Introduction
The genus Clavellotis (Castro-Romero & Baeza-Kuroki, Reference Castro-Romero and Baeza1984) currently contains 14 species occurring in marine fishes across the world (Castro-Romero et al., Reference Castro-Romero, Montes, Ӧktener, Ibañez-Shimabukuro, Theiller and Campos2025). However, only 1 congeneric species has been previously reported in Mexico, C. sebastidis Castro and González (Reference Castro and González2005), from the gills of Sebastes miniatus on the Pacific coast (Rodríguez-Santiago et al., Reference Rodríguez-Santiago, Rosales-Casián and Grano-Maldonado2014, Reference Rodríguez-Santiago, Gómez, Rosales-Casián and Grano-Maldonado2015, Reference Rodríguez-Santiago, Rosales-Casián and Grano-Maldonado2016; Raupach et al., Reference Raupach, Barco, Beermann, Laakmann, Mohrbeck, Neumann, Kihara, Pointner, Radulovici, Segelken-Voigt, Wesse and Knebelsberger2015). Still, there are no records of Clavellotis on marine fishes across the Atlantic coast of Mexico.
During the course of a survey on the metazoan parasites of marine and estuarine fish of the Yucatán Peninsula, specimens of a lernaeopodid copepod were collected from the gills of three species of sparids, i.e. Lagodon rhomboides (L.), Archosargus rhomboidalis (L.) and Archosargus probatocephalus (Walbaum) in 2 localities. These specimens were consistent with the concept of Clavellotis and represented an undescribed species. The new species is described herein by using morphological and molecular characters, and its position within the phylogenetic tree of the genus is discussed.
Materials and methods
Sample collection
Between 2022 and 2025, 120 individuals of 3 species of sparids, namely L. rhomboides (n = 60), A. rhomboidalis (n = 31), and A. probatocephalus (n = 29) were collected from 2 localities of the Yucatán Peninsula, La Carbonera lagoon (21°13’–21°14’ N; 89°52’–89°54’ W), and Celestún lagoon (21°13’54’’ N; 89°53’25’’ W); some specimens were also sampled from offshore Celestún (20°52’05’’ N; 90°29’08’’ W) (Figure 1). Fish were obtained from commercial capture and, in some cases, by using seine nets. In the latter case, fish were kept alive in containers with salt water and constant aeration until necropsied; these fish were euthanized by spinal severance (pithing), following the procedures accepted by the American Veterinary Medical Association (AVMA, 2020). All individuals were dissected, and gills were examined under a stereomicroscope. Specimens were preserved in 100% ethanol for further analysis. Morphological terminology follows Kabata (Reference Kabata1979), Castro-Romero and Baeza (Reference Castro-Romero and Baeza1984) and Boxshall and Halsey (Reference Boxshall and Halsey2004). Measurements of specimens are given in micrometres. Specimens were deposited in the Colección Nacional de Crustáceos (CNCR), Instituto de Biología, Universidad Nacional Autónoma de México, Mexico City.
Map of the Yucatán Peninsula, Mexico showing the sampling localities, represented with a red star.

DNA extraction, amplification and sequencing
Total genomic DNA was extracted from the egg sacs of individual copepods. Each egg sac was digested overnight at 56 °C in a solution containing 10 mM Tris-HCl (pH 7.6), 20 mM NaCl, 100 mM Na2 EDTA (pH 8.0), 1% sarkosyl and 0.1 mg mL−1 proteinase K. Total genomic DNA was extracted using DNAzol (Molecular Research Center, Cincinnati, OH) following manufacturer’s instructions. The target genes were amplified via Polymerase Chain Reaction (PCR). The partial cytochrome c oxidase subunit I (coxI) gene was amplified using the forward primer LCO1490 (5’-GGTCAACAAATCATAAAGATATTG-3’) and reverse primer HCO2198 (5’-TAAACTTCAGGGTGACCAAAAAATCA-3’) (Folmer et al., Reference Folmer, Black, Hoeh, Lutz and Vrijenhoek1994), using the following cycling conditions: 4 min initial denature at 94°C, followed by 34 cycles of 94 °C for 30 sec, 44–48 °C for 1 min and 72 °C for 1 min; and a final extension phase at 72 °C for 7 min. The large subunit of the nuclear ribosomal RNA gene (28S) was amplified using the forward primer 391 (5’-AGCGGAGGAAAAGAAACTAA-3’) (Nadler and Hudspeth, Reference Nadler and Hudspeth1998) and reverse primer 536 (5’-CAGCTATCCTGAGGGAAAC-3’) (García-Varela and Nadler, Reference García-Varela and Nadler2005). Sequencing internal primers were 502 plus 503 (Stock et al., Reference Stock, Campbell and Nadler2001; García-Varela and Nadler, Reference García-Varela and Nadler2005). Cycling conditions: 4 min initial denature at 95 °C followed by 30 cycles of 94 °C for 30 seg, 50 °C for 45 seg and 72 °C for 2 min; and a final extension phase at 72 °C for 7 min. Contigs were assembled and base-calling differences were resolved using Geneious Pro 4.8.4 (Biomatters Ltd). Sequences were compared to the GenBank library content with the BLAST tool, and were deposited at the GenBank database (Table 1).
Collection data and GenBank accession numbers of the copepods used in this study. Newly generated sequences are shown in bold

a Unpublished sequences available in the GenBank dataset.
Sequence comparison and phylogenetic analysis
Sequences were aligned using MAFFT v.7 (Katoh et al., Reference Katoh, Rozewicki and Yamada2019) via the online server. The best partitioning scheme and substitution model were selected under the Bayesian Information Criterion (BIC; Schwarz, Reference Schwarz1978) using the ‘greedy’ algorithm implemented in PartitionFinder v.1.1.1 (Lanfear et al., Reference Lanfear, Calcott, Kainer, Mayer and Stamatakis2014). The optimal nucleotide substitution models were SYM + I + G for the first, GTR + I + G for the second, GTR + G for the third codon positions of COI, and K80 + G for 28S. Phylogenetic reconstruction was performed with Bayesian Inference (BI) in MrBayes v.3.2.3 (Ronquist et al., Reference Ronquist, Teslenko, Van Der Mark, Ayres, Darling, Höhna, Larget, Liu, Suchard and Huelsenbeck2012). Two independent Metropolis-Coupled Markov Chain Monte Carlo (MCMC) analyses were run for 20 million generations, sampling every 1000 generations. Convergence was assessed by ensuring that the average standard deviation of split frequencies remained below 0.01, as recommended by Ronquist et al. (Reference Ronquist, Teslenko, Van Der Mark, Ayres, Darling, Höhna, Larget, Liu, Suchard and Huelsenbeck2012). Posterior Probabilities (PP) were used as measures of clade support. A majority-rule consensus tree with branch lengths was constructed after discarding the first 25% of sampled trees as burn-in. To estimate divergence among lineages, pairwise genetic distances (p-distance; Nei and Kumar, Reference Nei and Kumar2000) were calculated. Distance matrices were generated in MEGA X (Kumar et al., Reference Kumar, Stecher, Li, Knyaz and Tamura2018), using 1000 bootstrap replicates and assuming uniform nucleotide substitution rates (transition + transversions).
Results
Systematics
Class Copepoda (Milne Edwards, 1840)
Order Siphonostomatoida (Burmeister, 1835)
Family Lernaeopodidae (Milne Edwards, 1840)
Genus Clavellotis Castro-Romero & Baeza-Kuroki, Reference Castro-Romero and Baeza1984
Clavellotis mayae n. sp.
Description
Female: Measurements (based on the holotype and 23 paratypes specimens) in micrometres (Figures 2, 3, 5A, 5B, and 5C). Cephalothorax 2217–2914 (2436; n = 23) length by 206–548 (398; n = 23) width (Figure 2A). Dorsal shield well defined, tapering posteriorly (Figure 2B). Maxilla 563–1016 (729; n = 22) length by 178–473 (330; n = 22) width; shorter than cephalothorax, reaching ca. 28.9% of its length, separated, and fused only distally; each maxilla has 2 lobes; aliform processes oval, oblique, near base of maxilla and distant from excretory duct (Figures 2A, 2C, and 2D). Trunk 1265–1912 (1,663; n = 24) length by 636–984 (845; n = 24) width (Figure 2A); distal part wider than medial and basal regions; elongated, pyriform, with a conspicuous subcircular projection (flap) covering egg-sac insertion (Figure 3A). Genital process short 155–311 (253; n = 22) length by 116–195 (164; n = 22) width, blunt distally (Figure 3A). Egg sac 782–2,760 (1980; n = 17) length by 183–651 (392; n = 17) width (Figure 2A). Antennule short 127; (n = 1) length, 4-segmented; basal part widened (2 segments well defined), bearing a short whip (Figure 3B1). Distal 2 segments lacking solus; terminal segment with armature: setae 1, 2, 3 (spiniform), 4 (digitiform), 5 (bifid), 6 (longest) (Figure 3B2). Antenna 219 (n = 1) length; sympod strong, wide; exopod longer and wider than endopod, dorsal margin with 2 rows of denticles (Figure 3C1); endopod 2-segmented, narrower, distally with 3 equal spines (1–3) (Figure 3C2). Labrum subtriangular; rostrum with a central long seta, 4 short setae on each side (Figure 3D). Mandible blade with formula P1, P1, P1, B5; lacking secondary teeth; primary dentition subequal, first 2 basal teeth longer (Figure 3E). Maxillule 101 (n = 1) length, lobate; endite with 2 papillae each bearing long setae and 1 shorter dorsally; palp lateral, with 2 unequal short setae (Figure 3F). Maxilliped corpus wide 190 (n = 1); myxal area with 1 spine; distal margin with 2 small processes (1 subtriangular, 1 blunt); subchela strong (37.5% of corpus length) with proximal seta; claw slightly curved, with short accessory barb (one-third of claw length) (Figure 3G).
Drawings of the female of Clavellotis mayae n. sp. from the Yucatán Peninsula, Mexico. (A) Female, lateral view. (B) Dorsal shield. (C) Maxilla and aliform process, lateral view. (D) Maxilla and aliform process, dorsal view. Abbreviations: ap, aliform process; Ce, cephalothorax; es, egg sac; m, maxilla; lb, lateral lobes; t, trunk. Scale bars: A = 500 µm; B, C, and D = 100 µm.

Drawings of the female of Clavellotis mayae n. sp. from the Yucatán Peninsula, Mexico. (A) Trunk distal margin. (B1) Antennule entire; (B2) detail of distal armature. (C1) Antenna; (C2) endopod detail. (D) Labrum ventral view. (E) Mandible. (F) Maxillule. (G) Maxilliped. Abbreviations: ab, accessory barb; cl, claw; co, corpus; en, endopod; ex, exopod; f, flaps; gp, genital process; p, palp; r, rostral seta; 1, 6, 4, 5, 3, 2, antennule armature; 3, 2, 1, endopod setate; t, trunk; w, whip. Scale bars: A = 100 µm; B, C, D and E = 20 µm; F = 10 µm and G = 50 µm.

Male: (based on Allotype and 6 paratypes specimens) in micrometres (Figures 4, 5C, and 5D). Usually attached near female genital process (Figure 5C). Suborbicular, longer than wide 609–792 (549; n = 7) length by 508–558 (486; n = 7) width, trunk strongly curved; genital process short, disto-ventral, near maxilliped base (Figure 4A). Antennule 41 (n = 1) length, uniramous (Figure 4B1), 3-segmented; basal segment with distal whip; second with short solus; distal segment with armature: setae 1, 2, 3 (spiniform); 4 (digitiform); 5 (simple); 6 (longest) (Figure 4B2). Antenna 89 (n = 1) length; 2-segmented base (Figures 4B1 and B2); endopod and exopod are in line with the sympod, and endopod is longer than exopod; with strong dorsal curved spine, ventral adhesion pad densely spinulose and 1 seta; exopod unarmed (Figures 4C1 and C2). Mandible very thin blade, armed distally with 8 equal teeth, lacking secondary dentition (Figure 4D). Maxillule endite with 2 papillae each bearing 1 seta; palp lateral, with 2 short distal setae (Figure 4E). Maxilla first segment globose; distal part forming curved claw (Figure 4F). Maxilliped first segment rectangular, disto-ventrally with 3 teeth; second segment short, distally armed ventrally with 3 teeth (Figure 4G); both segments aiding attachment to female genital area.
Drawings of the male of Clavellotis mayae n. sp. from the Yucatán Peninsula, Mexico. (A) Male, entire, lateral view. (B1) Antennule entire; (B2) detail of distal armature. (C1) Antenna entire; (C2) detail distal of endopod. (D) Mandible. (E) Maxillule. (F) Maxilla. (G) Maxilliped dorsal and ventral view. Abbreviations: m, maxilla; mp, maxilliped; gp, genital process; en, endopod; ex, exopod; p, palp; 1, 6, 4, 5, 3, 2, antennule armature. Scale bars: A = 100 µm; B, C = 20 µm; D = 10 µm; E, F and G = 25 µm.

Photographs of Clavellotis mayae n. sp. from the Yucatán Peninsula, Mexico. (A) Female, entire lateral view. (B) Posterior end of female with egg sacs. (C) Male attached to the female genital process. (D) Male. Scale bars: A, B and C = 500 µm; D = 200 µm.

Taxonomic summary
Type host: Lagodon rhomboides (Linnaeus, 1766)
Other hosts: Archosargus rhomboidalis (Linnaeus, 1758)
Archosargus probatocephalus (Walbaum, 1792).
Type locality: La Carbonera lagoon, Yucatán Peninsula, Mexico, Gulf of Mexico
Other locality: Celestún, Yucatán
Site of infection: gills
Material examined: 32 specimens from L. rhomboides (22 female and 10 male, 10 fish infected of 60 examined, intensity: 1–7); 45 from A. rhomboidalis (41 female and 4 male, 13 fish infected of 31, intensity 1–7); and 8 from A. probatocephalus (7 female and 1 male, 4 fish infected of 29, intensity 1–9).
Etymology: The specific epithet, mayae refers to the Maya civilization that inhabited the Yucatán Peninsula, where the host species of the copepod are widely distributed.
Specimens deposited: Holotype female CNCR 38370, Allotype male CNCR 38369, paratype female and male of L. rhomboides CNCR 38371 from La Carbonera lagoon, paratype female of A. rhomboidalis CNCR 38372 and paratype female of A. probatocephalus CNCR 38373 from Celestún lagoon were deposited in the Mexican National Crustacean Collection (CNCR).
GenBank accession: PX802343–PX802346 for 28S; PX830982–PX830993 for cox1
ZooBank LSID: zoobank.org:act:08155F38-6947-4A6D-84D9-D0B980CEB8F2XX
Remarks
According to the diagnosis given by Castro-Romero and Baeza (Reference Castro-Romero and Baeza1984), Clavellotis mayae n. sp. belongs to the genus Clavellotis by possessing a lateral process at the base of cephalothorax and by having a trunk without posterior processes, but with a genital process.
Among the currently known species of Clavellotis, the new species can be readily distinguished from those exhibiting a subcircular trunk, i.e. C. dilatata (Krøyer, 1863), C. girellae Castro-Romero et al. (Reference Castro-Romero, Montes, Ӧktener, Ibañez-Shimabukuro, Theiller and Campos2025), C. tarakihi (Hewitt & Blackwell, 1987) and C. branchiostegui (Yamaguti, 1939) because they lack an elongated, and pyriform to sub-quadrangular trunk. In this respect, C. mayae n. sp. is more similar to C. briani Benmansour, Ben Hassine, Diebakate & Raibaut, 2001, C. dubius (Castro-Romero et al. Reference Castro-Romero, Montes, Ӧktener, Ibañez-Shimabukuro, Theiller and Campos2025), C. fallax (Heller, 1865), C. pagri (Krøyer, 1863), C. sargi (Kurz, 1877), C. sebastidis (Castro and González Reference Castro and González2005) and C. strumosa (Brian, 1906). However, most of these species can be readily distinguished. For instance, the trunk in C. fallax, C. strumosa, C. sargi and C. pagri is slightly longer than wide; the trunk in C. briani is subquadrangular, whereas in the new species the trunk is elongated, pyriform, and possess a conspicuous subcircular projection (flap) covering the egg-sac insertion. Thus, the new species is morphologically more similar to C. dubius and C. sebastidis by the shape of the trunk.
Comparison with C. dubius
In C. dubius, the maxilla is short, about half the trunk length, and lacks lateral lobes whereas in the new species, the maxilla bears 2 distinct lateral lobes. The aliform process is oval and simple in both species. The trunk of C. dubius lacks distal flaps over the egg-sac attachment, instead showing 2 lobes, while the new species bears subcircular flaps covering the egg-sac bases. The genital process is subrectangular and more developed in C. dubius, vs. short and blunt in the new species. The antennule is 4-segmented in both, but the armature differs (1, 3, 4, 5, 6 in C. dubius; 1, 2, 3, 4, 5, 6 in C. mayae n. sp.). In the antenna, C. dubius shows an exopod longer than the endopod, with few dorsal spines; the endopod bears 3 unequal setae. In contrast, the new species has 2 rows of dorsal spinules on the exopod and 3 subequal endopodal setae. The mandibular formula also differs: C. dubius shows P1S1, P1S1, P1S1, B5, while C. mayae n. sp. lacks secondary dentition, with the first 2 basal teeth larger than the others. The maxillule is similar in endite armature but differs in palp setation. The maxilliped of the new species bears 2-min distal processes absent in C. dubius. Males of C. mayae n. sp. are more distinctly oval (vs. subcircular in C. dubius), with maxilla and maxilliped positioned close together, and differs in claw shape and maxilliped armature. The species C. dubius was described from the pomacentrid Chromis crusma (Valenciennes, 1833) off the coast of Chile, in the Southeast Pacific, whereas the new species is described from sparids in the Gulf of Mexico.
Comparison with C. sebastidis
Clavellotis sebastidis possess an elongated trunk, approximately equal in length to the cephalothorax, lacking lobes at the egg-sac base; whereas in C. mayae n. sp., the cephalothorax is longer than the trunk and bears distinct flaps over the egg-sac base. The maxilla in C. sebastidis is as long as the trunk and lacks protuberances, whereas in the new species it is short (about one-third trunk length), and bears paired lateral lobes. The aliform process is circular in C. sebastidis, oval in the new species. The antennule is 3-segmented in C. sebastidis but 4-segmented in the new species, and the armature differs accordingly. The antenna is similar in both, but the mandible differs: C. sebastidis shows P1S1, P1S1, P1S1, B4, whereas C. mayae n. sp. lacks secondary dentition and has the first 2 basal teeth longer. The maxillule also differs in palp setation. The maxilliped is similar in general structure, but C. sebastidis lacks distal processes on the corpus whereas the new species possesses 2-min distal processes on the corpus absent; in addition, C. sebastidis has a spinulose ventral subchela margin, which is absent in the new species. Males of C. sebastidis are subcircular and possess a dorsal projection, while C. mayae n. sp. is oval-shaped and lacks dorsal projection. The genital process is shorter and more separated in C. sebastidis; in the new species, it is more developed and positioned near the maxilliped. Antennule armature, mandibular dentition and maxillule setation also different. Clavellotis sebastidis was originally described by Castro and González (Reference Castro and González2005) from the rockfish Sebastes oculatus (Valenciennes 1833) off the southern coast of Argentina, and was later reported from S. miniatus (Jordan & Gilbert 1880) along the Northwest Pacific coast of Mexico (Rodríguez-Santiago et al., Reference Rodríguez-Santiago, Rosales-Casián and Grano-Maldonado2014, Reference Raupach, Barco, Beermann, Laakmann, Mohrbeck, Neumann, Kihara, Pointner, Radulovici, Segelken-Voigt, Wesse and Knebelsberger2015, Reference Rodríguez-Santiago, Rosales-Casián and Grano-Maldonado2016).
All the differences discussed above clearly support the recognition of the specimens belonging to Clavellotis from sparid fish of the Yucatán Peninsula as a distinct species, herein described as Clavellotis mayae n. sp., representing the first record of the genus on the Atlantic coast of Mexico, parasitizing sparid fish (L. rhomboides, A. rhomboidalis and A. probatocephalus).
Molecular characterization
28S rDNA. Partial 28S rDNA sequences were generated for 4 specimens of C. mayae n. sp. (686–1063bp) parasitizing L. rhomboides collected in La Carbonera lagoon and A. rhomboidalis in Celestún lagoon. The newly generated sequences were compared with homologous sequences of Lernaeopodidae available in GenBank (Table 1), the alignment was 482 bp long and contained 22 taxa. Ergasilus sp. (KR048842) was used as an outgroup. The sequences of C. mayae n. sp. nested with all species of Clavellotis, a relationship supported by high posterior probability values (0.97) (Figure 6). The intraspecific genetic divergence among isolates of C. mayae n. sp. was low, ranging from 0.42% to 1%, whereas the interspecific divergence varied between 1% and 3% with respect to the other congeners (Table 2). In the 28S phylogenetic tree the new species was recovered as the sister species of a clade containing C. dubius plus C. girellae and C. dilatata, the only members of the genus for which 28S rDNA sequences are available.
Phylogenetic relationships of Clavellotis mayae n. sp. (Copepoda: Lernaeopodidae) from Lagodon rhomboides (L.) and Archosargus rhomboidalis (L.) from the Yucatán Peninsula, Mexico, and other Lernaeopodid copepod species, obtained from Bayesian Inference (20 000 000 generations) of the 28S rDNA sequences. Branch support values correspond to posterior probabilities ≥0.90. Newly generated sequences are shown in bold.

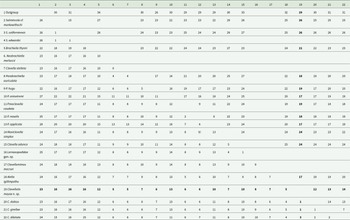
Interspecific genetic distances among species included in the phylogenetic analyses, 28S rDNA distances below the diagonal; cox1 distances above the diagonal; distances are expressed in percentages. Newly generated sequences are shown in bold

cox1 mDNA. Partial cox1 mDNA sequences were generated for 12 specimens of C. mayae n. sp. collected from L. rhomboides (579–676 bp), A. rhomboidalis (472–651 bp) and A. probatocephalus (663 bp). The newly generated sequences were analysed together with sequences of Lernaeopodidae available in GenBank (Table 1), the alignment included 121 isolates. Ergasilus sp. (KR048842) was used as an outgroup. The 12 isolates of C. mayae n. sp. clustered together in a well-supported clade (PP = 1), as the sister taxa of a clade containing 8 species of Clavellotis for which cox1 is currently available (Figure 7). The intraspecific genetic divergence among isolates varied between 1% and 3% (Table 2). Instead, the cox1 interspecific divergence between C. mayae n. sp. and all the other species of Clavellotis varied from 12% to 14% (Table 2).
Phylogenetic relationships of Clavellotis mayae n. sp. (Copepoda: Lernaeopodidae) from Lagodon rhomboides (L.), Archosargus rhomboidalis (L.) and Archosargus probatocephalus (Walbaum) from the Yucatán Peninsula, Mexico, and other Lernaeopodid copepod species, obtained from Bayesian Inference (20 000 000 generations) of the cox1 mtDNA sequences. Branch support values correspond to posterior probabilities ≥0.90. Newly generated sequences are shown in bold.

Discussion
Morphological and molecular data corroborated that Clavellotis mayae n. sp. represents a new species. Morphologically, the new species shows closer affinities with species bearing an elongated or pyriform trunk, but differing from them by possessing a subcircular, orbicular, or quadrangular trunk and notably, by having a suboval aliform process together with lobules at the base of the maxilla; this represents a unique combination of characters not observed in any other Clavellotis species. In addition, the bases of the egg sacs are protected by trunk protrusions resembling flaps, a condition that further distinguishes the new species.
These morphological differences were corroborated by molecular data. DNA sequences support that all isolates for both molecular markers reveal the presence of a single species irrespective of the species of sparid they infect; molecular data also corroborated the validity of the new species considering the phylogenetic position on the trees, and the interspecific genetic divergence with respect to all the congeneric species for which sequences of the 2 molecular markers are available. In this case, the cox1 tree resulted to be more informative for species recognition since a larger number of species of Clavellotis have been sequenced; however, higher rank phylogenetic relationships contrast with the 28S tree, particularly regarding the monophyly of genera such as Parabrachiella which appears to be paraphyletic in the cox1 tree, whereas in the 28S tree it is recovered as monophyletic, although with a lower number of available sequences (Figures 6 and 7).
With the inclusion of the new species described herein from sparid hosts from the Yucatán Peninsula, Mexico, the total number of recognized species in the genus rises to 14. Species in the genus are widely distributed across the globe. Six species have been reported from the Mediterranean Sea, namely C. briani, C. characis, C. fallax, C. sargi, C. strumosa and C. pagri. Interestingly, C. pagri has been also reported from Argentinean and Brazilian waters, in the Southwestern Atlantic (Cantatore et al., Reference Cantatore, Braicovich, Alarcos, Lanfranchi, Rossin, Vales and Timi2012). Another 3 species were reported from the South Pacific, including C. dilatata, C. dubius and C. girellae. Clavellotis sebastidis apparently possesses a wider distribution range, since it has been reported from the South Atlantic and North Pacific waters (Castro and González, Reference Castro and González2005; Rodríguez-Santiago et al., Reference Rodríguez-Santiago, Rosales-Casián and Grano-Maldonado2014, Reference Raupach, Barco, Beermann, Laakmann, Mohrbeck, Neumann, Kihara, Pointner, Radulovici, Segelken-Voigt, Wesse and Knebelsberger2015, Reference Rodríguez-Santiago, Rosales-Casián and Grano-Maldonado2016). In addition, C. bilobata Pillai, 1962 was described from the Indian Ocean, C. terakihi from Australian waters, and C. branchiostegui from Japanese waters (Walter and Boxshall, Reference Walter and Boxshall2025).
The validation of some species require further corroboration, such as the recently described Clavellotis helicoleni Izawa, 2025, since the male morphology does not conform with the diagnostic characters of Clavellotis and the female possesses small caudal rami which is absent in all other species. Also, some records from Japan require an integrative review, such as C. nodula Do & Ho, 1983, which has been proposed as a synonym of C. dilatata.
Acknowledgements
JEN thanks to Secretaría de Ciencia, Humanidades, Tecnología e Innovación (SECIHTI) for the postdoctoral fellowship ‘Estancias Posdoctorales por México Modalidad Académica’ (CVU 351170). We thank Maribel Badillo and Alfredo Gallardo from BioCon (Laboratorio de Biología de la Conservación, UNAM) and Norberto García Colin (Laboratorio de Biología Celular y Molecular, ENES Mérida) for providing lab facilities; Laura Márquez and Nelly López from the Laboratorio Nacional de Biodiversidad (LANABIO, Biology Institute, UNAM) for their help with DNA sequencing processin; Carlos Jiménez for his help fishing host; we thank M. Marcia Montes for the drawings of the new species. We also thank Laura Elena López León for providing English editing services for the manuscript, and the 2 anonymous reviewers for their valuable comments and suggestions which helped to improve it.
Author contribution
RCR BPO and JEN conceived and designed the study; RCR, JEN and BPO performed morphological analyses; BPO and MMM performed molecular analysis; BPO, JEN, PCR, MMM and GPP conducted data analyses, wrote and edited the paper.
Financial support
This study was partially funded by the Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica, PAPIIT UNAM IN200824 to GPPL.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
Specimens were sampled under the collecting permit issued by the Comisión Nacional de Acuacultura y Pesca to Alfredo Gallardo Torres (No. PPF/DGOPA-001/20). Fish were humanely euthanized following the protocols described by the 2020 edition of the AVMA Guidelines for the Euthanasia of Animals.