the metabolic pathway itself; the ability of the parasite to salvage (Coppens, Reference Coppens2013), hijack and remodel host SL; and developmental regulation during the parasitic life cycle, which adds another layer of intricacy rendering the deconvolution of any observed effects difficult to interpret. Fortunately, many of those problems can be now overcome with advances in technology. High resolution localization studies in protozoan parasites can benefit greatly from new microscopic techniques such as Airy-scan (Huff, Reference Huff2015), super-resolution microscopy (Florentino et al. Reference Florentino, Real, Bonfim-Melo, Orikaza, Ferreira, Pessoa, Lima, Sasso and Mortara2014) and upcoming technologies, e.g. phase-modulation nanoscopy (Pal, Reference Pal2015; Ward & Pal, Reference Ward and Pal2017), which can elucidate spatial arrangement of proteins of interest within the parasite to reveal potential interaction partners and shed light on mechanistic features. Similarly, new advances in chemical probes, and SL analogues in particular, such as bifunctional lipid technology (Haberkant & Holthuis, Reference Haberkant and Holthuis2014) coupled with high throughput proteomic (Ramaprasad et al. Reference Ramaprasad, Mourier, Naeem, Malas, Moussa, Panigrahi, Vermont, Otto, Wastling and Pain2015), could identify different interaction partners that would help map the biosynthetic pathway and its critical interactions. The effects of these probes on the parasite (and host) cell can now be comprehensively evaluated by monitoring the transcriptome, proteome, metabolomics (Watson, Reference Watson2010) and lipidome (Marechal et al. Reference Marechal, Riou, Kerboeuf, Beugnet, Chaminade and Loiseau2011). Such studies could reveal multiple windows of opportunity to exploit as potential drug targets. The targets identified in this way can now be rapidly genetically validated in the parasitic protozoa by applying modern gene editing technologies, such as CRISPR/Cas9 (Sugi et al. Reference Sugi, Kato and Weiss2016). Compared with the classical methodologies, this tool enables fast and efficient application for single gene (Serpeloni et al. Reference Serpeloni, Jimenez-Ruiz, Vidal, Kroeber, Andenmatten, Lemgruber, Morking, Pall, Meissner and Avila2016), and systematic genome-wide knockout generation (Sidik et al. Reference Sidik, Huet, Ganesan, Huynh, Wang, Nasamu, Thiru, Saeij, Carruthers, Niles and Lourido2016). Additionally, the development of novel orthogonal approach for conditional knockout strategies, e.g. tetracycline-induced gene disruption Tet-system (Meissner et al. Reference Meissner, Schlüter and Soldati2002), rapamycin-induced Cre recombinase-assisted gene excision (Andenmatten et al. Reference Andenmatten, Egarter, Jackson, Jullien, Herman and Meissner2013; Collins et al. Reference Collins, Das, Wong, Andenmatten, Stallmach, Hackett, Herman, Muller, Meissner and Blackman2013; Jimenez-Ruiz et al. Reference Jimenez-Ruiz, Wong, Pall and Meissner2014), has allowed testing of essential gene functionality, in Leishmania spp. (Duncan et al. Reference Duncan, Myburgh, Philipon, Brown, Meissner, Brewer and Mottram2016) and T. gondii (Pieperhoff et al. Reference Pieperhoff, Pall, Jimenez-Ruiz, Das, Melatti, Gow, Wong, Heng, Muller, Blackman and Meissner2015).
Aside from the increase ability to robustly validate targets such as SL biosynthesis, global collaboration between academia and pharmaceutical partners is expediting the process of drug discovery of new anti-protozoal drugs. For example, within the sphere of targeting SL biosynthesis in the protozoa, we have managed several projects with industrial partners, MRCT and Tres Cantos Open Lab Foundation (https://www.openlabfoundation.org, an initiative of GlaxoSmithKline), in the pursuit of identifying new compound scaffolds active against the Leishmania spp IPC synthase utilising yeast (Norcliffe et al. Reference Norcliffe, Alvarez-Ruiz, Martin-Plaza, Steel and Denny2014) as a vehicle for drug discovery (Denny & Steel, Reference Denny and Steel2015). The generated results and techniques could readily be translated to other disease targets. Other global initiatives include Open Innovation Drug Discovery, Eli Lilly, which is focused on cancer, cardiovascular disease, endocrine disorders, neuroscience and tuberculosis. The Centers for Therapeutic Innovation, facilitates Pfizer and academic researchers to work together in order to develop new biologics programs and WIPO Re:Search, provide participant researchers with access to patents and expertise related to drug discovery for 19 NTDs, malaria and tuberculosis (Sheridan, Reference Sheridan2011).
Finally, SL biosynthesis represents a gold mine for new drug targets alongside at least two axes, de novo synthesis and salvage and remodelling. On one hand, the protozoan de novo SL biosynthetic pathway comprises three key steps, and considering their divergence compared with the mammalian host, identifying specific inhibitors for those could open an opportunity for anti-protozoal drugs with synergistic effects and lower incidences of resistance. On the other hand, the nature of obligate intracellular parasites dictates that further efforts should be directed towards the catabolic/salvage pathway where parasite–host dependencies could be exploited in order to identify additional key steps, or host enzymes, where inhibitors would exert further synergism with the de novo inhibitors.
To summarize, the landscape of anti-protozoan drug discovery requires immediate attention: with the re-evaluation of knowledge gained, the application of recent technologies; and the support of coordinated global discovery efforts. The multifaceted effects of SLs as a dynamic matrix of interaction (spatial and temporal) and function makes SL biosynthesis highly alluring for drug intervention, after all, everybody needs SLs, right?
ACKNOWLEDGMENTS
We thank Dr Ehmke Pohl for helpful discussions.
FINANCIAL SUPPORT
JGM and PWD are supported by the Biotechnology and Biological Research Council (BB/M024156/1 and NPRONET awards).




