Introduction
Leishmaniasis is a complex infectious disease caused by unicellular parasites of the genus Leishmania and has become a huge burden on many of the undeveloped and developing tropical countries worldwide (Desjeux, Reference Desjeux1996; World Health Organisation, 2023). The microbial promastigote stage of this parasite is transmitted to mammalian hosts, including humans, by Phlebotomus or Lutzomyia sand flies (Killick-Kendrick, Reference Killick-Kendrick1999; Burza et al., Reference Burza, Croft and Boelaert2018). Leishmaniasis causes approximately 1–2 million cases and more than 20 000 deaths annually, while 350 million people are at risk (World Health Organization, 2010; Alvar et al., Reference Alvar, Vélez, Bern, Herrero, Desjeux, Cano, Jannin and de Boer2012; PAHO/WHO Leishmaniasis Fact Sheet, 2017; World Health Organisation, 2023). Impoverished people and dearth of healthcare facilities have become the major instigators of the disease, which have exacerbated the current risks of acquiring this disease to a significant level (Burza et al., Reference Burza, Croft and Boelaert2018; Selvapandiyan et al., Reference Selvapandiyan, Croft, Rijal, Nakhasi and Ganguly2019; World Health Organisation, 2023).
There are different clinical manifestations of leishmaniasis viz. visceral (VL), cutaneous (CL), and mucosal, whereas the most life-threatening and commonly reported cases are VL and CL forms, respectively (Burza et al., Reference Burza, Croft and Boelaert2018; PAHO/WHO Leishmaniasis Fact Sheet, 2017). VL, which is abundant in Africa, Brazil and India, causes severe damage to the reticuloendothelial system with dissemination of parasites, and more than 95% of cases ending up fatally, if left untreated (Aronson et al., Reference Aronson, Herwaldt, Libman, Pearson, Lopez-Velez, Weina, Carvalho, Ephros, Jeronimo and Magill2017; World Health Organisation, 2023). About 95% of CL occur in the Americas, Asia, and the Mediterranean basin, and the majority of mucocutaneous cases are reported in countries such as Ethiopia, Bolivia, Peru, and Brazil (Burza et al., Reference Burza, Croft and Boelaert2018). Thus, the widespread nature, and the potential risk of the outbreak of this disease necessitate expeditious disease control measures and prevention campaigns, along with rapid diagnosis, increased public awareness, and effective treatment strategies.
A number of different treatment strategies, like chemotherapy, thermotherapy, and cryotherapy are currently being used to treat leishmaniasis globally, but antimony (Sb) is the mainstay of treatment (Guerin et al., Reference Guerin, Olliaro, Sundar, Boelaert, Croft, Desjeux, Wasunna and Bryceson2002; Silva et al., Reference Silva, Liyanage, Deerasinghe, Chandrasekara, Chellappan and Karunaweera2021a, Reference Silva, Liyanage, Deerasinghe, Sumanasena, Munidasa, de Silva, Weerasingha, Fernandopulle and Karunaweera2021b; Madusanka et al., Reference Madusanka, Silva and Karunaweera2022). Sodium stibogluconate (SSG, pentostam) and meglumine antimoniate (MA, glucantime) are the 2 major medicaments of pentavalent Sb (Sb(V))based drugs in use (Guerin et al., Reference Guerin, Olliaro, Sundar, Boelaert, Croft, Desjeux, Wasunna and Bryceson2002; Haldar et al., Reference Haldar, Sen and Roy2011). The most effective dosage of Sb(V) is 20 mg kg−1 day−1 for 20–28 days, and the injections may cause localized pain (Madusanka et al., Reference Madusanka, Silva and Karunaweera2022). Initially, antimonials were tremendously successful; however, the responsiveness has dwindled over the decades of use, and their current therapeutic prospects appear dim (Silva et al., Reference Silva, Liyanage, Deerasinghe, Chandrasekara, Chellappan and Karunaweera2021a, Reference Silva, Liyanage, Deerasinghe, Sumanasena, Munidasa, de Silva, Weerasingha, Fernandopulle and Karunaweera2021b). The increased drug unresponsiveness in leishmaniasis is attributed to the inappropriate use of drug regimens, resulting in progressive drug tolerance in parasites (Sundar et al., Reference Sundar, Thakur, Tandon, Agrawal, Mishra, Mahaptra and Singh1994). Both the metal-containing-drug activity and the emergence of resistance in Leishmania are closely associated with the trypanothione-based thiol metabolism (Mukhopadhyay et al., Reference Mukhopadhyay, Dey, Xu, Gage, Lightbody, Ouellette and Rosen1996; Ouellette et al., Reference Ouellette, Drummelsmith and Papadopoulou2004; Krauth-Siegel and Comini, Reference Krauth-Siegel and Comini2008; Monte-Neto et al., Reference Monte-Neto, Coelho, Raymond, Légaré, Corbeil, Melo, Frézard and Ouellette2011), and the resistance has been particularly linked to increased Sb detoxification and sequestration (Moreira et al., Reference Moreira, Monte Neto, Andrade, Santi, Reis, Frézard and Murta2013; Gazanion et al., Reference Gazanion, Fernández-Prada, Papadopoulou, Leprohon and Ouellette2016; Dumetz et al., Reference Dumetz, Cuypers, Imamura, Zander, D'Haenens, Maes, Domagalska, Clos, Dujardin and De Muylder2018). Apart from that plethora of genes, protein functions and metabolic pathways are interconnected with the arousal of Sb unresponsiveness, which is of greatest concern for its epidemiology and threatens to undermine disease control efforts. The differential gene expression and genetic modifications are of paramount importance for Leishmania in bringing about drug resistance, and such discrepancies are informative in predicting possible drug responses (Carter et al., Reference Carter, Hutchison, Henriquez, Légaré, Ouellette, Roberts and Mullen2006; Kumar et al., Reference Kumar, Sisodia, Misra, Sundar, Shasany and Dube2010; Torres et al., Reference Torres, Adaui, Ribeiro-Alves, Romero, Arévalo, Cupolillo and Dujardin2010; Biyani et al., Reference Biyani, Singh, Mandal, Chawla and Madhubala2011; Adaui et al., Reference Adaui, Castillo, Zimic, Gutierrez, Decuypere, Vanaerschot, de Doncker, Schnorbusch, Maes, van der Auwera, Maes, Llanos-Cuentas, Arevalo and Dujardin2011a, Reference Adaui, Schnorbusch, Zimic, Gutirrez, Decuypere, Vanaerschot, De Doncker, Maes, Llanos-Cuentas, Chappuis, Arvalo and Dujardin2011b; Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018; Ghosh et al., Reference Ghosh, Verma, Kumar, Pradhan, Selvapandiyan, Salotra and Singh2020, Reference Ghosh, Kumar, Verma, Sharma, Pradhan, Selvapandiyan, Salotra and Singh2022). Often, the Sb resistance is accompanied by the transcriptional modifications of a certain set of genes that collaboratively interfere with the therapeutic effect of Sb detoxification through the incorporation of its active form into conjugates, and diminishing the intracellular Sb build up (Haimeur et al., Reference Haimeur, Brochu, Genest, Papadopoulou and Ouellette2000). For example, Patino et al. demonstrated the presence of more than 800 differentially expressed genes in Sb-resistant and -sensitive Leishmania (Patino et al., Reference Patino, Imamura, Cruz-Saavedra, Pavia, Muskus, Méndez, Dujardin and Ramírez2019). Moreover, a proteomic study quantitatively evaluated the Sb-resistant and -susceptible isolates of Leishmania donovani, whose genes were differentially expressed in relation to stress-related pathways, intracellular survival, and other key metabolic pathways (Biyani et al., Reference Biyani, Singh, Mandal, Chawla and Madhubala2011). Apart from that, studies have reported differential gene expression in Sb resistance (Walker et al., Reference Walker, Gongora, Vasquez, Drummelsmith, Burchmore, Roy, Ouellette, Gomez and Saravia2012; Das et al., Reference Das, Shah, Tandon, Yadav, Sahasrabuddhe, Sundar, Siddiqi and Dube2015; Andrade et al., Reference Andrade, Gonçalves, Liarte, Lima, Guimarães, de Melo Resende, Santi, de Oliveira, Velloso, Delfino, Pescher, Späth, Ruiz and Murta2020).
Although there are a multitude of findings published on Sb resistance-related gene expression, indicating both parallel and contradictory observations, an overall discussion based on the findings of individual studies is warranted to determine the collective scientific significance. In this review, we strived to summarize the variations of Sb resistance-related gene expressions in Leishmania with reference to their relative abundances in terms of mRNA or protein level fluctuations. Furthermore, this study will disclose intriguing areas related to the battle against Sb resistance, which would help in navigating future research towards more productive discoveries in disease control and prevention of leishmaniasis.
Molecular basis of Sb effect and resistance
For more than 6 decades, Sb was the first line of treatment against all forms of leishmaniasis that showed high efficacy (Haldar et al., Reference Haldar, Sen and Roy2011; Negera et al., Reference Negera, Gadisa, Hussein, Engers, Kuru, Gedamu and Aseffa2012). According to the pro-drug model, Sb(V) reduces to its active trivalent state (Sb(III)) by trypanothione (T(SH)2), the most effective intracellular thiol in Leishmania parasites (Dos Santos Ferreira et al., Reference Dos Santos Ferreira, Silveira Martins, Demicheli, Brochu, Ouellette and Frézard2003), either within the host cell (López et al., Reference López, Aguilar, Mercado, Bravo and Quiroz2015), prior to importation into the parasite, or within the parasite itself (Shaked-Mishant et al., Reference Shaked-Mishant, Ulrich, Ephros and Zilberstein2001; Denton et al., Reference Denton, McGregor and Coombs2004; Zhou et al., Reference Zhou, Messier, Ouellette, Rosen and Mukhopadhyay2004; Haldar et al., Reference Haldar, Sen and Roy2011). Moreover, host macrophage thiols like glutathione (GSH) and glycylcysteine are also known to achieve non-enzymatic Sb reduction (Dos Santos Ferreira et al., Reference Dos Santos Ferreira, Silveira Martins, Demicheli, Brochu, Ouellette and Frézard2003). Antimonials have been found to enter the parasite cells via phosphate transporters (Rosen, Reference Rosen2002). Most notably, this molecular reduction followed by the production of more toxic Sb(III), which exerts a lethal effect on Leishmania, is stage-specific as it predominantly takes place in the amastigotes compared to the promastigotes, which in turn elucidates the comparatively higher Sb(V) susceptibility of amastigotes (Callahan et al., Reference Callahan, Roberts, Rainey and Beverley1994; Ephros et al., Reference Ephros, Bitnun, Shaked, Waldman and Zilberstein1999; Shaked-Mishant et al., Reference Shaked-Mishant, Ulrich, Ephros and Zilberstein2001; Goyard et al., Reference Goyard, Segawa, Gordon, Showalter, Duncan, Turco and Beverley2003). The exact mechanism of the therapeutic action is an enigma, and it is believed that Sb induces parasite cell apoptosis (Fig. 1) through genomic DNA degradation, accumulation of reactive oxygen species (ROS) and nitric oxide, diminishing mitochondrial potential, and increasing intracellular Ca2+ (Sereno et al., Reference Sereno, Holzmuller, Mangot, Cuny, Ouaissi and Lemesre2001; Lee et al., Reference Lee, Bertholet, Debrabant, Muller, Duncan and Nakhasi2002; Sudhandiran and Shaha, Reference Sudhandiran and Shaha2003; Basu et al., Reference Basu, Mookerjee, Sen, Bhaumik, Sen, Banerjee, Naskar, Choudhuri, Saha, Raha and Roy2006; Vergnes et al., Reference Vergnes, Gourbal, Girard, Sundar, Drummelsmith and Ouellette2007; Moreira et al., Reference Moreira, Leprohon and Ouellette2011; Garg and Goyal, Reference Garg and Goyal2015). These impose a lethal stress on parasites by inhibiting macromolecular synthesis and energy metabolism to diminish their vital metabolic pathways, together with the interruption of glycolysis and fatty acid oxidation, which ultimately lead to the death of parasites (Berman et al., Reference Berman, Waddel and Hanson1985; Herman et al., Reference Herman, Gallalee and Best1987).
Sb metabolism and related gene expression in wild-type Leishmania amastigote. TR, trypanothione reductase; TDR1, thiol-dependent reductase 1; γ-GCS, gamma-glutamylcysteine synthetase; MRPA, multidrug-resistant protein A; AQP1, aquaglyceroporin 1; ODC, ornithine decarboxylase; TryP, tryparedoxin peroxidase; MAPK, mitogen-activated protein kinase; GSH, glutathione; T(SH)2, trypanothione; TS2, trypanothione disulphide; ROS, reactive oxygen species.

Sb resistance triggered gene expression
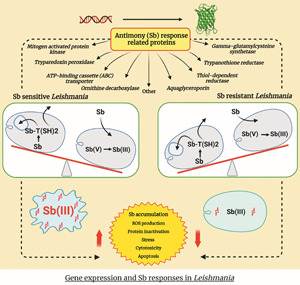
The exact molecular mechanisms and biochemistry of Sb resistance in Leishmania still remain ambiguous and yet to be expounded (Fernandez-Prada et al., Reference Fernandez-Prada, Sharma, Plourde, Bresson, Roy, Leprohon and Ouellette2018). However, the mostly argued mechanism of Sb resistance is linked to the increased Sb(III) detoxification and sequestration (Ashutosh et al., Reference Ashutosh, Sundar and Goyal2007; Garg and Goyal, Reference Garg and Goyal2015; Gazanion et al., Reference Gazanion, Fernández-Prada, Papadopoulou, Leprohon and Ouellette2016), which subsequently result in reduced Sb accumulation within parasites (Fig. 2) (Ouellette and Papadopoulou, Reference Ouellette and Papadopoulou1993; Ouellette et al., Reference Ouellette, Drummelsmith and Papadopoulou2004). Furthermore, the formation of Sb–trypanothione conjugates in the presence of excess trypanothione and its rapid extrusion are largely exploited for Sb resistance in Leishmania (Mukhopadhyay et al., Reference Mukhopadhyay, Dey, Xu, Gage, Lightbody, Ouellette and Rosen1996; Rai et al., Reference Rai, Bhaskar, Goel, Nath Dwivedi, Sundar and Goyal2013). There were consistently high thiol levels in Sb-resistant and genetically different clinical isolates of Leishmania (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022).
Comparison of Sb metabolism and related gene expression in Sb-resistant vs -sensitive amastigotes. The vertical red and blue arrows indicate gene upregulation and downregulation, respectively. The purple vertical arrows exhibit the increase/decrease of each component. TR, trypanothione reductase; TDR1, thiol-dependent reductase 1; γ-GCS, gamma-glutamylcysteine synthetase; MRPA, multidrug-resistant protein A; AQP1, aquaglyceroporin; ODC, ornithine decarboxylase; TryP, tryparedoxin peroxidase; MAPK, mitogen-activated protein kinase; T(SH)2, trypanothione; TS2, trypanothione disulphide; ROS, reactive oxygen species. The (i1), (i2), (i3), (a1), (a2), and (a3) are mechanisms that affect the intracellular Sb concentration (A): (i1) decreased Sb influx; (i2) decreased Sb(V) to Sb(III) conversion; (i3) decreased T(SH)2 synthesis; (ii) decreased Sb–thiol conjugate formation; (iii) increased Sb(III) accumulation; (iv) decreased cell apoptosis. (B) (a1) decreased Sb influx; (a2) increased Sb(V) to Sb(III) conversion; (a3) increased T(SH)2 synthesis; (b) increased Sb–thiol conjugate formation; (c) decreased Sb(III) accumulation; (d) increased cell apoptosis

Sb resistance in Leishmania is markedly associated with the expression of proteins related to Sb reduction, transport, and thiol synthesis (Table 1) (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022). Recent publications revealed 844 and 803 differentially expressed genes between Sb(SSG)-resistant and -sensitive Leishmania braziliensis and Leishmania panamensis, respectively, with over 100 genes showing ⩾2-fold change in resistant lines of each strain (Patino et al., Reference Patino, Imamura, Cruz-Saavedra, Pavia, Muskus, Méndez, Dujardin and Ramírez2019). Sb-resistant overexpression of transcripts assigned in the gene ontology categories such as ubiquitination, host–parasite interaction, protein phosphorylation, microtubule-based movement, and cellular process and the downregulated processes were rRNA processing, ribosome biogenesis, ribonucleoprotein complex, nucleosome assembly, and translation (Andrade et al., Reference Andrade, Gonçalves, Liarte, Lima, Guimarães, de Melo Resende, Santi, de Oliveira, Velloso, Delfino, Pescher, Späth, Ruiz and Murta2020). In addition, most of the differentially expressed proteins in sodium antimony gluconate (SAG)-sensitive isolate AG83-S vs SAG-resistant GE1-R were related to translation and metabolic enzymes (Biyani et al., Reference Biyani, Singh, Mandal, Chawla and Madhubala2011).
List of Sb resistance-related genes in Leishmania

Copy number variation (CNV) is among the crucial molecular mechanisms deployed by Leishmania to increase the transcript levels of resistance genes. CNV involves the duplication of either specific genomic region or complete chromosomes as intra- or extrachromosomal elements (Ullman, Reference Ullman1995; Leprohon et al., Reference Leprohon, Fernandez-Prada, Gazanion, Monte-Neto and Ouellette2015; Papadopoulou et al., Reference Papadopoulou, Ouellette, Laffitte and Leprohon2016). Moreover, gene amplification that occurs in a part of the amplicon or intrachromosomal level has also been observed in differential gene expression, which leads to metal-drug resistance in Leishmania including Sb stress (Grondin et al., Reference Grondin, Haimeur, Mukhopadhyay, Rosen and Ouellette1997; Leprohon et al., Reference Leprohon, Légaré, Raymond, Madore, Hardiman, Corbeil and Ouellette2009; Mukherjee et al., Reference Mukherjee, Boisvert, do Monte-Neto, Coelho, Raymond, Mukhopadhyay, Corbeil and Ouellette2013). The presence of single-nucleotide polymorphisms (SNPs) in genes encoding functional and structural proteins related to Sb resistance is known to regulate Leishmania resistance towards chemotherapy (Downing et al., Reference Downing, Imamura, Decuypere, Clark, Coombs, Cotton, Hilley, De Doncker, Maes, Mottram, Quail, Rijal, Sanders, Schönian, Stark, Sundar, Vanaerschot, Hertz-Fowler, Dujardin and Berriman2011; Coelho et al., Reference Coelho, Boisvert, Mukherjee, Leprohon, Corbeil and Ouellette2012; Rastrojo et al., Reference Rastrojo, García-Hernández, Vargas, Camacho, Corvo, Imamura, Dujardin, Castanys, Aguado, Gamarro and Requena2018). The presence of 3 SNPs in serine acetyltransferase, the protein involved in cysteine synthesis, exhibited increased Sb resistance in Leishmania infantum mutants having impaired adenosine triphosphate (ATP)-binding cassette (ABC) transporters (Douanne et al., Reference Douanne, Wagner, Roy, Leprohon, Ouellette and Fernandez-Prada2020). Further, an SNP that occurred in protein kinase in L. infantum has an influence on Sb resistance (Brotherton et al., Reference Brotherton, Bourassa, Leprohon, Légaré, Poirier, Droit and Ouellette2013). The resistance phenotype of Leishmania is a final product of various cellular mechanisms, including gene overexpression (supplementary material) accompanied by preadaptations like structural and functional modulations.
ABC transporter (multidrug-resistant protein A)
The ABC transporter superfamily consists of functionally Sb-resistant proteins of Leishmania that have exerted an Sb detoxification potential via direct membrane efflux of Sb to the extracellular milieu (El Fadili et al., Reference El Fadili, Messier, Leprohon, Roy, Guimond, Trudel, Saravia, Papadopoulou, Légaré and Ouellette2005; Singh et al., Reference Singh, Chatterjee and Sundar2014; Douanne et al., Reference Douanne, Wagner, Roy, Leprohon, Ouellette and Fernandez-Prada2020). Multidrug-resistant protein A (MRPA, also known as ABCC3) is one of the ABC transporters, formerly identified as a P-glycoprotein (Ouellette et al., Reference Ouellette, Légaré, Haimeur, Grondin, Roy, Brochu and Papadopoulou1998), and has been reported to be a Leishmania intracellular protein found in membrane vesicles near the flagellar pocket, at the sites of endo- and exocytosis of the parasite (Fig. 1) (El Fadili et al., Reference El Fadili, Messier, Leprohon, Roy, Guimond, Trudel, Saravia, Papadopoulou, Légaré and Ouellette2005; Ashutosh et al., Reference Ashutosh, Sundar and Goyal2007). Previous literature has amply demonstrated the inevitable role of MRPA in Sb resistance via intracellular sequestration of Sb–thiol conjugates into vesicles (Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007; Moreira et al., Reference Moreira, Monte Neto, Andrade, Santi, Reis, Frézard and Murta2013; Singh et al., Reference Singh, Chatterjee and Sundar2014; Gazanion et al., Reference Gazanion, Fernández-Prada, Papadopoulou, Leprohon and Ouellette2016). In addition, MRPA expression resulted in an increased drug resistance in L. donovani in relation to altered fluidity in the cell membranes and decreased drug accumulation (Bhandari et al., Reference Bhandari, Sundar, Dujardin and Salotra2014). Previously, MRPA was suggested as one of the 2 prediction models for determining Sb treatment failure that could predict the treatment outcome with high accuracy (Torres et al., Reference Torres, Ribeiro-Alves, Romero, Dávila and Cupolillo2013). Hence, MRPA expression could be a crucial characteristic of Sb resistance (Fig. 2). Interestingly, L. donovani was able to develop SSG resistance even under arsenic (As) stress because both Sb and As pressures could trigger the same overexpression of the ABC transporter (Perry et al., Reference Perry, Wyllie, Raab, Feldmann and Fairlamb2013).
Extrachromosomal amplification within circular amplicons of MRPA has been extensively studied in different Leishmania species (Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007; Leprohon et al., Reference Leprohon, Légaré, Raymond, Madore, Hardiman, Corbeil and Ouellette2009; Moreira et al., Reference Moreira, Monte Neto, Andrade, Santi, Reis, Frézard and Murta2013). Likewise, the adaptive gene amplification of MRPA observed in L. infantum during in vitro Sb(III) selection corroborates its significance in Sb tolerance (Ubeda et al., Reference Ubeda, Raymond, Mukherjee, Plourde, Gingras, Roy, Lapointe, Leprohon, Papadopoulou, Corbeil and Ouellette2014). More importantly, MRPA amplification confers the first line of defence against Sb(III) stress in Leishmania, providing the driving force for the inception of underlying molecular adaptations upon an infection and signalling pathways (Dumetz et al., Reference Dumetz, Cuypers, Imamura, Zander, D'Haenens, Maes, Domagalska, Clos, Dujardin and De Muylder2018). Therefore, its expression level during drug pressure could be a determinant of the parasites' destiny as well as providing strong insights upon the subsequent development of Sb resistance. Beyond Sb transportation, an indirect correlation was explored between Leishmania MRPA expression and cellular redox homoeostasis that was affected by glucose-6-phosphate dehydrogenase and trypanothione reductase (TR) interaction upon metalloid exposure, including Sb (Ghosh et al., Reference Ghosh, Saini, Das, Mandal, Sardar, Ansari, Abhishek, Kumar, Singh, Verma, Equbal, Ali and Das2017). Dumetz et al. demonstrated the importance of overexpression of the H locus, which harbours the MRPA gene, over M locus and increases the Sb resistance in 3-fold (Dumetz et al., Reference Dumetz, Cuypers, Imamura, Zander, D'Haenens, Maes, Domagalska, Clos, Dujardin and De Muylder2018).
MRPA was expressed in Leishmania with a direct correlation to Sb-resistant phenotype, and it was widely expressed in most of the parasite and clinical forms of leishmaniasis (Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007; Barrera et al., Reference Barrera, Rojas, Weiss, Fernandez, McMahon-Pratt, Saravia and Gomez2017; Fekrisoofiabadi et al., Reference Fekrisoofiabadi, Fekri, Moradabadi, Vahidi, Khaleghi, Ram and Dabiri2019). For example, the MRPA expression was found to be markedly augmented in Sb-resistant Indian isolates of L. donovani and Leishmania tropica, respectively, compared to their sensitive counterparts (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022). Furthermore, 2 independent methods viz. cDNA-amplified fragment length polymorphism approach and quantitative polymerase chain reaction (qPCR) analysis demonstrated approximately similarly augmented expressions (2–3-fold) of MRPA in Sb-resistant clinical anthroponotic cutaneous leishmaniasis (ACL) isolates of L. tropica compared to the sensitive counterparts (Kazemi-Rad et al., Reference Kazemi-Rad, Mohebali, Erfan, Saffari, Raoofian, Hajjaran, Hadighi, Khamesipour, Rezaie, Abedkhojasteh and Heidari2013; Mohebali et al., Reference Mohebali, Kazemirad, Hajjaran, Kazemirad, Oshaghi, Raoofian and Teimouri2019). An antibody assay recognized high MRPA levels in Leishmania guyanensis and Leishmania amazonensis-resistant lines, but the detection was not successful for the sensitive lines, which further demonstrated differential MRPA expression between 2 phenotypes (Moreira et al., Reference Moreira, Monte Neto, Andrade, Santi, Reis, Frézard and Murta2013). Moreover, a full genome microarray hybridization in L. amazonensis showed a robust (5-fold) MRPA expression in Sb-resistant parasites compared to its wild-type (Monte-Neto et al., Reference Monte-Neto, Coelho, Raymond, Légaré, Corbeil, Melo, Frézard and Ouellette2011). Interestingly, the obstruction of MRPA expression conferred increased drug susceptibilities in Leishmania; for instance, the MRPA null mutants of L. infantum promastigotes exhibited drastic declines (20-fold) in their half maximal inhibitory concentration (IC50) values against Sb(III), whereas the corresponding amastigotes showed increased sensitivity to Sb(V) compared to their wild-type (Douanne et al., Reference Douanne, Wagner, Roy, Leprohon, Ouellette and Fernandez-Prada2020). Appropriately, following the selection of Sb(III) resistance, the transcript levels of sensitive parasites in a study were elevated, reaching a 1.5–3.0-fold expression as same as the resistant parasites (Dumetz et al., Reference Dumetz, Cuypers, Imamura, Zander, D'Haenens, Maes, Domagalska, Clos, Dujardin and De Muylder2018). This observation affirms that the selective drug pressure is able to provoke Sb resistance in Leishmania through MRPA-mediated mechanisms as a preadaptation of parasites to harsh conditions, thus, it is salient to indicate that the overexpression of this protein may enable the parasites to withstand or weaken the Sb therapeutic effect as a successful counter mechanism. In addition, studies have reported adaptive expression of MRPA in an Sb concentration-dependent manner, whereas an initial elevation of the copy number was seen in all the Sb(III)-resistant mutants, and it was gradually decreased to the wild-type level in the subsequent several passages in the absence of drug pressure (Haimeur et al., Reference Haimeur, Brochu, Genest, Papadopoulou and Ouellette2000). This would further inform the drug pressure-induced overexpression and the significance of MRPA-mediated pathways to achieve Sb resistance in certain instances.
Contrariwise, MRPA-independent resistance mechanisms were also possible, in which it was not ubiquitously upregulated in all the Sb-resistant isolates of a study (Moreira et al., Reference Moreira, Anacleto and De Petrillo-Peixoto1998; Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007). Accordingly, MRPA-independent Sb resistance was accompanied by unchanged mRNA levels in resistant Leishmania, whereas it was sometimes considered a protein without an important role in Sb transportation (Dos Reis et al., Reference Dos Reis, do Monte-Neto, Melo and Frézard2017) or a non-essential transcript for Sb resistance; however, the disruption of this protein triggered Sb hypersensitivity in both amastigotes and promastigotes of L. infantum (Douanne et al., Reference Douanne, Wagner, Roy, Leprohon, Ouellette and Fernandez-Prada2020). The presence of a high resistance index was found to be essential for the upregulation of MRPA, while the energy-dependent Sb resistance pathway of resistant mutants did not rely on the upregulation of this gene (Rai et al., Reference Rai, Bhaskar, Goel, Nath Dwivedi, Sundar and Goyal2013; Dos Reis et al., Reference Dos Reis, do Monte-Neto, Melo and Frézard2017). In a study related to L. donovani, Kumar et al. observed upregulation of MRPA in 6 resistant and 1 sensitive isolate with no significant elevation of its expression in another 4 resistant isolates of the same species (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012). Moreover, the MRPA level of L. panamensis was augmented only in in vitro-adapted Sb-resistant strains, and no significant difference was observed in clinically resistant lines (Barrera et al., Reference Barrera, Rojas, Weiss, Fernandez, McMahon-Pratt, Saravia and Gomez2017). This was further corroborated by the studies that have reported no or negligible amplification of MRPA in glucantime-resistant clinical Leishmania isolates (Ullman et al., Reference Ullman, Carrero-Valenzuela and Coons1989; Moreira et al., Reference Moreira, Anacleto and De Petrillo-Peixoto1998; Gómez Pérez et al., Reference Gómez Pérez, García-Hernandez, Corpas-López, Tomás, Martín-Sanchez, Castanys and Gamarro2016). In addition, an MRPA amplification was observed only in 3 out of 4 SAG-resistant isolates, and the rest did not show any sign of amplification (Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007). Therefore, there must be multiple factors behind the MRPA expression under Sb exposure that can modulate its expression. Accordingly, MRPA was a candidate marker for drug resistance with 69% accuracy as a prediction model in determining the treatment outcome of clinical Sb-treatment upon L. braziliensis (CL) (Torres et al., Reference Torres, Ribeiro-Alves, Romero, Dávila and Cupolillo2013).
Of note, many studies have surveyed paradoxical results about MRPA expression linked to its expression in Sb-resistant and -sensitive parasites, which are not satisfactorily resolved yet. Although MRPA seemed to have wide expression in Leishmania, species-specific discrepancies of the expression levels were also possible in Sb resistance. For example, a quantitative reverse transcription (RT)-PCR analysis showed no differential expression in Sb-resistant L. infantum, in spite of 2-fold increased mRNA expression in Sb(III)-resistant isolates of L. guyanensis, L. braziliensis, and L. amazonensis compared to the susceptible lines (Moreira et al., Reference Moreira, Monte Neto, Andrade, Santi, Reis, Frézard and Murta2013). Moreover, increased MRPA levels were seen (1.5–25.2-fold) in Sb(III)-resistant L. braziliensis and Sb(V)-unresponsive L. tropica along with simultaneous expression in respective responsive lines, so it diminishes the possibility of the functional relevance of MRPA in resistance phenotypes (Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018; Rugani et al., Reference Rugani, Gontijo, Frézard, Soares and Do Monte-Neto2019). On the other hand, Victoria et al. demonstrated that sitamaquine can successfully circumvent the Sb resistance caused by MRPA expression (Pérez-Victoria et al., Reference Pérez-Victoria, Bavchvarov, Torrecillas, Martínez-García, López-Martín, Campillo, Castanys and Gamarro2011), and the MRPA-mediated Sb resistance was reverted by buthionine sulphoximine, a GSH biosynthesis-specific inhibitor (El Fadili et al., Reference El Fadili, Messier, Leprohon, Roy, Guimond, Trudel, Saravia, Papadopoulou, Légaré and Ouellette2005). Hence, the obstruction of the interaction between MRPA and GSH may be an effective approach for drug design (Fekrisoofiabadi et al., Reference Fekrisoofiabadi, Fekri, Moradabadi, Vahidi, Khaleghi, Ram and Dabiri2019).
Moreover, Callahan et al. demonstrated an oxidative state-dependent selective Sb resistance in Leishmania major, where the MRPA expression could manifest resistance to Sb(III) and not against Sb(V), albeit with clear evidence for the intracellular conversion of Sb(V) to its reduced form during its mode of action in the parasites (Callahan and Beverley, Reference Callahan and Beverley1991; Dos Santos Ferreira et al., Reference Dos Santos Ferreira, Silveira Martins, Demicheli, Brochu, Ouellette and Frézard2003). Moreover, significant MRPA expression was observed in promastigotes under increased Sb(III) stress compared to the intracellular amastigotes exposed to Sb(V) (Gazanion et al., Reference Gazanion, Fernández-Prada, Papadopoulou, Leprohon and Ouellette2016; Fernandez-Prada et al., Reference Fernandez-Prada, Sharma, Plourde, Bresson, Roy, Leprohon and Ouellette2018). Accordingly, the amastigotes must have evolved pathways to confer drug protection via MRPA-independent mechanisms as well. Collectively, the MRPA expression in Sb resistance is not a consistent event, which is broadly affected by multiple factors.
Aquaglyceroporin (AQP1)
Aquaglyceroporins are a subcategory of aquaporins that primarily involve water and glycerol transportation in mammalian cells (Verkman, Reference Verkman2008; Mukhopadhyay et al., Reference Mukhopadhyay, Bhattacharjee and Rosen2014). In Leishmania, AQP1 has been implicated as a protein that imports Sb(III) into the cells, and its decreased expression has been broadly discussed and attributed to Sb resistance in many studies (Fig. 2A) (Gourbal et al., Reference Gourbal, Sonuc, Bhattacharjee, Legare, Sundar, Ouellette, Rosen and Mukhopadhyay2004; Gómez Pérez et al., Reference Gómez Pérez, García-Hernandez, Corpas-López, Tomás, Martín-Sanchez, Castanys and Gamarro2016). It is predominantly found in the flagellums of the promastigote stage, which is then relocated to the parasite surface eventually after post-translational phosphorylation by the mitogen-activated protein kinase (MAPK) (Mandal et al., Reference Mandal, Sharma, Kruse, Sander-Juelch, Munro, Wang, Vilg, Tamás, Bhattacharjee, Wiese and Mukhopadhyay2012; Sharma et al., Reference Sharma, Mandal, Mandal, Bhattacharjee and Mukhopadhyay2015). Since the AQP1 expression is highly associated with the Sb accumulation in Leishmania, reduced expression or the perturbation of its gene expression has been extensively reported in relation to Sb resistance (Gourbal et al., Reference Gourbal, Sonuc, Bhattacharjee, Legare, Sundar, Ouellette, Rosen and Mukhopadhyay2004; Marquis et al., Reference Marquis, Gourbal, Rosen, Mukhopadhyay and Ouellette2005; Mukherjee et al., Reference Mukherjee, Boisvert, do Monte-Neto, Coelho, Raymond, Mukhopadhyay, Corbeil and Ouellette2013; Sharma et al., Reference Sharma, Mandal, Mandal, Bhattacharjee and Mukhopadhyay2015; Mohebali et al., Reference Mohebali, Kazemirad, Hajjaran, Kazemirad, Oshaghi, Raoofian and Teimouri2019; Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022).
AQP1 expression increased the Sb(III) accumulation in Leishmania compared to the untreated control (Sharma et al., Reference Sharma, Mandal, Mandal, Bhattacharjee and Mukhopadhyay2015); therefore, the suppression of its transcripts is much preferred by the resistant Leishmania and vice versa (Fig. 2) (Kazemi-Rad et al., Reference Kazemi-Rad, Mohebali, Erfan, Saffari, Raoofian, Hajjaran, Hadighi, Khamesipour, Rezaie, Abedkhojasteh and Heidari2013; Douanne et al., Reference Douanne, Wagner, Roy, Leprohon, Ouellette and Fernandez-Prada2020). Accordingly, single-allele disruption or subtelomeric deletion of AQP1 resulted in drastically reduced Sb accumulation in Leishmania, accompanied by prompt Sb resistance (Gourbal et al., Reference Gourbal, Sonuc, Bhattacharjee, Legare, Sundar, Ouellette, Rosen and Mukhopadhyay2004; Monte-Neto et al., Reference Monte-Neto, Laffitte, Leprohon, Reis, Frézard and Ouellette2015). Furthermore, terminally deleted mutants of AQP1 could restore their Sb(III) resistance following the episomal transfection of the gene, through which the IC50 of the mutants (by 20–50-fold) subsequently dropped in a rigorous decline (Mukherjee et al., Reference Mukherjee, Boisvert, do Monte-Neto, Coelho, Raymond, Mukhopadhyay, Corbeil and Ouellette2013). The AQP1 expression in Sb-resistant isolates of ACL showed greater suppression than that of the sensitive strains (Kazemi-Rad et al., Reference Kazemi-Rad, Mohebali, Erfan, Saffari, Raoofian, Hajjaran, Hadighi, Khamesipour, Rezaie, Abedkhojasteh and Heidari2013), and more significantly, a negative correlation was seen between AQP1 expression and the IC50 or time taken to cure ACL lesions in the responsive cases of natural isolates (Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018; Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022). Similarly, the decreased expression of AQP1 in the Sb-resistant clinical isolates of L. donovani and L. tropica was further affirmed by a negative correlation between IC50 and AQP1 expression (Mohebali et al., Reference Mohebali, Kazemirad, Hajjaran, Kazemirad, Oshaghi, Raoofian and Teimouri2019; Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022). Leishmania parasites of CL or post kala-azar dermal leishmaniasis (PKDL) had more robust antimonial accumulation than that of the VL and were more Sb-sensitive, which was rendered so by the elevated expression and mRNA stability of AQP1 (Mishra et al., Reference Mishra, Madhubala and Singh2013; Mandal et al., Reference Mandal, Mandal, Sharma, Charret, Papadopoulou, Bhattacharjee and Mukhopadhyay2015). A downregulation of AQP1 was seen in the majority of Sb(III)-resistant VL and PKDL-derived L. donovani isolates albeit with several exceptions (Mandal et al., Reference Mandal, Maharjan, Singh, Chatterjee and Madhubala2010). Not only that, an atypical form of tegumentary leishmaniasis caused by L. braziliensis showed an outstanding 65-fold downregulation of AQP1 in their clinical isolates than that of the reference strain (Rugani et al., Reference Rugani, Gontijo, Frézard, Soares and Do Monte-Neto2019). In addition, there was a comparable AQP1 expression in the Sb treatment failure isolates of L. major, which was found to be 58.71-fold less than that of the treatment-responsive isolates (Sharma et al., Reference Sharma, Mandal, Mandal, Bhattacharjee and Mukhopadhyay2015). More importantly, the 2 Sb transporters, MRPA and pentamidine resistance protein 1 (PRP1), were found to have increased their expression in clinically resistant parasites along with simultaneous suppression of AQP1 transcripts, with an emphasis on the increased Sb detoxification plus decreased influx (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022). Apart from that, the AQP1 expression in Sb(V)-resistant L. donovani isolated from Nepal and the AQP1 copy number derived from chromosome 31 in the resistant mutants of L. major were found to be lower than that of their sensitive strains (Decuypere et al., Reference Decuypere, Rijal, Yardley, De Doncker, Laurent, Khanal, Chappuis and Dujardin2005). Interestingly, the transfection of AQP1 followed by its increased expression in Leishmania tarentolae, L. major, and L. infantum developed hypersensitivity to metalloids such as As(III) and Sb(III) (Gourbal et al., Reference Gourbal, Sonuc, Bhattacharjee, Legare, Sundar, Ouellette, Rosen and Mukhopadhyay2004). Another study also revealed supportive evidence of similar hypersensitivity in L. major isolates of CL patients, and secondarily, their resistance emerged with the deletion, inactivation through mutation and reduced expression of AQP1 (Eslami et al., Reference Eslami, Hatefi, Ramezani, Tohidfar, Churkina, Orlov, Hosseini, Boozhmehrani and Vakili2021). Conversely, in vitro transfection failed to enhance Sb susceptibility of resistant promastigote lines as well as sensitive lines compared to their respective parent strains (Mandal et al., Reference Mandal, Maharjan, Singh, Chatterjee and Madhubala2010). Leishmania donovani clinical isolates of SAG-resistant and -sensitive parasites showed marked down- and upregulations respectively, whereas the expression difference was more prominent between the amastigote lines than the respective promastigotes. The Sb(V)-resistant L. donovani isolated from Nepal exhibited 6–7-fold significantly lower AQP1 expression than that in the sensitive strains (Decuypere et al., Reference Decuypere, Rijal, Yardley, De Doncker, Laurent, Khanal, Chappuis and Dujardin2005).
There are many disputes among research findings about the AQP1 expression in drug-resistant Leishmania. For instance, Maharjan et al. suggested that the downregulation of AQP1 was just one of the Sb-resistant mechanisms in Leishmania and that not all the resistant ones consistently downregulate it (Maharjan et al., Reference Maharjan, Singh, Chatterjee and Madhubala2008). Further, it was ascertained by the high AQP1 copy number observed in the resistant parasites compared to the sensitive ones, which was not in line with the reduced import of Sb(III) (Maharjan et al., Reference Maharjan, Singh, Chatterjee and Madhubala2008). It was also suggested that AQP1 is not an essential protein for the survival of Leishmania (Plourde et al., Reference Plourde, Ubeda, Mandal, Do Monte-Neto, Mukhopadhyay and Ouellette2015). In vitro-selected Sb-resistant mutants L. braziliensis, L. infantum, and L. guyanensis did not show a significant difference in AQP1 mRNA level compared to the control, in agreement with the absence of its function in Sb transportation in non-natural-resistant mutants (Torres et al., Reference Torres, Adaui, Ribeiro-Alves, Romero, Arévalo, Cupolillo and Dujardin2010; Moreira et al., Reference Moreira, Monte Neto, Andrade, Santi, Reis, Frézard and Murta2013; Dos Reis et al., Reference Dos Reis, do Monte-Neto, Melo and Frézard2017). Interestingly, a study on L. panamensis revealed decreased AQP1 levels of in vitro-adapted Sb-resistant strains, and no significant difference was observed in the clinically resistant lines (Barrera et al., Reference Barrera, Rojas, Weiss, Fernandez, McMahon-Pratt, Saravia and Gomez2017). Therefore, growing evidence has suggested the possibility of AQP1 neutral Sb resistance, and therefore, it has also hinted at the prevalence of many critical cellular functions of these transcripts other than the Sb influx.
In agreement with the wide array of functions achieved by the AQP1, a handful of reports indicate its upregulation without affecting the inherited Sb resistance and noticeably suggest an alternative mechanism of Sb resistance. Leishmania infantum amastigotes with Sb(III) resistance had increased AQP1 expression, which was reverted to the wild-type in the presence of drug pressure (Marquis et al., Reference Marquis, Gourbal, Rosen, Mukhopadhyay and Ouellette2005). Further, L. major parasites isolated from non-healing cases showed increased AQP1 expression (Eslami et al., Reference Eslami, Zarchi, Moradi, Hejazi, Sohrevardi, Vakili and Khamesipour2016; Alijani et al., Reference Alijani, Hosseini, Ahmadian, Boughattas, Eslami, Naderian and Ajamein2019). In a study aiming to investigate the biomarkers of Sb resistance, L. donovani showed a marked AQP1 upregulation in all the selected clinically Sb-sensitive isolates, in comparison to significant downregulation observed in only 30% of resistant ones, whereas others showed similar expression to the wild-type (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012). According to the available evidence on AQP1 expression, it may be a multifunctional protein in Leishmania that is also significantly involved in Sb resistance. Therefore, many elaborate studies are warranted to clearly understand the network of those functions.
Gamma-glutamylcysteine synthetase
Gamma-glutamylcysteine synthetase (γ-GCS, l-glutamate: l-cysteine γ-ligase) catalyses the rate-limiting step of GSH biosynthesis that leads to the trypanothione overexpression (Mukhopadhyay et al., Reference Mukhopadhyay, Dey, Xu, Gage, Lightbody, Ouellette and Rosen1996; Haimeur et al., Reference Haimeur, Brochu, Genest, Papadopoulou and Ouellette2000; Lu, Reference Lu2001). During its mode of action, firstly, γ-GCS triggers the covalent bond formation between glutamate and cysteine to synthesize gamma-glutamylcysteine, which in turn binds with glycine, resulting in GSH formation (Fig. 1) (Olin-Sandoval et al., Reference Olin-Sandoval, Moreno-Sanchez and Saavedra2012). Accordingly, the activity of this protein is controlled by the intracellular GSH levels, and the non-allosteric feedback, as well as the transcriptional and translational factors (Lu, Reference Lu2001). Moreover, γ-GCS overexpression was considered to confer increased virulence, cell viability, and drug resistance in parasites (Pérez-Rosado et al., Reference Pérez-Rosado, Gervais, Ferrer-Rodríguez, Peters and Serrano2002; González-Chávez et al., Reference González-Chávez, Vázquez, Mejia-Tlachi, Márquez-Dueñas, Manning-Cela, Encalada, Rodríguez-Enríquez, Michels, Moreno-Sánchez and Saavedra2019).
The γ-GCS has been reported to be upregulated in Sb-resistant Leishmania and implicated as a protein that triggers Sb-detoxification pathways (Fig. 2A) (Grondin et al., Reference Grondin, Haimeur, Mukhopadhyay, Rosen and Ouellette1997; Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007). For example, an elevated γ-GCS expression was observed in therapeutic failure in L. guyanensis in all the in vitro growth phases of the promastigote (Torres et al., Reference Torres, Adaui, Ribeiro-Alves, Romero, Arévalo, Cupolillo and Dujardin2010). Its expression in Sb-resistant L. major derived from CL patients was 20 times higher than that of the sensitive and it was suggested to be a possible biomarker in the identification of clinical resistance (Ghobakhloo et al., Reference Ghobakhloo, Motazedian and Fardaei2016). Fittingly, there was a positive correlation between γ-GCS expression and the IC50 values of Sb-resistant clinical kala-azar isolates of L. tropica and L. donovani with several-fold overexpression (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022). The γ-GCS was not merely associated with developing Sb resistance; however, its depleted expression was associated with adverse effects on parasites; for instance, the downregulation of these transcripts rendered decreased parasite oxidative defence that made the parasites more susceptible to drug effects, which was in line with the reported upregulation and downregulation in the majority of resistant isolates and the sensitive ones, respectively (Fig. 2). The RNA expression level of γ-GCS was 2.1 times higher in clinical Sb-resistant isolates of L. tropica compared to the sensitive isolates, but it was upregulated only in 70% of resistant isolates, whereas 75% of sensitive isolates experienced downregulations (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012). Additionally, γ-GCS expression was dependent on the host organ and the type of Leishmania strain, which informs about the influence of environmental factors that could govern its expression (Carter et al., Reference Carter, Hutchison, Henriquez, Légaré, Ouellette, Roberts and Mullen2006). Apart from that, γ-GCS expression has been attributed to rapid wound healing in ACL, which was ascertained by a negative correlation seen between γ-GCS expression and the time taken to cure lesions of the responsive cases of field isolates (Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018). Hence, this implicates the functional relevance of γ-GCS expression with a possible relation to GSH-dependent pathways to accelerate the healing process.
There were also discrepancies in γ-GCS expression levels in Leishmania in relation to Sb stress and parasite defence. The resistant Leishmania isolates, including the resistant standards, were neutral in γ-GCS expression, while the sensitive parasites showed inconsistencies of expression having either been upregulated (2.32-fold), downregulated (<0.6-fold), or unaltered (Mohebali et al., Reference Mohebali, Kazemirad, Hajjaran, Kazemirad, Oshaghi, Raoofian and Teimouri2019). Rai et al. suggested that γ-GCS is not consistently expressed, is not involved in naturally Sb-resistant Leishmania, or has a role only in highly resistant parasites (Rai et al., Reference Rai, Bhaskar, Goel, Nath Dwivedi, Sundar and Goyal2013). Furthermore, a pronounced downregulation was seen in Sb-resistant L. donovani (Decuypere et al., Reference Decuypere, Rijal, Yardley, De Doncker, Laurent, Khanal, Chappuis and Dujardin2005, Reference Decuypere, Vanaerschot, Rijal, Yardley, Maes, De Doncker, Chappuis and Dujardin2008). Even though γ-GCS has been studied as an inducer of thiol biosynthesis, γ-GCS-independent thiol elevations have also been characterized in natural Sb-resistant L. donovani. Furthermore, the γ-GCS amplification was found to be negligible in those parasites, showing that it was not directly involved in the thiol synthesis of that particular strain (Mittal et al., Reference Mittal, Rai, Ashutosh, Gupta, Sundar and Goyal2007). Based on the current evidence, thiol production may not be solely dependent on the γ-GCS activity, which may sometimes enable its differences in expression without interfering with Sb resistance, thus minimizing the likelihood of this protein being a potential expression marker of Sb resistance.
Mitogen-activated protein kinase
MAPKs are primarily involved in the phosphorylation of other proteins and are associated with cellular stress response, proliferation, infectivity, differentiation, and apoptosis (Wiese, Reference Wiese1998; Hindley and Kolch, Reference Hindley and Kolch2002). There are around 17 different MAPK proteins in Leishmania. MAPK3 and MAPK9 are exclusively expressed in the promastigote stage and are involved in flagellum maintenance (Bengs et al., Reference Bengs, Scholz, Kuhn and Wiese2005), whereas MAPK1 and MAPK2 are implicated in Sb resistance (Sharma et al., Reference Sharma, Mandal, Mandal, Bhattacharjee and Mukhopadhyay2015). The Sb resistance achieved through MAPK was found to have been associated with the modulation of AQP activity (Mandal et al., Reference Mandal, Sharma, Kruse, Sander-Juelch, Munro, Wang, Vilg, Tamás, Bhattacharjee, Wiese and Mukhopadhyay2012). Furthermore, the co-expression of MAPK with AQP1 increases Sb(III) uptake and drug sensitivity in L. major (Mandal et al., Reference Mandal, Sharma, Kruse, Sander-Juelch, Munro, Wang, Vilg, Tamás, Bhattacharjee, Wiese and Mukhopadhyay2012). Therefore, MAPK must have at least an indirect effect on the Sb transportation mechanisms of Leishmania cells (Fig. 1). Metal-based drugs like Sb(III) induce ROS production leading to subsequent cell apoptosis interconnected with activation of MAPK signalling cascade (Leonard et al., Reference Leonard, Harris and Shi2004; Mann et al., Reference Mann, Davison, Colombo, Colosimo, Diaz, Padovani, Guo, Scrivens, Gao, Mader and Miller2006; Garg and Goyal, Reference Garg and Goyal2015), which is why favourable downregulation of MAPK could be a promising adaptation to avert Sb cytotoxicity.
A several-fold decreased expression of MAPK1 was observed in Sb-resistant L. donovani compared to the sensitive reference that showed a slight increase, which is suggestive of the possible involvement of MAPK1 in triggering cell death pathways upon Sb exposure (Ashutosh et al., Reference Ashutosh, Garg, Sundar, Duncan, Nakhasi and Goyal2012). Furthermore, observation of reduced protein levels in those resistant strains further validated the aforementioned downregulation, and besides, the MAPK overexpression enabled cells to have 2–3-fold increased susceptibility to both Sb(V) and Sb(III) than the cells transfected with the empty vectors (Ashutosh et al., Reference Ashutosh, Garg, Sundar, Duncan, Nakhasi and Goyal2012). In addition, RT-PCR assays revealed a differential expression of MAPK with a suppression of its transcript levels in Sb-resistant L. major and L. tropica clinical isolates compared to the respective sensitive parasites (Kazemi-Rad et al., Reference Kazemi-Rad, Mohebali, Erfan, Saffari, Raoofian, Hajjaran, Hadighi, Khamesipour, Rezaie, Abedkhojasteh and Heidari2013; Sharma et al., Reference Sharma, Mandal, Mandal, Bhattacharjee and Mukhopadhyay2015). The deletion of MAPK2 in L. major resulted in reduced uptake of Sb(III) and slower healing (Mandal et al., Reference Mandal, Sharma, Kruse, Sander-Juelch, Munro, Wang, Vilg, Tamás, Bhattacharjee, Wiese and Mukhopadhyay2012) because of increased parasite viability following less Sb toxicity. Contrarily, MAPK transcripts were more abundantly expressed in 90% of SAG-resistant L. donovani clinical isolates, together with one of the sensitive lines (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012). Altogether, most of the time MAPK can show decreased expression as a preadaptation to Sb resistance, but the inconsistency of its abundance could be a negative factor for the suitability of this protein as a biological Sb marker. However, the positive regulation of MAPK accompanied by AQP1-mediated Sb accumulation would be an attractive phenomenon for drug designing (Fig. 2B) (Mandal et al., Reference Mandal, Sharma, Kruse, Sander-Juelch, Munro, Wang, Vilg, Tamás, Bhattacharjee, Wiese and Mukhopadhyay2012).
Ornithine decarboxylase
Ornithine decarboxylase (ODC) is the rate-limiting protein of the polyamine biosynthetic pathway, which is important for cell growth and proliferation, and its expression is mostly mediated by gene amplification (Haimeur et al., Reference Haimeur, Guimond, Pilote, Mukhopadhyay, Rosen, Poulin and Ouellette1999; Ilari et al., Reference Ilari, Fiorillo, Baiocco, Poser, Angiulli and Colotti2015). It is involved in spermidine biosynthesis as the final product (Fig. 1), and studies have revealed pronounced amplification attributed to both the Sb and As resistance in Leishmania (Haimeur et al., Reference Haimeur, Guimond, Pilote, Mukhopadhyay, Rosen, Poulin and Ouellette1999; Fonseca et al., Reference Fonseca, Comini, Resende, Santi, Zoboli, Moreira and Murta2017). Thus, the elevated ODC may functionally assist the parasites to alleviate the antiproliferative effect caused by metalloid drugs.
There was an overexpression of ODC in Sb-resistant L. donovani compared to the sensitive line; however, it was not expressed on extrachromosomal circles. Apart from that, it was further validated by the protein level overexpression in all the resistant isolates (Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007). Both the promastigotes and the amastigotes of L. donovani overexpressed their ODC levels as a self-protective mechanism against SAG, resulting in notable rises in their IC50 values than that of the wild-type strains (Singh et al., Reference Singh, Mukherjee, Khomutov, Persson, Heby, Chatterjee and Madhubala2007). Moreover, the gene transfection followed by ODC expression was able to increase Sb(III) resistance in 2-fold compared to the wild-type parasites or empty vector-transfected L. guyanensis. In fact, the parasites were more susceptible to the Sb effect with the inhibition of ODC, and the opposite was experienced with the overexpression. For example, α-difluoromethylornithine (DMFO) pre-treated L. guyanensis cells exhibited 648-fold susceptibility to Sb(III) for wild-type in comparison to the 1.5-fold observed in the ODC-transfected clones (Fonseca et al., Reference Fonseca, Comini, Resende, Santi, Zoboli, Moreira and Murta2017). More remarkably, the canine-infected L. infantum clinical isolates showed increased RNA expression in Sb-resistant lines compared to the susceptible, especially in the absence of γ-GCS and trypanothione synthetase (TryS) expression, indicating the functional relevance of ODC in elevating T[SH]2 levels in resistant lines (Gómez Pérez et al., Reference Gómez Pérez, García-Hernandez, Corpas-López, Tomás, Martín-Sanchez, Castanys and Gamarro2016). There was a noticeable difference in ODC gene expression between the natural resistance and resistant mutants since only the naturally resistant L. donovani parasites could augment the expression, while the mutants remained unchanged compared to the reference Dd8 strain (Rai et al., Reference Rai, Bhaskar, Goel, Nath Dwivedi, Sundar and Goyal2013). Furthermore, the ODC expression was amplified in both genetic and protein levels in Sb-resistant Indian L. donovani and Peruvian L. braziliensis, and on the contrary, a downregulation was observed in L. donovani isolates from Nepal (Decuypere et al., Reference Decuypere, Rijal, Yardley, De Doncker, Laurent, Khanal, Chappuis and Dujardin2005; Mukherjee et al., Reference Mukherjee, Padmanabhan, Singh, Roy, Girard, Chatterjee, Ouellette and Madhubala2007; Adaui et al., Reference Adaui, Castillo, Zimic, Gutierrez, Decuypere, Vanaerschot, de Doncker, Schnorbusch, Maes, van der Auwera, Maes, Llanos-Cuentas, Arevalo and Dujardin2011a, Reference Adaui, Schnorbusch, Zimic, Gutirrez, Decuypere, Vanaerschot, De Doncker, Maes, Llanos-Cuentas, Chappuis, Arvalo and Dujardin2011b), hence permitting queries about the exact biological role of this protein in Sb resistance.
Moreover, in L. panamensis, ODC failed to provoke Sb(III) resistance in laboratory-selected stains, owing to the possible hindrance of expression due to the activation of alternative polyamine synthesis pathways or the import of polyamines (Goyeneche-Patino et al., Reference Goyeneche-Patino, Valderrama, Walker and Saravia2008). The intracellular amastigotes of clinical isolates showed strikingly decreased ODC expression in Sb(V)-resistant lines than that of the sensitive ones (Decuypere et al., Reference Decuypere, Rijal, Yardley, De Doncker, Laurent, Khanal, Chappuis and Dujardin2005) which resulted due to the changes in ODC-mediated thiol-biosynthesis in such a way that facilitates parasite-friendly intracellular environment and arrested activation of the Sb(V) in amastigotes (Decuypere et al., Reference Decuypere, Rijal, Yardley, De Doncker, Laurent, Khanal, Chappuis and Dujardin2005). Supporting evidence was published on Sb-resistant L. guyanensis mutants, whose ODC level had no relative difference compared to the parental strain (Dos Reis et al., Reference Dos Reis, do Monte-Neto, Melo and Frézard2017). The prevalence of contradictory evidence of expressions related to Sb resistance raises doubt on the diagnostic use of ODC. Accordingly, previous analysis based on Youden's J statistics justified the foregoing results with the observation of only 50% specificity of ODC to detect the Sb(V) resistance in L. braziliensis clinical isolates (Adaui et al., Reference Adaui, Castillo, Zimic, Gutierrez, Decuypere, Vanaerschot, de Doncker, Schnorbusch, Maes, van der Auwera, Maes, Llanos-Cuentas, Arevalo and Dujardin2011a, Reference Adaui, Schnorbusch, Zimic, Gutirrez, Decuypere, Vanaerschot, De Doncker, Maes, Llanos-Cuentas, Chappuis, Arvalo and Dujardin2011b).
Trypanothione reductase
TR and thioredoxin peroxidase are among the several proteins involved in spermidine metabolism (Ilari et al., Reference Ilari, Fiorillo, Baiocco, Poser, Angiulli and Colotti2015), and it has become an attractive drug target since its unavailability in mammals (Krauth-Siegel and Inhoff, Reference Krauth-Siegel and Inhoff2003; Vázquez et al., Reference Vázquez, Paulino, Salas, Zarate-Ramos, Vera and Rivera2017). The protozoans have evolved a trypanothione/TR system instead of a GSH/glutathione reductase system that is found in mammalian cells (Fig. 1) (Baiocco et al., Reference Baiocco, Colotti, Franceschini and Ilari2009). Trypanothione forms a complex with Sb(III) and it is efficiently transported via plasma membrane vesicles via an ATP-coupled efflux pump, resulting in Sb resistance (Mukhopadhyay et al., Reference Mukhopadhyay, Dey, Xu, Gage, Lightbody, Ouellette and Rosen1996; Gómez Pérez et al., Reference Gómez Pérez, García-Hernandez, Corpas-López, Tomás, Martín-Sanchez, Castanys and Gamarro2016). In a study, TR conferred oxidative protection to both amastigotes and promastigote stages against Sb(V) and Sb(III) and consequently intensified the treatment failure capacity of the non-responders (Zabala-Peñafiel et al., Reference Zabala-Peñafiel, Dias-Lopes, Souza-Silva, Miranda, Conceição-Silva and Alves2023). Hence, it is rational that Sb(III) inhibits TR in a reversible manner so as to secondarily avert the reduction of trypanothione, leading to the accumulation of more disulphides, which would ultimately weaken the resistance (Fig. 2B) (Wyllie et al., Reference Wyllie, Cunningham and Fairlamb2004). Research findings corroborate the Sb(III)-mediated inhibition of TR in Leishmania, which eventually boosts the concentration of the disulphide forms of the intracellular trypanothione and glutathione, which perturbs the cellular thiol redox potential (Wyllie et al., Reference Wyllie, Cunningham and Fairlamb2004). For instance, inhibition of TR leads to apoptotic death of Leishmania parasites owing to instantaneous decline of thiol content (Ghosh et al., Reference Ghosh, Saini, Das, Mandal, Sardar, Ansari, Abhishek, Kumar, Singh, Verma, Equbal, Ali and Das2017), thus, TR expression is of greater importance for protozoans, not only during drug pressure but also for their survival in any circumstance.
The TR protein augmentation was proportional to its activity, with a more than doubled mean activity in non-responders vs responders, in the clinical isolates of L. tropica from ACL patients (Nateghi-Rostami et al., Reference Nateghi-Rostami, Tasbihi and Darzi2022). The high thiol levels observed in natural Sb-resistant L. donovani cells with concurrent amplification of TR and MRPA were not mediated by the γ-GCS (Mittal et al., Reference Mittal, Rai, Ashutosh, Gupta, Sundar and Goyal2007). Furthermore, a western blot analysis revealed 4-fold overexpression of TR in Sb(III)-resistant natural canine isolates of L. infantum vs sensitive (Gómez Pérez et al., Reference Gómez Pérez, García-Hernandez, Corpas-López, Tomás, Martín-Sanchez, Castanys and Gamarro2016). The TR expression seemed to be modulated during promastigote growth, and however, it was upregulated in most of the parasites of Sb-unresponsive isolates (9/10) along with some cured ones (3/11) (Adaui et al., Reference Adaui, Castillo, Zimic, Gutierrez, Decuypere, Vanaerschot, de Doncker, Schnorbusch, Maes, van der Auwera, Maes, Llanos-Cuentas, Arevalo and Dujardin2011a, Reference Adaui, Schnorbusch, Zimic, Gutirrez, Decuypere, Vanaerschot, De Doncker, Maes, Llanos-Cuentas, Chappuis, Arvalo and Dujardin2011b), and it pointed out a functionally irrelevant outcome.
Additionally, the TR expression was almost the same in between the Sb-unresponsive and -sensitive L. donovani clinical extracts, and it was suggested that TR had no clear role in Sb resistance in Leishmania (Nateghi-Rostami et al., Reference Nateghi-Rostami, Tasbihi and Darzi2022). Studies conducted by Wyllie et al. did not show any correlation between the TR activity and the clinical Sb resistance in L. donovani isolates, which is why they suggested its negligible involvement in resistance phenotype (Wyllie et al., Reference Wyllie, Mandal, Singh, Sundar, Fairlamb and Chatterjee2010). Likewise, TR showed a similar and consistently expressed pattern in both MA-responsive and -unresponsive L. tropica, whereas approximately 20% of samples from both types, TR was not affected by Sb (Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018). In summary, TR is an extremely important protein in Sb-resistant Leishmania, with mostly elevated expressions despite possible outliers that could diminish its functional relevance.
Thiol-dependent reductase 1
Thiol-dependent reductase 1 (TDR1) is a tetramer with a functional domain containing omega class glutathione-S transferases that involves the reduction of Sb(V) to Sb(III), with the help of GSH as the reductant (Fig. 1) (Denton et al., Reference Denton, McGregor and Coombs2004; Haldar et al., Reference Haldar, Sen and Roy2011). TDR1 was strikingly upregulated in the amastigotes than that of the promastigotes, and it was attributed to the TDR1-mediated reduction of Sb(V) to Sb(III), whereas promastigotes were prominently sensitive to Sb(V) than the amastigotes (Shaked-Mishant et al., Reference Shaked-Mishant, Ulrich, Ephros and Zilberstein2001; Denton et al., Reference Denton, McGregor and Coombs2004). All the MA non-responders in a study showed significantly elevated metabolic activity against hydrogen peroxide (H2O2) compared to the lower activity seen in responders, which was in line with the elevated trypanothione peroxidase levels in non-responders (Nateghi-Rostami et al., Reference Nateghi-Rostami, Tasbihi and Darzi2022). A 3–4-fold increased TDR expression was experienced in SSG-resistant parasites of L. donovani and L. tropica than in the sensitive line (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022).
Interestingly, the TDR1 expression was broadly variable in the L. tropica Sb-responsive isolates, with the majority of them having fold changes between 2.3 and 1124. In the meantime, the expression was not very prominent in the unresponsive lines, having high expression only in several isolates (Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018). Moreover, the TDR1 expression was several-fold downregulated in MA-unresponsive clinical L. tropica (Oliaee et al., Reference Oliaee, Sharifi, Afgar, Kareshk, Asadi, Heshmatkhah, Bamorovat, Jafarzadeh, Mohammadi and Daneshvar2018).
Tryparedoxin peroxidase
Tryparedoxin peroxidase (TryP) is a principal enzyme that provides parasites with antioxidant defence through the detoxification of harmful peroxides (Flohé et al., Reference Flohé, Budde and Hofmann2003; Iyer et al., Reference Iyer, Kaprakkaden, Choudhary and Shaha2008). Since the Sb treatment is largely associated with the accumulation of deadly ROS in Leishmania, the enhanced expression of these proteins envisages the role of antioxidant defence and detoxification in the emergence of resistance (Fig. 2) (Wyllie et al., Reference Wyllie, Cunningham and Fairlamb2004; Mandal et al., Reference Mandal, Wyllie, Singh, Sundar, Fairlamb and Chatterjee2007).
In Leishmania, the Sb stress elevated the TryP expression in both cytosol and mitochondria; however, the cytosolic expression was more remarkable than the mitochondrial counterpart (Wyllie et al., Reference Wyllie, Vickers and Fairlamb2008; Das et al., Reference Das, Giri, Sundar and Shaha2018). The overexpression of TryP resulted in decreased Sb(III) sensitivity in Leishmania, while the overexpression of the enzymatically inactive form failed to bring about resistance, which corroborates the fact that TryP-mediated Sb resistance was independent of sequestration or membrane efflux of Sb(III) (Wyllie et al., Reference Wyllie, Vickers and Fairlamb2008). A several-fold augmented TryP protein levels were reported in Sb-resistant Leishmania, accompanied by activated H2O2 metabolism as well as increased tolerance to exogenous H2O2 than the respective sensitive or parental lines (Andrade and Murta, Reference Andrade and Murta2014). Moreover, the elevation of TryP was also accompanied by a significant (>2 times average) TR expression in the resistant extracts, with an emphasis on the synergistic interactions of proteins involved in thiol-metabolism in gaining Sb resistance (Nateghi-Rostami et al., Reference Nateghi-Rostami, Tasbihi and Darzi2022). A 3 times TryP boost was observed in Sb-resistant L. tarentolae compared to that observed in lysates of revertant, along with a positive correlation to the subsequent peroxidase activity (Wyllie et al., Reference Wyllie, Vickers and Fairlamb2008). In addition, a comparative proteomic analysis has revealed a highly abundant TryP expression in Sb(III)-resistant L. braziliensis and L. infantum chagasi cells (Matrangolo et al., Reference Matrangolo, Liarte, Andrade, De Melo, Andrade, Ferreira, Santiago, Pirovani, Silva-Pereira and Murta2013). In contrast, the incorporation of TryP through transfection neither exhibited significant gene expression nor tolerance of oxidative stress in L. infantum compared to the parental lines (Andrade and Murta, Reference Andrade and Murta2014).
Heat shock proteins
The chemical inhibition of heat shock protein (HSP) was found to be a strategy to produce antileishmanial drugs (Das et al., Reference Das, Banerjee, Kamran, Ejazi, Asad, Ali and Chakrabarti2020). Heat shock protein-83 (HSP83) and heat shock protein-70 (HSP70) are involved in antileishmanial drug-mediated programmed cell death by interfering with mitochondrial membrane potential (Vergnes et al., Reference Vergnes, Gourbal, Girard, Sundar, Drummelsmith and Ouellette2007).
Matrangolo et al. demonstrated prominent expression profiles of HSP83, heat shock protein-60 (HSP60), and HSP70 in the resistant cell lines of both L. braziliensis and L. infantum chagasi, particularly the HSP70 equivalents, having boosted in both the cytoplasm and the mitochondria of the L. infantum chagasi lines (Matrangolo et al., Reference Matrangolo, Liarte, Andrade, De Melo, Andrade, Ferreira, Santiago, Pirovani, Silva-Pereira and Murta2013). HSP70 and HSP83 were reported to be overexpressed in the membrane-enriched fraction of the SAG-resistant L. donovani clinical isolates (Kumar et al., Reference Kumar, Sisodia, Misra, Sundar, Shasany and Dube2010), and in the meantime, the increased expression pattern of HSP60 and HSP70 has been observed in relation to Sb-resistant mechanisms in L. (Viannia) panamensis (Peláez et al., Reference Peláez, Muskus, Cuervo and Marín-Villa2012). The substantial abundance of HSP70 in in vitro selected Sb-resistant Leishmania of both amastigotes and promastigotes, and the corresponding revertant cells in the absence of drug pressure, could be suggestive of the stability and functional significance of this protein in Sb-resistant Leishmania (Brochu et al., Reference Brochu, Halmeur and Ouellette2004). Inversely, the Sb sensitivity was aligned with the downregulation of HSP83 in L. donovani (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012). In addition, the CL caused by naturally Sb-resistant L. tropica was associated with HSP70 differential expression (Özbilgin et al., Reference Özbilgin, Zeyrek, Güray, Çulha, Akyar, Harman, Özbel, Ertabaklar, Çavuş and Gündüz2021). The L. infantum promastigotes were superior to L. tarentolae in acquiring Sb(III) defence through robust HSP70 expression (Brochu et al., Reference Brochu, Halmeur and Ouellette2004). Nonetheless, a study proposed HSP70 to have only a 75% success rate as an effective candidate prediction model for determining Sb resistance of CL clinical cases caused by L. braziliensis (Torres et al., Reference Torres, Ribeiro-Alves, Romero, Dávila and Cupolillo2013).
However, a few of the findings possibly disprove the HSP-mediated Sb resistance in Leishmania as well as impose limitations on the suitability of this gene in predicting resistance. For instance, HSP83 was upregulated only in 40% of clinically resistant L. donovani lines compared to the LdAG83 reference strain (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012), and also the transfection of the HSP70 gene was unable to circumvent the Sb effect and confer direct resistance towards Sb (Brochu et al., Reference Brochu, Halmeur and Ouellette2004).
Other important genes with elevated expression in Sb resistance
Only a few studies have been conducted on histone modification and gene regulation in trypanosomatids. There are 2 types of histones in Leishmania; core histones like H2A, H2B, H3, H4, and linker histone H1 (Soto et al., Reference Soto, Requena, Gomez, Navarrete and Alonso1992, Reference Soto, Quijada, Alonso and Requena1997; Fasel et al., Reference Fasel, Robyr, Mauel and Glaser1993; Martínez et al., Reference Martínez, Thomas, Alonso, Carmelo, González, Del Castillo and Valladares2002). Histone expression in Leishmania has been implicated as a coupled mechanism to DNA replication that affects the level of translation via post-transcriptional mechanisms (Abanades et al., Reference Abanades, Ramírez, Iborra, Soteriadou, González, Bonay, Alonso and Soto2009), and a few studies have discussed the histone expression in Sb-resistant Leishmania. However, so far, histone expression has not been characterized as a direct contributor to drug resistance, but it may have a function in the epigenetic programming of resistance genes in Leishmania. Apart from that, histone epigenetic markers were essential for the survival of L. major (Anderson et al., Reference Anderson, Wong, Baugh, Ramasamy, Myler and Beverley2013; Afrin et al., Reference Afrin, Khan and Hemeg2019). Meanwhile, the studies aiming to explore the correlation between histone function and antimonial drug resistance are not well established yet.
Overexpression of H2A into L. donovani conferred decreased Sb susceptibility against not only SAG but also developed resistance against amphotericin-B and miltefosine (Singh et al., Reference Singh, Kumar, Duncan, Nakhasi and Salotra2010). H1 was found to have elevated expression in 9 out of 10 Sb-resistant L. donovani isolates, and H2A or H4 were upregulated in 50% of the SAG-resistant L. donovani clinical strains, however, it was found that there was a >2-fold downregulation in all the sensitive lines compared to the LdAG83-sensitive reference line (Kumar et al., Reference Kumar, Singh, Bhandari, Kulshrestha, Negi and Salotra2012). Moreover, H1 and H2A showed protein level upregulation in SAG-resistant L. donovani, kala-azar (Singh et al., Reference Singh, Kumar, Duncan, Nakhasi and Salotra2010). Based on the present data, the histone modification could be a potential activator of the resistant genes but has to be further characterized to reveal its exact molecular relationship to Sb resistance.
The novel Sb-resistant markers, ARM56 and ARM58 were significantly elevated in resistant Leishmania, although with unknown function or molecular mechanisms in this regard. ARM proteins contain conserved domains with hydrophobic amino acids and form transmembrane structures; particularly, ARM56 and ARM58 are located as a subtelomeric cluster in chromosome 34 and their co-expression has been implicated in Sb resistance (Nevado et al., Reference Nevado, Bifeld, Höhn and Clos2016). Especially, the overexpression of ARM58 was found to minimize the Sb effect by reducing its accumulation in Leishmania cells by at least 70% (Schäfer et al., Reference Schäfer, Nevado, Zander and Clos2014). Most importantly, following in vitro selection for Sb(III), the ARM58 mRNA level exhibited an 800% boost compared to the wild-type and was also conferred protection on amastigotes as well as promastigotes against both Sb(III) and Sb(V) (Nühs et al., Reference Nühs, Schäfer, Zander, Trübe, Tejera Nevado, Schmidt, Arevalo, Adaui, Maes, Dujardin and Clos2014). There was a clear correlation between Sb resistance and ARM56/ARM58 expression, whereas elevation of ARM58 was identified as an exclusive feature in Sb-resistant field isolates, hence urges for extensive studies before validating this protein as an Sb-resistant marker (Rugani et al., Reference Rugani, Gontijo, Frézard, Soares and Do Monte-Neto2019).
Another protein, PRP1, belongs to the ABC transporter superfamily and was initially considered to confer cross-resistance towards the antimony trichloride but not Sb(V) (Coelho et al., Reference Coelho, Beverley and Cotrim2003); nonetheless, a recent finding illustrated a significantly elevated expression in SSG-resistant L. tropica and L. donovani resulting in more than 4-fold expression (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022).
Parasite surface antigen-2 was consistently augmented in both the transcriptional and translational expression in SAG-resistant L. donovani clinical isolates resulting in more than 1.5-fold higher expression than the sensitive. Moreover, the overexpression of this protein could transform the sensitive strains to resistance with a decline of their Sb susceptibility level in >12-fold (Bhandari et al., Reference Bhandari, Kumar, Verma, Srividya, Negi, Singh and Salotra2013). Arsenate reductase-2 was a 3–4-fold expression in VL SSG-resistant L. donovani, and L. tropica compared to the sensitive reference strain AG83 (Khanra et al., Reference Khanra, Das, Sarraf, Datta, Das, Manna and Roy2022).
Conclusions
Sb resistance in Leishmania is mainly achieved by the expression modification of Sb transporter genes, Sb-reducing enzymes, and thiol-synthesizing enzymes. Of the genes studied in the present review, MRPA, γ-GCS, ODC, TR, TDR1, TryP, and HSP illustrated a general likelihood of upregulation, while AQP1 and MAPK showed a tendency to downregulate in the Sb-resistant Leishmania and vice versa, therefore, the resistance may have been orchestrated by the functional relevance of genes. Likewise, the relative gene expression in Sb resistance can exhibit similarities among different resistant isolates, but there are chances for deviations to be made from the mostly accepted phenomena. The reason for the presence of inconsistencies of protein functions may be due to molecular adaptations like polymorphism and post-translational modifications. Altogether, the gene expression-based confirmation of Sb resistance in Leishmania by examining the upregulation or downregulation of a particular gene compared to a control will be useful in scientific studies to investigate the underlying biology; however, its application in patient diagnosis of clinical resistance may not be reliable due to possible false-positive or false-negative results and in some cases, the inadequacy of research work. Nonetheless, it is possible to minimize the potential misinterpretations provided that multiple resistance gene expressions are included in the analysis and if their relative expressions are highly significant. The current review circumscribed the Sb resistance-related gene expression to facilitate future research that will fulfil the unmet need for the detection of biomarkers for Sb-resistant leishmaniasis, which remains an obvious need to achieve effective disease control and elimination.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182023001129
Author's contribution
Conceptualization: Rajamanthrilage Kasun Madusanka and Angamuthu Selvapandiyan; methodology: Rajamanthrilage Kasun Madusanka and Angamuthu Selvapandiyan; formal analysis and investigation: Rajamanthrilage Kasun Madusanka, Angamuthu Selvapandiyan, Hermali Silva and Nadira Karunaweera; writing – original draft preparation: Rajamanthrilage Kasun Madusanka; writing – review and editing: Rajamanthrilage Kasun Madusanka, Angamuthu Selvapandiyan, Hermali Silva and Nadira Karunaweera; funding acquisition: Nadira Karunaweera and Angamuthu Selvapandiyan]; supervision: Angamuthu Selvapandiyan, Nadira D. Karunaweera and Hermali Silva]. All authors read and approved the final manuscript.
Financial support
Indo–Sri Lanka fellowship was awarded to R. K. M. (INSA/DST-ISRF/2022/45). This research study was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number U01AI136033 and Indian Council of Medical Research under award number 63/8/2013-BMS. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health and Indian Council of Medical Research.
Competing interests
None.
Data availability statement
There is no extra data for this paper.