Introduction
It is estimated that over 60% of human infectious diseases are zoonotic in origin (Taylor et al., Reference Taylor, Latham and Woolhouse2001). However, most disease control programmes focus on humans only (Morse et al., Reference Morse, Mazet, Woolhouse, Parrish, Carroll, Karesh, Zambrana-Torrelio, Lipkin and Daszak2012), following single-pathogen single-host epidemiological theory (Antonovics, Reference Antonovics2017). In the last few decades, there has been an increase in the interaction and exposure among humans, livestock and wildlife due to increased intrusion by humans into wildlife habitat, which has led to transmission and evolution of pathogens, and the emergence of infectious diseases (Cleaveland et al., Reference Cleaveland, Laurenson and Taylor2001; Jones et al., Reference Jones, Patel, Levy, Storeygard, Balk, Gittleman and Daszak2008; Cole and Viney, Reference Cole and Viney2018). Shared habitats create pathways for gastrointestinal parasite transmission, primarily via faecal-oral and soil-transmitted routes, as parasite stages are shed in faeces and contaminate soil and water sources used by multiple host species (Brooker et al., Reference Brooker, Bethony and Hotez2004; Cibot et al., Reference Cibot, Guillot, Lafosse, Bon, Seguya and Krief2015; Obanda et al., Reference Obanda, Maingi, Muchemi, Ng’Ang’A, Angelone and Archie2019). Helminths such as strongylid nematodes and Strongyloides are multicellular worms, so-called soil-transmitted nematodes, that develop through distinct egg and larval stages in the environment and usually do not multiply within a single host, with the exception of S. stercoralis that can hatch larvae within the host, initiate autoinfection and progressively increase the burden. On the other hand, unicellular eukaryotic parasites such as Cryptosporidium, Giardia and microsporidia transmitted via faecal-oral way replicate within host cells and are transmitted via environmentally resistant oo/cysts or spores (CDC, 2024a). However, both intrinsic (host immune response, physiology, etc.) and extrinsic (rainfall, temperature, etc.) factors contribute to determining whether a parasite eventually establishes itself within a particular host species or population (Turner and Elena, Reference Turner and Elena2000; Duffy et al., Reference Duffy, Burch and Turner2007). While gastrointestinal parasites, including helminths and protozoans, are generally less fatal than many viral or bacterial pathogens, long-term infections can still cause significant health impacts and increase host morbidity (Alum et al., Reference Alum, Rubino and Ijaz2010).
Strongylid nematodes, a diverse group of gastrointestinal parasites, infect a wide range of mammalian hosts, including ruminants, carnivores, humans or non-human primates (Brooker et al., Reference Brooker, Bethony and Hotez2004; Van Wyk et al., Reference Van Wyk, Cabaret and Michael2004; Lichtenfels et al., Reference Lichtenfels, Kharchenko and Dvojnos2008; Ilík et al., Reference Ilík, Kreisinger, Modrý, Schwarz, Tagg, Mbohli, Nkombou, Petrželková and Pafčo2023). Although some strongylids exhibit certain host specificity, others are rather generalists and pose zoonotic risks in areas where the host species live in close proximity and share habitats and resources (Ghai et al., Reference Ghai, Chapman, Omeja, Davies and Goldberg2014; Hasegawa et al., Reference Hasegawa, Modrý, Kitagawa, Shutt, Todd, Kalousová, Profousová and Petrželková2014, Reference Hasegawa, Shigyo, Yanai, McLennan, Fujita, Makouloutou, Tsuchida, Ando, Sato and Huffman2017; Ota et al., Reference Ota, Hasegawa, McLennan, Kooriyama, Sato, Pebsworth and Huffman2015; Pafčo et al., Reference Pafčo, Kreisinger, Čížková, Pšenková-Profousová, Shutt-Phillips, Todd, Fuh, Petrželková and Modrý2019). For example, some strongylid nematodes in cattle such as Ostertagia ostertagi, Haemonchus placei or Cooperia oncophora (Charlier et al., Reference Charlier, Van der Voort, Hogeveen and Vercruysse2012; Jabbar et al., Reference Jabbar, Cotter, Lyon, Koehler, Gasser and Besier2014; Pafčo et al., Reference Pafčo, Nosková, Ilík, Anettová, Červená, Kreisinger, Pšenková, Václavek, Vyhlídalová, Ježková, Malát, Mihalca and Modrý2024) or Strongylus vulgaris in equids (Jürgenschellert et al., Reference Jürgenschellert, Krücken, Bousquet, Bartz, Heyer, Nielsen and von Samson-himmelstjerna2022) show strong host specificity, while Oesophagostomum spp. infect broad range of hosts. Zoonotic potential of O. bifurcum and O. stephanostomum has been demonstrated by detection of these parasites in humans and other primates in Africa (Pafčo et al., Reference Pafčo, Kreisinger, Čížková, Pšenková-Profousová, Shutt-Phillips, Todd, Fuh, Petrželková and Modrý2019; Ilík et al., Reference Ilík, Kreisinger, Modrý, Schwarz, Tagg, Mbohli, Nkombou, Petrželková and Pafčo2023), including in Uganda (Ghai et al., Reference Ghai, Chapman, Omeja, Davies and Goldberg2014; Cibot et al., Reference Cibot, Guillot, Lafosse, Bon, Seguya and Krief2015). Different Oesophagostomum spp. are commonly found in domestic animals such as ruminants or pigs (Sharma et al., Reference Sharma, Shukla, Dixit and Dixit2011; Mishra et al., Reference Mishra, Dohare, Pashupathi, Vijayasarathi, Yadav and Wodeyar2024). Typically, parasitic strongylid nematodes assemble into diverse communities that include multiple coexisting species and genera (Van Wyk et al., Reference Van Wyk, Cabaret and Michael2004; Lichtenfels et al., Reference Lichtenfels, Kharchenko and Dvojnos2008; Mclean et al., Reference Mclean, Kinsella, Chiyo, Obanda, Moss and Archie2012).
Strongyloides is a genus of nematodes with approximately 50 species of intestinal parasites that exclusively infect vertebrates such as amphibians, reptiles, birds and mammals (Viney and Lok, Reference Viney and Lok2015). Infection caused by Strongyloides is also frequently documented in both free-ranging and captive primates across all continents except Antarctica, with a possibility of missing data from captive primates in Australia (Nosková et al., Reference Nosková, Sambucci, Petrž Elková, Červená, Modrý and Pafčo2024). So far, 3 species have been described in primates; Strongyloides cebus has exclusively been identified in the Central and South American primates (Dorris et al., Reference Dorris, Viney and Blaxter2002; Mati et al., Reference Mati, Junior, Pinto and De Melo2013), with no records of human infections with this species. While it is believed that S. fuelleborni is the primary cause of infection in wild non-human primates, there have been reports of mixed infections involving both S. stercoralis and S. fuelleborni in wild central chimpanzee (Pan troglodytes troglodytes) in Tanzania (Hasegawa et al., Reference Hasegawa, Sato, Fujita, Nguema, Nobusue, Miyagi, Kooriyama, Takenoshita, Noda, Sato, Morimoto, Ikeda and Nishida2010), indicating that simultaneous infections may be relatively common (Hasegawa et al., Reference Hasegawa, Kalousova, McLennan, Modry, Profousova-Psenkova, Shutt-Phillips, Todd, Huffman and Petrzelkova2016a). Strongyloides fuelleborni was found mainly in humans that shared habitat with other primates, while S. stercoralis has worldwide distribution in humans (Nosková et al., Reference Nosková, Sambucci, Petrž Elková, Červená, Modrý and Pafčo2024). Strongyloides stercoralis can also infect dogs and cats (Felis catus) (Thamsborg et al., Reference Thamsborg, Ketzis, Horii and Matthews2017). In experimental settings, S. stercoralis has been effectively transmitted from humans to dogs (Jariwala et al., Reference Jariwala, Redding and Hewitt2017; Thamsborg et al., Reference Thamsborg, Ketzis, Horii and Matthews2017). However, recent comparative research using mitochondrial cox1 and nuclear 18S rDNA markers found the presence of 2 distinct genetic populations of S. stercoralis in dogs: type B seemingly exclusive to dogs, while type A is found in both dogs and primates including humans (Jaleta et al., Reference Jaleta, Zhou, Bemm, Schär, Khieu, Muth, Odermatt, Lok and Streit2017; Nagayasu et al., Reference Nagayasu, Aung, Hortiwakul, Hino, Tanaka, Higashiarakawa, Olia, Taniguchi, Win, Ohashi, Odongo-Aginya, Aye, Mon, Win, Ota, Torisu, Panthuwong, Kimura, Palacpac and Maruyama2017). Strongyloides spp. are also found in domestic animals worldwide such as S. westeri in foals, S. papillosus in cattle and small ruminants, S. ransomi in pigs and so on (Thamsborg et al., Reference Thamsborg, Ketzis, Horii and Matthews2017).
Cryptosporidium, Giardia and microsporidia are found in both humans and other animals; the infectious diseases caused by them (cryptosporidiosis, giardiasis and microsporidiosis respectively) have been documented in high-income countries as well as low- and middle-income countries (Fayer, Reference Fayer2010; Thellier and Breton, Reference Thellier and Breton2008; Xiao and Fayer, Reference Xiao and Fayer2008). There are currently 48 recognized species of Cryptosporidium (Stensvold et al., Reference Stensvold, Larsen, Grüttner, Nielsen, Engberg and Lebbad2024) with the literature describing over 150 genotypes, and in humans, at least 25 species and 3 genotypes have been detected, with C. hominis and C. parvum being the most common and responsible for diarrheal disease of varying severity (Fayer, Reference Fayer2010; Xiao, Reference Xiao2010; Stensvold et al., Reference Stensvold, Larsen, Grüttner, Nielsen, Engberg and Lebbad2024). In cattle, the dominant species are C. parvum, C. bovis, C. ryanae and C. andersoni, with C. parvum as the major cause of clinical disease in young calves (Thomson et al., Reference Thomson, Innes, Jonsson and Katzer2019). Goats commonly harbour C. parvum, C. ubiquitum and C. xiaoi (Ryan et al., Reference Ryan, Zahedi, Feng and Xiao2021), while in pigs, C. scrofarum and C. suis are most frequently reported, with C. parvum also present in some countries (Chen et al., Reference Chen, Qin, Wu, Xu, Huang, Li and Zhang2023). In dogs, C. canis is the primary species detected and can occasionally infect humans (Taghipour et al., Reference Taghipour, Olfatifar, Bahadory, Godfrey, Abdoli, Khatami, Javanmard and Shahrivar2020). In the 1920s, over 50 Giardia species were proposed, but current research focusing on trophozoite/cyst morphology acknowledges only 6 species (Ryan and Cacciò, Reference Ryan and Cacciò2013). Giardia intestinalis (syn. G. lamblia, G. duodenalis), the species infecting humans and most mammals, comprises 8 valid assemblages A–H, among which assemblages A and B are the primary causes of human giardiasis and are associated with diarrhoea, malabsorption and weight loss (Ryan and Cacciò, Reference Ryan and Cacciò2013; Mohammed, Reference Mohammed2024). In non-human primates such as vervet monkeys, infections typically involve assemblages A and B, mirroring human pathogenic patterns (Feng and Xiao, Reference Feng and Xiao2011). In dogs, assemblages C and D are most common and are generally host-adapted with variable pathogenicity, while occasional infections with zoonotic assemblage A also occur (Thompson, Reference Thompson, Sterling and Adam2004; Sprong et al., Reference Sprong, Cacciò and Van Der Giessen2009). In livestock, assemblage E predominates in goats, pigs and cattle, usually causing subclinical to mild gastrointestinal disease, although diarrheal outbreaks have also been documented (Feng and Xiao, Reference Feng and Xiao2011; Ryan and Zahedi, Reference Ryan and Zahedi2019).
Certain genotypes of microsporidian Enterocytozoon bieneusi, such as genotypes B and Peru7, are primarily host-specific, whereas other genotypes such as D and EbpC are considered zoonotic and have been detected in both humans and other animals, demonstrating cross-species transmission potential (Thellier and Breton, Reference Thellier and Breton2008). Cryptosporidium spp., G. intestinalis and several microsporidia that can be transmitted between animals and humans have been reported in humans, livestock, companion animals and wildlife (Thellier and Breton, Reference Thellier and Breton2008; Xiao and Fayer, Reference Xiao and Fayer2008; Ryan and Cacciò, Reference Ryan and Cacciò2013). Several studies have shown that non-human primates are potential sources of Cryptosporidium, Giardia and microsporidia infection for humans (Du et al., Reference Du, Zhao, Shao, Fang, Tian, Zhang, Wang, Wang, Qi and Yu2015); cases of human-pathogenic Cryptosporidium spp., G. intestinalis and E. bieneusi infections in wild, semi-wild and captive primates have also been reported in Asia, Africa and the Americas (Nizeyi et al., Reference Nizeyi, Cranfield and Graczyk2002; Ye et al., Reference Ye, Xiao, Ma, Guo, Liu and Feng2012, Reference Ye, Xiao, Li, Huang, Amer, Guo, Roellig and Feng2014; Sak et al., Reference Sak, Petrzelkova, Kvetonova, Mynarova, Shutt, Pomajbikova, Kalousova, Modry, Benavides, Todd and Kvac2013, Reference Sak, Petrželková, Květoňová, Mynářová, Pomajbíková, Modrý, Cranfield, Mudakikwa and Kváč2014; Karim et al., Reference Karim, Dong, Yu, Jian, Zhang, Wang, Zhang, Rume, Ning and Xiao2014; Mynářová et al., Reference Mynářová, Foitová, Kváč, Květoňová, Rost, Morrogh-Bernard, Nurcahyo, Nguyen, Supriyadi and Sak2016) owing to the close phylogenetic relationship between humans and primates (Du et al., Reference Du, Zhao, Shao, Fang, Tian, Zhang, Wang, Wang, Qi and Yu2015).
To this end, the present study addresses 2 central research questions: (1) How do parasite prevalence and species richness differ among sympatric hosts, including humans, vervet monkeys, livestock and dogs, at Lake Nabugabo, Uganda? and (2) Which parasite species from the order Strongylida, and the genera Strongyloides, Cryptosporidium, Giardia, Encephalitozoon and Enterocytozoon are shared among these host species?
Materials and methods
Study site
The study was conducted on the shores of Lake Nabugabo, Uganda (0°22′–12° S, 31°54′ E, 1136 m, Supplementary Figure 1), a satellite of Lake Victoria that is mostly surrounded by wetlands, grasslands, patches of degraded forest, farmers’ fields and small buildings (Chapman et al., Reference Chapman, Twinomugisha, Teichroeb, Valenta, Sengupta, Sarkar, Rothman and Waller2016). The area receives an average of 1348 mm of rain annually (Chapman et al., Reference Chapman, Twinomugisha, Teichroeb, Valenta, Sengupta, Sarkar, Rothman and Waller2016) in 2 seasons with high rainfall (March–May and September–November) and 2 seasons with low rainfall (December–February and June–August) (Schwegel et al., Reference Schwegel, Filazzola, Chapman and Schoof2023). The Nabugabo Research Site on the shores of the lake is home to long-term research in 3 neighbouring groups of vervet monkeys, which served as subjects for this study.
Sample collection
Between July and September 2022, 75 faecal samples (mean = 2.03, range = 1–4 per individual) from adult (>5 years or dispersal for males, after first birth for females) and subadult (4–5 years for males, 6 months before adulthood for females) vervet monkeys in 3 neighbouring groups were collected (25 samples from each group; N male = 30, N female = 45). We also non-invasively collected 25 faecal samples from different dogs, 25 individual faecal samples from different livestock (9 cows, 7 goats and 9 pigs), as well as samples from the 25 different pit latrines of households in Bbaale village nearby Lake Nabugabo, Uganda. The faecal samples from vervets, dogs and livestock were collected as soon as the individuals were observed defecating to avoid contamination with the environment/soil. The samples from the pit latrines were collected using a sludge nabber, which was properly washed and disinfected with 90% ethanol after each sample collection. All collected samples were fixed in 10% formalin for microscopic analysis and 96% ethanol for molecular analysis. The samples were kept at room temperature until they were transported to the labs for the analyses.
Microscopic analysis
The samples fixed in 10% formalin were analysed at the Central Diagnostic Laboratory, College of Veterinary Medicine Animal Resources and Biosecurity, Makerere University, Kampala, Uganda using formalin-ether sedimentation technique (Uga, Reference Uga2010). The parasites were identified to the family and genus level wherever possible (Soulsby, Reference Soulsby1982; Zajac and Conboy, Reference Zajac and Conboy2006).
DNA isolation and sequencing
Total faecal DNA was extracted for detection of strongylid nematodes and genus Strongyloides from ethanol-preserved faecal samples using the DNeasy PowerSoil Pro kit (Qiagen, Germany) after the aliquots were left overnight at 37 °C to evaporate the residual ethanol at the Institute of Vertebrate Biology, Czech Academy of Sciences. No modifications were made to the manufacturer’s protocol. Eluted DNA was stored at −20 °C until further analyses.
The internal transcribed spacer 2 (ITS2) of strongylid nematodes was amplified using primers Strongyl_ITS-2_F and Strongyl_ITS-2_R and high-throughput sequencing (HTS) (Pafčo et al., Reference Pafčo, Čížková, Kreisinger, Hasegawa, Vallo, Shutt, Todd, Petrželková and Modrý2018, Reference Pafčo, Kreisinger, Čížková, Pšenková-Profousová, Shutt-Phillips, Todd, Fuh, Petrželková and Modrý2019). Briefly, the HTS libraries were generated using a 2-step-PCR approach following the Nextera primer design. DNA extractions from strongylid negative human faeces and water were used as negative controls and synthetic DNA templates as positive controls. All samples were prepared in 2 technical PCR replicates with different tag primer barcodes. The final library was sequenced using MiSeq Reagent Kit v3 (2 × 300 bp pair-end reads, 600 cycles) by Illumina MiSeq platform.
Strongyloides spp. DNA was detected by real-time quantitative PCR (qPCR) at the Real-Time PCR LightCycler® (Roche, Switzerland) targeting a conservative region (101 bp) of the gene for 18S rRNA (18S rDNA) using primers Stro18S-1530F, Stro18S-1630R and the probe Stro-18S-1586T (Verweij et al., Reference Verweij, Canales, Polman, Ziem, Brienen, Polderman and van Lieshout2009). A subset of positive samples from each host species was then used for amplification of the hypervariable region IV of 18S rDNA (HVR-IV; ∼255 bp) in a conventional PCR assay using primers New_HVR-IV_F and New_HVR-IV_R (Barratt et al., Reference Barratt, Lane, Talundzic, Richins, Robertson, Formenti, Pritt, Verocai, De Souza, Soares, Traub, Buonfrate and Bradbury2019). The PCR amplification and subsequent product treatment were carried out as described previously (Nosková et al., Reference Nosková, Ilík, Stéphane Singa Niatou, Dumas, Fuh, Dicky, Kurucová, Baláž, Judita Petrželková and Pafčo2026). Purified products were sent for commercial sequencing to Macrogene (Macrogene Europe, Netherlands).
Total genomic DNA (gDNA) for detection of Cryptosporidium spp., G. intestinalis, E. bieneusi and Encephalitozoon spp. was isolated using the GeneAll® Exgene™ Stool DNA Mini Kit (GeneAll Biotechnology Co., Ltd, South Korea) according to the manufacturer’s guidelines, with the following modifications at the Institute of Parasitology, Biology Centre, Czech Academy of Sciences. The alcohol was evaporated from each sample at 65 °C overnight. A 100 μL of 0.5 mm glass beads and 1000 μL of lysis buffer were added to each tube and oo/cysts/spores were disrupted for 60 sec at 5.5 m s−1 using glass beads in a FastPrep®-24 Instrument (MP Biomedicals, USA). The extracted gDNA was then stored at −80 °C. Nested PCR protocols were used to identify Cryptosporidium spp. at partial sequences of gene encoding small subunit of rRNA (18S rDNA), G. intestinalis at partial sequences of gene encoding triosephosphate isomerase (TPI) and E. bieneusi and Encephalitozoon spp. at partial sequences of gene encoding the ITS of rRNA using established protocols and primers (Didier et al., Reference Didier, Vossbrinck, Baker, Rogers, Bertucci and Shadduck1995; Katzwinkel-Wladarsch et al., Reference Katzwinkel-Wladarsch, Lieb, Heise, Löscher and Rinder1996; Xiao et al., Reference Xiao, Escalante, Yang, Sulaiman, Escalante, Montali, Fayer and Lal1999; Buckholt et al., Reference Buckholt, Lee and Tzipori2002; Sulaiman et al., Reference Sulaiman, Fayer, Bern, Gilman, Trout, Schantz, Das, Lal and Xiao2003). PCR amplicons were purified by GenElute™ Gel Extraction Kit (Sigma, USA) according to the manufacturer’s instructions and sequenced in both directions (SeqMe, Czech Republic).
Bioinformatics and data assembly of strongylid nematodes
Gene-specific primers were trimmed using skewer (Jiang et al., Reference Jiang, Lei, Ding and Zhu2014). Next, low-quality reads (expected error rate >2) were eliminated, and the filtered data set was denoised using dada2 (Callahan et al., Reference Callahan, McMurdie, Rosen, Han, Johnson and Holmes2015) to obtain amplicon sequence variants (ASVs). ASV are unique DNA sequences generated from targeted amplification of specific regions in the genome, allowing for precise identification of distinct taxa. All ASV sequences are deposited in the GenBank (Supplementary Table 1). Naive Bayesian RDP classifier (Wang et al., Reference Wang, Garrity, Tiedje and Cole2007) implemented in the dada2 pipeline was used for taxonomic assignment of ASVs with 80% posterior confidence. The reference training data set was downloaded from the National Center for Biotechnology Information (NCBI) database (200 top blast hits for each of our ASVs, environmental samples were excluded). To avoid inflation of strongylid diversity due to PCR/sequencing artefacts that were not corrected by dada2, we considered only those ASVs that were consistently present in both technical PCR duplicates for a given sample (Pafčo et al., Reference Pafčo, Čížková, Kreisinger, Hasegawa, Vallo, Shutt, Todd, Petrželková and Modrý2018). We also excluded ASVs that did not correspond to the Strongylida order. Read counts for ASVs corresponding to the same genus were grouped within the resulting abundance matrix.
Phylogenetic analyses for Strongyloides and protists
We aligned sequences using Geneious Prime 2024.0.7 (www.geneious.com) and the phylogenetic trees were derived from IQ-tree (Minh et al., Reference Minh, Schmidt, Chernomor, Schrempf, Woodhams, von Haeseler and Lanfear2020) based on the maximum likelihood method with 1000 bootstrap replicates. The sequences from this study are deposited in GenBank (Supplementary Table 2).
Statistics
The prevalence of a parasite taxon was calculated by dividing the total number of individuals with a specific parasite taxon obtained from both microscopic and molecular methods by the total number of individuals of a specific host group sampled, and the parasite species richness as the number of parasite taxa found in each host’s faecal samples throughout the study period (Bush et al., Reference Bush, Lafferty, Lotz and Shostak1997). Parasite infection rate was calculated as the presence of at least 1 parasite in the faecal sample divided by the number of samples for a particular host category. Chi-square tests of independence, followed by pairwise comparisons, were conducted to test for significant differences in proportion of samples infected across different host groups as well as proportion of samples infected with each parasite taxa across different host groups.
Strongylid ASV prevalence was defined as the number of hosts infected with each strongylid ASV divided by the number of hosts sampled. Differences in number of reads per host type and relative abundances of strongylid nematode communities among host type were compared using generalized linear models (GLMs) followed by pairwise comparisons. Differences in alpha diversity of strongylid communities were investigated by using the number of ASVs per sample as a proxy measure. The number of different haplotypes per sample was highly correlated with the other commonly used alpha diversity indices, such as Shannon index (cor. coef. = 0.78), Simpson (cor. coef. = 0.63) or inverse Simpson (cor. coef. = 0.78), and thus we used it as the only alpha diversity index in our analyses. The differences in haplotype diversity among host categories were tested by GLM with a quasibinomial error distribution, followed by a Tukey post hoc test for pairwise comparison. Community composition was defined as prevalence and relative representation of ITS2 ASVs using 2 beta diversity metrics: Jaccard and Bray–Curtis ecological distances. Principal coordinate analysis (PCoA) was performed on both Jaccard and Bray–Curtis dissimilarities. Permutational analysis of variance (PERMANOVA) was performed, followed by analysis of similarity (ANOSIM), to test the interspecific differences in strongylid nematode community compositions among the hosts. Multivariate GLMs from the R package mvabund (Wang et al., Reference Wang, Naumann, Wright and Warton2012) were implemented to search for community-wide divergence and identification of significant ASVs that varied due to the different host species effect.
Results
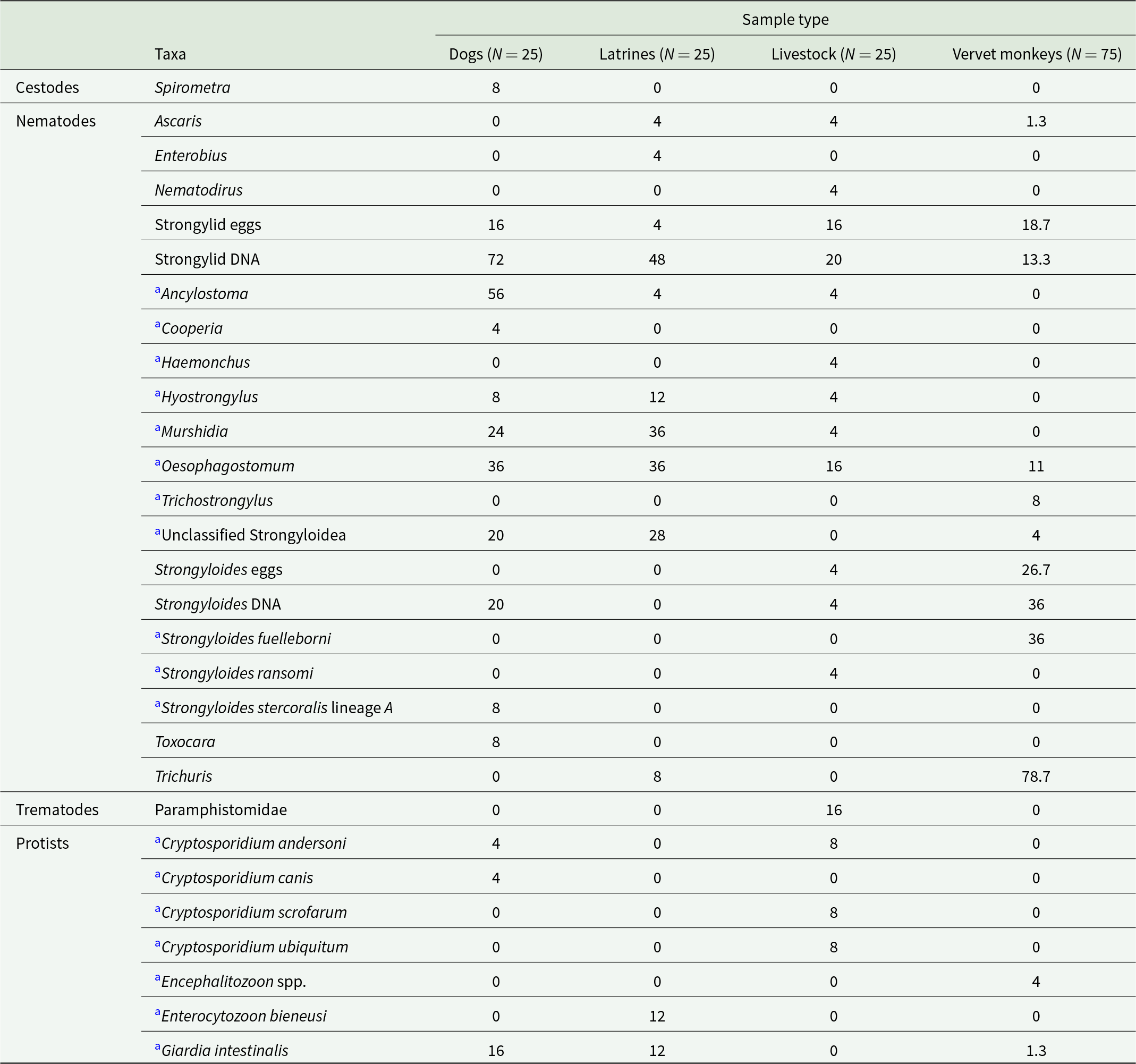
The total number of parasite taxa identified by microscopic and genetic analyses was 33 (Tables 1 and 2, Supplementary Table 3), with dogs harbouring 18 parasite taxa, livestock harbouring 14 parasite taxa, vervet monkeys harbouring 10 parasite taxa and in latrine samples 10 parasite taxa was detected. Using microscopy, cestodes, nematodes and trematodes were observed in 84 samples, whereas all protists were detected exclusively through molecular analyses (Table 1). We further focused on strongylid nematodes and Strongyloides. Based on microscopy alone, strongylid nematodes could not be differentiated and could not be reliably assigned to lower taxonomic level; the eggs were detected in 23 samples (dogs: 4, latrines: 1, livestock: 4, vervets: 14). In contrast, HTS enabled the discrimination of individual ASVs (Supplementary Table 1), with 45 samples (dogs: 18, latrines: 12, livestock: 5, vervets: 10) detected positive. Of the 23 samples identified positive by microscopy, only 7 samples were found positive by HTS sequencing. In contrast, HTS sequencing identified 38 positive samples that microscopy had identified as negative. Strongyloides eggs were identified in 21 (dogs: 0, latrines: 0, livestock: 1, vervets: 20) samples by microscopy, whereas 33 samples (dogs: 5, latrines: 0, livestock: 1, vervets: 27) tested positive using qPCR. Additionally, 15 samples that tested positive by microscopy were negative by qPCR, while 27 samples positive by qPCR were negative by microscopy.
Prevalence of parasite taxa detected by microscopic and genetic methods and their prevalence in the dogs, latrine samples, livestock and vervet monkeys at the shore of Lake Nabugabo, Uganda

a Parasite taxa identified from DNA sequencing.
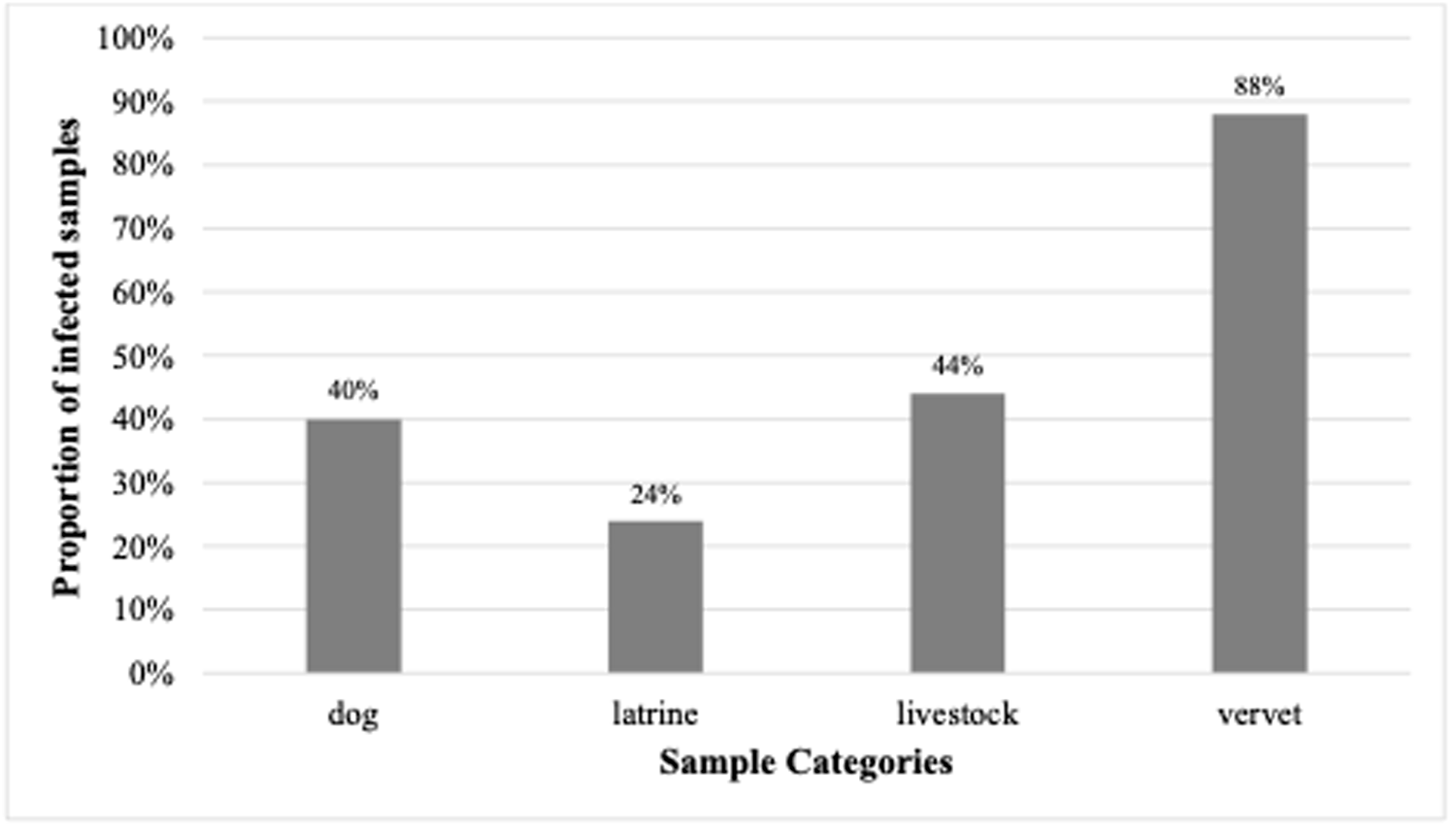
The microscopic analysis itself showed that the parasitic infection rate was 40% (10/25) in dogs, 24% (6/25) in latrine samples, 44% (11/25) in livestock and 88% (66/75) in vervet monkeys. Dogs (mean ± SE = 0.4 ± 0.1) were found to be more infected than latrine samples (mean ± SE = 0.24 ± 0.09) (χ 2 = 5.88, df = 1, p = 0.02; Fig. 1) and latrine samples were also less infected than livestock (mean ± SE = 0.44 ± 0.10) (χ 2 = 8.91, df = 1, p < 0.05; Fig. 1). All hosts were less infected than vervet monkeys (mean ± SE = 0.88 ± 0.04) (latrines: χ 2 = 83.12, df = 1, p < 0.001, dogs: χ 2 = 50, df = 1, p < 0.001, livestock: χ 2 = 43.14, df = 1, p < 0.001; Fig. 1). Other comparisons were not significant.
Proportion of infected samples from dogs (N = 25), latrines (N = 25), livestock (N = 25) and vervet monkeys (N = 75) at the shore of Lake Nabugabo, Uganda.

Strongylid nematodes
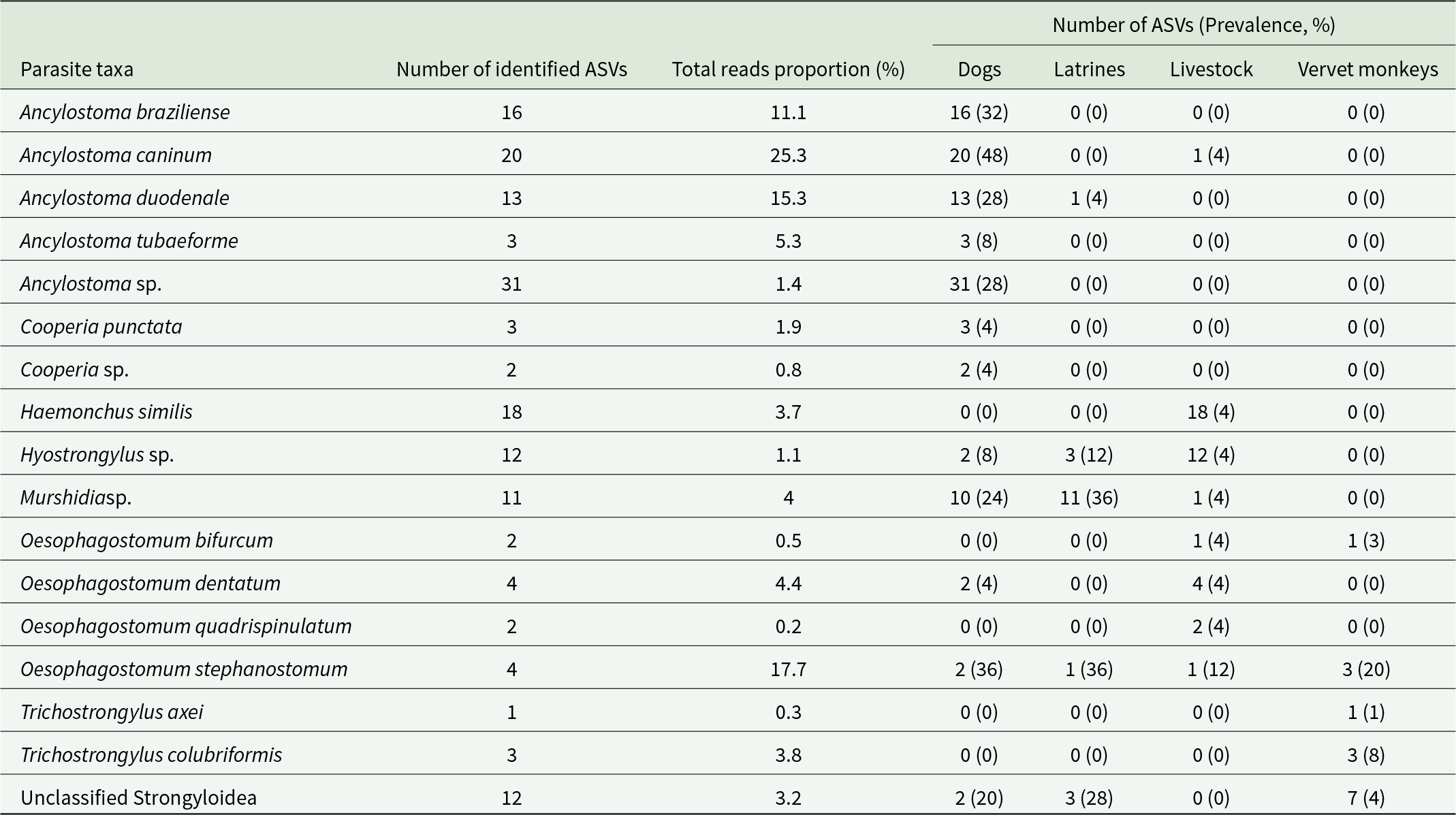
In total 1201 808 high-quality ITS2 reads were identified, with an average 22 675.62 reads (SD: 31 227.53; range: 223–166 772) and median sequencing depth of 12 422 reads per sample. In total, 157 individual ASVs were identified, including 7 strongylid genera namely Ancylostoma, Cooperia, Haemonchus, Hyostrongylus, Murshidia, Oesphagostomum and Trichostrongylus (Table 2). Of the known units, the highest number of ASVs belonged to Ancylostoma caninum in dogs, followed by Haemonchus similis in livestock (Table 2). In addition, 57 ASVs could not be identified to the species level (present in 16% of the samples), out of which 31 ASVs belonged to genus Ancylostoma, 2 ASVs to genus Cooperia. 12 ASVs to Hyostrongylus and 12 ASVs to order Rhabditida whose further taxonomy could not be assigned, and these are referred to as ‘Unassigned Strongyloidea’ further along.
Number of identified amplicon sequence variants (ASVs), their proportion of total reads, number of ASVs in each sample type and ASVs prevalence among host species

The shaded cells represent the taxa with ASVs detected in multiple sample types.
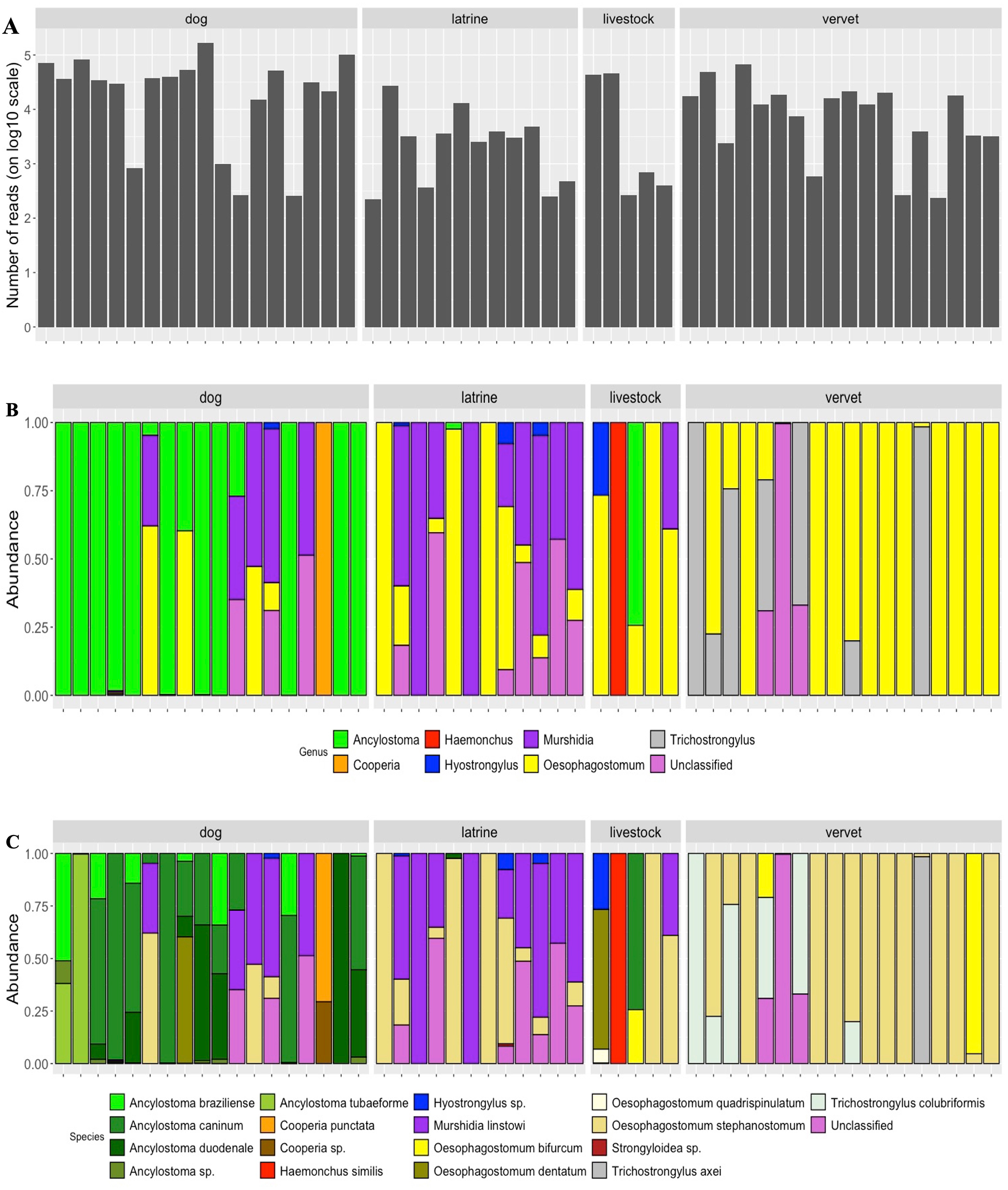
The number of reads varied significantly across sample type (χ 2 = 15.63, df = 3, p = 0.001) (Fig. 2A), with dogs having a higher number of reads compared to latrines (z = 37845, SE = 10439, df = 49, t = 3.63, p = 0.004) and vervet monkeys (z = 27881, SE = 9337, df = 49, t = 2.98, p = 0.02). The most prevalent variants belonged to 3 genera: Ancylostoma, Murshidia and Oesophagostomum (Table 2). There was a significant interspecific difference in the relative abundances of strongylid nematode communities among sample type (genus: χ 2 = 8.79, df = 3, p = 0.03; species: χ 2 = 24.24, df = 3, p < 0.001) (Fig. 2B). At the genus level, latrine samples harboured fewer genera compared to vervet monkeys (z = −0.28, SE = 0.10, df = 96, t = −2.77, p = 0.03; Fig. 2B). At the species level, vervet monkeys harboured more species compared to dogs (z = −0.37, SE = 0.08, df = 124, t = −4.85, p < 0.001) and latrine samples (z = −0.27, SE = 0.09, df = 124, t = −3.03, p = 0.02; Fig. 2C).
Bar plots showing (A) number of reads in each individual sample on a log10 scale, (B) relative community composition of strongylid nematodes in examined samples at the genus level, (C) relative community composition of strongylid nematodes in examined samples at the species level. Each column represents a sample. Numbers of reads (A)/relative abundances (B, C) of reads are depicted as colour panels.

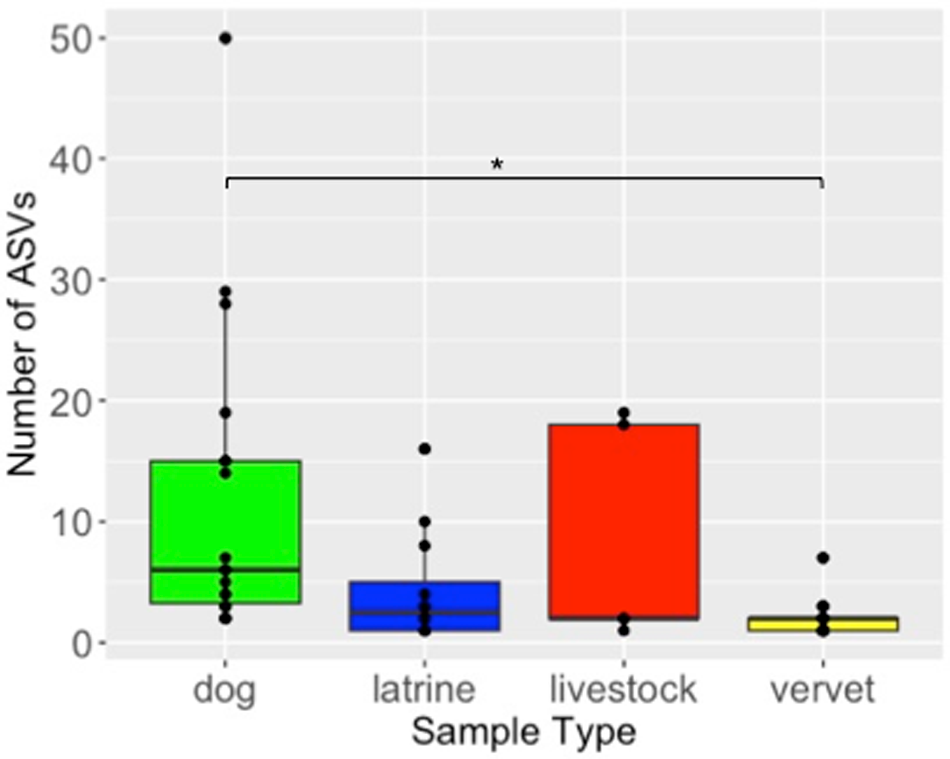
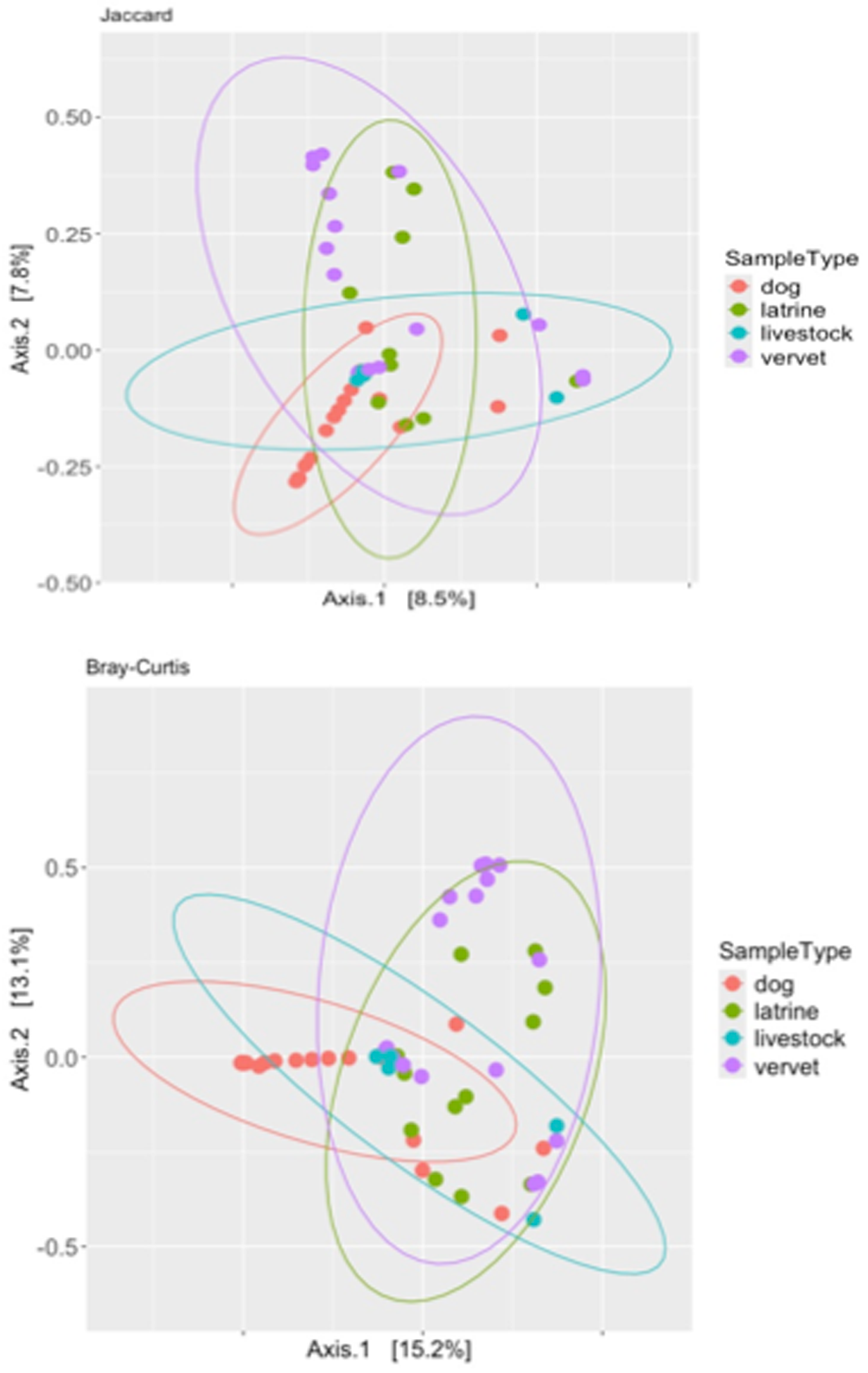
Variant diversity (explored as number of ASVs) differed significantly among sample type (GLM: F (3, 53) = 21.59, p < 0.001). Variant diversity in vervet monkeys was lower compared to dogs (z = 1.78, SE = 0.48, t ratio = 3.71, p = 0.001) (Fig. 3), while there was no evidence of significant differences between all other hosts (Fig. 3). PCoA diagrams based on both Jaccard and Bray–Curtis ecological distances confirmed clear differences among studied hosts in both composition and relative abundance of strongylid ASVs (Fig. 4). Significant differences between different sample types in the composition of their strongylid nematode communities were further confirmed by PERMANOVA (Jaccard: F (3,53) = 1.71, p < 0.001; Bray–Curtis: F (3,53) = 2.76, p < 0.001) and ANOSIM (Jaccard: R = 0.32, p = 0.001; Bray–Curtis: R = 0.32, p = 0.001) tests. Tukey post hoc testing revealed significant differences between dogs and latrine samples, dogs and vervets, and latrine samples and vervets for both Jaccard and Bray–Curtis distances (p < 0.05), while all other pairwise comparisons were not statistically significant.
Boxplot of amplicon sequence variants (ASVs) counts according to sample types.

PCoA ordination diagrams of beta diversity of strongylid nematode communities based on Jaccard ecological distance (presence/absence of amplicon sequence variants (ASVs)); Bray–Curtis ecological distance (relative abundances of reads).

Mvabund testing confirmed the interspecific differences (mvabund: ΔDF = 3, χ 2 = 749.1, p < 0.05) and identified 11 ASVs with whose different relative abundances were the main driving force of diversity among different host species in contrast to shared haplotypes (Supplementary Figure 2). Differences among hosts were mainly due to greater frequencies of A. caninum, A. duodenale, A. braziliense in dogs, Murshidia sp. in latrine samples and O. stephanostomum in vervets (Supplementary Figure 1).
Strongyloides
Molecular analysis revealed that 27 vervet monkey samples were infected with S. fuelleborni, whose HVR-IV sequences clustered with the isolates on GenBank sampled from non-human primates and humans from East and Central Africa as well as East Asia (Supplementary Figure 3). Dogs were infected with S. stercoralis lineage A. This was confirmed by the phylogenetic analysis, as the obtained sequences clustered with S. stercoralis lineage A sequences available in GenBank from humans, primates and dogs across Africa, Europe and Southeast Asia (Supplementary Figure 3). Three dogs were also found to be infected with S. ransomi, most likely a spurious parasitism as a consequence of eating pig faeces. One pig was also found to be infected with S. ransomi. Strongyloides ransomi found in dogs and pig clustered with the isolates of S. ransomi on GenBank sampled from wild boar in Brazil and the isolates of S. venezuelensis from brown rat (Rattus norvegicus) in Japan (Supplementary Figure 3).
Protists
Out of the 25 faecal samples collected from livestock, 24% tested positive for Cryptosporidium spp., with infection rates ranging from 0 to 8% (Table 1). In dogs (n = 25), 8% were positive, with infection rates from 0 to 4% (Table 1). In this study, 4 Cryptosporidium spp., namely C. andersoni in 1 dog and 2 cows, C. canis in 1 dog, C. scrofarum in 2 pigs and C. ubiquitum in 2 goats were identified. Cryptosporidium 18S rDNA sequences from a dog clustered with the isolates of C. canis on GenBank sampled from dogs in Europe and racoon dogs (Nyctereutes procyonoides), minks, blue foxes (Vulpes lagopus) and dogs in East Asia (Supplementary Figure 4). The Cryptosporidium 18S rDNA sequences from a dog also clustered with C. canis found in water in Spain (Supplementary Figure 4). The Cryptosporidium 18S rDNA sequences obtained from goats clustered with the isolates of C. ubiquitum on GenBank sampled from goats and Tibetan sheep (Ovis ammon hodgsoni) in China, hosts such as roe deer (Capreolus capreolus), common ostrich (Struthio camelus), nutria (Myocastor coypus) in Europe, alpaca (Vicugna pacos) in South America, long-tailed chinchilla (Chinchilla lanigera) in East Asia and eastern grey squirrel (Sciurus carolensis) and American red squirrel (Tamiasciurus hudsonicus) in North America (Supplementary Figure 4).
For Giardia intestinalis assemblage A, 16% of dog samples, 12% of latrine samples and 1.3% of vervet monkey samples were positive (Table 1). The TPI sequences obtained from dogs, latrines and vervet monkeys clustered with the isolates of G. intestinalis assemblage A on GenBank sampled from rhesus macaque (Macaca mulatta), goat, sheep (Ovis aries), François’ leaf monkey (Trachypithecus francoisi), cattle in China, sheep in USA, alpaca in Australia and humans in China, Australia and Malaysia (Supplementary Figure 5).
Enterocytozoon bieneusi was detected in 12% of human household samples (Table 1). The ITS sequences obtained from 2 latrine samples clustered with the isolates of Enterocytozoon bieneusi genotype Peru8 on GenBank sampled from human, guereza colobus (Colobus guereza), rhesus and Assamese macaques (Macaca assamensis) in China and humans in Peru (Supplementary Figure 6). Similarly, the ITS sequence obtained from 1 latrine sample clustered with the isolates of Enterocytozoon bieneusi genotype D on GenBank sampled from dogs, golden snub-nosed monkeys (Rhinopithecus roxellana) and northern white-cheeked gibbons (Nomascus leucogenys) in China, humans in China and Egypt, foxes (Vulpes vulpes) in Poland and mountain gorillas (Gorilla beringei beringei) in Rwanda (Supplementary Figure 6).
Out of 75 faecal samples collected from vervet monkeys, only 4% tested positive for Encephalitozoon species (Table 1). One of the ITS sequences obtained from vervet samples clustered with the isolates of Encephalitozoon intestinalis on GenBank from red panda (Ailurus fulgens) in China, Central African chimpanzee, moustached guenon monkey (Cercopithecus cephus) in Cameroon and humans in Russia and Iran (Supplementary Figure 7). The other 2 ITS sequences obtained from vervet samples were found 86.2% homologous with Encephalitozoon cuniculi on GenBank from red panda in China, 83.7% homologous with E. cuniculi from mountain gorilla in Rwanda, 82.9% homologous with E. cuniculi from cat in Austria and 82.6% homologous with E. cuniculi from rabbit (Oryctolagus cuniculus) in Japan (Supplementary Figure 7).
Shared parasites
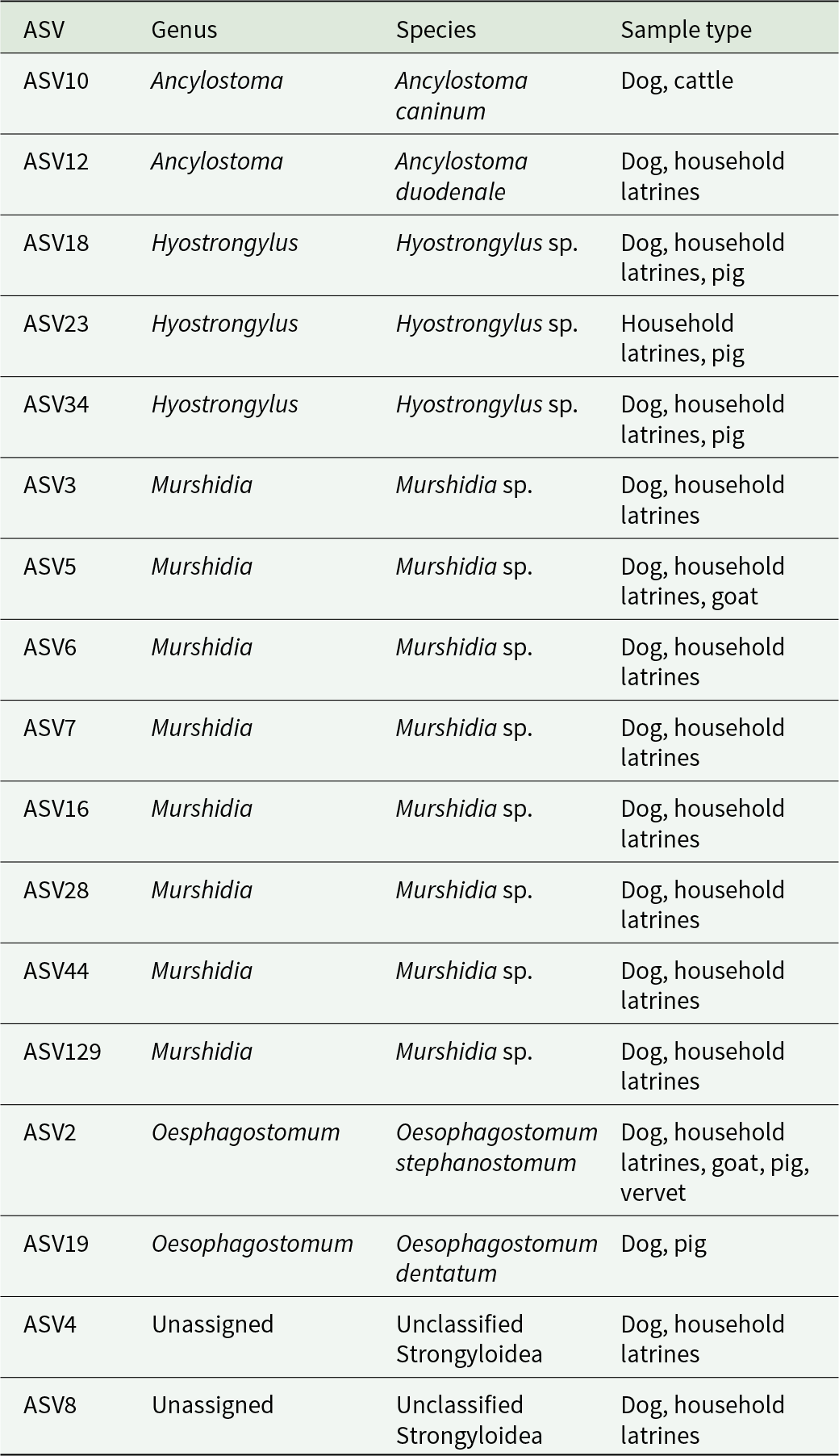
Overall, several parasite taxa were shared among multiple sample types (Tables 1 and 2), indicating potential cross-species transmission or overlapping environmental exposure. Eggs of Ascaris sp. were detected across latrine samples, pigs and vervets, whereas eggs of Trichuris sp. were detected in latrine samples and vervet monkeys (Table 1). Hyostrongylus was present in dogs, latrine samples and a pig, whereas Murshidia was shared among dogs, latrine samples and a goat (Table 2). Oesophagostomum bifurcum was shared between cattle and vervets, and O. dentatum was identified in both a dog and a pig (Table 2). Notably, O. stephanostomum was the most widespread, occurring in vervets, pigs, latrine samples and dogs (Table 2). Haplotypes of O. stephanostomum were shared across a wide range of hosts, including dogs, latrines, goats, pigs and vervets. Murshidia haplotypes were detected in dogs, goats and latrines while A. caninum was shared between dogs and cattle. Ancylostoma duodenale appeared in both dogs and latrine samples. Hyostrongylus haplotypes were found in dogs, latrines and pigs and O. dentatum was shared between dogs and pigs (Table 3). Cryptosporidium andersoni occurred in dogs and cattle (Table 1). Similarly, G. intestinalis was shared among dogs, latrine samples and vervets (Table 1).
Amplicon sequences variants (ASVs) shared among different sample types

Discussion
In this study, parasite prevalence, species richness and infection rate in dogs, latrine samples, livestock and vervet monkeys at Lake Nabugabo, Uganda were assessed by employing both microscopy and molecular techniques, identifying a total of 33 parasite taxa across the 4 sample types. The findings highlight important differences in parasite diversity and prevalence, shedding light on potential parasite transmission dynamics among wildlife, domestic animals and humans, along with species-specific differences in susceptibility or exposure (Fenton and Pedersen, Reference Fenton and Pedersen2005; Bielby et al., Reference Bielby, Fisher, Clare, Rosa and Garner2015).
A higher diversity of parasites was found in dogs, with 18 parasite taxa identified, compared to the other host species, likely due to coprophagy, close contact with humans and exposure to various parasites in domestic environments. Free-roaming dogs at Nabugabo may also be more prone to encountering contaminated areas and shared habitat with wildlife. In general, dogs have been linked to emerging human diseases like eosinophilic enteritis and alveolar echinococcosis and thus may pose risks for people (Salb et al., Reference Salb, Barkema, Elkin, Thompson, Whiteside, Black, Dubey and Kutz2008). Our findings are consistent with established research, which highlights dogs as a major source of zoonotic parasites, facilitating the transmission or transport of these parasites between livestock, wildlife and humans.
Strongylid nematodes
Among the 7 genera of strongylid nematodes found in this study, the most prevalent in dogs was Ancylostoma, while it was Oesophagostomum in vervets and latrine samples. These strongylids are commonly reported in a wide variety of hosts (Brooker et al., Reference Brooker, Bethony and Hotez2004; Ghai et al., Reference Ghai, Chapman, Omeja, Davies and Goldberg2014; Cibot et al., Reference Cibot, Guillot, Lafosse, Bon, Seguya and Krief2015; Pafčo et al., Reference Pafčo, Kreisinger, Čížková, Pšenková-Profousová, Shutt-Phillips, Todd, Fuh, Petrželková and Modrý2019; Nezami et al., Reference Nezami, Blanchard and Godoy2023). Interestingly, Hyostrongylus and Murshidia were also prevalent genera in latrine samples; however, these have never been reported from human samples before.
Ancylostoma caninum, one of the most common parasitic nematodes in dogs globally (Nezami et al., Reference Nezami, Blanchard and Godoy2023), was found in 12 dogs and in 1 cow, suggesting a possible passage through contaminated soil or water or faeces in that case. Ancylostoma duodenale was found in 7 dogs and 1 latrine sample. Although A. duodenale was detected in only 1 latrine sample, it is unclear how many individuals used that latrine, meaning that several users could have been infected and the true prevalence in the local population may be higher. Other studies have reported high prevalence of this parasite in humans, clearly confirming humans as its host (Adenusi and Ogunyomi, Reference Adenusi and Ogunyomi2003; Pafčo et al., Reference Pafčo, Kreisinger, Čížková, Pšenková-Profousová, Shutt-Phillips, Todd, Fuh, Petrželková and Modrý2019). One previous study documented its presence in humans in the Dzanga‐Sangha Protected Areas of the Central African Republic while the human samples were dominantly infected by Necator (Pafčo et al., Reference Pafčo, Kreisinger, Čížková, Pšenková-Profousová, Shutt-Phillips, Todd, Fuh, Petrželková and Modrý2019). Although A. duodenale often coexists with Necator americanus in many endemic areas (Adenusi and Ogunyomi, Reference Adenusi and Ogunyomi2003), we did not detect any Necator sequences in this study. Ancylostoma duodenale is considered a human-specific hookworm, with humans recognized as the only natural definitive host (CDC, 2024b) and its detection in the canine samples is likely a result of coprophagy of human faeces. Nevertheless, dogs may represent an important reservoir contributing to parasite transmission, as has been previously demonstrated for N. americanus (Boyko et al., Reference Boyko, Marie Harrison, Humphries, Galvani, Townsend, Otchere, Wilson and Cappello2020).
Dogs in this study were also infected with 3 additional species of Ancylostoma: A. braziliense, A. tubaeforme and an unclassified Ancylostoma species. While A. tubaeforme is typically considered a feline parasite, it is genetically and biologically related to dog hookworms such as A. caninum and A. braziliense. Notably, A. braziliense has been previously reported in cats in the United States of America, demonstrating its ability to infect multiple host species (Liotta et al., Reference Liotta, Koompapong, Yaros, Prullage and Bowman2012). Therefore, although presence of the feline species A. tubaeforme in dogs could be a consequence of coprophagy, it may also be a case of potential cross-species transmission. These findings underscore the diversity of hookworm infections in dogs and the potential for interspecies overlap among Ancylostoma species. Moreover, presence of these Ancylostoma species represents a risk for public health as canine hookworms cause larva migrans cutanea in humans (Nezami et al., Reference Nezami, Blanchard and Godoy2023).
Murshidia was found in dogs, latrine samples and livestock (goats). Murshidia species are primarily known to infect elephants (Loxodonta spp.), warthogs and other herbivores (Chel et al., Reference Chel, Iwaki, Hmoon, Thaw, Soe, Win, Bawm, Htun, Win, Oo, Masum, Ichii, Nakao, Nonaka and Katakura2020; Malviya et al., Reference Malviya, Vijayasarathi, Dohare, Raju, Yadav, Author and Gopika2024). To our knowledge, this is the first report of Murshidia in goats and dogs, although in the latter case is most likely a spurious parasite. The detection of Murshidia in human samples requires further consideration. It is possible that the presence of the parasite in human latrine samples was influenced by the method of sample collection. Specifically, the samples were obtained from pit latrines, where various materials, including livestock faeces, could have been discarded. This creates a risk of cross-contamination, which might explain the unexpected finding of Murshidia in human samples. It is also important to note that multiple individuals may have used the same pit latrine, and/or a single individual may have used more than 1 pit latrine, potentially resulting in multiple ‘pit latrine’ infections.
This study identified 4 species of Oesophagostomum across different host species, highlighting the broad distribution of these parasites in both domestic and wild animals, as well as humans. A single pig sample was found to be infected with O. stephanostomum, O. quadrispinulatum and O. dentatum. Similar taxa were reported in Tibetan pigs (Li et al., Reference Li, Lan, Luo, Shahzad, Zhang, Wang, Zhang, Liu, Liu, Hao, Sizhu and Li2017), although the identification of O. stephanostomum in that study remains uncertain due to the low BLAST sequence identity and incongruent phylogenetic placement relative to authentic O. stephanostomum references. Given that pigs are not typical hosts for O. stephanostomum, its occurrence here could represent a spurious infection or an unrecognized cross-host transmission event, both of which warrant further molecular investigation.
Two Oesophagostomum species are commonly found in both human and non-human primates in Africa: O. stephanostomum mostly in great apes and occasionally in humans (Cibot et al., Reference Cibot, Guillot, Lafosse, Bon, Seguya and Krief2015; Narat et al., Reference Narat, Guillot, Pennec, Lafosse, Grüner, Simmen, Bokika Ngawolo and Krief2015; Ilík et al., Reference Ilík, Kreisinger, Modrý, Schwarz, Tagg, Mbohli, Nkombou, Petrželková and Pafčo2023) and O. bifurcum in humans and various monkeys (Ziem et al., Reference Ziem, Magnussen, Olsen, Horton, Asigri and Polderman2006; Ghai et al., Reference Ghai, Chapman, Omeja, Davies and Goldberg2014; Sirima et al., Reference Sirima, Bizet, Hamou, Červená, Lemarcis, Esteban, Peeters, Mpoudi Ngole, Mombo, Liégeois, Petrželková, Boussinesq and Locatelli2021). We detected both species in vervet monkeys with O. stephanostomum having higher prevalence and this species was also found in human latrine samples. While there is not much information available on pathogenicity of O. stephanostomum in humans, O. bifurcum is known cause of potentially severe illness in some areas of West Africa (Ziem et al., Reference Ziem, Magnussen, Olsen, Horton, Asigri and Polderman2006). Their presence in the area deservers further attention, which should focus on a detailed epidemiology and clinical picture of the infection in humans. The finding that O. dentatum was shared between pigs and dogs is consistent with previous studies suggesting that Oesophagostomum species can be maintained in multiple definitive host species (Ghai et al., Reference Ghai, Chapman, Omeja, Davies and Goldberg2014; Cháves-González et al., Reference Cháves-González, Morales-Calvo, Mora, Solano-Barquero, Verocai and Rojas2022), further complicating the ecological dynamics of these parasites; that said, the presence of O. dentatum in dogs is more likely a result of coprophagy.
Among other strongylid nematodes detected in various hosts, genus Hyostrongylus was found in human, dog and pig samples. Hyostrongylus is typically found in pigs (Kouam and Ngueguim, Reference Kouam and Ngueguim2022), and the presence of Hyostrongylus in dogs could be a spurious infection, while its presence in humans could be a result of sampling method from pit latrines. Thus, this finding requires further examination. In this study, variants of Cooperia punctata and a Cooperia sp. were found in dogs, which could be the result of coprophagy. The genus Cooperia belongs to Trichostrongyloid nematodes that are important in veterinary medicine, as they play a role in mixed-species helminth infections, leading to production losses in livestock worldwide (Stromberg et al., Reference Stromberg, Gasbarre, Waite, Bechtol, Brown, Robinson, Olson and Newcomb2012).
Dogs exhibited higher strongylid alpha diversity than vervets. This high diversity among dogs was attributed primarily to the prevalence and dominance of A. caninum variants, which not only exhibited high infection rates but also contributed substantially to the overall relative abundance of strongylid nematodes in the host population. Specifically, dogs were found to harbour 11 distinct species of strongyle nematodes, raising the possibility of dogs as a major reservoir of these parasites. However, it should be highlighted that most of these infections are likely due to coprophagy of infected faeces from animals or humans. That said, dogs can still spread eggs or larvae that may remain infectious to other hosts (e.g. in case of N. americanus (Boyko et al., Reference Boyko, Marie Harrison, Humphries, Galvani, Townsend, Otchere, Wilson and Cappello2020)) which warrants further investigation. On the other hand, the strongylid communities of humans, livestock and vervets were dominated by variants belonging to O. stephanostomum, T. colubriformis and Murshidia sp. As previously noted, however, the detection of Murshidia sp. in human samples warrants careful consideration.
Strongylid nematodes are capable of surviving in their hosts for years and infection typically do not lead to death (Ilík et al., Reference Ilík, Kreisinger, Modrý, Schwarz, Tagg, Mbohli, Nkombou, Petrželková and Pafčo2023). In fact, none of the sampled animals displayed obvious symptoms of severe infections such as respiratory difficulties, anaemia, weight loss, diarrhoea, worsened quality of hair, apathy, inappetence, etc. (Ilík et al., Reference Ilík, Kreisinger, Modrý, Schwarz, Tagg, Mbohli, Nkombou, Petrželková and Pafčo2023). Dogs, which showed the highest strongylid diversity in this study, could, therefore, contribute to environmental contamination even when infections are subclinical. Similarly, vervets and livestock, especially goats and pigs, harboured several taxa usually linked to other host species, suggesting they may sustain these parasites without noticeable pathology. Such asymptomatic infections can support cross-species transmission and complicate our understanding of local parasite dynamics and zoonotic risk. Moreover, even if the infection will not establish and the parasites stages just pass through the digestive tract of the host, these individuals then serve as vectors and may spread the stages in the environment (Boyko et al., Reference Boyko, Marie Harrison, Humphries, Galvani, Townsend, Otchere, Wilson and Cappello2020).
Strongyloides
Despite identifying S. fuelleborni in vervet monkeys and S. stercoralis in dogs, both of which are Strongyloides species that have been previously found in humans (Hasegawa et al., Reference Hasegawa, Hayashida, Ikeda and Sato2009, Reference Hasegawa, Kalousova, McLennan, Modry, Profousova-Psenkova, Shutt-Phillips, Todd, Huffman and Petrzelkova2016b; Schär et al., Reference Schär, Guo, Streit, Khieu, Muth, Marti and Odermatt2014; Thanchomnang et al., Reference Thanchomnang, Intapan, Sanpool, Rodpai, Tourtip, Yahom, Kullawat, Radomyos, Thammasiri and Maleewong2017), we did not detect any Strongyloides infections in human latrine samples at Lake Nabugabo, Uganda. Of particular concern is S. stercoralis, a soil-transmitted helminth known to infect humans, non-human primates and dogs (Olsen et al., Reference Olsen, van Lieshout, Marti, Polderman, Polman, Steinmann, Stothard, Thybo, Verweij and Magnussen2009). Globally, S. stercoralis represents a significant public health concern because it can establish lifelong infections through autoinfection, often remaining undetected for decades, and can progress to hyperinfection or disseminated strongyloidiasis in immunocompromised individuals (Keiser and Nutman, Reference Keiser and Nutman2004; Buonfrate et al., Reference Buonfrate, Requena-Mendez, Angheben, Muñoz, Gobbi, Van Den Ende and Bisoffi2013). In humans, clinical manifestations range from mild or asymptomatic infection to gastrointestinal, respiratory and dermatological symptoms and in cases of hyperinfection, patients may develop severe disseminated disease and sepsis, carrying more than 60% mortality rates (Buonfrate et al., Reference Buonfrate, Requena-Mendez, Angheben, Muñoz, Gobbi, Van Den Ende and Bisoffi2013; Geri et al., Reference Geri, Rabbat, Mayaux, Zafrani, Chalumeau-Lemoine, Guidet, Azoulay and Pène2015). Some estimates suggested that S. stercoralis infects over 600 million people and is the main species responsible for strongyloidiasis (Buonfrate et al., Reference Buonfrate, Bisanzio, Giorli, Odermatt, Fürst, Greenaway, French, Reithinger, Gobbi, Montresor and Bisoffi2020). Although widespread community outbreaks are uncommon, S. stercoralis clusters have been documented in institutional settings, including long-term facilities, mental health institutions and among populations with poor sanitation (Schär et al., Reference Schär, Trostdorf, Giardina, Khieu, Muth, Marti, Vounatsou and Odermatt2013).
While human infections with S. fuelleborni have also been reported, these are rare and are typically associated with individuals who frequent areas inhabited by non-human primates, where S. fuelleborni is believed to originate (Hasegawa et al., Reference Hasegawa, Sato, Fujita, Nguema, Nobusue, Miyagi, Kooriyama, Takenoshita, Noda, Sato, Morimoto, Ikeda and Nishida2010). Nevertheless, there are rare cases of imported and asymptomatic S. fuelleborni infection in people. For instance, a Belgian student who returned from the Democratic Republic of the Congo was treated at the Institute of Tropical Medicine in Antwerp for fever, rash and diarrhoea, where stool microscopy revealed Trichuris trichiura and Strongyloides eggs, leading to a molecularly confirmed diagnosis of S. fuelleborni infection (Potters et al., Reference Potters, Micalessi, Van Esbroeck, Gils and Theunissen2020). Although we did not detect human infections in this study, the presence of this species in vervet monkeys and dogs indicates a potential risk of zoonotic transmission, which has been confirmed in humans, dogs and gorillas in the Central African Republic (Nosková et al., Reference Nosková, Ilík, Stéphane Singa Niatou, Dumas, Fuh, Dicky, Kurucová, Baláž, Judita Petrželková and Pafčo2026). This is especially concerning given the close proximity between humans and these animals, as well as the possibility of human exposure to environmental contamination with parasite larvae. This makes S. stercoralis a major cause of long-term health issues and suffering (Vadlamudi et al., Reference Vadlamudi, Chi and Krishnaswamy2006; Prendki et al., Reference Prendki, Fenaux, Durand, Thellier and Bouchaud2011; Schär et al., Reference Schär, Trostdorf, Giardina, Khieu, Muth, Marti, Vounatsou and Odermatt2013). A pig sample and 3 dog samples were found to be infected with S. ransomi. Strongyloides ransomi is known to infect pigs globally (Giang et al., Reference Giang, Nguyen, Hoan, Thu Huyen, Kim Lan and Doanh2017); thus, finding pig and dog samples infected with S. ransomi was not surprising. However, dogs might have been positive because of consumption of pig faeces, resulting in spurious parasitism.
Protists
Cryptosporidium, Giardia. and microsporidia species are capable of infecting a broad range of hosts and crossing species barriers, owing to their low host specificity (Thompson, Reference Thompson, Sterling and Adam2004; Franzen, Reference Franzen2008). Despite numerous reports of Cryptosporidium spp. in primates (Lim et al., Reference Lim, Ngui, Shukri, Rohela and Mat Naim2008; Karim et al., Reference Karim, Dong, Yu, Jian, Zhang, Wang, Zhang, Rume, Ning and Xiao2014; Sak et al., Reference Sak, Petrželková, Květoňová, Mynářová, Pomajbíková, Modrý, Cranfield, Mudakikwa and Kváč2014; Butel et al., Reference Butel, Mundeke, Drakulovski, Krasteva, Ngole, Mallié, Delaporte, Peeters and Locatelli2015; Mynářová et al., Reference Mynářová, Foitová, Kváč, Květoňová, Rost, Morrogh-Bernard, Nurcahyo, Nguyen, Supriyadi and Sak2016), our study did not detect any Cryptosporidium infections in either latrine samples or vervet monkeys. Cryptosporidium andersoni, a common species in ruminants (Lindsay et al., Reference Lindsay, Upton, Owens, Morgan, Mead’ and Blagburnc2000; Kváč et al., Reference Kváč, Kouba and Vítovec2006), was detected only in 2 cows in this study. The finding that only a small number of host samples tested positive for other Cryptosporidium species – such as C. andersoni and C. canis in 2 dogs, C. scrofarum in 2 pigs and C. ubiquitum in 2 goats suggests that these species may be present at a low prevalence in the studied populations. Although the prevalence across all host species was low, the presence of these Cryptosporidium species remains epidemiologically relevant. As many domestic animals can shed oocysts even in the absence of clinical symptoms, the infected cows, dogs, pigs and goats identified here could act as asymptomatic reservoirs (Hatam-Nahavandi et al., Reference Hatam-Nahavandi, Ahmadpour, Carmena, Spotin, Bangoura and Xiao2019). This has potential implications for environmental contamination, particularly in shared water sources or areas where humans and wildlife overlap (Berhanu et al., Reference Berhanu, Ayana, Megersa, Ashenafi and Waktole2022). At Nabugabo, cows are often brought to the lake to drink where they likely defecate in water that is sometimes used for laundry or swimming. Thus, even low-level circulation of these species may contribute to transmission risk for more vulnerable hosts, including young animals, immunocompromised individuals and wildlife species with higher susceptibility (Berhanu et al., Reference Berhanu, Ayana, Megersa, Ashenafi and Waktole2022).
Among the 2 assemblages (A and B) of G. intestinalis most commonly reported in humans, dogs, cats, livestock and wild animals including non-human primates (Feng and Xiao, Reference Feng and Xiao2011; Cama and Mathison, Reference Cama and Mathison2015; Kváč et al., Reference Kváč, Hofmannová, Ortega, Holubová, Horčičková, Kicia, Hlásková, Květoňová, Sak and McEvoy2017), our study identified assemblage A in latrine samples, dogs and a vervet monkey, suggesting a potential for zoonotic transmission. Giardia, primarily assemblages A and B, is among the most prevalent intestinal parasites affecting humans, with an estimated 200 million people in Asia, Africa and Latin America suffering from symptomatic infections (Yason and Rivera, Reference Yason and Rivera2007). The presence of assemblage A in both latrine samples and animals at Lake Nabugabo underscores the potential role of animals as reservoirs for human infections, particularly given that there is habitat overlap between human populations, domestic animals and vervet monkeys.
In this study, E. bieneusi was detected exclusively in latrine samples, which aligns with previous research indicating that this species is the primary causative agent of microsporidiosis in humans. Enterocytozoon bieneusi is responsible for over 90% of human microsporidiosis cases worldwide, making it the most significant species of microsporidia in terms of public health impact (Xu et al., Reference Xu, Ma, Zhang, Zhang, Zhao, Ba, Rui-Du, Xing, Wang and Zhao2016; Khanduja et al., Reference Khanduja, Ghoshal, Agarwal, Pant and Ghoshal2017; Qiu et al., Reference Qiu, Xia, Li, Ping, Ding and Liu2019; Tao et al., Reference Tao, Ni, Du, Jiang, Li, Qiu, Ye-Li and Zhang2020). However, one surprising and somewhat unexpected finding of this study is that only 12% of latrine samples tested positive for E. bieneusi. This low prevalence within latrine samples contrasts with the global distribution of E. bieneusi infections and raises important questions about the factors influencing the parasite’s transmission in this region. Given the established zoonotic potential of E. bieneusi and its known presence in a variety of hosts, including livestock, pets and wildlife (Tao et al., Reference Tao, Ni, Du, Jiang, Li, Qiu, Ye-Li and Zhang2020), one might expect a higher prevalence of infection within households where humans often interact with animals. One possibility is that household-level transmission may be less common in this region, either due to effective sanitation practices or the fact that E. bieneusi may primarily be transmitted through waterborne routes, as is commonly seen in other endemic areas (Xu et al., Reference Xu, Ma, Zhang, Zhang, Zhao, Ba, Rui-Du, Xing, Wang and Zhao2016); while humans sometimes use lake water for laundry and swimming, well-water is available. In humans, E. bieneusi infections are often asymptomatic or mild, especially in immunocompetent individuals, but when symptoms occur, they typically include watery diarrhoea, abdominal discomfort and malabsorption (Nourrisson et al., Reference Nourrisson, Lavergne, Moniot, Morio and Poirier2024). From a veterinary perspective, E. bieneusi is also important because it infects a wide range of domestic and wild animals, and these animals can act as reservoirs for human infection (Taghipour et al., Reference Taghipour, Bahadory, Khazaei, Zaki, Ghaderinezhad, Sherafati and Abdoli2022). Another potential explanation for the low prevalence of E. bieneusi in household samples could be related to the diagnostic methods employed in this study. While molecular techniques such as PCR are highly sensitive, the detection of E. bieneusi can sometimes be limited by the timing of sample collection or the presence of low parasite loads. Additionally, because of the use of pooled pit latrine samples, it is not possible to determine the exact number of infected people or track individual infection dynamics. Therefore, it is possible that the 12% prevalence reflects only the most overt cases of infection, while subclinical or lower-level infections may have gone undetected.
Finally, a microsporidium closely related to Encephalitozoon intestinalis was detected only in 4% (N = 3) of vervet monkey samples. Although we did not detect this microsporidia or E. intestinalis in other host species, including human latrine samples, previous research has shown that E. intestinalis does infect humans in other regions. Studies have reported E. intestinalis infections in humans, particularly in immunocompromised individuals, such as those with HIV/AIDS or undergoing chemotherapy (Sokolova et al., Reference Sokolova, Demyanov, Bowers, Didier, Yakovlev, Skarlato and Sokolova2011; Tavalla et al., Reference Tavalla, Mardani-Kateki, Abdizadeh, Nashibi, Rafie and Khademvatan2017). The low prevalence of E. intestinalis observed in vervet monkeys in this study may reflect similarly low prevalence rates in other animal species in the region, which could be influenced by factors such as local environmental conditions and hygiene practices.
Limitations
This study has a few important limitations. Because we relied on pooled pit latrine samples, we could not link parasite detections to specific individuals, and heavily infected people may have contributed disproportionately to what we found. Some degree of cross-contamination within latrines is also possible. Pit latrines are common areas where human waste is discarded, but they can also contain other materials, including livestock and dog faeces. The mixing of human and animal waste in these environments could raise the risk of cross-contamination, particularly with parasites that rely on livestock or dogs as their definitive and/or reservoir hosts, which may compromise the accuracy of the findings. Another limitation of our study is that our molecular approach detects parasite DNA but cannot tell whether it reflects an active infection or simply the transient passage of ingested material, which is especially relevant for unexpected host–parasite matches. Furthermore, our sampling was cross-sectional, it captures only a single point in time and does not allow us to determine transmission direction or seasonal patterns. Additionally, we were not able to sample all potential hosts or environmental sources, such as soil or water, which limits how fully we can describe the transmission system. Finally, discrepancies between microscopy and molecular detection were observed. DNA-positive microscopy-negative cases are consistent with the higher sensitivity of molecular methods, which can detect low-intensity infections that may be missed under the microscope. In contrast, microscopy-positive DNA-negative results may stem from occasional difficulties in distinguishing similar morphological features or from the uneven distribution of parasites stages within faecal samples. These differences reflect the inherent strengths and limitations of each diagnostic method and should be taken into account when interpreting the findings. Future work with longitudinal sampling, more detailed genotyping and environmental monitoring will be important for resolving transmission pathways and better understanding the associated risks.
Conclusion
This study highlights the diverse and complex patterns of parasitic infections found in humans, domestic animals and wildlife, with dogs and livestock showing the highest diversity, and vervet monkeys and livestock showing the highest infection rate. The 33 parasite taxa identified across different sample types emphasize the ecological connections among the host species and the potential zoonotic risks involved. Our findings carry important implications for both public health and local livelihoods. Several parasites detected can cause disease even in healthy individuals, highlighting real risks for nearby communities. Livestock parasites like Trichostrongylus and Haemonchus also suggest potential production losses in goats and cattle, which can impact households that depend on small-scale farming. Because humans, livestock, dogs and vervet monkeys live in close contact around Lake Nabugabo, this area functions as a shared transmission hotspot where zoonotic and anthroponotic parasites can circulate easily. Although we did not formally assess parasite-prevention practices, several local factors likely affect transmission. Dogs roam freely, and most livestock, except pigs, graze in open areas, increasing environmental contamination. Water is mainly drawn from wells (shaduf) or the lake. Some residents boil water or wear footwear, but it is unclear how consistently these practices are followed or whether they are used specifically for parasite prevention. These conditions suggest that exposure to contaminated soil and water is common and may help explain some of our results.
Supplementary material
The supplementary material for this article can be found at https://figshare.com/s/56daa2a3c059b68ae1d8.
Data availability statement
Data are available at: https://figshare.com/s/7183b93db8e369344c6a.
Acknowledgements
We thank the Uganda Wildlife Authority (UWA), the Uganda National Council for Science and Technology (UNCST), the Makerere University School of Health Sciences Research Ethics Committee (MAKSHSREC) and the School of Veterinary Medicine and Animal Resources Institutional Animal Care and Use Committee (SVAR IACUC) for granting permission to conduct this research. We thank Dr Dennis Twinomugisha, Matovu Ponsyano, Katwere Livingstone and Justine Namuyomba for logistical support and assistance with the data collection. We also extend our sincere appreciation to the residents of Bbaale village for their willingness to participate and provide samples. Finally, we thank Lordrick Alinaitwe and Ester Nansubuga at Makerere University, Uganda for conducting the microscopic analysis of the samples. OpenAI GPT-5 (i.e. ChatGPT) was used for light editing (e.g. grammar verification).
Author contributions
PU conceived and designed the study, collected the samples and conducted statistical analyses. BC, MK, KS, BP, EN, VI and NH conducted genetic analyses. PU, BC, MK, KS, BP, EN, VI and VAMS wrote and edited the manuscript. PU and VAMS acquired the funding.
Financial support
Funding for the research was provided by the Anne Innis Dagg Grant from the Canadian Society of Zoologists (PU), NSERC and New Frontiers in Research Fund (VAMS). The work of BC and BP was funded by the Czech Science Foundation (22-16475S).
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
Research was approved by York University (ACC-2016-7W, ACC-2020-06; HPRC-E2019-191), Uganda Wildlife Authority (UWA EC 361 and COD-96-05), Uganda National Council for Science and Technology (NS349ES) and School of Veterinary Medicine and Animal Resources, Institutional Animal Care and Use Committee (SVAR_IACUC/125-2022).