Introduction
Site-specific weed management (SSWM) involves adapting weed management strategies to match weed variation within a given field (Fernández-Quintanilla et al. Reference Fernández-Quintanilla, Peña, Andújar, Dorado, Ribeiro and López-Granados2018). In agriculture, weeds often grow in distinct patches rather than uniformly across a field (Maxwell and Luschei Reference Maxwell and Luschei2005); as a result, broadcast herbicide applications often treat areas of the field where no weeds are present. In theory, using SSWM could result in increased herbicide savings, decreased herbicide expenses, and decreased environmental contamination (Arsenijevic et al. Reference Arsenijevic, de Avellar, Butts, Arneson and Werle2021; Barroso et al. Reference Barroso, Fernandez-Quintanilla, Maxwell and Rew2004; dos Santos Ferreira et al. 2019). An additional benefit is that SSWM could allow for the economical application of multiple herbicide mechanisms of action (MOAs) so that more expensive chemistries could be applied only where needed. Not only would this be less expensive for the farmer, but applications with diversified MOAs help to slow the development of herbicide-resistant weeds (Evans et al. Reference Evans, Tranel, Hager, Schutte, Wu, Chatham and Davis2015). In modern agriculture, successful weed control can be very difficult due to increases in herbicide-resistant weed cases, rising costs of herbicides, and shortages of crop protection products brought on by economic consequences of the COVID-19 pandemic (Dayan Reference Dayan2021; Mordor Intelligence 2022). As such, strategies such as SSWM that aim to reduce input quantities and costs could potentially benefit farmers and contribute to the sustainability of cropping systems around the world (Bongiovanni and Lowenberg-Deboer Reference Bongiovanni and Lowenberg-Deboer2004).
The key component to SSWM involves the accurate detection of weed positions within a given field, but the development of a robust and accurate detection system for field conditions remains a challenge (Gao et al. Reference Gao, French, Pound, He, Pridmore and Pieters2020). One of the ways this challenge is being addressed is by applying artificial intelligence using convolutional neural networks (CNNs). Convolutional neural networks are a type of deep neural network that excel at pattern recognition and can be utilized in a variety of tasks ranging from image analysis to audio file analysis (Albawi et al. Reference Albawi, Mohammed and Al-Zawi2017). The most common use of CNNs in the agricultural sector involves image analysis; CNNs analyze the textural, spectral, and spatial features of images and can extract features unseen by the human eye (Albawi et al. Reference Albawi, Mohammed and Al-Zawi2017; Sapkota et al. Reference Sapkota, Singh, Cope, Valasek and Bagavathiannan2020). Fruit counting, weed detection, disease detection, and grain yield estimation are ways that CNNs have been used in agriculture (Biffi et al. Reference Biffi, Mitishita and Liesenberg2021; Hussain et al. Reference Hussain, Farooque, Schumann, McKenzie-Gopsill, Esau, Abbas, Acharya and Zaman2020, Reference Hussain, Farooque, Schumann, Abbas, Acharya, McKenzie-Gopsill, Barrett, Afzaal, Zaman and Cheema2021; Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020; Yang et al. Reference Yang, Shi, Han, Zha and Zhu2019).
Two approaches to documenting and treating weeds in the field are real-time in situ weed detection and herbicide application and scouting and developing weed maps to guide SSWM (Cardina et al. Reference Cardina, Johnson and Sparrow1997; Somerville et al. Reference Somerville, Sønderskov, Mathiassen and Metcalfe2020). In situ weed detection involves recognizing weeds in real time and can lead to plants being treated in a timelier manner. Platforms that have been developed to detect weeds in situ include “smart” sprayers, autonomous weeding robots, and unmanned aerial spraying vehicles (Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020), most of which use some type of CNN technology. CNNs have been shown to be accurate in tasks such as segmentation (dividing images into regions based on pixel similarities), image classification (assigning a label to an image based on the objects present), and object detection (identifying objects within an image) (Biffi et al. Reference Biffi, Mitishita and Liesenberg2021; Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020; Stanford 2022; ThinkAutomation 2022). Object detection CNNs are typically at the forefront of in situ weed detection, as there is often greater value in detecting and localizing agricultural pests as opposed to assigning labels to images with pests located in them (Chen et al. Reference Chen, Lin, Cheng, Hung, Lin and Chen2021).
In recent years, open-source object detection algorithms have become available, such as those from the TensorFlow Object Detection API (Huang et al. Reference Huang, Rathod, Sun, Zhu, Korattikara, Fathi, Fischer, Wojna, Song, Guadarrama and Murphy2017), the “You Only Look Once” (YOLO) algorithm series (first introduced by Redmon et al. Reference Redmon, Divvala, Girshick and Farhadi2016), and the Detectron algorithm series (Lin et al. Reference Lin, Goyal, Girshick, He and Dollár2018). These object detectors have been used to implement a variety of computer vision tasks, including cancer cell detection (Al Zorgani et al. Reference Al Zorgani, Mehmood and Ugail2022), facial recognition (Mattman and Zhang Reference Mattman and Zhang2019), underwater fish detection (Xu and Matzner Reference Xu and Matzner2018), and projects related to the development of self-driving vehicles (Kulkarni et al. Reference Kulkarni, Dhavalikar and Bangar2018). Open-source algorithms are typically pretrained on very large data sets, such as the Microsoft COCO (Common Objects in Context) data set (Lin et al. Reference Lin, Maire, Belongie, Bourdev, Girshick, Hays, Perona, Ramanan, Zitnick and Dollár2014). Through utilizing a process called transfer learning, pretrained algorithm parameters can be fine-tuned to detect custom objects. Transfer learning involves using information learned from one object detection algorithm and applying this information to identify different, yet related, objects (Ghazi et al. Reference Ghazi, Yanikoglu and Aptoula2017). This eliminates the need to train algorithms from scratch, which is a very computationally expensive and time-consuming process (Ruder Reference Ruder2021). Open-source algorithms fine-tuned to identify agricultural crops and weeds have been used in a variety of studies, including late-season species detection in soybean [Glycine max (L.) Merr.] of Palmer amaranth (Amaranthus palmeri S. Watson), waterhemp (Amaranthus tuberculatus (Moq.) Sauer], common lambsquarters (Chenopodium album L.), velvetleaf (Abutilon theophrasti Medik.), and Setaria spp. (Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020); detection of wild radish (Raphanus raphanistrum L.) and capeweed [Arctotheca calendula (L.) Levyns] in barley (Hordeum vulgare L.) (Thanh Le et al. Reference Thanh Le, Truong and Alameh2021); and weed detection in a variety of crops, including lettuce (Lactuca sativa L.) (Osorio et al. Reference Osorio, Puerto, Pedraza, Jamaica and Rodriguez2020), carrots (Daucus carota L. var. sativus Hoffm.) (Ying et al. Reference Ying, Xu, Zhang, Shi and Liu2021), corn (Zea mays L.) (Ahmad et al. Reference Ahmad, Saraswat, Aggarwal, Etienne and Hancock2021), and onions (Allium cepa L.) (Parico and Ahamed Reference Parico and Ahamed2020).
For this study, we focused on detecting A. palmeri in soybean-cropping systems using open-source object detection algorithms. Amaranthus palmeri has been designated the most problematic weed in the United States (WSSA 2016), and it can reduce soybean yields by as much as 68% (Klingaman and Oliver Reference Klingaman and Oliver1994; Kumar et al. Reference Kumar, Liu, Peterson and Stahlman2021). Therefore, controlling this weed is very important for United States soybean producers. Large numbers of training images are necessary to train object detection algorithms to identify custom objects (Pokhrel Reference Pokhrel2020); however, nonproprietary image databases of A. palmeri are often unavailable. In addition, even though algorithms have been previously trained on A. palmeri in the midwestern United States (Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020), we did not find many studies that investigated the relationship of model evaluation metrics as influenced by A. palmeri growth features, including canopy diameter, plant height, percent ground cover, and weed density. Understanding the relationship between algorithm evaluation metrics and A. palmeri growth features could benefit precision weed applications. For example, future databases could focus on collecting images of A. palmeri plants with growth features best detected by the algorithm. In addition, farmers and agricultural professionals could gain a better understanding of which field conditions would benefit the most from deploying these algorithms. For instance, A. palmeri infestations with large plants and high populations may not be the best environments to use this technology for site-specific applications.
We hypothesized that as weed diameter and height increase, object detection algorithms will be better able to identify A. palmeri plants; however, as A. palmeri density and ground cover increases, ability to identify will decrease. Object detection algorithms can have difficulty both detecting small objects (Li et al. Reference Li, Liang, Wei, Xu, Feng and Yan2017) and detecting all object occurrences if objects are present in high densities in an image (Sun et al. Reference Sun, Dai, Zhang, Chang and He2022). The specific objectives of this study were (1) to develop an annotated image database of A. palmeri and soybean with multiple weed densities and soybean row spacings that can be used to fine-tune object detection algorithms, (2) compare multiple open-source algorithms’ effectiveness in detecting A. palmeri, and (3) evaluate the relationship between A. palmeri growth features (diameter, height, density, and ground cover) and A. palmeri detection ability.
Materials and Methods
Image Acquisition
To establish conditions representative of multiple A. palmeri densities and soybean-cropping systems, field locations were identified at the Kansas State University, Department of Agronomy Ashland Bottoms Research Farm near Manhattan, KS (39.122°N, 96.635°W) and at the Lund Research Farm near Gypsum, KS (38.797°N, 97.448°W) in 2021. At each location, 24 plots of soybeans were planted at a seeding rate of 331,000 seed ha−1: 12 plots were planted at 38-cm-wide row spacing, and 12 plots were planted at 76-cm-wide row spacing; plot dimensions were 3.1-m wide and 9.1-m long. Both field sites had a naturally occurring population of A. palmeri that was allowed to germinate and grow with the soybeans. These field locations allowed multiple densities of A. palmeri to be photographed while growing among soybean in different row spacings, providing a greater diversity of field situations to be “seen” by each algorithm.
The training database was built with 1,500 images taken of A. palmeri only, soybean only, or both species between May 27 and July 27 (Table 1). Imagery was taken both with a TG-610 handheld camera (OM Digital Solutions, Hachioji-City, Tokyo, Japan), and with a DJI Inspire 1 unmanned aerial vehicle (UAV) equipped with a Zenmuse X5R RAW camera (DJI, Shenzhen, China). The TG-610 has a sensor size of 28 mm2, whereas the Zenmuse X5R has a larger sensor size of 225 mm2. To increase the variability of the photographed vegetation, aboveground altitudes at which the images were taken varied from 1.5 m to 8 m, often with the minimum height chosen on any given day determined by vegetation height. For example, as the plants increased in height, it became necessary to increase the UAV flight altitude to prevent propeller downdraft from collapsing the plants. To add an additional source of variability, images were also collected under a variety of lighting conditions.
Dates, number of images, platform used, and height above ground for image collection at Manhattan and Gypsum, KS, field locations in 2021.

a HH, handheld Olympus TG-510 digital camera; UAV, unmanned aerial vehicle with Zenmuse camera.
b Total pixel dimensions per image: A = 4,608 × 3,456; B = 4,288 × 3,216.
Field-collected Data
To model algorithm evaluation metrics related to A. palmeri growth features, plant height, canopy diameter, and density were taken weekly between the middle rows of each plot from 1 to 4 wk after planting (WAP) and 1 to 5 WAP for the Manhattan and Gypsum plots, respectively. Amaranthus palmeri density was measured each week in a 0.25-m2 quadrat placed at random within these rows. A total of four height and diameter measurements were recorded from random plants within the quadrats. Within this study, A. palmeri height was defined as the tallest measurable structure, and diameter was defined as the widest portion of the plant within the top 20 cm. In plots with a total of fewer than four A. palmeri plants observed, height and diameter measurements corresponded to the total number of observable plants. Data were taken on A. palmeri plants after the formation of the first true leaf and any growth stage afterward, as cotyledons proved to be too difficult to annotate on the image data set. On these same dates, five photos were taken within the middle of each plot with the handheld camera, approximately 1.5 m above the canopy. Five photos were taken to provide a representative sample of the plot, as plots were 9.1 m in length. These images were kept separate from the training data set and were used to evaluate algorithm performance within each plot.
Image Processing and Data Annotation
Raw image outputs from the handheld and the Zenmuse cameras produced images with dimensions that were too large and would exceed processor memory capacity. To begin, every input image was cropped to dimensions of 2,880 × 2,880 pixels to remove the “fish-eye” effect that often accompanies aerial imagery (Gurtner et al. Reference Gurtner, Walker and Boles2007). Next, these images were tiled into smaller dimensions of 1,024 × 1,024 pixels using Python 3.9.7 (Python Software Foundation 2022) and the Pillow module (Clark Reference Clark2022). This allowed images large enough to retain features necessary for labeling, but small enough so as to not exhaust processor memory during training. Each input image was tiled into 20 new images of 1,024 × 1,024 pixels for a total of 30,000 images, which allowed for more images to be added to the training database. Images in which no plant features were visible or those of poor quality were simply discarded and not labeled; in all, 4,492 images were selected for labeling.
Images were labeled using the annotation tool LabelImg (Tzutalin 2015), which allows users to draw rectangular bounding boxes around objects within imagery and assign classes to each box. In many cases, the presence of multiple classes of objects can lead to better detection results due to the presence of multiple feature gradients (Oza and Patel Reference Oza and Patel2019). Therefore, we chose to annotate two classes for this study: A. palmeri and soybean. Using the same methods described by Sivakumar et al. (Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020), bounding boxes were drawn over both individual and patches of A. palmeri and soybean. In the event of irregularly shaped plants or patches of plants, multiple bounding boxes were drawn to encompass the entirety of the plant features. Labeling partial sections of irregularly shaped plants has been shown to be beneficial to object detectors (Sharpe et al. Reference Sharpe, Schumann and Boyd2018, Reference Sharpe, Schumann and Boyd2020a; Zhuang et al. Reference Zhuang, Li, Bagavathiannan, Jin, Yang, Meng, Li, Li, Wang, Chen and Yu2022), so these irregular features were not ignored. In any given image, both plant species could be present, so they were labeled accordingly. Amaranthus palmeri plants were labeled at all growth stages ranging from the formation of the first true leaf through inflorescence, and soybeans were labeled from the VE-VC stage through the R2 stage (Fehr et al. Reference Fehr, Caviness, Burmood and Pennington1971) (Figure 1). Amaranthus palmeri plants visible within each labeled image ranged from no plants (only soybean labeled) to roughly 115 labeled specimens. Throughout the image data set, soybean growth stage was homogeneous on each plant date, but A. palmeri could have multiple growth stages visible due to its ability to emerge throughout the growing season (Jha and Norsworthy Reference Jha and Norsworthy2009; Shyam et al. Reference Shyam, Chahal, Jhala and Jugulam2021). Figure 2 illustrates the image labeling process.
Examples of images collected for soybean in the VE-VC (A) and R2 (B) growth stages.

Illustration of the annotation process. Amaranthus palmeri and soybean plants are labeled in this figure with orange and white boxes, respectively. Bounding boxes overlap with neighboring bounding boxes when plant features are irregular. In cases where a single bounding box could not encompass a plant without including a plant of another species, multiple irregular bounding boxes were drawn on a single specimen.

The images that were selected for labeling contained a total of 10,494 and 10,312 A. palmeri and soybean annotations across all growth stages, respectively. The data set was then divided into 90% training and 10% test images, used to train and evaluate the algorithms, respectively. The training and test data sets consisted of 4,042 and 450 images, respectively. The aforementioned images taken over the plots for analysis of A. palmeri growth features and algorithm evaluation metrics were not included in the training or testing data sets but were kept separate for further analysis.
Algorithm Selection
Three open-source algorithm architectures were used in this study: faster regional convolutional neural network (Faster R-CNN), single-shot detector (SSD), and two YOLO algorithms. Faster R-CNN algorithms are a modern development of what are called regional-based CNNs, first proposed by Girshick et al. (Reference Girshick, Donahue, Darrell and Malik2014). This approach was revolutionary, in that it was one of the first large-scale successful approaches to addressing the task of object localization and detection (Balasubramanian Reference Balasubramanian2021). As input images are fed into the algorithm, areas of interest based on groups of pixels are extracted from the image and fed into the neural network (Oinar Reference Oinar2021). The architecture has been updated with the development of Fast R-CNN in 2015 (Girshick Reference Girshick2015) and finally with Faster R-CNN in 2016 (Ren et al. Reference Ren, He, Girshick and Sun2017), with each version being faster in detection speed than the previous. Faster R-CNN is known as a two-stage object detector, in that it first extracts regions of interest where it is likely that the objects will be and then classifies these regions of interest (Du et al. Reference Du, Zhang and Wang2020). Consequently, Faster R-CNN is known to perform better in terms of detection accuracy but has slower detection speeds (Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020).
SSD and YOLO algorithms were proposed by Liu et al. (Reference Liu, Anguelov, Erhan, Szegedy, Reed, Fu and Berg2016) and Redmon et al. (Reference Redmon, Divvala, Girshick and Farhadi2016), respectively. Considered single-stage object detectors, they are generally faster and less computationally expensive than Faster R-CNN algorithms (Liu et al. Reference Liu, Anguelov, Erhan, Szegedy, Reed, Fu and Berg2016), allowing for faster detection and suitable for real-time detection applications. Instead of extracting regions of interest as R-CNN algorithms do, they accomplish object localization and classification in one forward pass of the neural network (Forson Reference Forson2017). As in Sivakumar et al. (Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020), Faster R-CNN was chosen for this project due to its detection performance, whereas SSD and YOLO algorithms were chosen due to their inference speeds.
The backbone models refer to the specific neural networks behind the architectures and allow for feature extraction from the input image (Shimao 2019). These networks are interchangeable, with multiple networks able to be used as backbone models (Li et al. Reference Li, Wang, Xie, Liu, Zhang, Li, Wang, Zhou and Liu2020). For this study, the Faster R-CNN architecture backed with the ResNet (He et al. Reference He, Zhang, Ren and Sun2016) network was chosen with multiple layers, including ResNet-50, ResNet-101, ResNet-152, and Inception ResNet-V2 (Szegedy et al. Reference Szegedy, Ioffe, Vanhoucke and Alemi2017). Additionally, ResNet-50 and ResNet-152 were also chosen as backbone models for the SSD architecture.
For the YOLO algorithms, YOLOv4 (Bochkovskiy et al. Reference Bochkovskiy, Wang and Liao2020) and YOLOv5 (Jocher et al. Reference Jocher, Stoken and Borovec2020) were used, both running on the Cross Stage Partial (CSP) Darknet53 (Bochkovskiy et al. Reference Bochkovskiy, Wang and Liao2020) network. YOLOv5 also implements a Path Aggregation Network, allowing for both increased propagation of lower-level features and improvements in using localization signals (Carlos and Ulson Reference Carlos and Ulson2021). This allows for an increase in accuracy when localizing an object (Carlos and Ulson Reference Carlos and Ulson2021). Additionally, the YOLOv5 algorithm consists of four releases: YOLOv5s, YOLOv5m, YOLOv5l, and YOLOv5x (ultralytics 2022b). YOLOv5x was chosen for this experiment, as it is considered the most accurate object detector of the four (Carlos and Ulson Reference Carlos and Ulson2021). Both YOLO algorithms were obtained from their respective GitHub repositories (Alexey Reference Alexey2022; Jocher Reference Jocher2022a). YOLO algorithms down-sample input images by a factor of 32 when training (Hui Reference Hui2018), so input images with width and height dimensions divisible by 32 are necessary. Our input image dimensions of 1,024 × 1,024 pixels fit this criterion. All Faster R-CNN and SSD algorithms were obtained from the TensorFlow 2 Detection Model Zoo (TensorFlow 2021); the respective algorithms with input dimension requirements of 1,024 × 1,024 pixels were chosen. All YOLO and TensorFlow models selected were pretrained, thus eliminating the need to train from scratch (Ruder Reference Ruder2021).
Training
All algorithms were trained on a virtual Ubuntu 18.04 computer available on Paperspace, a virtual machine learning platform (Paperspace Cloud Computing, https://www.paperspace.com). The computer was equipped with an Intel® Xeon® E5-2623 v4 processor (Intel Technologies, Santa Clara, CA, USA) equipped with 16 CPU cores and 60 GB of RAM. To increase training speed, training was done utilizing a NVIDIA P6000 Graphics Processing Unit (GPU) with 24 GB of RAM (NVIDIA, Santa Clara, CA, USA).
For all algorithms, the default training hyperparameters were accepted, except for the learning rates for both the TensorFlow algorithms and batch sizes for all algorithms. As algorithm loss was monitored during training, learning rate had to be lowered below default settings for most Faster R-CNN and SSD algorithms due to an issue with exploding gradients. When doing so, we also lowered the warm-up learning rate to a value below the learning rate to avoid errors during training. As input images were large, batch sizes were reduced to prevent exhausting the GPU’s memory capacity. Larger batch sizes were possible with smaller algorithms (i.e., Faster R-CNN ResNet-52 and Faster R-CNN ResNet-101), but batch sizes had to be reduced for larger algorithms (i.e., Faster R-CNN Inception ResNet-V2) to avoid the ResourceExhaustedError (TensorFlow 2017) indicating that the GPU was out of memory. Because batch size has been said to not be a significant factor in affecting algorithm performance (Ghazi et al. Reference Ghazi, Yanikoglu and Aptoula2017), we did not expect this to affect the outcome of our algorithms. Algorithm training information is presented in Table 2.
Training information and hyperparameters used in this study.

a Faster R-CNN, faster regional convolutional neural network; SSD, single-shot detector; YOLO, “You Only Look Once.”
During the image annotation process, all annotations were saved in Pascal VOC format. Pascal VOC format is compatible with TensorFlow algorithms, but not with YOLO algorithms. Therefore, before training the YOLO algorithms, copies of the annotations were saved in a separate folder and converted to YOLO format using Python 3.9.7. The script that was used can be obtained on the Convert PascalVOC Annotations to YOLO GitHub website (vdalv 2017).
All algorithms were trained for the default 100,000 steps, except for the Faster R-CNN Inception Resnet-V2, YOLOv4, and YOLOv5 algorithms. The default number of training steps for the Faster R-CNN Inception ResNet-V2 algorithm is 200,000, but upon monitoring the loss, it was determined that no further increases in algorithm training were being made, and training was terminated early. YOLOv4 training involves iterations, thereby defining a batch size before training and an iteration as complete when the algorithm has processed the number of images specified in the batch size. Finally, YOLOv5 output metrics were reported after each completed epoch, which is defined as one iteration through the entire training data set (Brownlee Reference Brownlee2018). Upon viewing an output of the evaluation metrics for each epoch, we terminated algorithm training after 41 epochs, and the best weights were automatically saved for analysis.
Image augmentation is an important aspect of model training, as it allows for a more comprehensive set of images to be passed through the algorithm and reduce overfitting (Shorten and Khoshgoftaar Reference Shorten and Khoshgoftaar2019). Each of the algorithms contained code to automatically augment images during algorithm training, according to each algorithm’s default settings. The Faster R-CNN architecture augmentations included random horizontal flips, hue adjustments, contrast adjustments, saturation adjustments, and random image cropping. The SSD architectures used random horizontal flips and random image cropping augmentations. For the YOLO algorithms, the YOLOv4 augmentations included random image saturation, exposure, and hue adjustments (Alexey Reference Alexey2020). Finally, the YOLOv5 model used random mosaicking. This process involves combining an input image with three random images from the training data set. The new mosaic is then passed through the algorithm for training (Jocher Reference Jocher2022b).
Algorithm Evaluation and Statistical Analysis
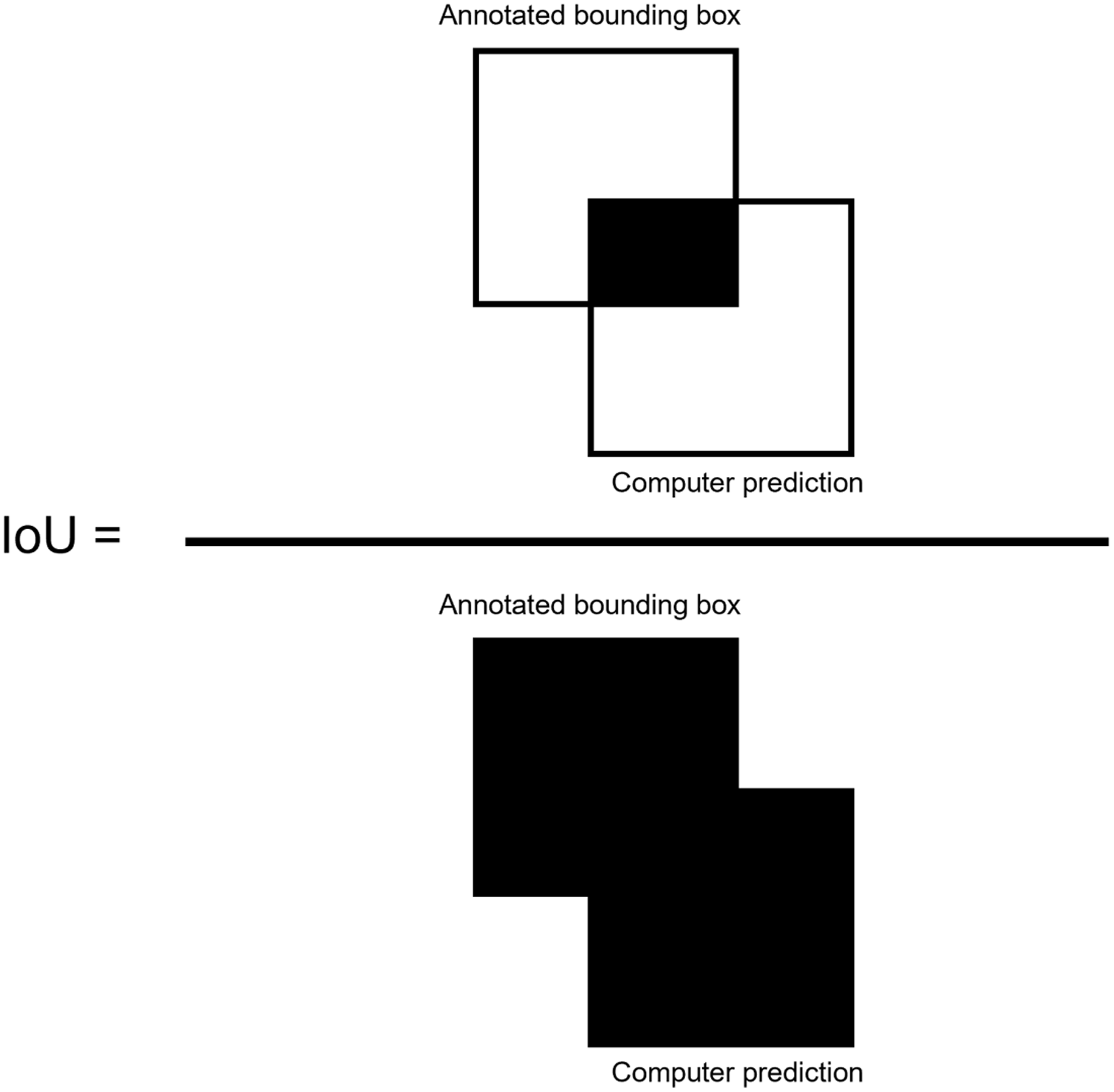
To measure overall performance of the algorithms, the metrics of precision, recall, and F1 score were computed for the test data set (Shung Reference Shung2018). Using an intersect over union (IoU) threshold of 0.5 (50%) between the annotated objects and the predicted bounding boxes (Henderson and Ferrari 2017), true-positive (TP) and false-positive (FP) detections are determined. IoU is defined as the overlap between the ground truth bounding boxes drawn during annotation and the predicted bounding box determined by the computer (Jin et al. Reference Jin, Sun, Che, Bagavathiannan, Yu and Chen2022), divided by the total area of each bounding box (Figure 3). IoU values greater than or equal to 0.5 were considered TP, and values less than 0.5 were considered FP predictions (Henderson and Ferrari 2017).
Intersection over union (IoU) equation, defined as the overlap between the ground truth annotation and the computer prediction bounding box, divided by the total area of the two bounding boxes. IoU overlaps greater than 0.5 were considered true-positive predictions, whereas overlaps less than 0.5 were considered false-positive predictions.
Precision is the ratio between the number of TP predictions and the total number of positive predictions, with the lowest value being 0 and the highest value being 1 (Hussain et al. Reference Hussain, Farooque, Schumann, Abbas, Acharya, McKenzie-Gopsill, Barrett, Afzaal, Zaman and Cheema2021). Precision is reduced when an algorithm makes many incorrectly positive, or FP, predictions or a low number of TP predictions but is increased by larger numbers of correct predictions and low FP detections (Gad Reference Gad2021). Precision was computed with the following equation:
Recall, also referred to as the TP rate (Hussain et al. Reference Hussain, Farooque, Schumann, Abbas, Acharya, McKenzie-Gopsill, Barrett, Afzaal, Zaman and Cheema2021), is a measure of how well a given algorithm identifies TP predictions (Huilgol Reference Huilgol2020). Also ranging from 0 to 1, a higher recall indicates better TP predictions. Recall was computed as follows:
where FN denotes false-negative detections.
The F1 score is the harmonic mean between precision and recall (Zhong et al. Reference Zhong, Hu and Zhou2019), with the best score being 1 and the worst being 0 (Hussain et al. Reference Hussain, Farooque, Schumann, Abbas, Acharya, McKenzie-Gopsill, Barrett, Afzaal, Zaman and Cheema2021). It was calculated as:
The average precision (AP) for each class is determined by graphing a precision–recall curve for each test image for both classes and computing the area beneath each curve (Henderson and Ferrari 2017). The AP is then used to find the mean AP (mAP) of the algorithm, which was calculated from an IoU threshold of 0.5 in this study using the following equation (Jin et al. Reference Jin, Sun, Che, Bagavathiannan, Yu and Chen2022):
where N corresponds to the total number of object classes. Values for mAP range from 0 to 1, with higher values corresponding to larger areas beneath the curve. For this study, the algorithm with the largest mAP was selected to analyze the model evaluation metrics for the photos taken above the individual plots.
For each measurement date, an average for each plot was determined for all field-collected data (A. palmeri height, diameter, and density). Amaranthus palmeri coverage was computed by multiplying the average canopy area by the average density per plot, assuming a circular shape:
where T is the average total A. palmeri coverage (m2), r is the average A. palmeri radius (m), and n is the average weed density (plants m−2).
Each test image taken over the plots was passed through the best-performing algorithm, and precision, recall, and F1 scores were generated. Each evaluation metric was averaged within its respective plot. Statistical analysis was done using R v. 4.1.2 (R Core Team 2021). Regression models (see “Results and Discussion” section) were used to test whether field-measured data (A. palmeri density, height, and coverage) significantly predicted algorithm evaluation metrics. Data were combined over all collection dates and locations for this analysis, as regression assumptions were checked visually for each location and were determined to meet all assumptions (data not shown) (Osborne and Waters Reference Osborne and Waters2002). Best regression models were selected based on the Akaike information criterion (AIC) values, such that the model with the lowest AIC value was selected for each evaluation metric. AIC values and weights (indicating the total predicative power among all tested models) were found using the AICcmodavg package in R (Mazerolle Reference Mazerolle2020). Density and coverage were found to be highly collinear (data not shown), so these variables were never included in the same model together.
Results and Discussion
Algorithm Comparison
The results of algorithm training are presented in Figure 4. After training, it was shown that YOLOv5 achieved the highest overall mAP value of 0.77. YOLOv4 and Faster R-CNN ResNet50 both achieved acceptable results with mAP values of 0.70, followed by SSD ResNet152 and Faster R-CNN Inception ResNet(v2) with values of 0.68. In most cases, the mAP values for the single-stage detectors (YOLO and SSD) on this data set were equal or superior to those of the two-stage detectors. Faster R-CNN models are generally considered accurate object detectors (Sivakumar et al. Reference Sivakumar, Li, Scott, Psota, Jhala, Luck and Shi2020), but they can be sensitive to background noise and often have difficulty detecting small objects (Amin and Galasso Reference Amin and Galasso2017; Roh and Lee Reference Roh and Lee2017). Our test data set contained images of multiple A. palmeri and soybean growth stages, including very small plants of both species. This could explain why the single-stage object detectors often outperformed the Faster R-CNN models in this study. Additionally, we were not surprised to see the higher performances of YOLOv4 and YOLOv5 algorithms, as previous versions of YOLO have been reported to detect weeds faster than Faster R-CNN algorithms and with greater accuracy than SSD algorithms (Ahmad et al. Reference Ahmad, Saraswat, Aggarwal, Etienne and Hancock2021). Given these results, YOLOv5 was selected for further analysis.
Mean average precision (mAP) results of each model after training. YOLOv5 was considered the best-performing algorithm of each tested model with a mAP of 0.77.

The YOLOv5 algorithm in this study was trained for 41 epochs (Figure 5), taking approximately 15 h to complete. Training was monitored based on the changes in mAP per epoch. Algorithm training was terminated after mAP values were seen to “plateau,” thus indicating no further meaningful gains in algorithm performance were expected. During training, YOLOv5 saved the best-performing weights, which were used to compute all further algorithm evaluations. In the precision–recall curve for YOLOv5 (Figure 6), the AP of A. palmeri (0.788) is greater than that of soybean (0.756), indicating that detection was slightly better for A. palmeri than for soybean. This could be related to differences in the way that the species were annotated. As soybean increased in size, larger bounding boxes were drawn over multiple plants, as there was a high level of overlap between individual plants (Figure 7). Separating out the individual plants would have been both difficult and time-consuming. On the other hand, although there were some overlaps observed with A. palmeri, these were much less pronounced and allowed for more individual weed plants to be annotated. Therefore, individual A. palmeri plants were presumably easier for the YOLOv5 algorithm to identify.
Change in mean average precision (mAP) @ 0.5 over each epoch during training. mAP was reported after the completion of each epoch. Training was terminated after visual inspection of curve and when mAP @ 0.5 curve was seen to “plateau.”

Precision–recall curve for YOLOv5. Amaranthus palmeri achieved a slightly higher average precision (AP) (0.788) than soybean. Solid blue line represents mean average precision (mAP) computed on the test data set. The AP for each class and the mAP for the overall algorithm were representative of the area of the graph under each respective curve.
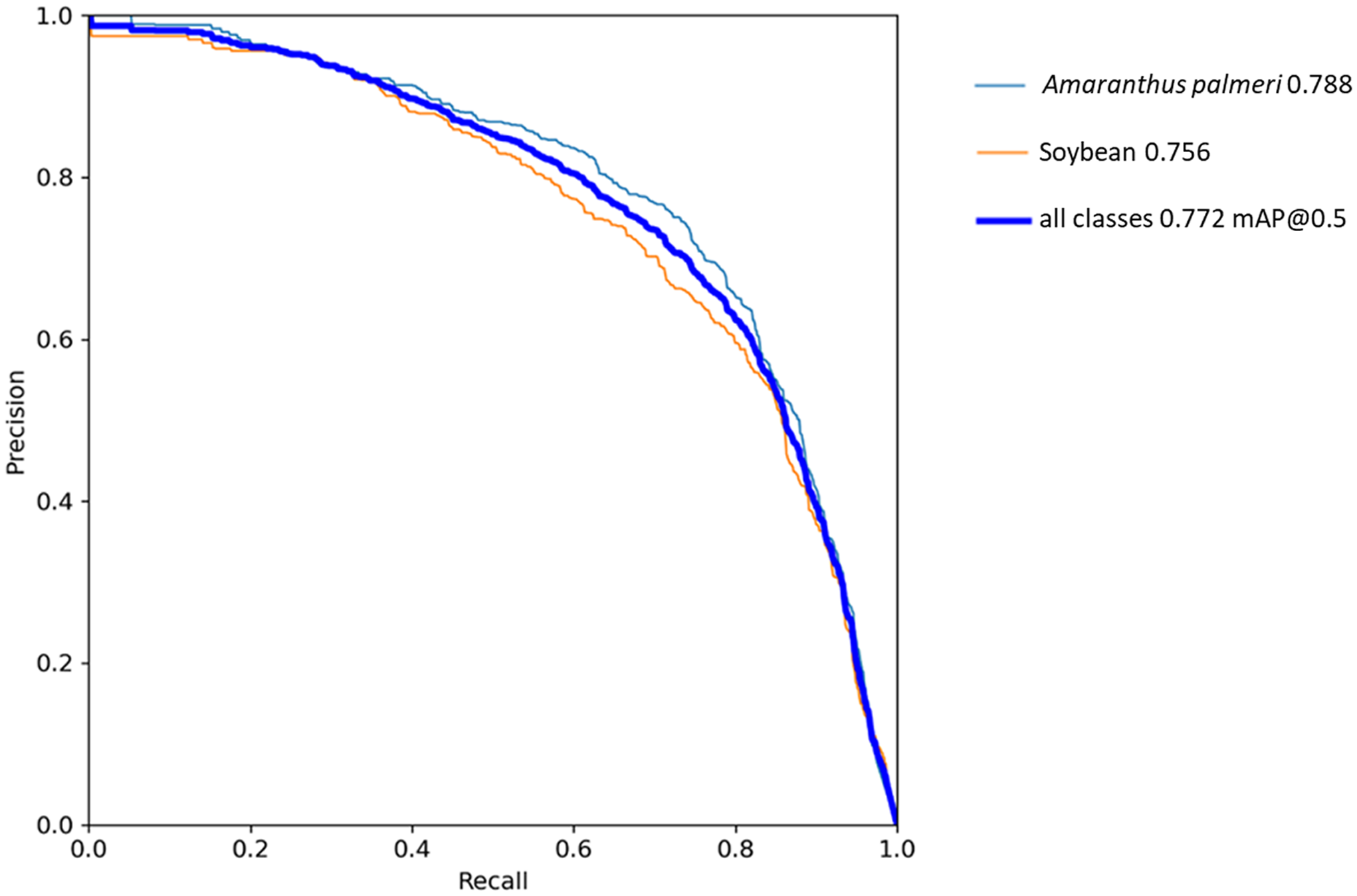
Image annotation of soybean at the R2 growth stage. As soybean populations were much higher than Amaranthus palmeri populations, there was a high level of soybean overlap. Therefore, it was necessary to include multiple soybean plants in each image. However, A. palmeri plants typically did not have as much overlap, and in most cases, it was much easier to identify and label individual plants.

When using YOLOv5 for detection, users can specify a confidence threshold as an input parameter to the detection script (ultralytics 2022a). This confidence threshold acts to limit the number of FP scores displayed in the final prediction (Wenkel et al. Reference Wenkel, Alhazmi, Liiv, Alrshoud and Simon2021). In our algorithm, lower confidence thresholds increased the likelihood of detecting either an A. palmeri or soybean plant, but the FP detection rates increased as a result. Figure 8 illustrates the F1 scores calculated on the test image data set. Interpretation of Figure 8 indicates that at a confidence threshold of 0.298, the highest F1 score (0.72) was achieved for both classes. This indicates that at this threshold, both precision and recall will be optimized for best detection results. As this confidence threshold is a recommended value, users still have the option to set the threshold to a value of their choosing.
F1 scores for YOLOv5 indicating the harmonic mean between precision and recall scores. Data indicated that detection results for both species would be best at a confidence threshold of 0.298.

There were differences between detections using lower (0.15) and higher (0.70) confidence thresholds (Figure 9). With multiple objects present in this image, YOLOv5 confidence in object detections ranged from low (0.36) to high (0.86); values are displayed immediately following the class prediction on each box. If this algorithm were to be used by a precision ground sprayer or a similar platform, we propose that using a lower confidence threshold would result in more A. palmeri plants being identified and treated. Consequently, the likelihood of soybean being incorrectly detected as A. palmeri would increase, but in soybean-cropping systems with herbicide-tolerant traits, this would not result in crop damage, assuming all current labels for such applications were followed. An increase in A. palmeri FP detections would likely lead to more herbicide being applied to the field. Interestingly, A. palmeri FP detections were not just limited to soybean; other broadleaf weeds such as carpetweed (Mollugo verticillata L.) and A. theophrasti were sometimes detected as A. palmeri with lower confidence intervals (Figure 10). This suggests that some weeds with features similar to A. palmeri plants would be sprayed if confidence intervals were lowered upon deployment. Regardless, further research is needed to determine which threshold would be optimal to reduce the volume of herbicides applied, while still achieving acceptable weed control.
YOLOv5 detection results for Amaranthus palmeri and soybean using confidence thresholds of 0.15 (A) and 0.70 (B). The likelihood of false-negative (FN) detections increases as confidence thresholds increase, as can be seen in B. Objects assigned a confidence interval of less than 0.70 are not detected in B. FN A. palmeri and soybean detections in B are indicated by the orange and white arrows, respectively.

Detection results for YOLOv5 with a confidence interval of 0.15. False-positive detections of Mollugo verticillata and Abutilon theophrasti as Amaranthus palmeri are denoted by arrows pointing from “A” and “B,” respectively.

Modeling YOLOv5 Evaluation Metrics
Only data for the A. palmeri class were used to model A. palmeri physical characteristics in relation to YOLOv5 evaluation metrics. For all regression models analyzed, the model P-values were significant, indicating that all ground-measured variables affected the evaluation metrics (Table 3). However, it was determined that model 5, which included the main effects and interaction effect of A. palmeri density and height, was the model that best fit the data. For precision, recall, and the F1 score, the model carried 72%, 86%, and 91% of the weights from the models that were compared, respectively. Therefore, this model was selected to describe the relationship between model evaluation metrics for density and height.
Regression models used to evaluate the effect of Amaranthus palmeri morphological parameters on model evaluation metrics and Akaike information criterion (AIC) used for model selection to detect A. palmeri. Bold type indicates that model 5 best fit the data.

a y em = evaluation metric (precision, recall, F1 score), x 1 denotes density (plants m−2), x 2 denotes height (cm), and x 3 denotes coverage (m2). β0, intercept; β1, coefficient for A. palmeri density (plants m−2); β2, coefficient for height (cm); β3, coefficient for coverage (m2); β4, interaction coefficient for density and height; β5, interaction coefficient for height and coverage; β6, coefficient for the square of density; β7, coefficient for the square of height; β8, coefficient for the square of coverage; β9, coefficient for the log of density; β10, coefficient for the log of height; β11, coefficient for the log of coverage.
b AIC weight, indicating the total predictive power among all tested models.
For all evaluation metrics analyzed with model 5, the interaction coefficient for density and height was significant (P = 0.049, 0.016, and 0.010 for precision, recall, and F1 score, respectively) (Table 4). As smaller weeds are generally more susceptible to control (Kieloch and Domaradzki Reference Kieloch and Domaradzki2011), we chose to model our results with four heights representing A. palmeri plants at early growth stages. The results of these predictions by the YOLOv5 algorithm indicated that smaller, younger A. palmeri plants growing at lower densities were detected better than taller plants (Figure 11). The results were as anticipated for density, but the prediction suggesting that YOLOv5 detection ability was greater for smaller plants was surprising, as species identification is often easier on larger plants with distinctive features. This may have been because there were a greater number of A. palmeri annotations of younger, smaller plants as opposed to larger, more mature plants. Additionally, larger plants have canopies that overlap with each other, making it difficult to distinguish and label large individual plants. Regardless, these predictions were encouraging, as algorithms that can detect smaller weeds are of more practical use, because weeds can be controlled much more easily when they are younger and smaller in size (Naghashzadeh and Beyranvand Reference Naghashzadeh and Beyranvand2015).
Linear regression results (model 5) for Amaranthus palmeri density (plants m−2) and height (cm) regressed against model evaluation metrics.

a Significant at: *P < 0.05; ***P < 0.001.
b RMSE, root mean-square error.
YOLOv5 precision (A), recall (B), and F1 score (C) changes as a function of Amaranthus palmeri density (plants m−2).

In relation to weed density affecting algorithm performance, our study conflicts with Yu et al. (Reference Yu, Schumann, Sharpe, Li and Boyd2020), in that they found that images with higher weed densities generally led to better algorithm detection results than those with lower weed densities. However, the study by Yu et al. (Reference Yu, Schumann, Sharpe, Li and Boyd2020) utilizes image classification rather than object detection. Rather than localizing the weeds within the image, the entire input image was classified with the weed species that were visible. Image classification for weed detection comes with some disadvantages, however, as the location of individual weeds was not provided, and multiple weed species within an image were not able to be detected (Ahmad et al. Reference Ahmad, Saraswat, Aggarwal, Etienne and Hancock2021). With object detection algorithms, object localization within the image allows for weeds to be located and controlled where they occur. However, we hypothesize that this application is best suited for postemergence applications (in the case of using precision herbicide application technology) where weed density is relatively low. Fields with very high weed densities would likely not benefit from a site-specific herbicide application, as the volume of herbicide needed for control would likely not be statistically different from a whole-field broadcast application. Further research is needed to determine the optimum weed density beyond which precision weed control has no economic or environmental benefits.
The overall precision, recall, and F1 scores computed for the 450 images in the test data set were 0.71, 0.70, and 0.71, respectively (data not shown). A precision of 0.71 indicates that the YOLOv5 algorithm was 71% accurate in successfully predicting A. palmeri and soybeans. Likewise, a recall of 0.70 indicates that the algorithm correctly predicted 70% of the plants belonging to either class (Jin et al. Reference Jin, Sun, Che, Bagavathiannan, Yu and Chen2022). These results were lower than previously reported by other YOLO weed detectors (Jin et al. Reference Jin, Sun, Che, Bagavathiannan, Yu and Chen2022; Zhuang et al. Reference Zhuang, Li, Bagavathiannan, Jin, Yang, Meng, Li, Li, Wang, Chen and Yu2022). However, our “test” data set consisted of images randomly selected from the large-input database and had a variety of A. palmeri growth stages and population densities. When evaluating YOLOv5 evaluation metrics on images taken within the plots with lower A. palmeri densities and shorter plant heights, the algorithm precision, recall, and F1 scores greatly improved. Based on the regression model fit to the data, A. palmeri plants 2-cm tall and growing at a density of 1 plant m−2 would be detected with precision, recall, and F1 scores of 0.87, 0.93, and 0.89, respectively.
YOLO algorithms have been used previously for weed detection. Ahmad et al. (Reference Ahmad, Saraswat, Aggarwal, Etienne and Hancock2021) achieved an overall mAP score of 0.543 when using YOLOv3 (Redmon and Farhadi Reference Redmon and Farhadi2018) to detect redroot pigweed (Amaranthus retroflexus L.), giant ragweed (Ambrosia trifida L.), common cocklebur (Xanthium strumarium L.), and green foxtail [Setaria viridis (L.) P. Beauv.]. Hussain et al. (Reference Hussain, Farooque, Schumann, McKenzie-Gopsill, Esau, Abbas, Acharya and Zaman2020) developed an in situ sprayer using both YOLOv3 and YOLOv3-tiny (Adarsh et al. Reference Adarsh, Rathi and Kumar2020) as backbone algorithms to detect C. album, achieving mAP scores of 0.932 and 0.782, respectively. Sharpe et al. (Reference Sharpe, Schumann, Yu and Boyd2020b) achieved good detection results when training the YOLOv3 algorithm to identify general classes of grasses, broadleaves, and sedge species; further, they found that including multiple classes (as opposed to a single class) in their algorithm increased precision, recall, and F1 metrics. Hu et al. (Reference Hu, Tian, Li and Ma2021) used YOLOv3 and YOLOv4 to detect 12 different weed species common to rice (Oryza sativa L.) and found that YOLOv4 achieved a mAP score that was 0.116 higher than YOLOv3. Our best mAP score was slightly lower than some of these YOLO weed detectors; however, it must be mentioned that this data set was collected with multiple cameras covering a variety of A. palmeri densities and growth stages. Data sets such as those collected by Jin et al. (Reference Jin, Sun, Che, Bagavathiannan, Yu and Chen2022) and Zhuang et al. (Reference Zhuang, Li, Bagavathiannan, Jin, Yang, Meng, Li, Li, Wang, Chen and Yu2022) consisted of a handheld camera taking multiple images at a consistent height. In this study, we collected imagery ranging from 1.5 m to 8 m above ground level. While many data sets collected consist of “ideal” specimens including plants grown in greenhouses or photos of individual plants, our data set was based on what a field sprayer or application UAV would observe in the field. As a result, during the labeling process, several overlapping bounding boxes had to be drawn, and it was impossible for each image to contain labels for individual plants.
In conclusion, this research demonstrated that YOLOv5, a free and open-source object detection algorithm, can detect A. palmeri in soybean-cropping systems. As site-specific herbicide applications become more widespread due to the potential for herbicide savings and environmental benefits, open-source algorithms such as YOLOv5 could enable increased development and adoption of precision weed detectors. Furthermore, this research suggests that our algorithm may be better at detecting smaller as opposed to larger A. palmeri plants. Upon further refinement and training of the algorithm, it may be of great use to growers, as smaller weeds are much more susceptible to control than larger ones.
Future research and improvements to our model will include adding more images to the data set. We included different imagery heights in this study to create a data set that could be utilized by multiple precision agriculture platforms such as precision ground sprayers and pesticide application UAVs. In the future, construction of specialized data sets that consist of imagery for each type of platform would be collected. For instance, imagery collected to train an algorithm for a precision ground sprayer should be at a height consistent with the sensors on the sprayer itself, that is, 60 cm above the target canopy. In this experiment, we trained the object detectors to identify two species (A. palmeri and soybean), and a future goal is to expand the number of weed species that can be detected by the YOLOv5 model. An increase in both the number of images and number of annotated weed species in these specialized data sets would increase the mAP of the YOLOv5 algorithm and reduce errors in object detection (Linn et al. Reference Linn, Mink, Peteinatos and Gerhards2019). Equal distribution of annotations among species is important when collecting these images. With further improvements to the algorithm, field tests will need to be carried out to both optimize weed detection and to treat weeds in real time using a precision ground sprayer with high-resolution cameras oriented close to plant canopies.
Acknowledgments
This research received no specific grant from any funding agency or the commercial or not-for-profit sectors. No conflicts of interest have been declared. This is contribution no. 23-042-J from the Kansas Agricultural Experiment Station.

















