Introduction
The crustose lichen genus Bacidia De Not. s. lat. is distributed worldwide and includes up to 230 species (Lücking et al. Reference Lücking, Hodkinson and Leavitt2017; Wijayawardene et al. Reference Wijayawardene, Hyde, Dai, Sanchez-Garcia, Goto and Magurno2022). Bacidia diversity is most comprehensively documented in Europe due mainly to the accessibility and long tradition of lichenology in the region. Species have been recorded in many European lichen floras, for example, in areas of Germany, the British Isles, and the Iberian Peninsula (e.g. Llop Reference Llop2007; Wirth et al. Reference Wirth, Hauck and Schultz2013; Cannon et al. Reference Cannon, Ekman, Kistenich, LaGreca, Printzen, Timdal, Aptroot, Coppins, Fletcher and Sanderson2021), and Bacidia diversity is also widely studied in North America (Ekman Reference Ekman1996). However, the diversity of Bacidia in Asia and the Caucasus remains largely unknown.
The Caucasus region harbours a rich, relict tertiary flora due to its unique environmental conditions that have remained stable for a long time. One of the first lichen checklists for the Caucasus region was made by Vainio (Reference Vainio1899) based on the Caucasian collection of Déchy and Lojka, which included several specimens of Bacidia s. lat. In the 20th century, extensive research on the Caucasus flora was carried out by Barkhalov (Reference Barkhalov1975, Reference Barkhalov1983), and by Vězda who worked particularly in the Caucasian reserve on the Black Sea coast and Abkhazia (Vězda Reference Vězda1983); these accounts also documented several species of Bacidia s. lat. More recently, Bacidia species have been reported in many lichenofloristic papers and checklists of the Caucasus covering northern (Urbanavichus & Urbanavichene Reference Urbanavichus and Urbanavichene2002, Reference Urbanavichus and Urbanavichene2017b, Reference Urbanavichus and Urbanavichene2018; Urbanavichene & Urbanavichus Reference Urbanavichene and Urbanavichus2019; Urbanavichus et al. Reference Urbanavichus, Urbanavichene, Vondrák and Ismailov2021), north-western (Otte Reference Otte2001, Reference Otte2004, Reference Otte2007a, Reference Otteb; Blinkova & Urbanavichus Reference Blinkova and Urbanavichus2005; Urbanavichus & Urbanavichene Reference Urbanavichus and Urbanavichene2014, Reference Urbanavichus and Urbanavichene2017a), western (Urbanavichene & Urbanavichus Reference Urbanavichene and Urbanavichus2016; Urbanavichus et al. Reference Urbanavichus, Vondrák, Urbanavichene, Palice and Malíček2020), south-western (Urbanavichene & Urbanavichus Reference Urbanavichene and Urbanavichus2014), north-eastern (Urbanavichus & Ismailov Reference Urbanavichus and Ismailov2013), central (Urbanavichene & Urbanavichus Reference Urbanavichene and Urbanavichus2018), eastern (Ismailov et al. Reference Ismailov, Urbanavichus, Vondrák and Pouska2017), and southern (Harutyunyan et al. Reference Harutyunyan, Wiesmair and Mayrhofer2011; Alverdiyeva & Novruzov Reference Alverdiyeva and Novruzov2014; Gasparyan & Sipman Reference Gasparyan and Sipman2016; Inashvili et al. Reference Inashvili, Kupradze and Batsatsashvili2022) parts of the region.
Urbanavichus (Reference Urbanavichus2010) was the first to compile data on lichens known for the Russian territory (incorporating the Caucasus), including 18 species of Bacidia s. lat. in the checklist currently known from the Caucasus. In subsequent studies, nearly half of these species were transferred to other genera, such as Aquacidia, Bacidina, Bellicidia, Biatora, Bibbya, Catillaria, Scutula and Toniniopsis, and several new species were later described or recorded for the region (Urbanavichus & Urbanavichene Reference Urbanavichus and Urbanavichene2014; Urbanavichene & Urbanavichus Reference Urbanavichene and Urbanavichus2016; Kistenich et al. Reference Kistenich, Timdal, Bendiksby and Ekman2018; Malíček et al. Reference Malíček, Palice, Vondrák, Łubek and Kukwa2018; Cannon et al. Reference Cannon, Ekman, Kistenich, LaGreca, Printzen, Timdal, Aptroot, Coppins, Fletcher and Sanderson2021; Gerasimova et al. Reference Gerasimova, Urbanavichene, Urbanavichus and Beck2021a). In addition, several species have been synonymized and/or recognized as belonging to other genera, such as Arthrorhaphis, Haematomma, Lecania and Scoliciosporum (Davydov & Printzen Reference Davydov and Printzen2012; Gerasimova & Ekman Reference Gerasimova and Ekman2017). Yet, the taxonomic position of some species is still unknown, such as Bacidia freshfieldii (Vain.) Zahlbr., which appears to be closely related to Catillaria (Gerasimova & Ekman Reference Gerasimova and Ekman2017). As such, at the beginning of our investigation 11 species of Bacidia s. str. were known from the Caucasus: Bacidia absistens (Nyl.) Arnold, B. albogranulosa Malíček et al., B. arceutina (Ach.) Th. Fr., B. biatorina (Körb.) Vain., B. fraxinea Lönnr., B. herbarum (Stizenb.) Arnold, B. laurocerasi (Delise ex Dube) Zahlbr., B. polychroa (Th. Fr.) Körb., B. rosella (Pers.) De Not., B. rubella (Hoffm.) A. Massal. and B. suffusa (Fr.) A. Schneid. However, a revision of the genus integrating molecular and morphological analysis was needed to comprehensively document the diversity of Bacidia in the region. Therefore, our research aimed to investigate the diversity of Bacidia s. str. in the Caucasus by applying an integrative approach, including morphological, anatomical, and molecular analyses.
Material and Methods
Study area and sampling
The Caucasus is located between the Caspian and the Black Seas and is bounded on the north by Russia (Kumo-Manych Depression) and on the south by Georgia, Armenia and Azerbaijan (Brummitt et al. Reference Brummitt, Pando, Hollis and Brummitt2001). The climatic conditions of the Caucasus range from warm and moist in the western Colchic region to hot and dry (Kura valley) in the east, spanning eight of the ten oceanity levels of the Northern Hemisphere, compared to only four observed in the Alps, according to the system of Jäger (Reference Jäger1968).
The specimens of Bacidia were mainly collected in the National Parks and Nature Reserves of the Northern Caucasus (Russia), namely in and around the Nature Reserve Bol'shoy Tkhach and the Caucasian Biosphere Reserve (1999–2019), Utrish (2001–2020), Erzi (2018) and Samursky National Park (2017), but also in Georgia (2012, 2015), Azerbaijan (2013) and Armenia (2015).
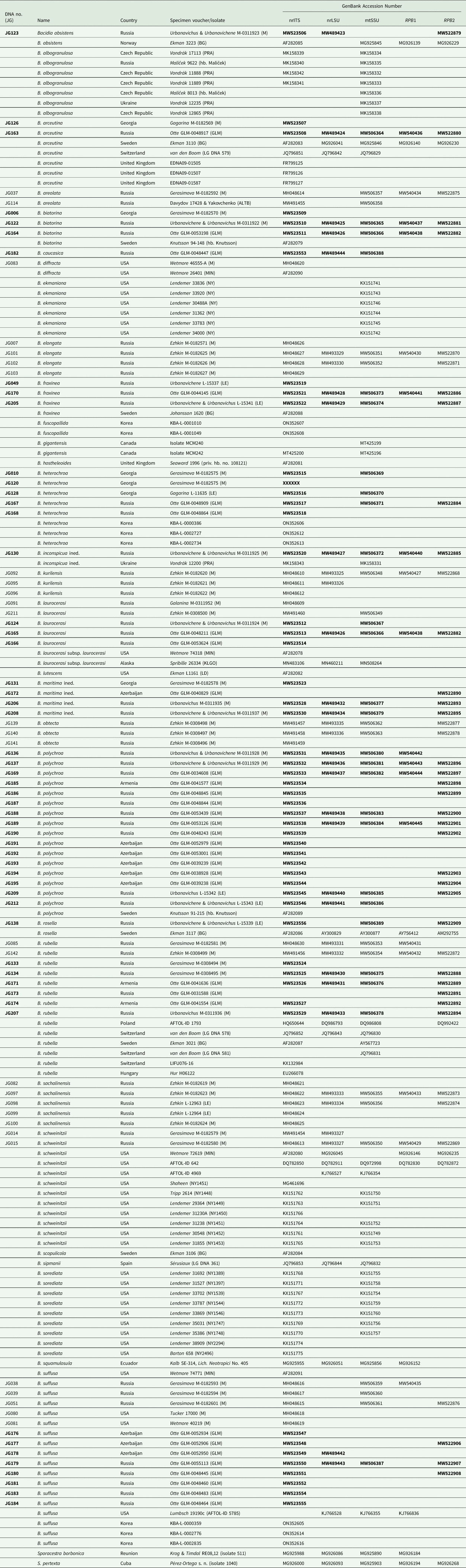
We studied the morphology of 237 specimens of Bacidia s. str. collected in the Caucasus (Supplementary Material File S1, available online). Of these, we obtained molecular sequences from 54 specimens belonging to 10 species and two putatively introduced, provisional taxa (B. inconspicua ined. and B. maritima ined.) of Bacidia s. str. that were representative of the known species diversity and inter- and infra-specific morphological variability. The only exception to this was B. herbarum, which we were not able to include in the molecular analysis as it is known only from one herbarium specimen (GLM-L-0054141, collected in Krasnodarskiy Krai at 2115 m a.s.l.). The specimens sampled for molecular analysis were collected in the northern part of the Caucasus (64.2%), Azerbaijan (20.7%), Georgia (9.4%), and Armenia (5.7%) from the bark of various phorophytes (Table 1).
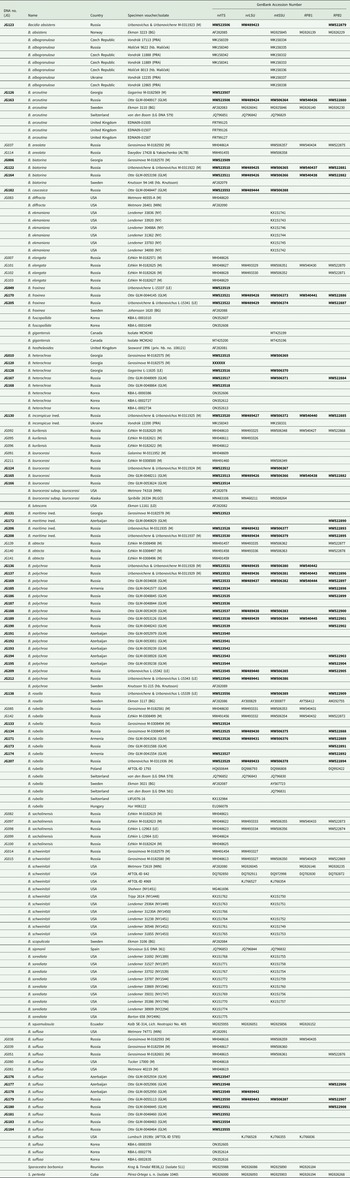
DNA numbers and specimen information for Bacidia species used in this study, with their respective GenBank Accession numbers. New sequences are in bold.
Morphology
Microscopic observations were made using a Zeiss Axioplan (Oberkochen, GmbH) light microscope equipped with differential interference contrast (DIC). Cross sections of apothecia were made on a Leica Jung Histoslide 2000 Mikrotom (Heidelberg, GmbH), with a thickness of 8–10 μm. Micrographs of cross-sections were taken on a Zeiss Axioplan with an attached AxioCam 512 Color camera, and images were processed with Zeiss ZEN v. 2.3 (blue edition). Macrographs of external characters were taken on a Leica Z6 Apo microscope (with a 2.0× Planapo lens; Leica, Germany) with a Sony Alpha 6400 camera (Sony, Japan) attached and equipped with a Stack Shot Rail macro rail (Cognisys, USA). A single image was mounted from 30–40 serial images using Helicon Focus v. 7 (Helicon, USA).
Measurements are given as (min–) x̄ ± SD (–max) (SD = standard deviation, n 1 = number of all observations, n 2 = number of specimens observed). We provide a detailed description of specimens using traditional microscopic techniques following Smith et al. (Reference Smith, Aptroot, Coppins, Fletcher, Gilbert, James and Wolseley2009) and the subdivision scheme of the proper exciple according to Ekman (Reference Ekman1996), differentiating the following structures: rim, lateral part, and medullary part. We used the following diagnostic characters to delimit the species lineages: 1) thallus structure; 2) colour of disc and margin of apothecium; 3) hypothecium colour; 4) colour and structure of exciple; 5) shape and size of ascospores. The standard reagents were used to study the colour reaction of the apothecia cross-sections and crystals solubility: a solution of 10% potassium hydroxide (KOH) in water, abbreviated K, and 50% solution of nitric acid (HNO3), abbreviated N. Pigment characterizations follow Meyer & Printzen (Reference Meyer and Printzen2000).
DNA extraction, PCR amplification and DNA sequencing
DNA extraction was carried out using the Stratec Invisorb Spin Plant Mini Kit (Stratec Molecular GmbH, Berlin) following the manufacturer's instructions. Five to eight apothecia were used from fresh material not older than five years, and thallus fragments were removed to minimize the risk of contamination by, for example, lichenicolous fungi. The same five target loci (three RNA-coding genes (nrITS, nrLSU, and mtSSU) and two protein-coding genes (RPB1 and RPB2)) as in Gerasimova et al. (Reference Gerasimova, Urbanavichene, Urbanavichus and Beck2021a) have been selected for PCR; amplification, purification and sequencing were performed as described in Gerasimova et al. (Reference Gerasimova, Ezhkin and Beck2018). Cycling conditions included initial denaturation at 95 °C for 2 min, 5 cycles of 95 °C for 40 s, 54 °C for 60 s, 72 °C for 90 s, 33 cycles of 95 °C for 40 s, 54 °C for 60 s, 72 °C for 90 s, and a final extension step at 72 °C for 7 min. In cases where the concentration of PCR product was not sufficient, a second PCR with a reduced number of cycles was conducted: denaturation at 95 °C for 2 min, 5 cycles of 95 °C for 40 s, 54 °C for 60 s and 72 °C for 90 s, 22 cycles of 95 °C for 40 s, 54 °C for 60 s and 72 °C for 90 s, with a final extension step at 72 °C for 7 min. We used five pairs of primers: ITS1F (Gardes & Bruns Reference Gardes and Bruns1993) and ITS4m (Beck & Mayr Reference Beck and Mayr2012), LR0R (Rehner & Samuels Reference Rehner and Samuels1994) and LR5 (Vilgalys & Hester Reference Vilgalys and Hester1990), mtSSU1 and mtSSU3R (Zoller et al. Reference Zoller, Scheidegger and Sperisen1999), fRPB2-5F and fRPB2-7cR (Liu et al. Reference Liu, Whelen and Hall1999) and newly designed primers gRPB1AFba (GAGTGYCCGGGACATTTTGG) and fRPB1cRba2 (GSCCRGCAATRTCGTTATCCA) for Bacidia.
Alignment and phylogenetic analyses
We obtained 142 sequences of Bacidia s. str. from the Caucasus, augmented the dataset with sequences of Bacidia s. str. from GenBank (Table 1) and included Sporacestra borbonica comb. ined. and S. pertexta (Nyl.) Stapnes & Timdal as outgroup species based on the results of Kistenich et al. (Reference Kistenich, Timdal, Bendiksby and Ekman2018).
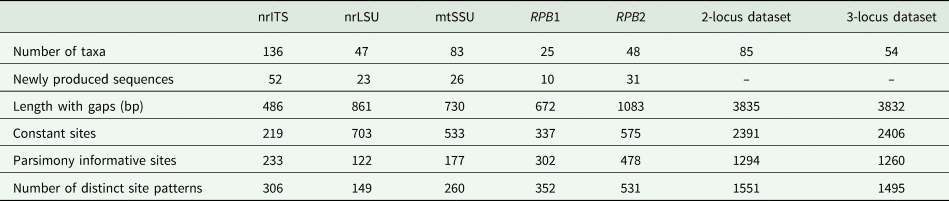
BLAST searches in GenBank were performed to detect and exclude sequences from accessory and lichenicolous fungi and contaminants. We performed phylogenetic analyses on each locus separately (‘single-locus’ analyses) and on concatenated alignments of loci. In the concatenated analyses, we assembled one alignment with a minimum of two out of the five target loci (herein referred to as the ‘two-locus’ analysis) to retain a higher number of samples (88 samples). A second concatenated alignment was also assembled to test the effect of decreased missing data on the analyses; a minimum of three loci were included, but only 54 samples were retained in this alignment (herein referred to as the ‘three-locus’ analysis). An overview of the taxa number and newly produced sequences for each genetic marker and concatenated alignments are summarized in Table 2. Due to the higher number of samples included, we mainly focus on the results of the nrITS and two-locus analyses in this paper. Alignment from the single and concatenated datasets is available as Supplementary Material File S2 (available online).
Overview of the number of taxa and newly produced sequences for each genetic marker and concatenated alignments (excluding outgroup).
Each locus was aligned using MUSCLE v. 3.8.31 using the default settings (Edgar Reference Edgar2004) implemented in the program PhyDE-1 v. 0.9971 and optimized manually. Ambiguous regions of nrITS1 were aligned using MAFFT v. 7.505 (Katoh & Standley Reference Katoh and Standley2013). Sites with more than 95% gaps were excluded, and alignments for the two-locus and three-locus analyses were concatenated manually. Substitution models for the concatenated and single locus datasets were selected using jModelTest v. 2 (Guindon & Gascuel Reference Guindon and Gascuel2003; Darriba et al. Reference Darriba, Taboada, Doallo and Posada2012) for BI, and ModelFinder (Kalyaanamoorthy et al. Reference Kalyaanamoorthy, Minh, Wong, von Haeseler and Jermiin2017) for ML analyses using IQ-TREE. Partition models were implemented for the concatenated alignments, allowing each partition to have its substitution model. Identical sequences were excluded from the subsequent analyses but are listed in Table 1.
We performed Bayesian inference (BI) and maximum likelihood (ML) analyses in RAxML and IQ-TREE on the single-locus and concatenated alignments of nrITS, nrLSU, mtSSU, RPB1 and RPB2. Bayesian inference was carried out using the Markov chain Monte Carlo method (MCMC) using MrBayes v. 3.2.6 (Ronquist et al. Reference Ronquist, Teslenko, van der Mark, Ayres, Darling, Höhna, Larget, Liu, Suchard and Huelsenbeck2012). A GTR substitution model with gamma-distributed rate variations across sites and a proportion of invariable sites was selected based on the result of jModelTest. Two parallel runs were performed (two cold chains), with a single tree saved every 10th generation for a total of 1 000 000 generations. The convergence of the Markov chain was examined according to the trends in likelihood values; the initial 10% was discarded as burn-in, and the results were summarized as a 50% majority-rule consensus tree.
ML analysis was performed with RAxML v. 8.2.4 following a GTRGAMMA model of molecular evolution with bipartitions drawn onto the most likely tree topology using multiple non-parametric bootstraps (Stamatakis Reference Stamatakis2014) on the CIPRES web portal (Miller et al. Reference Miller, Pfeiffer and Schwartz2010).
Further tree reconstruction using ML analysis was performed in IQ-TREE v. 1.6.12 using standard bootstrap approximation with 1000 bootstraps (Felsenstein Reference Felsenstein1985; Nguyen et al. Reference Nguyen, Schmidt, von Haeseler and Minh2015). Substitution models for concatenated (partitioned) and single-locus datasets were selected using ModelFinder (Kalyaanamoorthy et al. Reference Kalyaanamoorthy, Minh, Wong, von Haeseler and Jermiin2017).
The ML trees based on the different substitution models from single and concatenated datasets were congruent and in accordance with the Bayesian tree topology. Therefore, only the RAxML tree for the nrITS and concatenated dataset are shown, with RAxML bootstrap values (BSr), Bayesian posterior probabilities (PP), and IQ-TREE bootstrap values (BSi) used. The phylogenetic trees were visualized using FigTree v. 1.4.2 (Rambaut Reference Rambaut2009). Only clades that received BSr ≥ 70%, PP ≥ 0.95 and BSi ≥ 80% were considered highly supported and given in bold. The concatenated and individual gene trees obtained from RAxML, MrBayes and IQ-TREE are provided in Supplementary Material Figs S1–8 (available online).
Results
Morphology and taxonomy
Revision of the 237 herbarium specimens showed that the most common species of Bacidia s. str. in our Caucasus collection is B. polychroa (16.5%), while B. fraxinea (7%), B. arceutina (5.5%), B. laurocerasi (3.8%), B. absistens (c. 1%) and B. biatorina (c. 1%) are least frequent. Only one herbarium specimen of B. herbarum was studied, which was collected in Krasnodarskiy Krai at 2115 m a.s.l. (GLM-L-0054141).
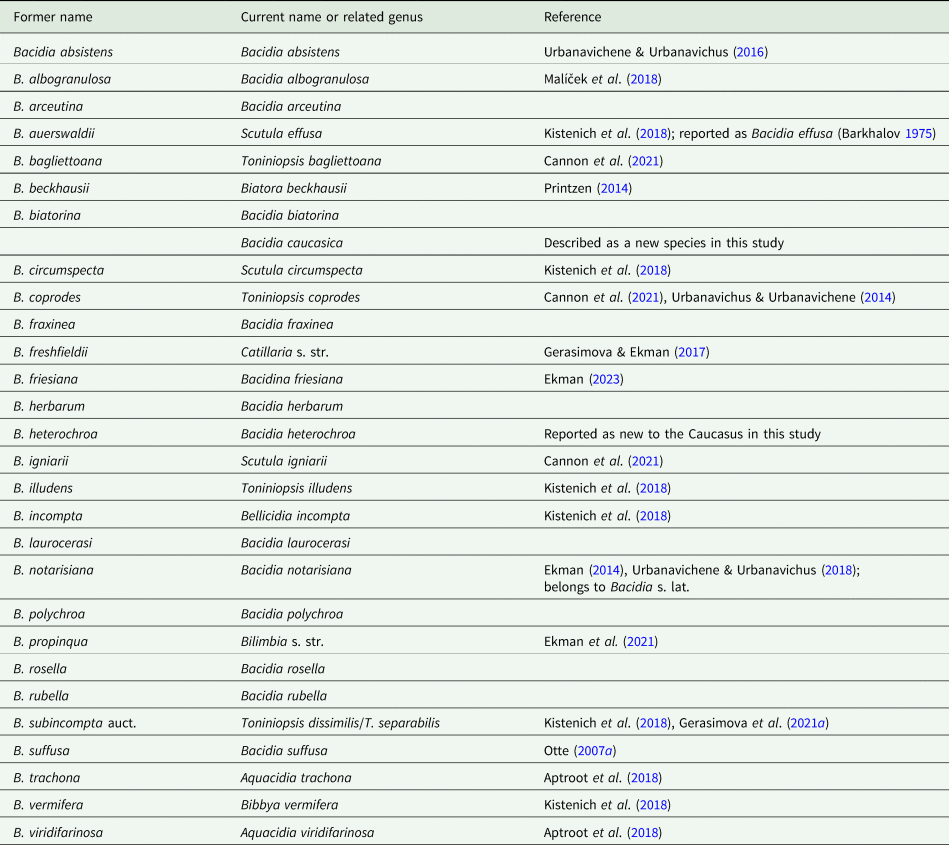
Our examination of herbarium material found the first record of Bacidia heterochroa (Müll. Arg.) Zahlbr. for Caucasus and Russia. Based on morphological and anatomical analyses, one new species, Bacidia caucasica sp. nov. (Suffusa group) was described (see ‘Taxonomy’), and two putative taxa were defined: Bacidia inconspicua ined. (Rubella group), and B. maritima ined. (Rubella group). The systematic revision of the Bacidia species list provided by Urbanavichus (Reference Urbanavichus2010) and their current taxonomic status are given in Table 3. Observations of the complex morphology of the Rubella group are summarized in Table 4.
Revised Bacidia species list based on the checklist of Urbanavichus (Reference Urbanavichus2010), including recently found or newly described species.
Main characters separating taxa of Bacidia rubella s. lat. group. Information for Bacidia fraxinea and B. rubella are given by Ekman (Reference Ekman1996), Ekman & Nordin (Reference Ekman and Nordin1993), Smith et al. (Reference Smith, Aptroot, Coppins, Fletcher, Gilbert, James and Wolseley2009) and Llop (Reference Llop2007); B. iberica and B. parathalassica are from Llop (Reference Llop2007) and Aragón & Martínez (Reference Aragón and Martínez2003); B. thyrrenica from Llop et al. (Reference Llop, Ekman and Hladun2007); B. obtecta and B. elongata are from Gerasimova et al. (Reference Gerasimova, Ezhkin and Beck2018, Reference Gerasimova, Ezhkin, Davydov and Beck2021b). Measurements for Caucasian taxa are newly provided (see taxa with ‘Caucasus’ in brackets after the species name. Measurements from the present study are given as (min–) mean ± SD (–max), and are otherwise are taken from the relative references. Taxa are ordered according to their morphological similarity.

Ecology
All studied specimens were collected mainly in old mixed coniferous-broad-leaved forest communities in floodplains and river valleys as well as in the drier habitats such as fruit orchards, deciduous forests without conifers, and recently clearcut young coppices. The main phorophytes included Abies nordmanniana, Acer campestre, Carpinus betulus, Quercus spp. (Q. pubescens, Q. petraea and Q. robur), Juniperus spp. (J. excelsa, J. oxycerdrus and J. foetidissima), and Pistacia mutica. Due to the proximity of the Caspian and Black Seas, the dense river system and the high groundwater level, the average relative humidity in all the studied areas was very high. The average annual rainfall ranged from 400 mm (Samursky National Park) and 600 mm (Utrish Nature Reserve) to 2000–3000 mm in the forest belt of the Caucasian Reserve. Most of the Bacidia species in the Caucasus are confined to humid environments, except B. rubella and B. fraxinea, which occur in a broader range of habitats from dry and warm to humid subtropics with high levels of sunlight. While both species could be present in dry areas, Bacidia rosella is one of the most hygrophilous species. It has been collected in the western and north-western Caucasus in old-growth forest habitat with Abies nordmanniana, Acer trautvetteri and Fagus orientalis at 1060–1600 m a.s.l., the most humid and shaded areas of all the studied localities due to the predominance of Abies nordmanniana.
Phylogeny
The BI and ML analyses for the single and concatenated datasets recovered highly concordant topologies of the phylogenetic groups (with a few exceptions discussed below). Both two-locus and three-locus trees resulted in well-supported nodes throughout the tree (the three-locus tree is provided in Supplementary Material Fig. S2, available online). In total, we produced 142 new sequences for this study for the various genetic markers. The nrITS comprised 40% of the alignment of a total dataset of 340 sequences, mtSSU comprised 24.7%, nrLSU 14%, RPB2 14%, and RPB1 7.3%.
All the sequences from the Caucasus obtained for this study nested in the correspondent clades recovered in the last multilocus phylogeny of Bacidia s. str. (Gerasimova et al. Reference Gerasimova, Urbanavichene, Urbanavichus and Beck2021a). Bacidia heterochroa from the Caucasus formed a new group (Figs 1 & 2). The two-locus and nrITS trees were congruent with the topology of the three-locus phylogeny, but while we retrieved high support in the nrITS tree for nodes towards the tips, the backbone in the multilocus phylogenies was better supported (Figs 1 & 2; Supplementary Material Fig. S2).
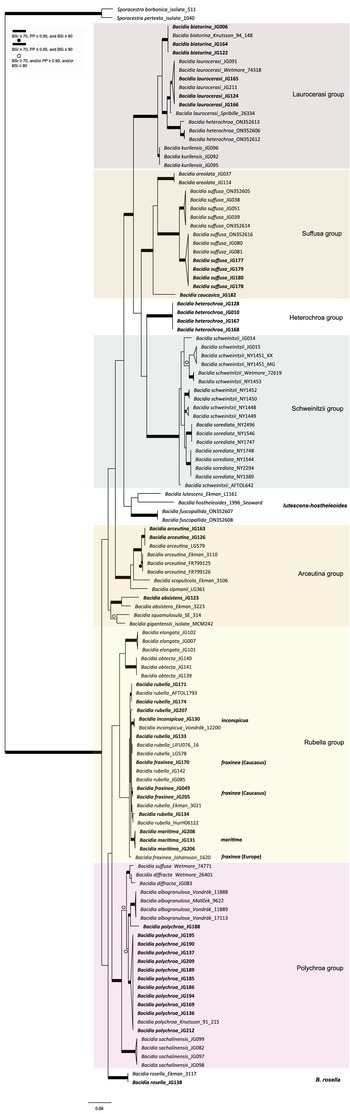
Maximum likelihood (ML) tree of Bacidia s. str. resulting from a RAxML analysis of the concatenated multilocus dataset with a minimum of two loci included (out of nrITS, nrLSU, mtSSU, RPB1 and RPB2). RAxML bootstrap values (BSr), Bayesian posterior probabilities (PP) and IQ-TREE bootstrap values (BSi) are indicated. Highly supported branches with BSr ≥ 70%, PP ≥ 0.95, and BSi ≥ 80% are marked in bold; strongly supported branches with BSr ≥ 70% and BSi ≥ 80% are also marked in bold with a dot above the branch; branches with BSr ≥ 70%, and/or PP ≥ 0.95, and/or BSi ≥ 80% are marked with a white dot. Major groups within clades are indicated, as are species within or outside groups. New sequences are in bold. For further information about sequences, see Table 1. Single phylogenetic trees resulting from the concatenated multilocus datasets from RAxML, BI, and IQ-TREE analyses are in Supplementary Material Figs S1 and S2, (available online). In colour online.
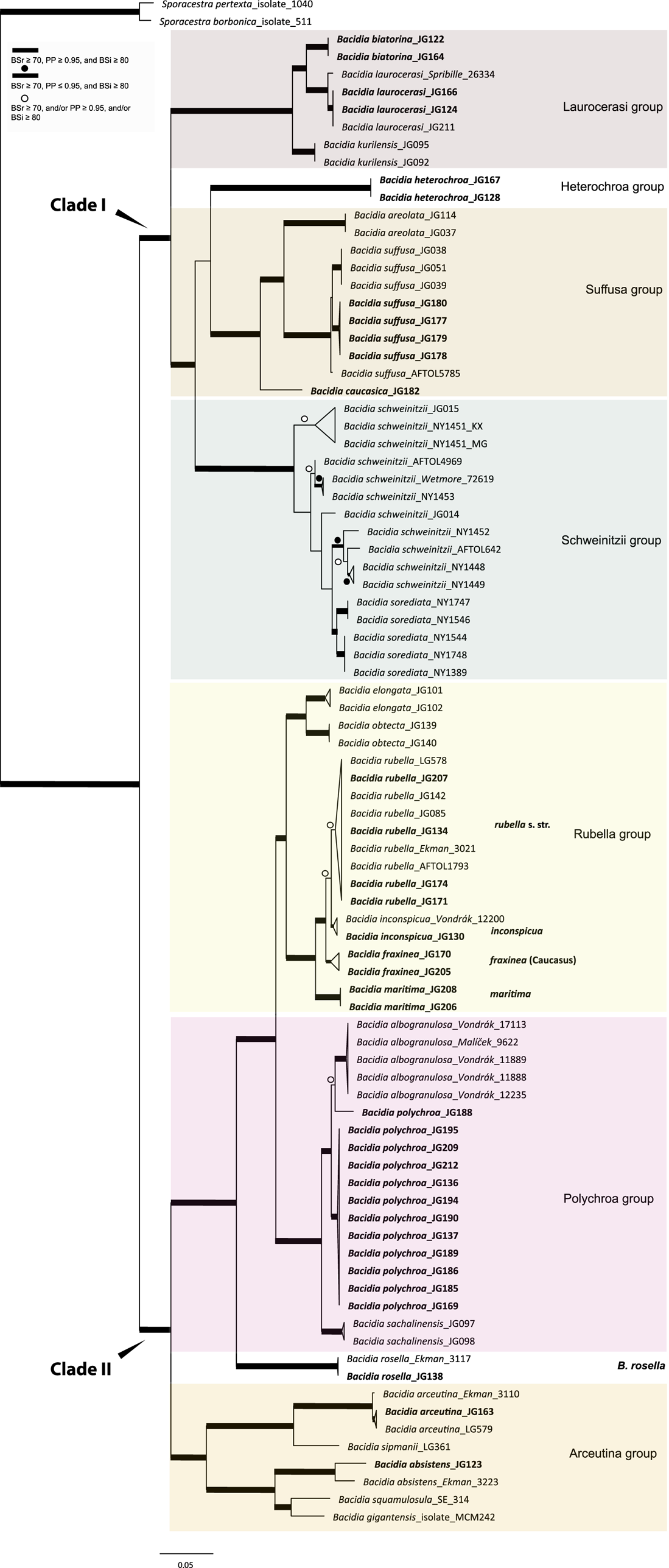
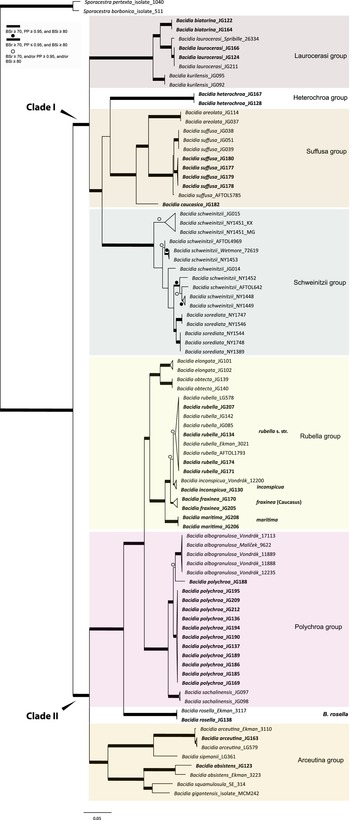
Maximum likelihood (ML) nrITS tree of Bacidia s. str. resulting from a RAxML analysis. RAxML bootstrap values (BSr), Bayesian posterior probabilities (PP), and IQ-TREE bootstrap values (BSi) are indicated. Highly supported branches with BSr ≥ 70%, PP ≥ 0.95, and BSi ≥ 80% are marked in bold; strongly supported branches with BSr ≥ 70% and BSi ≥ 80% are also marked in bold with a dot above the branch; branches with BSr ≥ 70%, and/or PP ≥ 0.95, and/or BSi ≥ 80% are marked with a white dot. Major groups within clades are indicated, as are species within or outside groups. New sequences are in bold. For further information about sequences, see Table 1. Single phylogenetic trees resulting from the concatenated multilocus datasets from RAxML, BI, and IQ-TREE analyses are in Supplementary Material S4 (available online). In colour online.
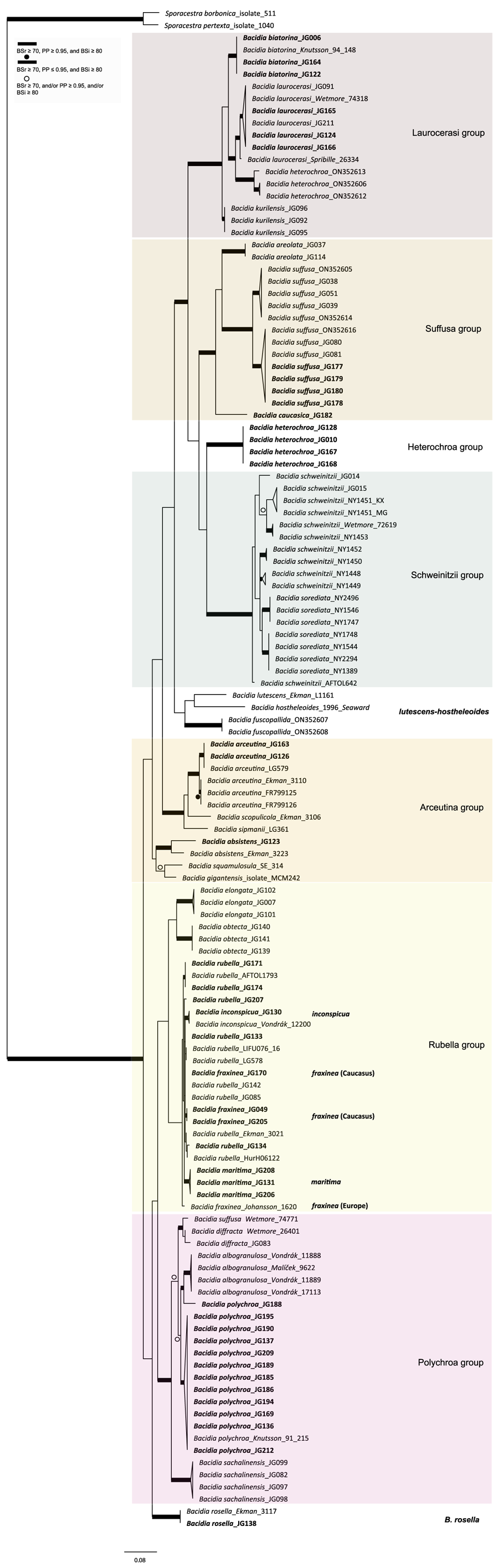
The main groups obtained from single and concatenated data in both ML and BI analyses are congruent and correspond to those obtained from the previous phylogenies (Ekman Reference Ekman2001; Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018, Reference Gerasimova, Ezhkin, Davydov and Beck2021b). Furthermore, similar to previous results, our phylogenies revealed two highly supported major clades in the two- and three-locus phylogenies. Clade I included the Laurocerasi, Schweinitzii, Suffusa, and Heterochroa groups from the Caucasus (BSr/PP/BSi: 100/1/100 in both phylogenies). Several sequences of B. heterochroa from South Korea formed a sister clade to B. laurocerasi in the nrITS phylogeny but with low support (BSr/PP/BSi: 44/0.75/43) (see ‘Discussion’). Clade II included the Arceutina (BSr/PP/BSi: 75/1/87, and 73/1/75), Rubella (BSr/PP/BSi: 85/1/82, and 86/1/88), Polychroa (BSr/PP/BSi: 100/1/100 in both phylogenies), and B. rosella groups (BSr/PP/BSi: 100/1/100 in both phylogenies). Bacidia lutescens Malme (referred to as B. thiersiana Lendemer by Lendemer (Reference Lendemer2020)), B. hostheleoides (Nyl.) Zahlbr., and recently described B. fuscopallida B.G. Lee & T.I. Heo formed a separate group within Clade II but without support in the nrITS phylogeny (Fig. 2). All clades in the multilocus phylogenies received strong support throughout the tree (i.e. with BSr ≥ 70%, PP ≥ 0.95 and BSi ≥ 80%). However, in the multilocus trees poor support was retrieved for the sister relationship of the Suffusa and Heterochroa groups (BSr/PP/BSi: 62/0.92/65, and 71/0.94/72), the subclades of the Schweinitzii group (not present in the Caucasus, discussed in detail in Gerasimova et al. (Reference Gerasimova, Ezhkin, Davydov and Beck2021b)), the position of the Bacidia rubella s. str. clade (BSr/PP/BSi: 72/0.55/66, and 54/-/37), the placement of B. inconspicua ined. (BSr/PP/BSi: 79/0.74/73, and 55/-/36), and the retrieval of the B. albogranulosa clade as sister to B. polychroa (JG188) (BSr/PP/BSi: 77/0.91/56 in the two-locus tree).
The retrieved groups are discussed in detail in previous studies (Gerasimova et al. Reference Gerasimova, Ezhkin, Davydov and Beck2021b); therefore, we focus on those with Caucasian representatives below. As phylogenies are congruent in the topology of the groups, the Results and Discussion are based on the two-locus and nrITS trees, as those are the most species-inclusive. Trees from the single and concatenated phylogenies not shown in the manuscript can be found in Supplementary Material Figs S1–8 (available online).
Laurocerasi group
In concatenated phylogenies, B. biatorina, B. laurocerasi and B. kurilensis Gerasimova, A. Ezhkin & A. Beck (Sakhalin endemic) formed one clade with high support of all subclades (Fig. 1). Two sequences of B. biatorina from the Caucasus formed a highly supported clade (BSr/PP/BSi: 100/1/100), including a sequence from Sweden (Fig. 2).
The sequences from the Caucasus formed a well-supported clade in the multilocus phylogenies and formed a clade together with the representatives from the Far East and the USA in the two-locus and nrITS phylogenies (Fig. 1 & 2). In all phylogenies, the sequence from Alaska (Spribille 26334) was placed as sister to all others with maximum support in the multilocus phylogenies (BSr/PP/BSi: 100/1/100) and high support in the nrITS tree (BSr/PP/BSi: 92/0.99/99).
Bacidia heterochroa clades from South Korea and the Caucasus
Bacidia heterochroa inhabits tropical and subtropical areas worldwide (Ekman Reference Ekman1996) and was recently found in South Korea (Lee & Hur Reference Lee and Hur2022). To include nrITS sequences of B. heterochroa from South Korea and keep the informative part of the nrITS alignment, we constrained our alignment to Bacidia Clade I (Supplementary Material Fig. S3, available online). Three sequences of B. heterochroa from Korea (GenBank ON352606, ON352612 and ON352613) formed a sister clade to B. laurocerasi, but with support in RAxML and BI analyses only (BSr/PP/BSi: 74/1/65). Instead, Bacidia heterochroa sequences from the Caucasus formed a separate clade sister to the Suffusa and Schweinitzii groups in all the phylogenies but with uncertain sister relationships (Fig. 2). In the multilocus phylogenies, it was sister to the Suffusa group with low support (BSr/PP/BSi: 62/0.92/65, and 71/0.94/72; Fig. 1 & Supplementary Material Fig. S2). In the nrITS tree, it was sister to the Schweinitzii group on a long branch and with low support (BSr/PP/BSi: 63/0.91/61; Fig. 2).
Suffusa group
The Bacidia suffusa sequences from the Caucasus formed a clade together with those from North America (JG080 and JG081; Figs 1 & 2) and a sequence from South Korea (ON352616) in the nrITS phylogeny (Fig. 2). They formed a sister clade to the representatives from the Far East and South Korea with strong support values in all phylogenies. The sequences from the Caucasus differ from the Far East individuals by 3% (up to 15 nucleotides); however, as we could not observe any strong morphological differences between the specimens from these two clades, we chose not to recognize these individuals as new species.
One sequence from the Caucasus (JG182) formed a separate clade in all phylogenies with strong support. Based on this phylogenetic evidence and its distinct morphology, a new species, Bacidia caucasica, is described (see ‘Taxonomy’).
Schweinitzii group
The taxa from the Schweinitzii group are mostly known from North America and the coast of the Russian Far East (see details in Gerasimova et al. (Reference Gerasimova, Ezhkin, Davydov and Beck2021b)) and have not been found in the Caucasus to date.
Rubella group
The highly supported Rubella group (BSr/PP/BSi: 85/1/82) forms four distinct clades comprising sequences of Bacidia fraxinea, B. rubella, B. elongata Gerasimova & A. Beck, B. obtecta Gerasimova et al., B. inconspicua ined. and B. maritima ined. (Fig. 1). Bacidia rubella sequences from the west coast of the Caspian Sea (JG172, JG206), and the north-east coast (JG208) and south-east coast (JG131) of the Black Sea formed a sister clade to Bacidia rubella s. lat. clades in all reconstructed phylogenies with maximum support; these are provisionally defined as Bacidia maritima ined. here. However, JG131 and JG172 are represented by only one sequence: nrITS and RPB2, respectively. The difference of nrITS sequences of JG206 and JG208 is almost 3% compared to other sequences from the B. rubella s. str. clade; however, this difference does not seem to be correlated with a difference in studied morphological characters (see ‘Discussion’, Table 4, and Fig. 3).
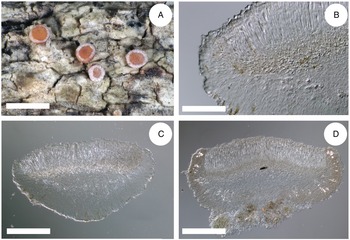
Thallus structure and apothecia variability of the most typical representatives of the Rubella group. A, Bacidia rubella s. str. from Caucasus (JG171, GLM-0041636). B, B. maritima ined. (JG206, M-0311935). C, B. inconspicua ined. (JG130, M-0311925). D, B. fraxinea from Caucasus (JG170, GLM-0044145). E, B. elongata (JG101, M-0182625). F, B. obtecta (JG141, M-0308496—holotype). Scales: A–E = 1 cm; F = 0.5 cm. In colour online.

Sequences from the Caucasus (JG130) and Ukraine (Vondrák 12200, PRA; identified and published as Bacidia cf. rubella by Malíček et al. (Reference Malíček, Palice, Vondrák, Łubek and Kukwa2018)) formed a highly supported clade sister to B. rubella s. str. in the two-locus phylogeny (BSr/PP/BSi: 90/1/87; Fig. 1), and are provisionally placed in Bacidia inconspicua ined. The nrITS sequences alone are insufficient to resolve the species relationships within B. rubella, thus appearing paraphyletic but without support for the paraphyly (Fig. 2).
The nrITS sequence of Bacidia fraxinea from Sweden (Johansson 1620) and one from Caucasus (JG170) were nested in a clade of B. rubella sequences in the nrITS phylogenies (Fig. 2), revealing paraphyletic status of the species as two other B. fraxinea sequences from the Caucasus formed a clade with high support in the two- and three-locus phylogenies (BSr/PP/BSi: 85/1/86, and 87/-/73).
Polychroa group
The Polychroa group comprises a highly supported clade of Bacidia polychroa, B. sachalinensis Gerasimova et al., B. diffracta S. Ekman and B. albogranulosa in the two-locus trees (Figs 1 & 2). All Caucasian B. polychroa sequences formed one clade with high support, including a sequence from Sweden in the nrITS phylogeny (BSr/PP/BSi: 93/1/98).
One sequence from Caucasus (JG188) was placed as a sister to the B. albogranulosa clade in the two-gene phylogeny with low support (BSr/PP/BSi: 77/0.91/56) but with high support in the nrITS tree (BSr/PP/BSi: 89/0.99/99).
Rosella group
The sequences from the Caucasus (JG138) and Norway (Ekman 3117) formed a highly supported clade in all phylogenies. In the two- and three-gene phylogenies, these sequences are retrieved in Clade II with maximum support, together with the Polychroa and Rubella groups (BSr/PP/BSi: 100/1/100 in both phylogenies).
Arceutina group
The B. arceutina sequences from the Caucasus (JG126 and JG163) and Switzerland (AFTOL LG579) formed a well-supported clade sister to the B. arceutina clade from Sweden in the two- and three-locus trees (BSr/PP/BSi: 100/1/100, and 99/1/98). The sequences differ by c. 2.5% (7–8 nucleotides) in nrITS. Bacidia arceutina is distributed in the western part of the Caucasus (viz. Krasnodarskiy Krai, Republic of Adygea, Georgia), with its southern-most limit recorded in a valley in the Lenkaran district of Azerbaijan (GLM-L-529756). Throughout its range, it is found at elevations of up to 1000 m in warm, well-lit and relatively dry lowlands and habitats with very high rainfall (from 1500 to 3000 mm per year). The B. absistens sequence from the Caucasus (JG123) was placed with the sequence from Norway as a sister to B. squamulosula (Nyl.) and B. gigantensis Lendemer et al., with high support in the multilocus phylogenies (BSr/PP/BSi: 100/1/99, and 73/1/75). Bacidia absistens in the Caucasus is recorded from shaded habitats at elevations of 700 m a.s.l. with very high humidity (rainfall of c. 1000 mm per year).
Discussion
Two main clades correlated with apothecia pigmentation
Our phylogenies of Bacidia with 142 additional sequences from the Caucasus were congruent with previous results based on nrITS and multilocus phylogenies (Ekman Reference Ekman2001; Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018, Reference Gerasimova, Ezhkin, Davydov and Beck2021b). In line with previous studies, we retrieved two large clades: Clade I includes the Laurocerasi, Schweinitzii and Suffusa groups, as well as the newly defined Heterochroa group (that consists of representatives from the Caucasus), and Clade II includes the Rubella, Polychroa and Arceutina groups. In a previous work, phylogenies mainly included specimens from temperate regions (Gerasimova et al. Reference Gerasimova, Ezhkin, Davydov and Beck2021b), while in this study, we also included representatives from the subtropics. Only the nrITS phylogeny included B. lutescens (B. thiersiana), B. hostheleoides, and the recently described B. fuscopallida (Lee & Hur Reference Lee and Hur2022), which formed an unsupported group nested in Clade II. The former two taxa are widespread in south-eastern North America and the Neotropics (Malme Reference Malme1935; Ekman Reference Ekman1996; Lendemer Reference Lendemer2020), while B. fuscopallida is known only from the Gangwon Province in South Korea (Lee & Hur Reference Lee and Hur2022), characterized by a moist, warm, temperate climate (Sayre et al. Reference Sayre, Karagulle, Frye, Boucher, Wolff, Breyer, Wright, Martin, Butler and Van Graafeiland2020). Further analyses are necessary to clarify the phylogenetic position of this group.
Similar to previous results, two main clades can be separated based on apothecial pigment. Specimens from Clade I have dark brown, red-brown or green pigments (Laurocerasi-brown and Bagliettoa-green), or a combination thereof, in the upper part of the hymenium and lateral exciple. The herein-defined Heterochroa group from the Caucasus also supports this differentiation; the Caucasian specimens are characterized by a dark purplish epithecium corresponding to the type specimen of B. heterochroa. The specimens from South Korea have a dark brown epithecium corresponding to the Laurocerasi group, where South Korean sequences were nested. In contrast, representatives from Clade II have a mixture of yellow, orange and/or brown apothecial pigments (Arceutina-yellow, Polychroa-brown and Rubella-orange) in the upper part of hymenium and lateral exciple. The specimens from the Caucasus in Polychroa and Rubella groups also supported the apothecial pigment differentiation. The representatives from the Lutescens-Hostheleoides group have been characterized by almost colourless or faintly and diffusely pigmented internal apothecial structures (Ekman Reference Ekman1996; Gerasimova et al. Reference Gerasimova, Ezhkin, Davydov and Beck2021b). In contrast, recently described B. fuscopallida is characterized by the clearly pigmented orange-brown to brown hypothecium (Lee & Hur Reference Lee and Hur2022). However, this clade was inferred to have very long branches in our analyses, suggesting that their relationship warrants further investigation in future studies, including additional samples.
The clade containing Bacidia absistens, B. gigantensis and B. squamulosa is exceptional because it has highly variable pigmentation. This evidence may be related to the exceptional diversity of secondary compounds in this group detected by TLC, such as 4-O-methylcryptochlorophaeic and homosekikaic acids (Tønsberg et al. Reference Tønsberg, Culberson and Johnson1995; Lendemer Reference Lendemer2020). This combination is so far unknown for species in other Bacidia s. str. groups, which contain atranorin as the main known secondary compound (Ekman Reference Ekman1996; Gerasimova et al. Reference Gerasimova J, Beck, Werth and Resl2022).
As a result of morphological and/or phylogenetic analyses, the current list of Bacidia s. str. in Caucasus includes 13 species, namely B. absistens, B. albogranulosa, B. arceutina, B. biatorina, B. caucasica, B. fraxinea, B. herbarum, B. heterochroa, B. laurocerasi, B. polychroa, B. rosella, B. rubella and B. suffusa, which is almost 68.4% of the 19 species of Bacidia s. str. known from Russia (Ekman Reference Ekman2009; Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018).
Laurocerasi group
The known distribution of Bacidia biatorina in Russia includes the European part, Far East, and Caucasus (Gerasimova Reference Gerasimova2016; Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018). However, only a small number of herbarium specimens were confirmed to belong to the species; therefore, its distribution in Russia remains insufficiently studied. Bacidia laurocerasi has a cosmopolitan distribution, having been recorded in Russia (Caucasus, Ural, Siberia, and the Far East), Europe, Macaronesia, Africa, North and South America, Asia, Australia, and New Zealand (Smith et al. Reference Smith, Aptroot, Coppins, Fletcher, Gilbert, James and Wolseley2009; Urbanavichus Reference Urbanavichus2010). Both B. biatorina and B. laurocerasi are restricted to forests with high humidity in the Caucasus. Bacidia biatorina is mainly found in undisturbed, beech-dominated, humid, shaded forests of the middle mountain forest belt of the north-western Caucasus, while B. laurocerasi is found in cooler, humid middle and even upper beech- and fir-dominated mountain forest belts but also inhabits stream valleys and lowlands.
Suffusa group: several species or different populations?
Bacidia suffusa is mainly known from the eastern temperate region of North America, the Caucasus and Russian Far East (Ekman Reference Ekman1996; Otte Reference Otte2007a; Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018), and it was also recently recorded in South Korea (Lee & Hur Reference Lee and Hur2022). The closely related B. areolata Gerasimova & A. Beck is known only from the Far East and is thus still considered endemic. The finding of B. suffusa in the Caucasus was surprising, as it indicates the species has a disjunct distribution across eastern North America and parts of Eurasia, including the Caucasus, but is absent in Europe (Otte Reference Otte2007a). The species inhabits lower mountain belts of oak and beech, indicating it could be restricted to Tertiary relict floras that are common in parts of the Caucasus.
To confirm the relationship between North American, Far Eastern and Caucasian representatives, we included several specimens from different parts of the Caucasus in our phylogeny (Table 1). Our results indicate that sequences of B. areolata and B. suffusa form a strongly supported clade. In previous work, single-locus and combined nrLSU, mtSSU and RPB1 phylogenies indicate that Far Eastern B. suffusa and North American B. suffusa sequences are not monophyletic (Gerasimova et al. Reference Gerasimova, Ezhkin, Davydov and Beck2021b). Intriguingly, in our analyses, the newly sequenced Caucasian specimens of B. suffusa form a clade with the North American individuals that is sister to the Far Eastern B. suffusa clade. The nrITS sequences in these two clades differed by 3% (up to 15 nucleotides), suggesting substantial genetic differentiation; however, no significant morphological differences were found between them. Therefore, we suggest clades containing sequences from 1) the Far East and South Korea and 2) Caucasus, North America and South Korea are two populations of B. suffusa on the cusp of divergence into separate species.
Importantly, one of the sequences (JG182) was consistently retrieved as a sister to B. areolata and B. suffusa subclades in all phylogenies with the highest support. Morphological examination indicates clear differentiation from B. areolata and B. suffusa s. lat., strongly supporting the recognition of this individual as a new species. As a result, a new species, Bacidia caucasica, is described below (see ‘Taxonomy’).
Rubella group: the complex morphology of taxa
Bacidia rubella is one of the most widespread species of Bacidia s. str., occurring in the Holarctic and known from Europe, Macaronesia, Africa, Asia and North America (Ekman Reference Ekman1996; Llop Reference Llop2007; Smith et al. Reference Smith, Aptroot, Coppins, Fletcher, Gilbert, James and Wolseley2009). In the Caucasus, it occurs in habitats with a wide range of humidity levels and light intensities, from dry, subtropical coastlines with Juniperus, to humid, dense, mid-mountain forests with Abies nordmanniana. It occurs in open upper mountain forest belts and has also been recorded from humid stream valleys, with evidence of natural or anthropogenic disturbance. Similar to B. rubella, B. fraxinea occurs across a wide range of habitats, found in the open, Mediterranean-like, drought-adapted forest vegetation of the Black Sea coast, in forest patches throughout agricultural areas of the forest-steppe region, and in lower mountain forest belts and higher-humidity habitats, in broad-leaved mesophilic forests. In lower mountain forest belts, B. fraxinea usually occurs in areas of relatively high humidity but is especially abundant in logged forests. Interestingly, the provisionally defined B. inconspicua ined. was also collected in young, dry, and warm secondary forest. This evidence leads to the conclusion that species from the Rubella group are most adapted to dry conditions, contrary to other Bacidia species.
This study included 14 additional specimens of both Bacidia rubella s. lat. and B. fraxinea s. lat. for sequencing and phylogenetic analysis. Whereas in the nrITS phylogeny, the Rubella group renders paraphyletic (viz. nesting B. rubella, B. fraxinea, and B. inconspicua ined.), the two- and three-locus phylogenies reveal six supported clades: 1) a large B. rubella s. str. clade comprising sequences from North America, Caucasus, Northern and Central Europe, and the Far East; 2) a newly defined B. maritima ined. from Caucasus (JG206 and JG208); 3) a clade of B. fraxinea from Caucasus (JG170 and JG205); 4) a clade of newly defined B. inconspicua ined. species from Caucasus (JG130) and Ukraine (Vondrák 12200, PRA); 5) a clade of endemic B. elongata from the Far East; 6) a clade of B. obtecta, endemic to the Far East (Figs 1 & 2). As we were not able to study the type material of the Mediterranean-European specimens of Bacidia rubella s. lat. and there are no sequence data available, we intend to obtain and include this data in our future studies. Nevertheless, we discuss the morphology of the Mediterranean representatives based on the literature summarized in Table 4.
Morphologically, B. fraxinea was mainly separated from B. rubella by thallus structure (Ekman & Nordin Reference Ekman and Nordin1993). This separation was supported in the first phylogenetic study on Bacidiaceae, where the two species formed two sister clades (Ekman Reference Ekman2001), with a 1% (4 nucleotides) difference in the nrITS sequences. More recent studies of the nrITS2 secondary structure of B. fraxinea (Johansson 1620) and B. rubella (Ekman 3021) revealed one hemi-compensatory base change (hemi-CBC) in the structurally conserved regions of helix III (Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018), also supporting the distinction of the species. However, subsequent single-gene and multilocus phylogenetic analyses resulted in a paraphyletic Bacidia rubella clade, where B. fraxinea (Johansson 1620) was frequently nested within the B. rubella s. str. clade (Gerasimova et al. Reference Gerasimova, Ezhkin and Beck2018, Reference Gerasimova, Ezhkin, Davydov and Beck2021b); this result is consistent with our results from the analysis of an enlarged dataset.
As previously mentioned, the thallus structure is an essential character for distinguishing species in the Rubella group more broadly (Table 4). Thus, B. fraxinea is characterized by a thin, inconspicuous to thick, verrucose, wrinkled, and warted thallus (Ekman & Nordin Reference Ekman and Nordin1993). In contrast, B. rubella s. str. is characterized by a granular thallus with isidia- to coral-like structures. Further delimitation based on thallus characteristics in later studies led to a description of two new taxa from the Mediterranean area: Bacidia iberica Aragón & I. Martínez and B. parathalassica Llop & Gómez-Bolea (Llop & Gómez-Bolea Reference Llop and Gómez-Bolea1999; Aragón & Martínez Reference Aragón and Martínez2003). The morphology of B. iberica and B. parathalassica is similar to B. rubella, but B. iberica has a thallus formed by the adpressed squamules, and B. parathalassica has a continuous, smooth to warted thallus, somewhat closer to B. fraxinea (Llop & Gómez-Bolea Reference Llop and Gómez-Bolea1999; Aragón & Martínez Reference Aragón and Martínez2003).
In addition to the thallus structure, the presence and distribution of crystals were used by Llop et al. (Reference Llop, Ekman and Hladun2007) to separate the taxa in the Rubella group, leading to the description of a new species, B. thyrrenica Llop. The authors distinguished two main groups: the first included B. thyrrenica, B. rosella and B. rubella, and the second was composed of B. fraxinea, B. iberica and B. parathalassica. The first group have crystals equally distributed in the upper part of the exciple, dissolving in K. However, sometimes B. rubella has been observed to have scarce crystals restricted to the uppermost part of the exciple (Llop et al. Reference Llop, Ekman and Hladun2007). A second group have clusters of colourless to pale yellow crystals in the medullary part of the exciple that dissolve in acid solution (HNO3). In all specimens, we observed colourless crystals, except for JG142 (Far East, B. rubella s. str. clade) and B. inconspicua ined. (Vondrák 12200, PRA) with yellow crystals in the exciple. Bacidia obtecta is also characterized by having crystals in the upper part of the hymenium and exciple, but these are colourless. However, as crystals are only occasionally present, we suggest this character is not consistent enough to differentiate the Rubella group taxa.
In addition to crystals, B. fraxinea, B. iberica, B. rubella and B. parathalassica are characterized by having one layer of more or less globose cells along the exciple margin. In the provisionally defined B. inconspicua ined. specimens, this character varies in two specimens (Table 4). In contrast, B. elongata and B. obtecta are characterized by having four layers of enlarged lumina cells along the exciple margin. Additionally, all species differ based on characteristics of spore size, hymenium height and apothecia colour (Table 4).
The newly introduced Bacidia inconspicua ined
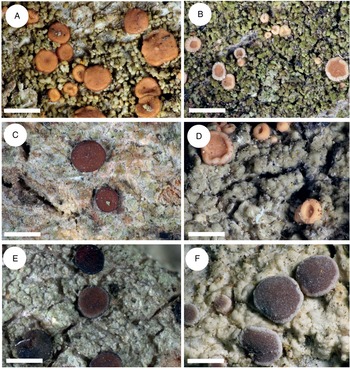
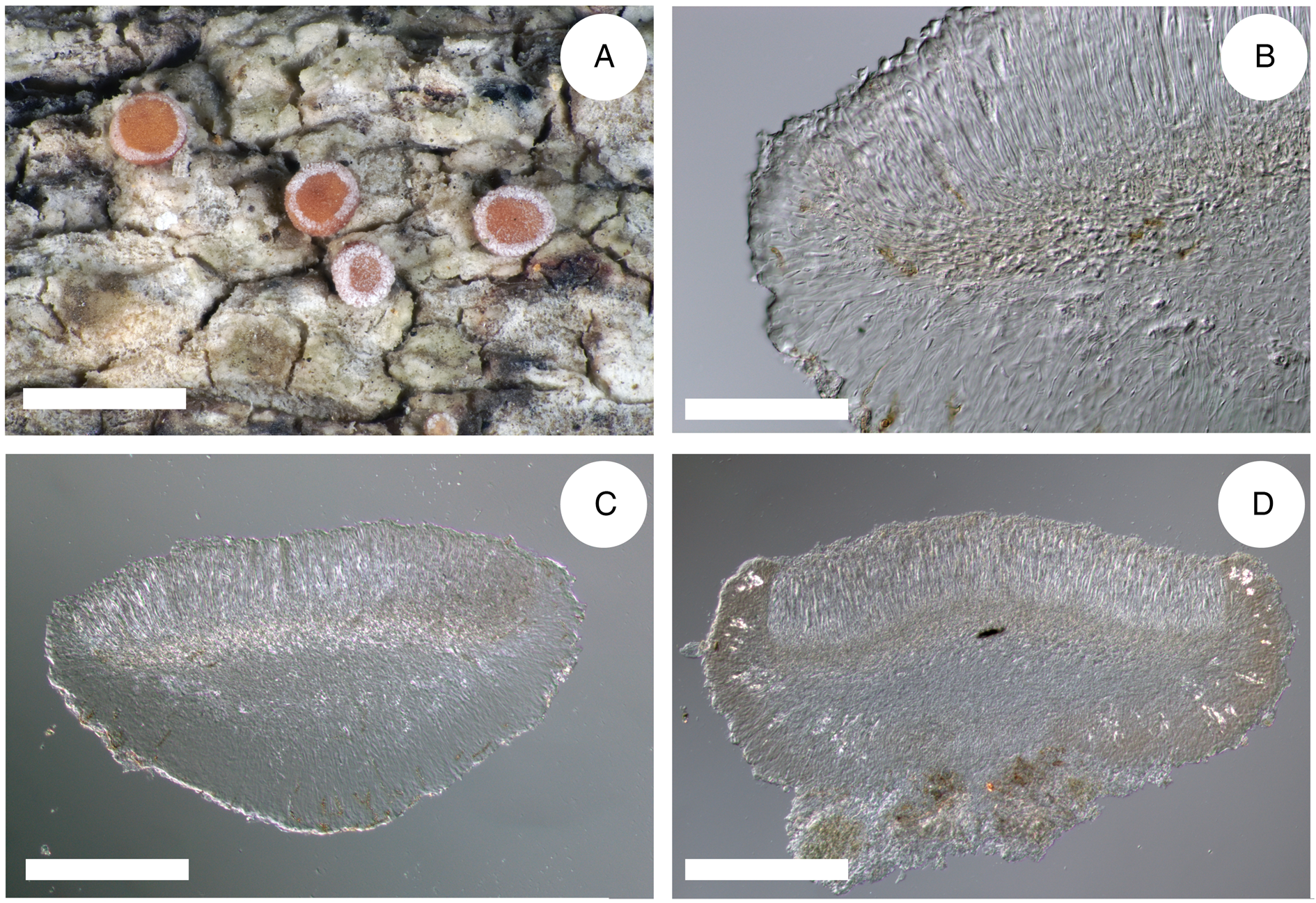
This is represented by two specimens: one collected in the Caucasus (JG130) and one in Ukraine (Vondrák 12200, PRA). The B. inconspicua ined. specimen was collected in a young, dry, and warm secondary forest with pine trees and Cladonia rangiferina undergrowth; the original primary forest of oak and hornbeam, probably logged and replaced by ash and pine plantations. Bacidia inconspicua ined. is characterized by a prosoplectenchymatic exciple without enlarged cells along the exciple rim (JG130), with sometimes up to three layers of enlarged lumina cells (Vondrák 12200, PRA), in contrast to B. iberica, B. fraxinea s. str., B. parathalassica and B. rubella s. str., which are characterized by one layer of more or less globose cells (Table 4 and references therein, Fig. 4). Based on thallus structure and phylogenetic evidence using a multilocus dataset, it seems that B. inconspicua ined. may represent a separate species belonging to the Rubella group. However, as only two specimens were studied and in light of the high plasticity of the phenotypic characters in the Rubella group, we refrain from describing a new species and thus define B. inconspicua provisionally. A detailed description of all distinguishing morphological characters for B. inconspicua ined. is given in Table 4 and Fig. 4.
Cross-sections of apothecia and thallus structure of Bacidia inconspicua ined. (A–C, M-0182578; D, J. Vondrák 12200, PRA). A, smooth, inconspicuous thallus with orange pruinose apothecia. B, cross-section of apothecium with detailed exciple structure. C, cross-section of apothecium viewed using a polarized filter. D, cross-section of apothecium with yellow clusters of crystals arranged in the lateral part of the exciple viewed using a polarized filter. Scales: A = 1 cm; B–D = 100 μm. In colour online.
The newly defined Bacidia maritima ined
This is known from four specimens: JG131, JG172, JG206, and JG208 (Figs 1 & 2); although, from the first two specimens, a sequence could be obtained from only one locus (nrITS and RPB2, respectively). Interestingly, most specimens were collected near the Black and Caspian Seas coasts, suggesting that the species may be confined to the maritime zone. In more detail, JG206 was collected on the west coast of the Caspian Sea, c. 3 km from the coast, in a shaded forest very rich in epiphytic lichens (82 species in 1 ha). JG208 was collected on a south-east slope, c. 750 m from the Black Sea coast in a Juniperus-Pistacia forest in dry subtropic conditions, in a sunny, warm locality but apparently influenced by the proximity of the sea. The fog and high humidity favoured a rich epiphytic composition of lichens in this locality (71 species for 10–12 trees). JG131 was collected in a shaded pine forest also close to the south-east coast of the Black Sea. However, JG172, collected in the southern Caucasus region of Azerbaijan, was found at a site much further inland, seemingly without maritime influence, in a pastured, coppice-like mixed forest. However, the number of specimens of Bacidia maritima ined. known to date is too small to characterize its overall distribution with certainty; it seems that the affinity for coastal sites is more strongly expressed in the somewhat harsher climate of the northern Caucasus.
Morphological examination indicates B. maritima ined. has a similar thallus structure and coloration of the upper and inner apothecia to B. rubella. However, B. maritima ined. has shorter spores than B. rubella, similar to B. iberica and B. parathalassica. The mean spore size of B. rubella from the Caucasus is 47.8 ± 6.6 μm (up to 75 μm) × (1.5–)2.7 ± 0.4(–4) μm wide, and they have up to 12 septa. The mean spore size of European B. rubella individuals is 40–70 μm (up to 84 μm) × 2.5–3(–4) μm wide with up to 13 septa, and of B. rubella North American individuals is 44–63 μm (up to 104 μm) × (2.1–)2.4–2.7–3.2(–4.3) μm wide with up to 13 septa (Table 4). On the contrary, the spores of B. maritima ined. are 48 ± 6.7 μm (up to 65 μm) × (2–)2.65 ± 0.3(–3.5) μm wide with up to 10 septa. Nevertheless, to clearly distinguish between the taxa, additional specimens are necessary for a more detailed description and measurement of spores; therefore, we refrain from describing a new species here.
Polychroa group: the presence of an unpigmented member
Bacidia polychroa is another widespread species of Bacidia s. str., occurring in Europe, North and South America and Asia (Ekman Reference Ekman1996; Llop Reference Llop2007; Smith et al. Reference Smith, Aptroot, Coppins, Fletcher, Gilbert, James and Wolseley2009). Bacidia albogranulosa is known from the Czech Republic, Poland, Ukraine and the Caucasus (Malíček et al. Reference Malíček, Palice, Vondrák, Łubek and Kukwa2018). Conversely, Bacidia diffracta and B. sachalinensis are known only as endemics from North America and the Russian Far East, respectively. Bacidia polychroa is particularly common in stream valleys but also inhabits humid lower and middle mountain forest belts of the north-western Caucasus. It is the most abundant Bacidia species in these forest belts of the north-western Caucasus and is usually found in shaded places with the young regrowth of coppiced trees.
We included 16 specimens of B. polychroa from the Caucasus in our phylogeny, encompassing its observed morphological variation. The nrITS analysis also included a European sequence (Fig. 2) and resulted in a well-supported clade of B. polychroa in the multilocus phylogeny. All taxa within the Polychroa group share the pigment Polychroa-brown in the hypothecium, exciple and hymenium, and a distinguishing K+ purplish reaction in apothecial cross-sections. This purple reaction is unknown for B. albogranulosa since it is known only in its sterile form (Malíček et al. Reference Malíček, Palice, Vondrák, Łubek and Kukwa2018) and was also not observed in the sister JG188, represented by pale or almost unpigmented apothecia. The specimen JG188 is characterized by its warted thallus similar to B. polychroa, but has very bright apothecia, which do not have a K+ purplish reaction. The lack of a K+ purplish reaction makes it easily confused with B. fraxinea. Other specimens of B. polychroa from the large clade also contained pale or unpigmented apothecia lacking the characteristic K+ purplish reaction. Therefore, despite distinct morphological characters, we did not find enough evidence to support the circumscription of a new species in this study.
Taxonomy
Bacidia caucasica Gerasimova, Otte & A. Beck sp. nov.
MycoBank No.: MB 847536
Similar to Bacidia suffusa but differs by abundant colourless crystals above the exciple edge and upper hymenium, and the prominent yellow coloration of the apothecia with darker, almost black, thinner margin.
Type: Russia, Adygea, Maykopskiy Rayon, im Sachraital, 44.12ʹN, 40.33ʹE, 715 m a.s.l., on the bark of Corylus avellana, 17 September 2015, V. Otte s. n. (GLM-L-0048447—holotype).
(Fig. 5)
Cross-section of apothecia and thallus structure of Bacidia caucasica (GLM-0048447, holotype). A, general overview of apothecia and thallus structure with the distinct black prothallus. B, detail view of apothecia and thallus structure, yellow apothecia with a dark pruinose margin. C, clusters of crystals in the upper part of hymenium. D, cross-section of apothecia with detailed exciple structure. E, acicular multiseptate spore. Scales: A = 3 cm; B = 1 cm; C–D = 100 μm; E = 10 μm. In colour online.
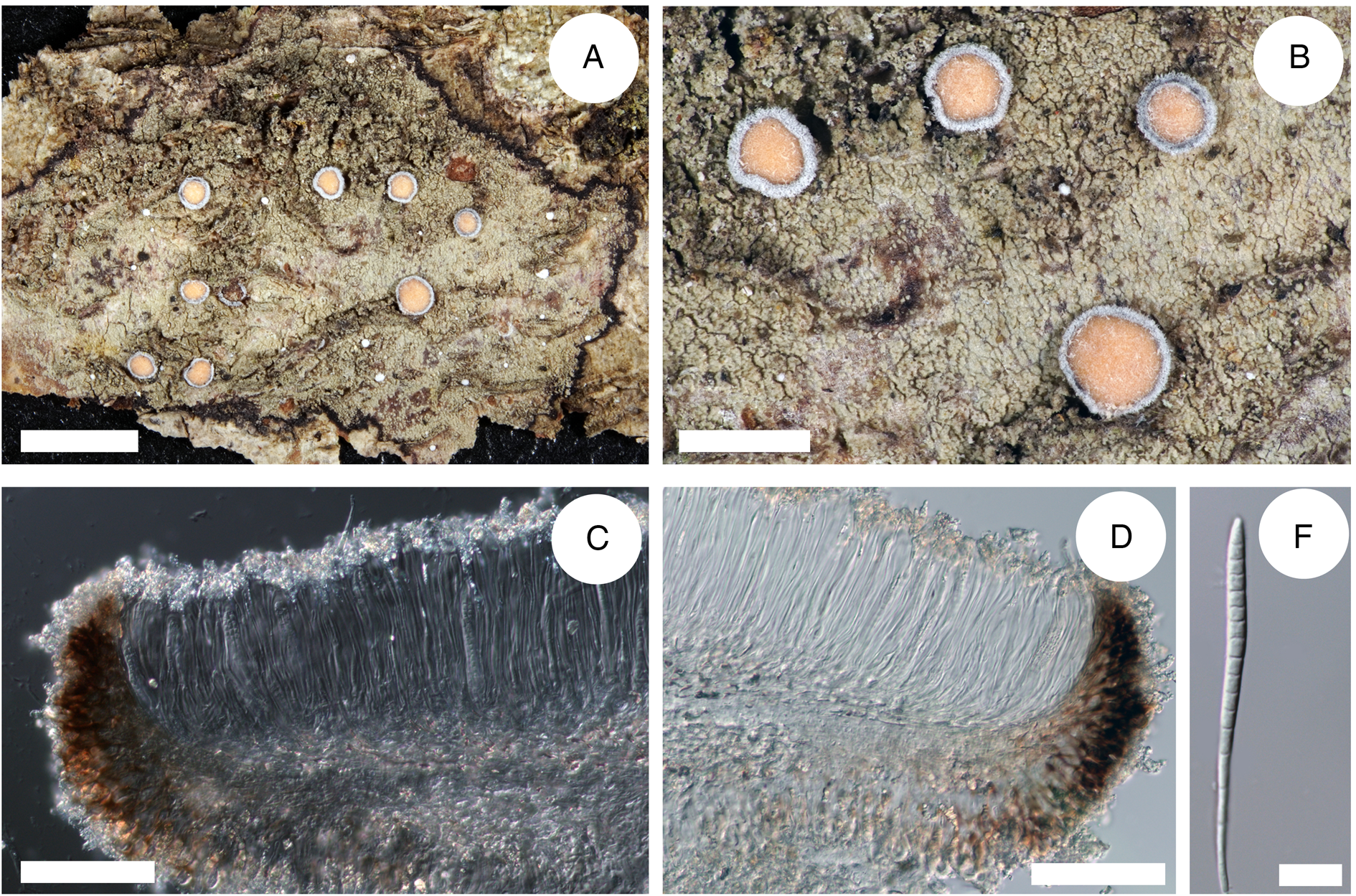

Thallus indeterminate, thin, rimose, partly smooth, mainly consisting of single or contiguous, more or less roundish or irregularly shaped warts. Warts ±flat, adnate to and only slightly raised above the surface; when spreading on mosses, the thallus is more or less granular; green-grey to dark green. Prothallus presents as a black line bordering the thallus. Photobiont chlorococcoid.
Apothecia 0.7–1.1 mm diam. (n 1 = 1, n 2 = 8), ±flat or with a margin slightly above the disc. Disc dark yellow to orange, slightly pruinose. Margin dark brown to black, covered by thick white pruina. Hymenium 84–125–150 μm tall (n 1 = 1, n 2 = 5), with crystals in the upper part (from pruina) not dissolving in K and N. Epithecium pale orange, almost colourless. Hypothecium straw-coloured, almost colourless. Exciple 37–42.5 μm wide (n 1 = 1, n 2 = 4), without or with minor crystals along the rim (from pruina) not dissolving in K and N. Rim dark brown, with two layers of enlarged more or less globose lumina cells, 5–8 μm wide and 7–10 μm long (n 1 = 55, n 2 = 10); the lateral part brown to dark brown, paler closer to hymenium, with crystals less than 0.5 μm or up to 5 μm, not dissolving in K and N. Medullary part under hypothecium pale straw to almost colourless. Paraphyses simple, 1–2.5 μm wide with apices ±clavate or not swollen. Ascospores acicular, (48–)57.8 ± 9.6(–71) μm long and (2–)3.3 ± 0.6(–5) μm wide (n 1 = 1, n 2 = 44), with (3–)10 ± 4(–15) septa (n 1 = 1, n 2 = 44).
Chemistry
Hypothecium and exciple K+ yellow; brown parts of exciple N+ purplish brown.
Pigments
Rubella-orange in epithecium; Laurocerasi-brown in rim and lateral part of exciple.
Etymology
The epithet ‘caucasica’ refers to the locality where the species was collected.
Habitat and distribution
The specimen was collected in a moist, mixed forest in a valley on the bark of Corylus avellana. The species is so far known only from a single collection in the Caucasus area.
Comments
Bacidia caucasica differs from the closely related B. suffusa primarily by abundant colourless crystals along the exciple edge and upper hymenium, the prominent yellow coloration of the apothecia with darker, almost black, thinner margin, and a thallus consisting of single or contiguous, more or less roundish or irregularly shaped warts (Fig. 5).
Bacidia heterochroa (Müll. Arg.) Zahlbr.
Bacidia heterochroa is distributed throughout the tropics and subtropics (Ekman Reference Ekman1996). It is recorded in India, Central and South America, Thailand, Macaronesia and North America (Ekman Reference Ekman1996; Breuss Reference Breuss2001, Reference Breuss2018; Aptroot et al. Reference Aptroot, Saipunkaew, Sipman, Sparrius and Wolseley2007; Diederich & Lawrey Reference Diederich and Lawrey J2007; Joseph & Sinha Reference Joseph and Sinha G2012; Etayo & Berger Reference Etayo and Berger2013; Aptroot & Spielmann Reference Aptroot and Spielmann2020), and it has recently been found in South Korea (Lee & Hur Reference Lee and Hur2022). Here, we report B. heterochroa for the Caucasus and Russia for the first time, representing the northernmost subtropical locality for the species. The individuals of B. heterochroa from the Caucasus were collected in the warm-wet Colchic region in a humid floodplain locality. Interestingly, these individuals were nested in the clade, sister to the Suffusa and Schweinitzii groups, which are also particularly abundant in the oceanic and temperate warm climates of Asia and North America (Ekman Reference Ekman1996; Lendemer et al. Reference Lendemer, Harris and Ladd2016).
Bacidia heterochroa collected in the present study is characterized by having one layer of enlarged lumina cells along exciple edge (up to 6 × 8 μm); a thallus whitish, grey to green-grey, consisting of thin adnate warts, rimose; apothecia black or dark brown with a margin of the same colour, paler below; epithecium dark brown, almost black; exciple edge dark red-brown or brown; lateral part pale brown, yellow-brown, brown to reddish brown; medullar part yellow to nearly colourless, paler than hypothecium; hypothecium yellow; hymenium (61.5–)85.6 ± 16.1(–122.5) μm; exciple (49–)68.4 ± 13.4(–88) μm. The characteristics of thallus and apothecia correspond to those provided by Ekman (Reference Ekman1996) for North America. However, the ascospores are shorter (22–56 × 2.5–4.3 μm) with up to 11 septa compared to ascospores of B. heterochroa from North America that are longer (32–73 × 2.5–4.3 μm) with up to 15 septa.
The studied type material of B. heterochroa is characterized by the dark brown, brown-purplish epithecium with a distinct K+ purplish reaction. In addition, the dark brown to black apothecia have brighter brown margins, at least in the lower part towards the base. These characteristics match those of B. heterochroa from the Caucasus. The specimens of B. heterochroa from South Korea have a dark brown epithecium (K+ purplish) and dark brown apothecia with concolourous margins (Lee & Hur Reference Lee and Hur2022). These characters resemble the closely related B. laurocerasi s. str. and probably represent a separate species. Nevertheless, B. heterochroa from the Caucasus was retrieved on a very long branch, suggesting that additional lineages may be missing from the analysis and that further detailed examination of specimens is necessary.
Additional specimens examined
Georgia: Ochamchirskiy Rayon: the Abkhazian Research Forest Experimental Station, 22 viii 2012, J. Gerasimova (LE L-11663, LE L-12975, LE L-12974, M-0182575); ibid., 22 viii 2012, L. Gagarina (LE L-11635, LE L-11621, LE L-11625, LE L-11626, LE L-11979); ibid., 23 viii 2012, L. Gagarina (LE L-11980). Gudautskiy Rayon: Ritsa Strict Nature Reserve, 14 viii 2012, J. Gerasimova (LE L-11656).—Russia: Krasnodarskiy Krai: Schachetal oberhalb von Solochaul, 24 viii 2016, V. Otte (GLM-L-0048864, GLM-L-0048863); ibid., 25 viii 2016, V. Otte (GLM-L-0048909).
Acknowledgments
We are grateful to Elizabeth Joyce (LMU, Munich) for improving the English text. IU and GU thank Nikolay Eskin (Caucasian Nature Biosphere Reserve), Olga Bykhalova (Utrish Nature Reserve) and Liza Khaykharoeva (Erzi Nature Reserve) for supporting their field research. VO is grateful to Vladimir I. Karataev (Maykop) for two decades of field assistance. JG was supported by a BAYHOST fellowship from the Bayerische Staatsministerium für Bildung und Kultus, Wissenschaft und Kunst. The herbarium work at Senckenberg Museum für Naturkunde Görlitz (GLM) was supported by the Senckenberg Taxonomy Grant (2018). Molecular work was supported by a grant to AB from the Bayerische Staatsministerium für Wissenschaft und Kunst via the Staatliche Naturwissenschaftliche Sammlungen Bayerns within the ‘SNSBinnovativ’ framework. Fieldwork was supported by the institutional research project ‘Flora and systematics of algae, lichens, and bryophytes of Russia and phytogeographically important regions of the world’ (no. 121021600184-6) of the Komarov Botanical Institute RAS to IU, and by a grant from the Russian Foundation for Basic Research to GU (no. 15-29-02396). The herbarium collections of the Utrish Reserve (LE) were revised with financial support to IU from the Ministry of Education and Science (Agreement N 075-15-2021-1056).
Author Contribution
AB and JG designed the study; JG performed the experiments and wrote the first draft of the manuscript; VO, IU and GU provided specimens; all authors contributed to the draft and revised the manuscript.
Author ORCIDs
Julia V. Gerasimova, 0000-0002-3212-3596; Irina N. Urbanavichene, 0000-0002-5492-5215; Gennadii P. Urbanavichus, 0000-0003-3222-5151; Andreas Beck, 0000-0003-2875-4464.
Supplementary Material
The Supplementary Material for this article can be found at https://doi.org/10.1017/S0024282923000385.