Introduction
The family Diplozoidae Palombi, 1949 (Monogenea) are highly specialized ectoparasitic flatworms that live almost exclusively on the gills of freshwater cyprinid fishes (Wu et al., Reference Wu, Lang and Wang2000; Valigurová et al., Reference Valigurová, Hodová, Sonnek, Koubková and Gelnar2011; Přikrylová et al., Reference Přikrylová, Mašová, Gelnar, Matla, Tavakol and Luus-Powell2018; Benovics et al., Reference Benovics, Koubková, Civáňová, Rahmouni, Čermáková and Šimková2021). They exhibit a unique life cycle, with life stages including eggs, diporpa, an X-shaped juvenile, and an X-shaped adult (Pečínková et al., Reference Pečínková, Matějusová, Koubková and Gelnar2007a; Avenant-Oldewage and Milne, Reference Avenant-Oldewage and Milne2014; Shen et al., Reference Shen, Zhao, Jiang, Xu, Tian, He, Jia, Xu, Meng and Fan2025).
Like most monogeneans, the taxonomy of diplozoids is based on the sclerotized structures of the haptor, particularly the clamps and central hooks, which provide the primary morphological features for species differentiation (Gussev and Kulemina, Reference Gussev and Kulemina1971; Khotenovsky, Reference Khotenovsky1985; Šebelová et al., Reference Šebelová, Kuperman and Gelnar2002; Pečínková et al., Reference Pečínková, Vøllestad, Koubková and Gelnar2007b). In the genus Paradiplozoon Achmerov, 1974, species identification predominantly relies on the length of the central hooks and the morphology of certain sclerotized elements of the clamps. The combination of the median sclerite and anterior joining sclerite shapes aids in more precise species differentiation (Konstanzová et al., Reference Konstanzová, Koubková, Kašny´, Ilgová, Dzika and Gelnar2017; Jirsová et al., Reference Jirsová, Ding, Civánová, Jirounková, Ilgová, Koubková, Kasny and Gelnar2018). However, these morphological measurements often exhibit high interspecific similarity and can vary significantly depending on factors such as host species, water temperature, or the geographical origin of the parasites (Matějusová et al., Reference Matějusová, Koubková, Gelnar and Cunningham2002). Additionally, the size and shape of these sclerotized structures may be influenced by fixation and preparation methods, potentially leading to misinterpretation (Huang et al., Reference Huang, Zhou, Yuan and Ding2023). Consequently, recent studies have increasingly adopted molecular approaches to complement traditional taxonomy. Among various molecular markers, the internal transcribed spacer 2 (ITS2) region of ribosomal DNA has been widely employed as a genetic marker for species delimitation and phylogenetic analyses of diplozoids. To date, around 60 out of over 80 described species of Diplozoidae have been sequenced for ITS2. While ITS2 has proven useful for species differentiation, it has limitations in distinguishing closely related species, and many originally described species remain unsequenced (Matějusová et al., Reference Matějusová, Koubková, D’Amelio and Cunningham2001, Reference Matějusová, Koubková and Cunningham2004; Sicard et al., Reference Sicard, Desmarais and Lambert2001; Dos Santos and Avenant-Oldewage, Reference Dos Santos and Avenant-Oldewage2020).
In recent decades, 9 Paradiplozoon species have been described from cyprinid fishes across Africa and Eurasia (Dos Santos et al., Reference Dos Santos, van Vuuren and Avenant-Oldewage2015; Shimazu et al., Reference Shimazu, Kobayashi, Tojo, Besprozvannykh and Ogawa2015; Benovics et al., Reference Benovics, Koubková, Civáňová, Rahmouni, Čermáková and Šimková2021; Nejat et al., Reference Nejat, Benovics, Řehulková, Vukić, Šanda, Kaya, Tarkan, Abdoli, Aksu and Šimková2023), with 3 species known from China (Fan et al., Reference Fan, Meng, Bai, Xu and Wang2018; Arken et al., Reference Arken, Hao, Guo, Zhang, Rong, Kamal, Tian, Kadir and Yue2022; Huang et al., Reference Huang, Zhou, Yuan and Ding2023). The diverse freshwater habitats in China support a rich diplozoid fauna. Nevertheless, little is known about diplozoid parasites of the genus Garra Hamilton, 1822 (Cyprinidae). To date, only Paradiplozoon bingolensis Civáňová, Koyun & Koubková, 2013 has been reported from Garra rufa Heckel, 1843 in Turkey (Civáňová et al., Reference Civáňová, Koyun and Koubková2013), and no species have been recorded from Chinese Garra species. The present study reports and describes a new Paradiplozoon species from Garra surgifrons Zhou and Sun, 2018, a native fish species in China, representing the first record of a diplozoid parasitizing this host genus in China. The type specimen of G. surgifrons was collected from the Longchuang River, an upper tributary of the Irrawaddy River drainage in Tengchong, Yunnan (Zhou & Sun, 2018). Further inquiries with aquaculture personnel revealed that the G. surgifrons individuals in the farm were introduced from the wild, and the aquaculture water source is drawn from the Irrawaddy River. Morphological characteristics, together with molecular evidence based on the ITS2 region, were employed to confirm its distinct taxonomic status and phylogenetic position within Diplozoidae. Furthermore, the combined morphological, molecular and biogeographic analyses provide new insights into the evolutionary relationships and potential origin of Paradiplozoon species in East and Southeast Asia.
Materials and methods
Sample collection
In August, September and December 2025, a total of 25 G. surgifrons (Cyprinidae, Labeoninae), with a size range of 13–15 cm in standard length, were obtained from a fish farm (102°52’37”N, 25°03’54”E) in Kunming, Yunnan, China. The collected fish were euthanized, and the gills of each fish were subsequently removed and examined using an Olympus SZ-X7 dissecting microscope (Olympus, Japan). The structures of the freshly collected parasite specimens were observed using an Olympus CX41 light microscope (Olympus, Japan) and photographed with the Capture 3.0 software (Tucsen Photonics Co., Ltd). Subsequently, some adult specimens were fixed in 70% ethanol for staining, while the remaining specimens (both larval and adult) were preserved in 95% ethanol for molecular analyses.
Morphological methods
Specimens preserved in 70% ethanol were rehydrated through a graded ethanol series and soaked in distilled water overnight. Then fixed in Bouin’s solution for 5–7 h, rinsed in distilled water and stained with alum carmine for 12 h. Differentiation was performed in acid alcohol for 20–30 sec, followed by dehydration through a graded ethanol series and clearing in cedarwood oil for 12 h. Finally, specimens were mounted in neutral balsam (Bai et al., Reference Bai, Wang, Li, Xu and Fan2014). Stained specimens were photographed using an Olympus CX-41 light microscope. Morphometric analyses of parasite structures were conducted using Capture 3.0, and morphological diagrams of the body, clamps, central hooks and eggs were drawn by tracing from existing photographs using a digital drawing board (Wacom, Saitama, Japan, Intuos Pro).
The dimensions of whole body, oral sucker, pharynx, central hooks and clamps characterizing parasites were measured by the straight-line distances between extreme ends. Since the adult body forms an X-shaped fusion body, the body length and width of an individual adult were measured as the straight-line distances between the end-points of the two fused individuals, and the average value was then taken.
Molecular methods
A total of 7 specimens (3 larvae and 4 adults) were used for molecular analyses, and the remaining specimens were preserved in 95% ethanol. Genomic DNA of the parasites was extracted using the TIANamp Micro DNA Kit (TIANGEN Biotech, Beijing, China) following the manufacturer’s instructions. The ITS2 region of the genomic DNA was amplified using universal primers (Matějusová et al., Reference Matějusová, Koubková, D’Amelio and Cunningham2001). Forward primer D: 5′–GGCTYRYGGNGTCGATGAAGAACGCAG–3′; Reverse primer B1: 5′–GCCGGATCCGAATCCTGGTTAGTTTCTTTTCC–3′. Polymerase chain reaction (PCR) was carried out in a 50 μL reaction mixture containing 2 μL DNA template, 19 μL reaction buffer containing dNTPs, 10 × buffer, and Taq polymerase, 2 μL of each primer, and 25 μL double-distilled water. The PCR conditions were as follows: initial denaturation at 90°C for 10 min, followed by 30 cycles of 95 °C for 30 sec for denaturation, 55 °C for 30 sec for annealing and 72 °C for 75 sec for extension, with a final extension at 72 °C for 10 min.
PCR products were verified by 1% agarose gel electrophoresis and stained with GoodView (Tanon, China) to visualize the bands. The amplified DNA fragments were sequenced, and the resulting sequences were submitted to the National Center for Biotechnology Information (NCBI) database and compared using BLAST. A total of 47 diplozoid species sequences were retrieved from NCBI, together with the two sequences obtained in this study, were used for phylogenetic analysis.
Base composition, parsimony analysis and nucleotide substitution rates under p-distance were calculated using MEGA 11.0 (Koichiro et al., Reference Koichiro, Glen and Sudhir2021). Phylogenetic trees were constructed using both Bayesian Inference (BI) and Maximum Likelihood (ML) methods in Phylosuite v2 (Zhao et al., Reference Zhao, Tong, Fangluan, Jakovlić, Qiong, Yindong, Xiang, Song, Liu, Lian, Zou, Li, Wang, Zeng and Zhang2025). The BI analysis was performed with 1 000 000 generations, and ML was conducted using the GTR + G model with 5000 bootstrap replicates. Tree topology robustness was assessed using both BI posterior probabilities and ML bootstrap support. Neoheterobothrium hirame Ogawa, 1999 (AB162613) was used as the outgroup for the phylogenetic analysis. The resulting phylogenetic tree was visualized using the Interactive Tree of Life (iTOL) platform (Letunic and Bork, Reference Letunic and Bork2021), with the aid of dataset files generated by Phylosuite v2.
Results
Family: Diplozoidae Palombi, 1949
Genus: Paradiplozoon Achmerov, 1974
Type species: Paradiplozoon garra n. sp.
ZooBank LSID: urn:lsid:zoobank.org:pub:5CCFCE3B-C6A0-44F6-9232-DB60A36FEF48
Type-host: G. surgifrons Zhou & Sun, 2018 (Cyprinidae, Labeoninae) (Figure 1A)
Site: Gills
Infection intensity: 1–2 parasites per infected fish
Infection rate: 1.60, 32 parasites in 20 fish
Type locality: a fish farm in Kunming, Yunnan, China. (102°52’37”N, 25°03’54”E)
Type material: Holotype (H2025090401), 6 paratypes (H2025090402, H2025090403, H2025090404, H2025080505, H2025120306, H2025120307), deposited in the Museum of Yunnan Normal University, Yunnan, China.
Etymology: The specific name garra refers to the host genus G. surgifrons.
Description
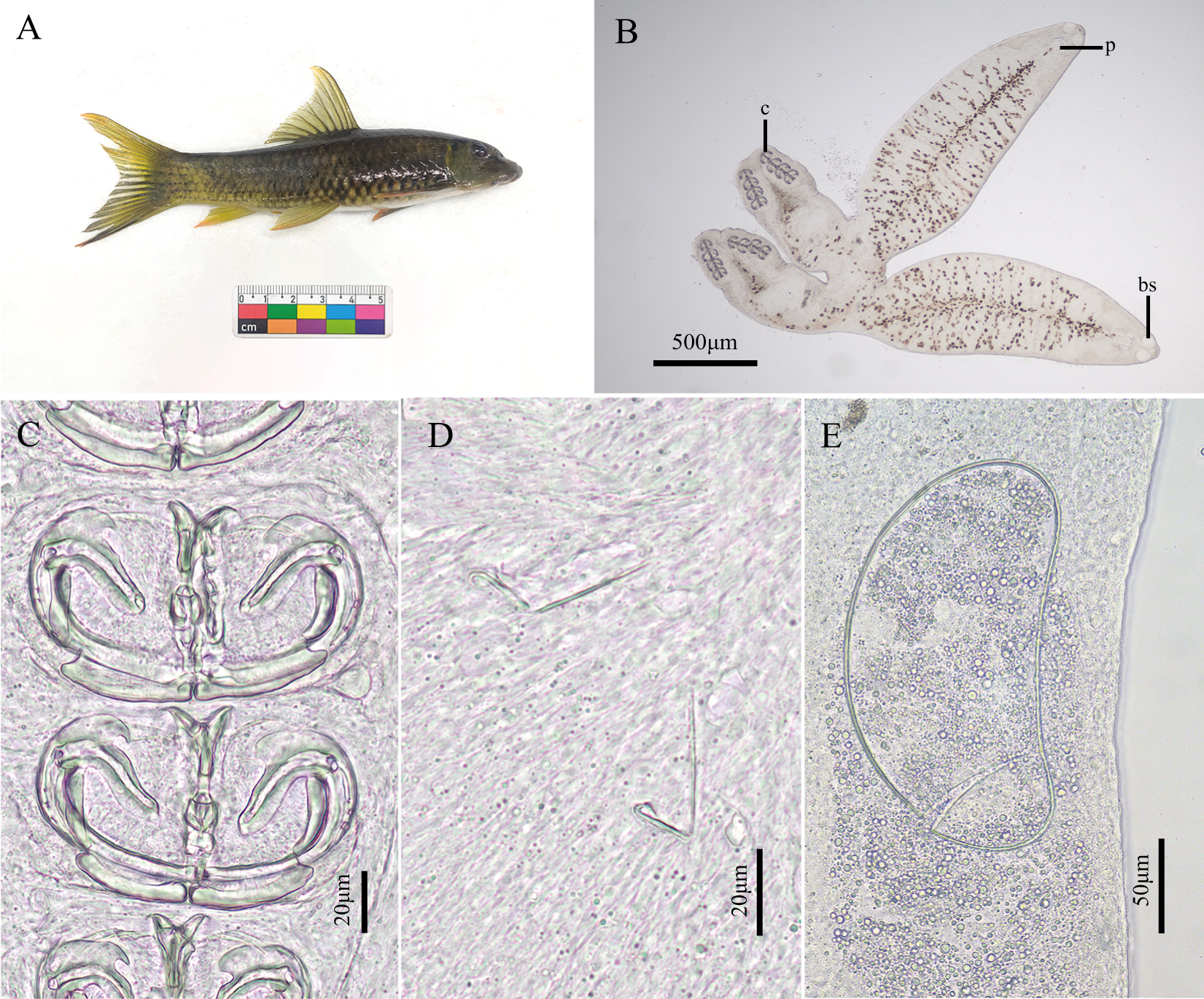
Description based on observation of 11 adult specimens. Body surfaces smooth, without folds. Anterior region of buccal suckers lacks round glands. No disc-shaped widened area with disciform structure between posterior and reproductive fusion areas, nor any special lobed enlargement (Figures 1B and 3A).
Photographs of host fish and Paradiplozoon garra n. sp. (A) Garra surgifrons Zhou & Sun, 2018; (B) whole body of worm; (C) clamps; (D) central hooks; (E) egg. (bs, buccal suckers; p, pharynx; c, clamps).

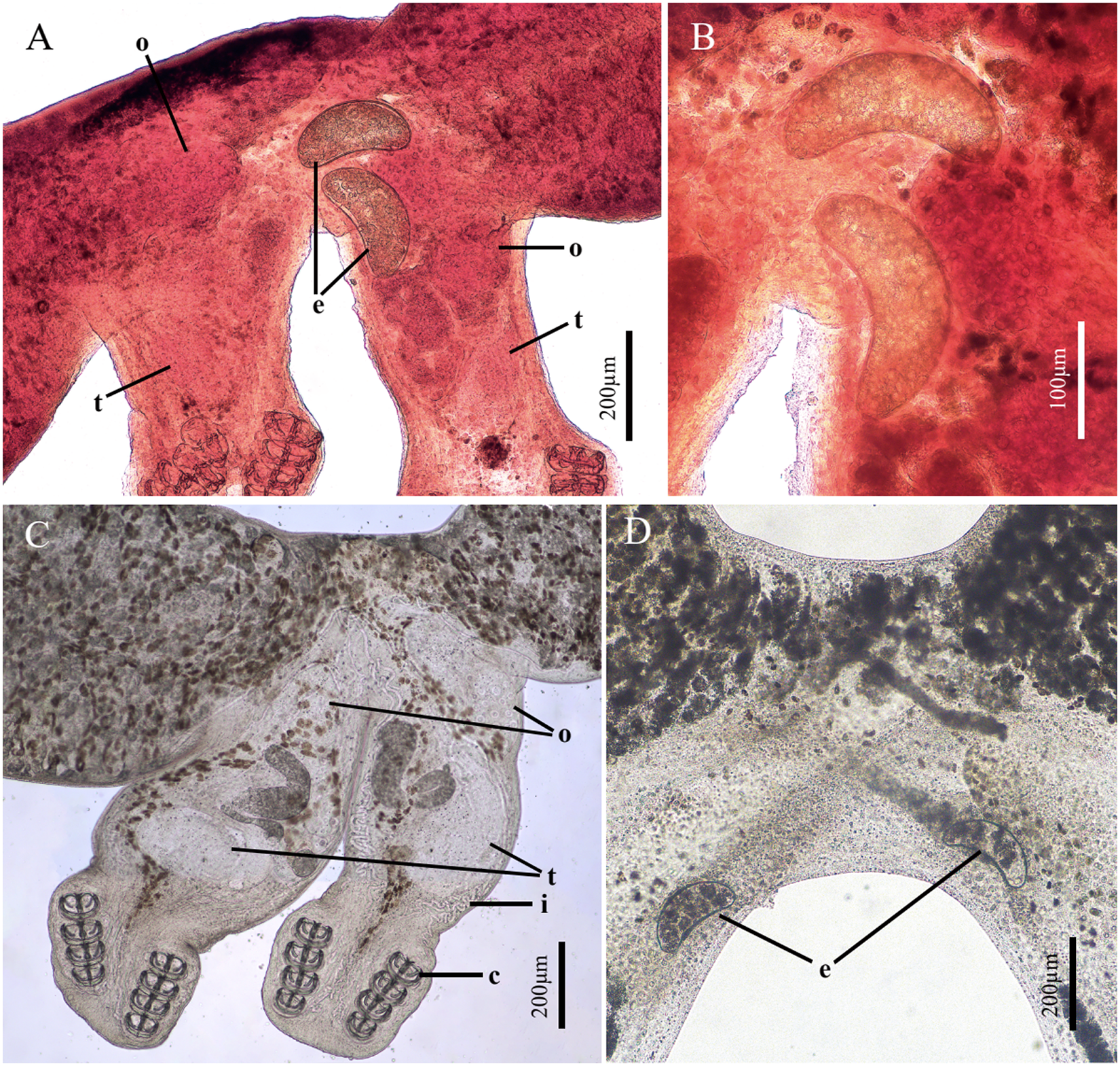
Adult specimens measure 4.610 (2.373–6.941, n = 8) mm in length and 0.724 (0.520–0.955, n = 8) mm in width. Intestine located centrally, extending to body fusion area. Vitelline follicles numerous, distributed at anterior part of body. Reproductive glands extend from fusion area to posterior part of body. Ovary single, 0.286 (0.253–0.335, n = 3) × 0.206 (0.134–0.283, n = 3) mm, situated in fusion area. Testis single, 0.223 (0.187–0.315, n = 3) × 0.189 (0.149–0.232, n = 3) mm, compact, unlobed, positioned posterior to ovary (Figures 2A and C).
Reproductive system of Paradiplozoon garra n. sp. (A) Reproductive system; (B) The eggs; (C) Reproductive system without eggs; (D) Reproductive system with eggs. (o, ovary; t, testis; e, eggs; i, intestine; c, clamps). A and B show stained specimens; C and D show specimens mounted in water.

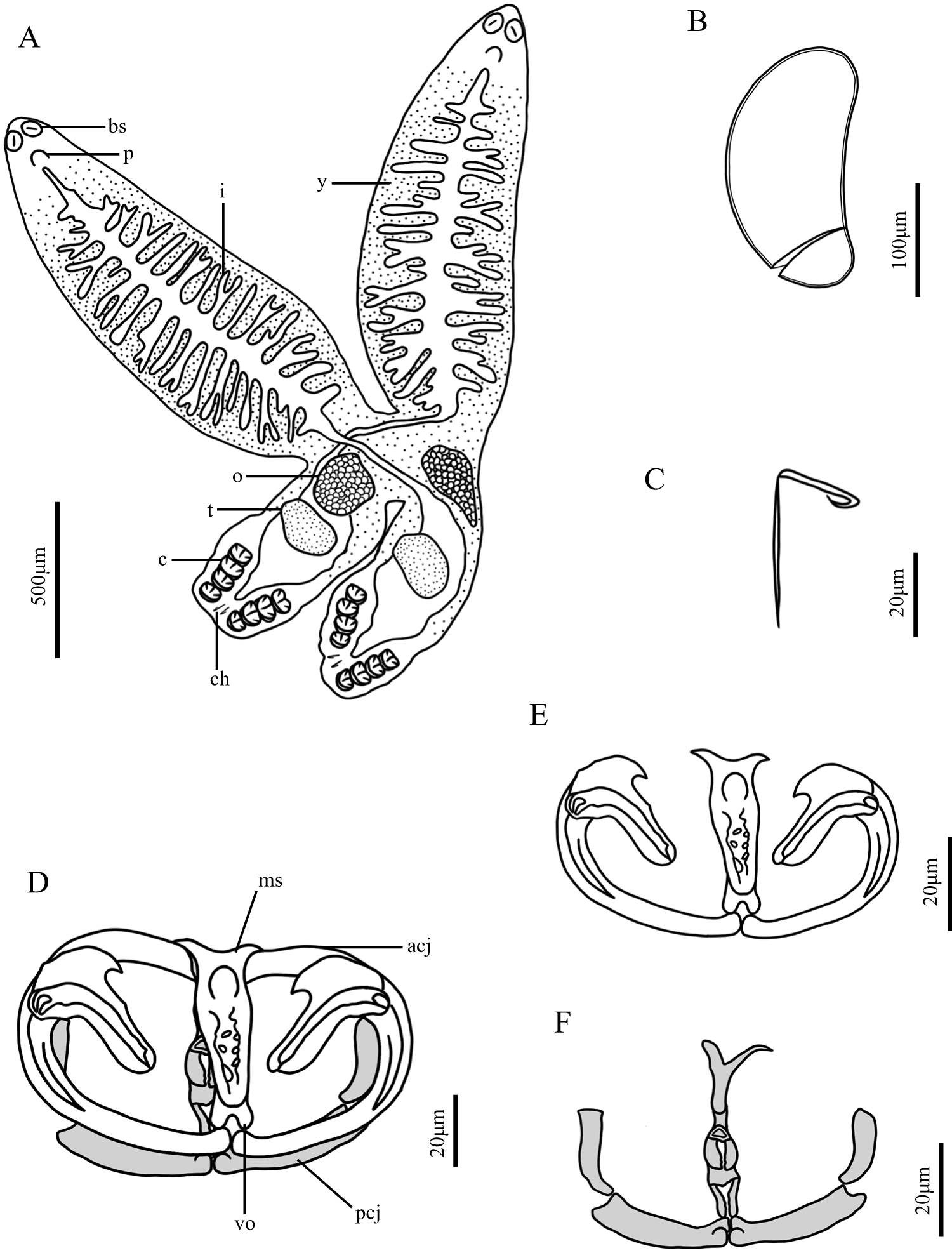
Eggs elliptical, lacking filaments attached to operculum (Figures 1E, 2B, 2D and 3B), 0.101 (0.094–0.110, n = 3) × 0.213 (0.205–0.221, n = 3) mm. Haptors measure 0.324 (0.258–0.369, n = 8) mm in length and 0.387 (0.316–0.493, n = 8) mm in width, bear four pairs of clamps and one pair of central hooks. Clamp I smallest, 0.062 (0.041–0.077, n = 15) × 0.090 (0.065–0.124, n = 15) mm. Clamp II 0.064 (0.050–0.080, n = 15) × 0.103 (0.077–0.130, n = 15) mm. Clamp III largest, 0.066 (0.053–0.079, n = 16) × 0.104 (0.074–0.127, n = 16) mm. Clamp IV 0.064 (0.052–0.075, n = 16) × 0.097 (0.068–0.124, n = 16) mm. Anterior clamp jaw composed of two curved sclerites. Posterior clamp jaw consists of medial and lateral parts. Both anterior arch of anterior clamp jaw and medial part of posterior jaw lack cross-striation. Spurs of anterior jaws form thin, elongated, hook-shaped structures. Anterior half of median plate shows thickened margins with irregularly arranged circular perforations; anterior joining sclerites short and V-shaped. Posterior half of median plate presents irregular, short trapezoid-like spur and pair of elongated, rod-shaped joining sclerites (Figures 1C and 3D–F).
Central hooks positioned between terminal protrusion of haptor and first pair of clamps. Hooks formed by handle and sickle connected together. Sickle curves toward handle, ending in winged tip curling toward connection. Central hook sickle 0.016 (0.012–0.021, n = 15) mm, handle 0.037 (0.029–0.045, n = 15) mm (Figures 1D and 3C).
Digital drawing of Paradiplozoon garra n. sp. (A) Adult; (B) egg; (C) Central hooks; (D) Clamp; (E) Clamp in anterior view; (F) Clamp in posterior view. (bs, buccal suckers; p, pharynx; i, intestine; y, vitellarium; o, ovary; t, testis; c, clamps ch, central hooks; ms, median sclerite; acj, anterior clamp jaw; pcj, posterior clamp jaw; vo, V-outgrowth).

Diagnosis
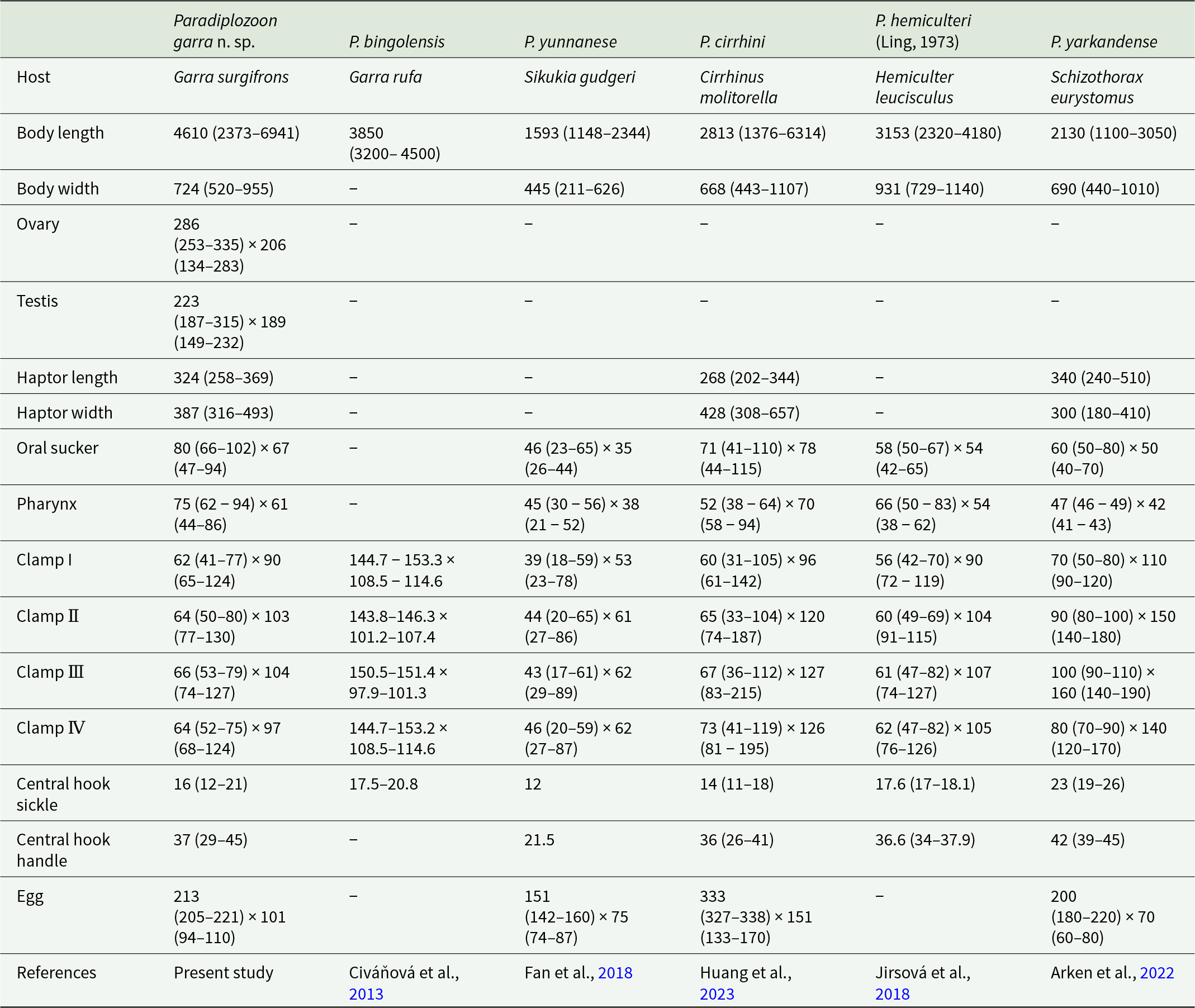
The new diplozoid species described from G. surgifrons differs markedly from P. bingolensis, collected from another congeneric host (G. rufa), in both morphological characteristics and molecular sequences. The main distinguishing features of P. bingolensis include: a rectangular anterior joining sclerite connected to the proximal tip of the anterior jaw. The posterior end of the median plate narrowing and terminating with a wide, rounded sclerite with an opening, and has a short posterior joining sclerite. In contrast, P. garra is characterized by thin, elongated, hook-shaped spurs on the anterior jaws. The anterior half of the median plate bears short V-shaped joining sclerites, while the posterior half features an irregular, short trapezoid-like spur with a pair of elongated, rod-shaped joining sclerites. Morphometric comparisons of P. garra with other closely related species of the genus are summarized in Table 1.
Measurements of Paradiplozoon garra n. sp. and closely related Paradiplozoon species

The values of measurement in micrometre (µm).
Molecular analyses
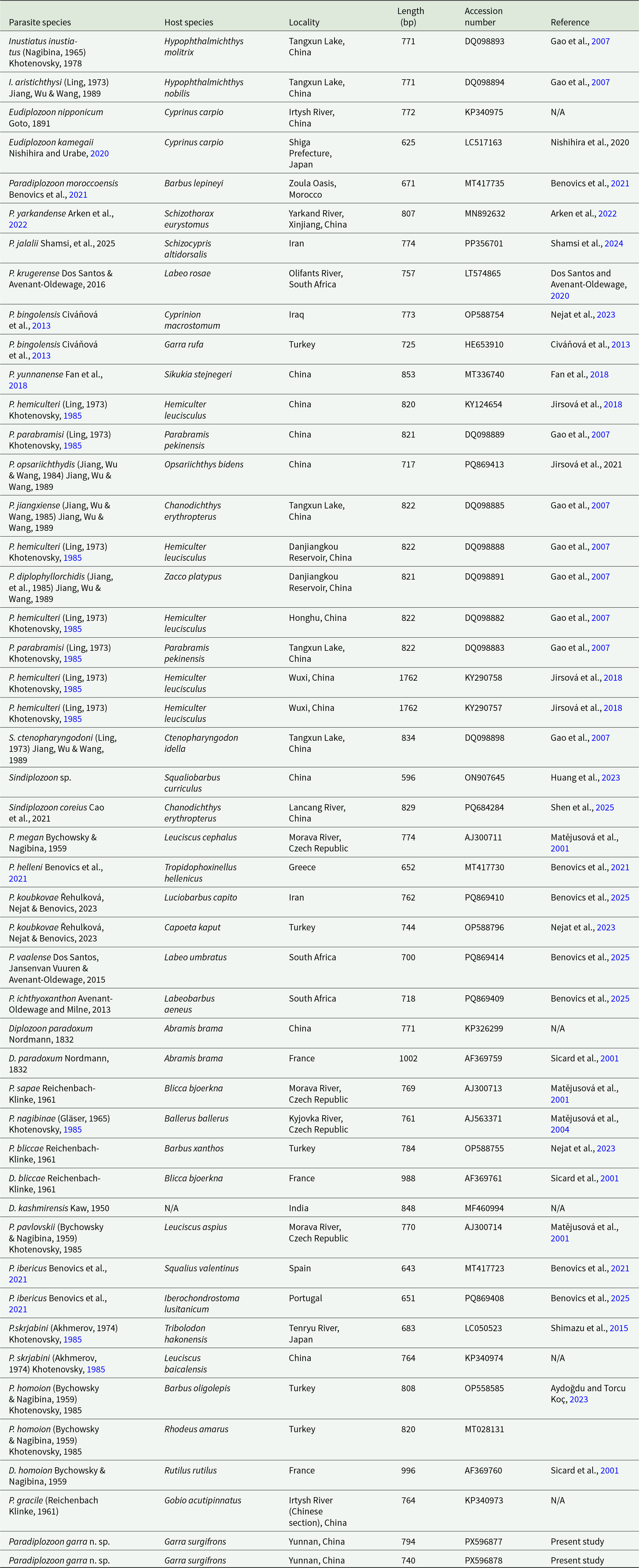
Sequence comparison of the ITS2 region from all seven specimens revealed identical sequences. Two representative specimens, one adult and one larva, were selected for subsequent ITS2 phylogenetic analysis, yielding sequences of 794 bp and 740 bp, respectively. The resulting sequences were submitted to GenBank under accession numbers PX596877 and PX596878. A total of 47 representative ITS2 sequences of diplozoid species were selected from different geographical regions, with detailed species information provided in Table 2. Alignment of the 49 ITS2 sequences yielded a total sequence length of 738 bp, comprising 215 conserved sites (29.1%), 523 variable sites (70.9%) and 433 parsimony-informative sites (58.7%). The relatively high proportion of variable and parsimony-informative sites indicates substantial sequence divergence among taxa.
List of diplozoid species, host species, locality of collection and GenBank accession numbers for DNA sequences used in phylogenetic analyses

BLAST analysis showed that, when compared with published ITS2 sequences of diplozoid species, sequence homology ranged from 78.20% to 81.36% within 98–100% coverage. The highest similarity was observed with Paradiplozoon parabramisi (Ling, 1973) Khotenovsky, 1985 (DQ098883) and P. opsariichthydis (Jiang, Wu & Wang, 1984) Jiang, Wu & Wang, 1989 (DQ098890), both exhibiting 81.36% similarity. The genetic p-distance (Supplementary materials) between P. garra n. sp. and other diplozoid species ranged from 16.42% to 33.68%. The species most closely related to P. garra n. sp. were P. hemiculteri (Ling, 1973) Khotenovsky, 1985 (KY124654), P. parapeleci Jiang et al., 1984 (DQ098882), and P. parabramisi (DQ098883), with genetic p-distance of 16.42%, 16.55% and 16.55%, respectively (see Supplementary materials).
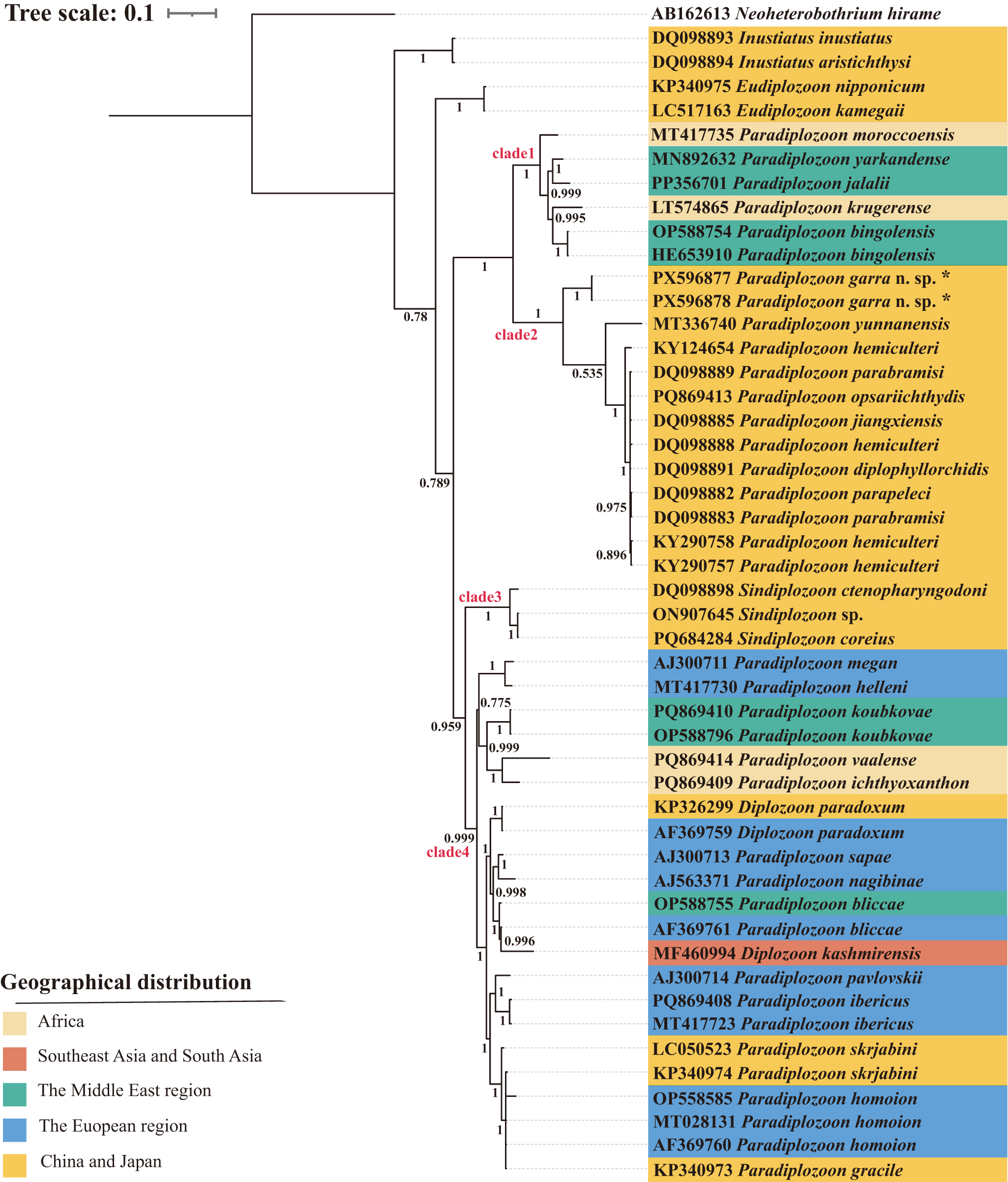
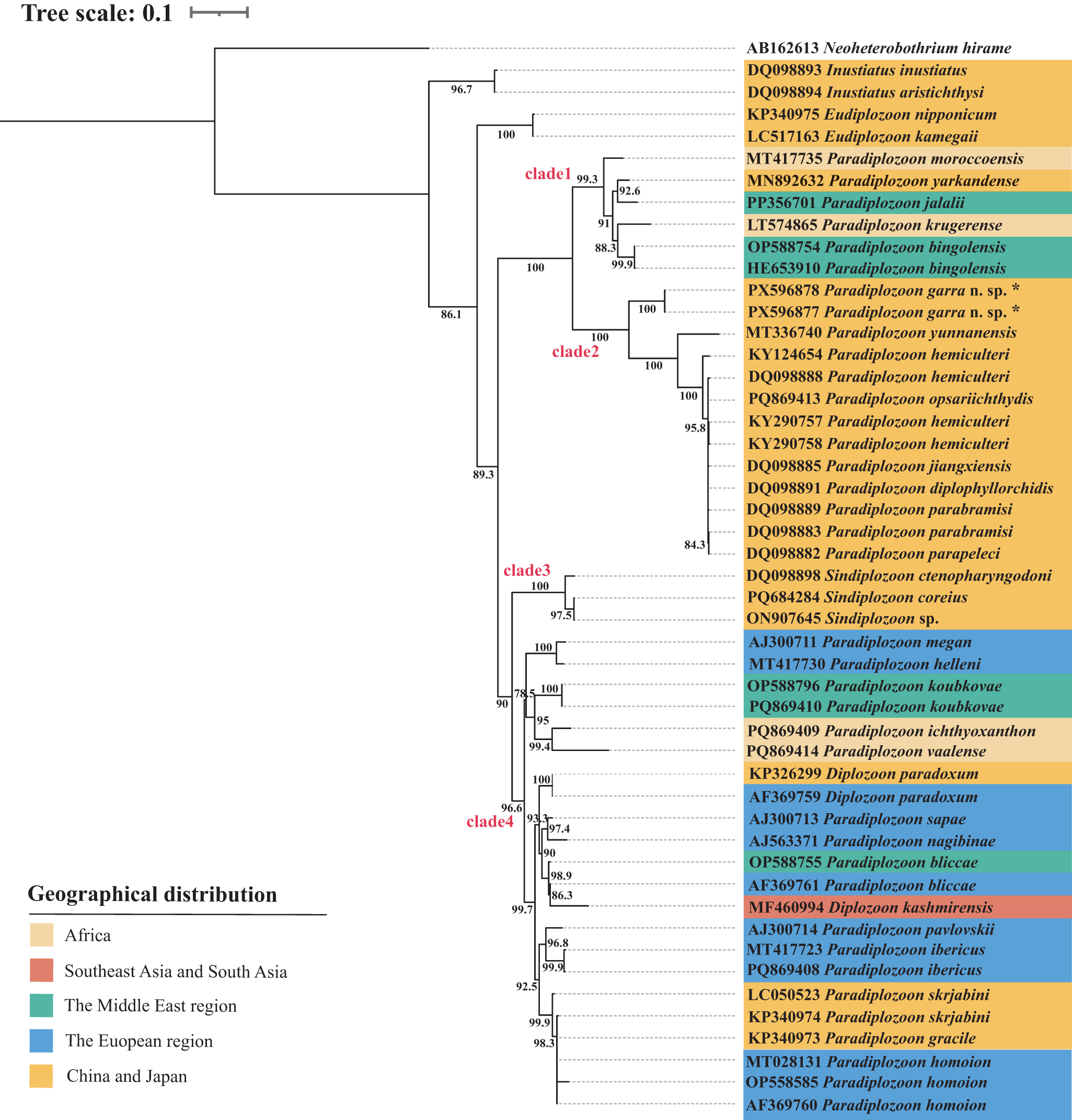
The phylogenetic tree showed that within the Diplozoidae, species of the genus Inustiatus Khotenovsky, 1978 diverged first, followed by the genus Eudiplozoon Khotenovsky, 1985. The remaining genera, Paradiplozoon, Sindiplozoon Khotenovsky, 1981, and Diplozoon von Nordmann, 1832, formed a large clade that was sister to Eudiplozoon. This large clade was further subdivided into two distinct subclades. Within one of the subclades, the two sequences of the newly described species (P. garra n. sp. PX596877, PX596878) formed a highly supported monophyletic group and clustered with other Paradiplozoon species from China forming clade 2. Clade 2 was sister to clade 1, which comprised Paradiplozoon species from the Middle East, China, Europe and South Africa, including P. krugerense Dos Santos & Avenant-Oldewage, 2016. In the other subclade, three Sindiplozoon species (DQ098898, ON907645, PQ684284) from China formed a highly supported group, designated as clade 3.
Clade 4 was sister to clade 3 and included Paradiplozoon and Diplozoon species from Europe, Paradiplozoon species from the Middle East, South Africa, China and Japan, as well as Diplozoon paradoxum von Nordmann, 1832 (KP326299) from China and D. kashmirensis Kaw, 1950 (MF460994) from India. While D. kashmirensis from India (MF460994) clustered with P. bliccae Reichenbach-Klinke, 1961 (AF369761) from France.
Discussion
Members of the genus Paradiplozoon exhibit distinctive morphological traits, making their identification at the genus level relatively straightforward. However, species-level identification remains challenging. In the present study, an integrative approach combining morphological and molecular analyses led to the identification of a new species, Paradiplozoon garra, collected from the gills of G. surgifrons in Kunming, southwest China.
In November 2024, a large number of Sindiplozoon coreius Cao, 2022 (Shen et al., Reference Shen, Zhao, Jiang, Xu, Tian, He, Jia, Xu, Meng and Fan2025) were collected from cultured Anabarilius grahami Regan, 1908 at the same fish farm where G. surgifrons samples were obtained. Notably, G. surgifrons was not co-cultured with A. graham and originated from a different source (with A. grahami introduced from Dali, Yunnan, and G. surgifrons from the Irrawaddy River), indicating that the farming conditions at this site are highly conducive to diplozoids proliferation, potentially detrimental to fish health. Other fish species co-cultured with G. surgifrons, including Barbodes hexagonolepis McClelland, 1839, Phoxinus lagowskii Dybowski, 1869 and Hemibarbus maculatus Bleeker, 1871, showed no signs of diplozoids infection, suggesting that P. garra exhibits a certain level of host specificity.
Morphologically, P. garra produces eggs without attached filaments, which reduces the likelihood that its eggs can remain stably adhered to gill filaments after being laid. Its attachment structures (central hooks and clamps) are relatively small compared to related species (Table 1), which is unfavourable for attachment. While the pronounced spurs on the clamps, distinct from other Paradiplozoon species, may enhance attachment efficiency. These morphological features, particularly the small size of central hooks and clamps combined with the unique spur on the clamps and the filament-free eggs, distinguish P. garra from all other known Paradiplozoon species and support its recognition as a new species. These observations suggest that its parasitic ability may be relatively unaffected by environmental conditions, possibly adapting to slower-flowing natural waters or resource-rich aquaculture environments, and adopting a relatively gentle attachment strategy. However, no wild populations of P. garra have yet been found, and future studies are needed to verify these hypotheses regarding its morphology and survival strategies.
Phylogenetic analyses (Figures 4 and 5), together with previous studies (Huang et al., Reference Huang, Zhou, Yuan and Ding2023), indicate that Chinese Paradiplozoon species belong to a relatively derived clade within Diplozoidae (Clade 2). Within this clade, P. yunnanense (Fan et al., Reference Fan, Meng, Bai, Xu and Wang2018; GenBank accession no. MT336740) from Yunnan occupies a basal position among Chinese taxa. Based on these findings, we speculate that Paradiplozoon populations distributed in southwestern China and extending into Southeast Asia may represent more evolutionarily primitive lineages, suggesting that Southeast Asia could be a potential origin centre for Paradiplozoon in East Asia. The newly described species in this study forms a distinct branch with high support (posterior probability = 1). It occupies a basal branch within Clade 2 and may represent a relatively ancestral lineage. Considering the natural distribution of its host in Yunnan (Lianghe County, Mengyang Town; Tengchong City, Mangbang Town) (Sun et al., Reference Sun, Li, Zhou and Li2018), it is reasonable to infer that natural populations of this parasite may also occur in adjacent regions of Myanmar and Laos, consistent with the geographic pattern shown in the phylogenetic tree. Compared with its close relatives in Clade 2, P. garra differs in host species (representing the first record of a diplozoid infecting a Garra species in China), geographic distribution (potentially occurring in the Irrawaddy River basin), and key morphological characters such as clamp and egg morphology, reinforcing the phylogenetic distinctions observed in our ITS2 analyses. Combined with the evidence, the species in this study is recognized as a new species. However, due to limited research on diplozoid parasites in Yunnan and Southeast Asia (Bai et al., Reference Bai, Wang, Li, Xu and Fan2014; Fan et al., Reference Fan, Meng, Bai, Xu and Wang2018), current evidence is insufficient to fully support this hypothesis regarding their geographic distribution and origin, and further data are needed for validation.
The rooted condensed tree of the parasite geographical distribution based on the BI analysis of ITS2 sequences of diplozoids. The numbers at nodes indicate posterior probabilities (%). The monogenean Neoheterobothrium hirame Ogawa, 1999 was used as the outgroup. Species marked with an * in the figures represent sequences from this study.

The rooted condensed tree of the parasite geographical distribution based on the ML analysis of ITS2 sequences of diplozoids. The numbers at nodes indicate bootstrap values (%). The monogenean Neoheterobothrium hirame Ogawa, 1999 was used as the outgroup. Species marked with an * in the figures represent sequences from this study.

The phylogenetic tree also shows that P. yarkandense from China clusters with P. moroccoensis from Morocco, along with several diplozoid species from the Middle East and Europe, forming Clade 1. This clustering supports the hypothesis that P. yarkandense, distributed in the Yarkand River in northwest China (Xinjiang), is geographically and phylogenetically close to Middle Eastern diplozoids. Similarly, P. moroccoensis from North Africa, near Europe, clusters with European diplozoid species.
While ITS2 is widely applied for species identification across monogeneans, including Paradiplozoon, its resolution for distinguishing closely related species is limited, and it may not accurately reflect deeper phylogenetic relationships. Therefore, ITS2 should ideally be complemented with additional markers and morphological analyses to achieve a robust understanding of species diversity.
In conclusion, this study improves our understanding of species diversity, host specificity and phylogenetic relationships within Diplozoidae. The discovery of P. garra from G. surgifrons represents the first record of a diplozoid infecting this host genus in China, indicating that diplozoid diversity in the region is likely greater than currently known. Phylogenetic analyses suggest that Chinese Paradiplozoon species form a derived lineage, with P. garra and P. yunnanensis occupying the base of this lineage, implying that southwestern China may be an important centre of origin and diversification for these parasites. Future studies should expand sampling across Yunnan and neighbouring Southeast Asian regions to further elucidate the evolutionary history and biogeographic patterns of diplozoid parasites. Moreover, in high-density, resource-rich aquaculture environments, diplozoids can proliferate rapidly, causing significant fish mortality, highlighting the urgent need for effective management strategies to mitigate infection risks.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182026102005.
Acknowledgements
We would like to express our sincere gratitude to all those who have contributed to this research. Special thanks go to Yuan-Wei Zhang, Xiao-Ai Wang and Jun-Xing Yang of Kunming Institute of Zoology Chinese Academy of Sciences for their invaluable guidance and support throughout the project.
Author contributions
LS participated in the sample collection, molecular lab work, data analysis and manuscript preparation. LXF and FYM participated in the revisions and additions to the article.
Financial support
This work was supported by the National Natural Science Foundation of China (32060115, 31260507 and 31560589) and Yunnan Province Basic Research Program Project (202301AT070087). This article is also supported by the Open Fund of the College of Life Sciences and the Doctoral Start-up Fund of Yunnan Normal University.
Competing interests
The authors declare no competing interests.
Ethical standards
All procedures contributing to this work comply with the ethical standards of the relevant national and institutional guides on the care and use of laboratory animals, and the study was approved by the Animal Care and Use Committee of Yunnan Normal University. The ethics review form number: ynnuethic2025-072.