Refine listing
Actions for selected content:
Contents
Research Article
Stability of Soft Quasicrystals in a Coupled-Mode Swift-Hohenberg Model for Three-Component Systems
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 559-581
-
- Article
- Export citation
Automated Parallel and Body-Fitted Mesh Generation in Finite Element Simulation of Macromolecular Systems
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 582-602
-
- Article
- Export citation
Development of a Combined Compact Difference Scheme to Simulate Soliton Collision in a Shallow Water Equation
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 603-631
-
- Article
- Export citation
Numerical Methods for Solving the Hartree-Fock Equations of Diatomic Molecules II
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 632-647
-
- Article
- Export citation
A Multigrid Method for Ground State Solution of Bose-Einstein Condensates
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 648-662
-
- Article
- Export citation
A 3D Multi-Phase Hydrodynamic Model for Cytokinesis of Eukaryotic Cells
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 663-681
-
- Article
- Export citation
Simulating Biofilm Deformation and Detachment with the Immersed Boundary Method
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 682-732
-
- Article
- Export citation
A Second-Order Finite Difference Method for Two-Dimensional Fractional Percolation Equations
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 733-757
-
- Article
- Export citation
The Hamiltonian Field Theory of the Von Mises Wave Equation: Analytical and Computational Issues
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 758-769
-
- Article
- Export citation
Efficient Implementation of Smoothed Particle Hydrodynamics (SPH) with Plane Sweep Algorithm
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 770-800
-
- Article
- Export citation
A Semi-Lagrangian Approach for Dilute Non-Collisional Fluid-Particle Flows
- Part of:
-
- Published online by Cambridge University Press:
- 16 March 2016, pp. 801-840
-
- Article
- Export citation
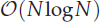 neighbour search method based on the plane sweep (PW) algorithm with
neighbour search method based on the plane sweep (PW) algorithm with