Introduction
Pathogenic free-living amoebae (pFLA) are single-celled organisms found worldwide in soils and freshwater. The pFLA are classified as amphizoic amoebae being capable of both free-living and parasitic lifestyles (Schuster and Visvesvara, Reference Schuster and Visvesvara2004). Members of this group, including Acanthamoeba spp., Balamuthia mandrillaris, and Naegleria fowleri, can cause rare, yet often fatal, central nervous system (CNS) infections in humans. A fourth amoebae, Sappinia pedata, has been identified as the cause of a nonlethal case of amoebic encephalitis with CNS symptoms (Gelman et al., Reference Gelman, Rauf, Nader, Popov, Borkowski, Chaljub, Nauta and Visvesvara2001).
N. fowleri infection causes primary amebic meningoencephalitis (PAM), an acute, fulminant, and rapidly fatal disease. Infection occurs when amoebae-contaminated freshwater is introduced into the nasal passages. The pathogen then migrates along the olfactory nerves to the cribriform plate and gains access to the brain. Once in the brain, the amoebae feed and proliferate. They trigger host immune-mediated inflammation that, in combination with feeding amoebae, leads to tissue destruction and death of the infected individual within 7–10 days (da Rocha-azevedo et al., Reference da Rocha-azevedo, Tanowitz and Marciano-Cabral2009). Current treatment of PAM involves a combination of agents, including amphotericin B, which has become a cornerstone of treatment. Other anti-infectives used in these cocktails include fluconazole, miconazole, miltefosine, azithromycin and rifampin (Grace et al., Reference Grace, Asbill and Virga2015). PAM is characterized by a high mortality rate (>97%) that is exacerbated by two treatment challenges. First, timely diagnosis is difficult, relying on visual identification of amoebae in cerebral spinal fluid or molecular diagnostic tests that are not widely available. Second, the efficacy of current treatments is poor, which may be partly due to delays in diagnosis preventing a timely initiation of drug treatment. While these issues are multifaceted, they highlight the need for new, rapidly acting treatments (Guemez and Garcia, Reference Guemez and Garcia2021).
B. mandrillaris and several species of Acanthamoeba cause granulomatous amebic encephalitis (GAE), a rare opportunistic infection. Like PAM, these infections are typically fatal, with mortality rates above 90%. B. mandrillaris and Acanthamoeba spp. are found in freshwater and soil and can be introduced through skin lesions or inhalation of cysts (Krol-Turminska and Olender, Reference Krol-Turminska and Olender2017). GAE caused by Acanthamoeba mostly occurs in immunocompromised individuals, while Balamuthia GAE can occur in immunocompetent individuals (Kalra et al., Reference Kalra, Sharma, Shyam, Tejan and Ghoshal2020). Because of nonspecific symptoms and limited technical expertise, GAE is often misdiagnosed as bacterial or viral encephalitis (Krol-Turminska and Olender, Reference Krol-Turminska and Olender2017).
Current therapeutic regimens for GAE consist of a variety of drugs, including pentamidine, sulfadiazine, flucytosine, fluconazole and miltefosine. In B. mandrillaris GAE, a macrolide may also be included in the treatment regimen. Despite these treatments, mortality associated with the disease remains high, perhaps due to the low efficacy and high toxicity of the recommended compounds (Spottiswoode et al., Reference Spottiswoode, Haston, Hanners, Gruenberg, Kim, DeRisi and Wilson2024). Recently, nitroxoline was used to successfully treat Balamuthia GAE, suggesting that the agent could be a new addition to the cocktail used to treat those infections (Spottiswoode et al., Reference Spottiswoode, Pet, Kim, Gruenberg, Shah, Ramachandran, Laurie, Zia, Fouassier, Boutros, Lu, Zhang, Servellita, Bollen, Chiu, Wilson, Valdivia and DeRisi2023).
In addition to GAE, Acanthamoeba species can cause an infection of the cornea known as Acanthamoeba keratitis (AK), which can lead to vision loss (Lorenzo-Morales et al., Reference Lorenzo-Morales, Khan and Walochnik2015). Along with freshwater sources like pools and hot tubs, the amoebae can also proliferate in inappropriately maintained contact lens solution. AK largely impacts contact lens wearers, particularly those who do not maintain proper lens hygiene or who swim or shower wearing their lenses. Treatment approaches often drive the amoebae to differentiate into an environmentally resistant cyst stage that is refractory to many drugs. Successful management of an eye infection requires elimination of both trophozoites and cysts, a challenge that requires prolonged and repeated administration of topical antimicrobial agents (Lorenzo-Morales et al., Reference Lorenzo-Morales, Khan and Walochnik2015).
Drug discovery is limited by the lack of approaches to validate potential targets
Two approaches have been used to identify drugs that are effective against amoebae. The first method uses phenotypic screens to identify agents that kill or impair amoebae growth (Rice et al., Reference Rice, Colon, Chen, Hull and Kyle2020). Further, these screens are typically designed to exclude agents that demonstrate unacceptable toxicity towards human cells. Promising hits are often subject to structure–activity relationship (SAR) studies to improve their drug-like properties and refine their specificity towards amoebae and away from human cell lines. Phenotypic screens have been used to identify many of the agents that constitute the current treatment cocktails (Laurie et al., Reference Laurie, White, Retallack, Wu, Moser, Sakanari, Ang, Wilson, Arkin and DeRisi2018). The second route, target-based drug discovery, involves developing inhibitors to cellular targets anticipated to be critical for amoebae survival, including, but not limited to, structural proteins, metabolic enzymes, or DNA replication machinery (Debnath et al., Reference Debnath, Calvet, Jennings, Zhou, Aksenov, Luth, Abagyan, Nes, McKerrow and Podust2017). Knowledge about the potential target allows biased inhibitor development using structure-guided synthesis (SGS). SGS, in tandem with SAR studies and counter screens against human homologs and cell lines, can result in highly specific inhibitors of essential amoebae cellular processes with limited toxicity against human cell lines. As an example of such an approach, we have found that inhibitors of the glucose metabolism enzyme enolase have proven to be potent anti-amebic agents with promising activity in rodent models of disease (Milanes et al., Reference Milanes, Yan, Pham, Muller, Kwain, Rees, Dominy, Whitehead, Millward, Bolejack, Shek, Tillery, Phan, Staker, Moseman, Zhang, Ma, Jebet, Yin and Morris2024).
In target-based drug development campaigns, typically one of the first steps is to confirm that the putative cellular target is critical for pathogen viability. This is usually tested using genetic approaches to ablate (by knockout) or limit expression (by RNAi or similar approach) of the target gene followed by assessment of the impact on amoebae health. If a target gene is essential, then it is likely that ablation or knockdown of the protein would be detrimental to the pathogen, justifying the pursuit of the development of inhibitors as lead compounds for drug discovery (Chatterjee-Kishore and Miller Reference Chatterjee-Kishore and Miller2005; Xu et al., Reference Xu, Wang, Ma, Dong, Ashby and Hao2024). Additionally, genetic approaches are frequently used to confirm that observed pathogen phenotypes are due to on-target action of an inhibitor. This may involve assays that demonstrate engineered cells that no longer express a target protein, or that express a mutated target protein that no longer engages the inhibitor, lose sensitivity to the agent. The limited tools that allow for genetic manipulation of gene expression in the pFLA is a major hurdle in the validation of compounds that have shown promising activity against amoebae in culture or animal models.
Hurdles for genetic approaches in pFLA
The ability to modify gene expression in the pFLA remains challenging, with no such tools reported for B. mandrillaris or N. fowleri. Some progress has been made in Acanthamoeba spp., with Peng et al. (Reference Peng, Omaruddin and Bateman2005) describing the transfection of cells using an ectopic vector that expressed GFP and G418 resistance from the TATA-binding protein promoter. Two other transient transfection approaches have been described that used the polyubiquitin promoter to drive expression, resulting in ∼50% transfection efficiency in each case (Hu and Henney, Reference Hu and Henney1997; Kong and Pollard, Reference Kong and Pollard2002).
While tools for use in N. fowleri have not been described, approaches have been developed for the manipulation of gene expression in the non-pathogenic species Naegleria gruberi. For example, stable expression of GFP and antibiotic resistance has been described (Faktorova et al., Reference Faktorova, Nisbet, Fernandez Robledo, Casacuberta, Sudek, Allen, Ares, Areste, Balestreri, Barbrook, Beardslee, Bender, Booth, Bouget, Bowler, Breglia, Brownlee, Burger, Cerutti, Cesaroni, Chiurillo, Clemente, Coles, Collier, Cooney, Coyne, Docampo, Dupont, Edgcomb, Einarsson, Elustondo, Federici, Freire-Beneitez, Freyria, Fukuda, Garcia, Girguis, Gomaa, Gornik, Guo, Hampl, Hanawa, Haro-Contreras, Hehenberger, Highfield, Hirakawa, Hopes, Howe, Hu, Ibanez, Irwin, Ishii, Janowicz, Jones, Kachale, Fujimura-Kamada, Kaur, Kaye, Kazana, Keeling, King, Klobutcher, Lander, Lassadi, Li, Lin, Lozano, Luan, Maruyama, Matute, Miceli, Minagawa, Moosburner, Najle, Nanjappa, Nimmo, Noble, Novak Vanclova, Nowacki, Nunez, Pain, Piersanti, Pucciarelli, Pyrih, Rest, Rius, Robertson, Ruaud, Ruiz-Trillo, Sigg, Silver, Slamovits, Jason Smith, Sprecher, Stern, Swart, Tsaousis, Tsypin, Turkewitz, Turnsek, Valach, Verge, von Dassow, von der Haar, Waller, Wang, Wen, Wheeler, Woods, Zhang, Mock, Worden and Lukes2020). In this work, the authors describe transfection of the flagellated form of the amoebae, a life form that is not found in Acanthamoeba spp. or B. mandrillaris. Transfection of N. gruberi trophozoites has also been reported (Nguyen et al., Reference Nguyen, Chapman, Mullican and Drescher2024). In that study, amoebae were transiently transfected with a modified version of the closed circular extrachromosomal ribosomal DNA element (CERE). Authentic CERE is maintained at ∼4000 copies per cell and could be a useful starting point for the development of artificial chromosomes as platforms for transgene expression.
Efforts towards transgene expression in N. fowleri have not progressed as far and approaches developed in N. gruberi have not yet been successfully adapted to the pathogen (Palmentiero, Roster, McKeon, Morris, unpublished). Gene silencing in N. fowleri following transfection of an episome driving dsRNA expression has been reported, but a lack of clarity over experimental details has hampered efforts to adapt these findings to general transfection approaches (Jung et al., Reference Jung, Kim, Lee, Song, Kim, Park, Im and Shin2008, Reference Jung, Kim, Song, Lee, Kwon, Kim, Park, Im and Shin2009). Transient transfections, using a plasmid that drives eYFP expression from a putative ubiquitin promoter, have been successful (Figure 1), with fluorescence persisting for 8 days before detectable transgene expression is lost (Palmentiero, McKeon, Morris, unpublished).
Transient transfection of N. fowleri (strain NfTY), with pPACeYFP.V2 yields detectable fluorescence for up to eight days before transgene expression was lost. This plasmid harbours a puromycin acetyltransferase gene (pac) and eYFP. Both are flanked on the 5′ side by a 1090 bp fragment amplified from immediately upstream of the N. fowleri ubiquitin ORF. The pac is flanked on the 3′ side with a 349 bp fragment of the N. fowleri actin UTR that is immediately downstream of the actin stop codon while eYFP is flanked by a 1000 bp of the N. fowleri ubiquitin gene UTR. Supercoiled plasmid (5 µg) was assembled into polyethylenimine nanoparticles and was transfected into ~2 × 104 trophozoites. Briefly, the plasmid was mixed with either PEI-40 or PEI-25 at 1:1 ratio (w/w) then added to the parasites. After 48 h, cells were selected by addition of 20 µg/mL puromycin, and fluorescence was observed over time. These images were captured 8 days after transfection and 6 days after initiation of selection. The scale bar is 50 µm.

Together, these studies highlight the limited number of tools currently available to manipulate gene expression in the pFLA. The development of a genetic toolbox to modify the pFLA is essential for the identification and characterization of novel drug targets. Here, we will address potential challenges and assess the possibility for genetic manipulation in the pFLA through interrogation of their genomes for proteins known to be involved in stable maintenance of transgenes, RNAi, and gene editing in other systems. The presence of these genes does not guarantee their role in transgenesis but may guide approach modifications required for successful gene expression manipulation in the organisms.
Barriers to DNA delivery
The first challenge to overcome in generating transgenic amoebae is identifying methods for breaching cellular membranes to deliver cargo. One approach is not uniformly suitable for all cell types, as cellular differences can impact the efficiency of transfection approaches. For example, electroporation transfection efficiency is impacted by membrane composition. In electroporation, electric pulses are used to generate transient pores or holes in membranes that allow nucleic acids to enter the cell. Membrane fluidity, which is dependent on membrane composition, can impact pore size and pore duration.
Transfection approaches that employ cationic particles harbouring foreign DNA can also be influenced by the plasma membrane composition. Additionally, the chemical makeup and size of the lipoplexes bearing the nucleic acids can also dramatically alter transfection efficiency, further complicating optimization of approaches using cationic particles (Ramezani et al., Reference Ramezani, Khoshhamdam, Dehshahri and Malaekeh-Nikouei2009). Like the cell outer membrane, the nuclear membrane composition can impact the delivery of transfected nucleic acid to the nucleus. This membrane is selectively permeable, typically blocking large DNA plasmids from accessing the nuclear interior.
Cellular responses to foreign DNA
Integration of foreign nucleic acid into a target cell is typically achieved using transfection or viral transduction to deliver transgenes into the cell. The fate of the nucleic acid at that point can vary, with cellular responses to nucleic acid acquired from the environment eliciting a spectrum of responses. In the following subsections, we consider the role of human proteins known to be altered in response to transfection with a focus on the potential for homologs in the pFLA.
Transfection can trigger the upregulation of mechanisms aimed at preventing viral infection or viral replication (Warga et al., Reference Warga, Anderson, Tucker, Harris and Elmer2023). The nature of the nucleic acid impacts this response, with size and single- or double-stranded nature being influential in cellular responses. For example, dsRNAs that are >30 bp are known to elicit the double-stranded RNA-dependent protein kinase (PKR) response in mammalian cells, triggering apoptosis (Lemaire et al., Reference Lemaire, Anderson, Lary and Cole2008).
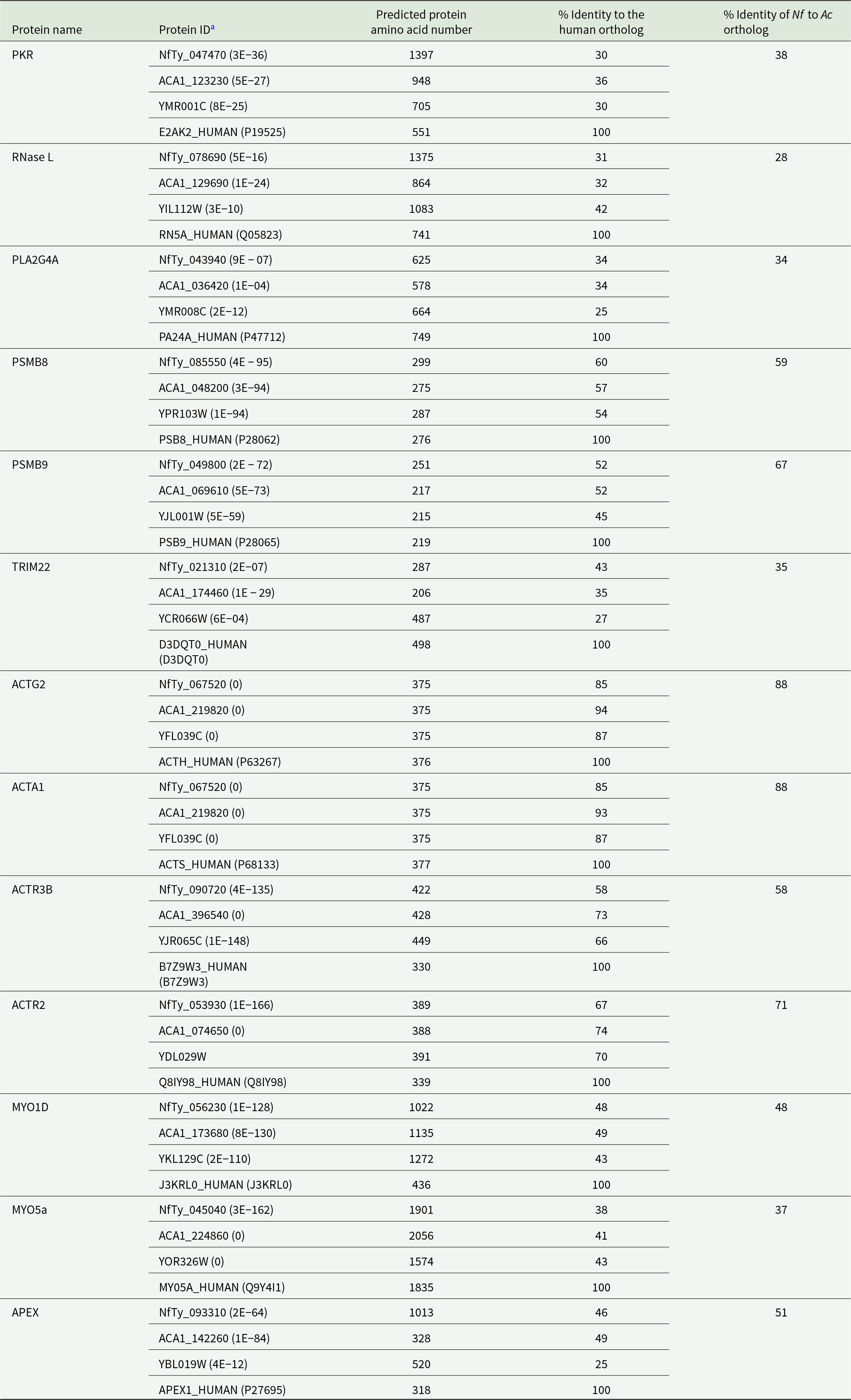
At this point, it is unclear if such a mechanism, which would likely impact cell culture health, is at work in the pFLA. N. fowleri and Acanthamoeba spp. may harbour a similar response, though homologous proteins have limited identity (30% and 36%, respectively) and have been annotated as putative translation initiation factors (Table 1). RNase L, another host enzyme that can respond to viral RNA, destroys both viral and host RNA while inducing type 1 interferon signalling. Again, possible homologs in the pFLA have limited identity, raising the possibility that this activity is absent in the amoebae.
pFLA orthologs of human and yeast proteins known to have changes in expression in response to transfections

a Proteins identified with lowest E-value after query using human orthologs against AmoebaDB (https://amoebadb.org/amoeba/app/) or FungiDB (https://fungidb.org/fungidb/app/). E-values are in parentheses.
The pFLA probably lack other nucleic acid responses found in mammals, including those that are elicited by the accumulation of DNAs in the cytoplasm. DNA sensors in mammalian cells can detect cytosolic DNA and produce cyclic GMP-AMP as a second messenger that leads to type I interferon production (Sun et al., Reference Sun, Wu, Du, Chen and Chen2013), which then activates antiviral cellular responses. The gamma-interferon-inducible protein 16 (IFI16) recognizes viral dsDNA and recruits factors to induce interferon-beta expression and subsequent antiviral activity (Thompson et al., Reference Thompson, Sharma, Atianand, Jensen, Carpenter, Knipe, Fitzgerald and Kurt-Jones2014). The absence of clear homologs in the pFLA to these proteins (and the interferons that are involved in the pathways) suggests the pFLA lack the canonical interferon response observed in mammals or elicit a response by distinct mechanisms.
The vehicle that is used to transfect foreign DNA can also elicit host cell responses. For example, in mammals, PLA2G4A, a phospholipase that hydrolyses phospholipids to generate the important inflammatory molecule arachidonic acid, can be activated in response to lipid introduction because of some lipofectamine transfection methods (Sarkar et al., Reference Sarkar, Jones, Hegdekar, Thayer, Kumar, Faden, Kane and Lipinski2020). The pFLA harbour predicted proteins with limited similarity to the human protein raising the possibility that the pFLA may harbour a similar activity. Additionally, the pFLA are noted for possessing a diverse arsenal of enzymes for the degradation of prey, so it is formally possible that one or more of these is impacting the vehicles used to deliver nucleic acids.
Proteosome-associated proteins
Foreign proteins that can be introduced with DNA during transfection are often degraded by the immunoproteasome complex. PSMB8 and PSMB9 are key components of that complex, and N. fowleri, Acanthamoeba spp. and S. cerevisiae likely have this cellular defense mechanism given the presence of predicted homologs (Table 1; Warga et al., Reference Warga, Anderson, Tucker, Harris and Elmer2023). TRIM22, a tumour suppressor that can influence targeting of cellular proteins to the proteosome to ultimately increase reactive oxygen species (ROS), was also upregulated in mammals in response to lipofectamine-based transfection. The pFLA have predicted protein homologs with limited (43% and 35%) identity, though the polypeptides are predicted to be less than one-half the size of the human enzyme, raising the possibility of distinct cellular function.
Delivery of cargo to the nucleus
Multiple proteins that impact transfection in mammalian cells are associated with cellular cargo movement, potentially reflecting the action of these proteins in the delivery of lipoplexes to the nucleus. These include actin and actin-associated proteins (e.g, ACTG2, ACTA1, ACTR3B and ACTR2) and myosin proteins like MYO1D and MYO5a. The pFLA amoebae have homologs of these critical proteins, but their role in transfection is unresolved (Table 1).
DNA repair
The apurinic/apyrimidinic endonuclease (APEX), a DNA repair enzyme that removes damaged or mismatched nucleotides in base excision repair, has been implicated as having a role in mammalian transfection, including repair of abasic sites that result from transfection reagents and maintaining recipient cell genome integrity (Lopez et al., Reference Lopez, Rodriguez and Banuelos2021). This is a conserved cellular DNA repair mechanism, so it is not surprising that the pFLA harbour putative proteins with moderate identity to the mammalian enzyme (46% and 49% for N. fowleri and A. castellanii homologs, respectively) (Table 1).
Possible solutions to this potential barrier in the pFLA
Our understanding of the cellular repertoire of pFLA proteins that could be creating barriers to transfection through responses to foreign nucleic acids and/or vehicles used to deliver those DNAs is poor. This makes forecasting the utility of various approaches challenging and initially suggests that breakthroughs will most likely occur after rigorous testing of a breadth of transfection approaches. These, tested in combinations of conditions known to improve DNA uptake in other systems (e.g., stresses akin to serum starvation in human cells, which can improve uptake (Wallenstein et al., Reference Wallenstein, Barminko, Schloss and Yarmush2010)), will likely be required to solve this key step.
Maintenance of foreign DNAs
Maintenance of transgenes requires parental cells to replicate and distribute the foreign nucleic acid to daughter cells. Heritable maintenance is commonly achieved through the delivery of dsDNAs as extrachromosomal episomes bearing elements required for duplication and distribution to daughter cells. Alternatively, stable transgenesis can be established through integration of the transgene into the chromosome, where it is replicated and distributed as part of the normal complement of DNA.
Episomes and maintenance of extrachromosomal DNAs
Nuclear extrachromosomal circular DNA (eccDNA) is nearly ubiquitous amongst eukaryotes and is commonly generated through homologous recombination (HR), non-homologous end joining (NHEJ), and double-strand breaks (DSBs) (Zhao et al., Reference Zhao, Yu, Zhang, Su and Zhou2022). Amongst the pFLA, only N. fowleri is known to maintain nuclear eccDNA, the closed CERE, which harbours the organism’s ribosomal RNA genes (Mullican et al., Reference Mullican, Drescher, Chapman and Tracy2020). Little is known about the replication and maintenance of the CERE, with findings in the non-pathogenic species N. gruberi providing insight into mechanisms likely found in N. fowleri. In N. gruberi, a single origin of replication for CERE has been described from a 2.1 kb fragment of non-ribosomal DNA (Mullican et al., Reference Mullican, Chapman and Tracy2019). Additionally, in N. gruberi CERE is replicated through a theta-type replication mechanism (Mullican et al., Reference Mullican, Chapman and Tracy2019). However, the N. fowleri reference strain, the TY strain, has a CERE that shares limited similarity to the sequenced N. gruberi and N. fowleri CERE in the non-ribosomal sequence (43.3%) and has a high similarity to the ribosomal DNA (91.7%). The limited sequence conservation between the N. gruberi and N. fowleri CERE non-ribosomal sequences makes origin identification by sequence comparison challenging (Nguyen et al., Reference Nguyen, Chapman, Tracy and Drescher2021).
Possible solutions to transgenesis using episomal DNAs in the pFLA
CERE may be a means of generating an artificial chromosome for transgene expression, like those that have been used in other systems (yeast, humans, trypanosomes (Murray and Szostak, Reference Murray and Szostak1983; Patnaik et al., Reference Patnaik, Axelrod, der Ploeg Lh and Cross1996; Harrington et al., Reference Harrington, Van Bokkelen, Mays, Gustashaw and Willard1997)). Others have cloned the entire N. gruberi CERE into a bacterial vector and reintroduced the chimeric CERE into amoebae, finding that this supported maintenance of the bacterial sequence. While it is unclear if the DNA remained in the cells as episomes, the bacterial sequences could be detected by PCR after seven cell passages in the absence of selection of the transgene (Nguyen et al., Reference Nguyen, Chapman, Mullican and Drescher2024). To date, the origin of replication from N. gruberi CERE has not supported episome maintenance in N. fowleri. Once the origin/autonomously replicating sequence (ARS) in the N. fowleri CERE is resolved, plasmids bearing this sequence may be a useful means of stable introduction of foreign DNA.
Transgene integration into the genome
Linear dsDNAs bearing transgenes can be incorporated into the genome by HR, NHEJ, or microhomology-mediated end joining (MMEJ) during chromosome replication or repair. HR requires a homologous template, usually a sister chromatid, to repair breaks, while canonical NHEJ is template-independent, requiring little to no sequence homology (<4 nt) between broken DNA ends for repair. MMEJ requires short stretches of microhomology (2–20 nt) to mediate DNA repair (Ranjha et al., Reference Ranjha, Howard and Cejka2018). Targeted gene integration likely requires HR, while NHEJ and MMEJ are sufficient for random integration, suggesting that the presence of any of these pathways could be exploited to generate transgenic amoebae by incorporation of foreign DNA into breaks during repair.
DSBs are made due to endogenous processes as well as exposure to exogenous factors. DSBs can form in unperturbed cells due to collisions between the replication and transcription machinery, nucleotide imbalances, or due to reactions with by-products of cellular metabolism (Yu and Anderson, Reference Yu and Anderson1997). Exposure to exogenous mutagens like ultraviolet (UV) irradiation and industrial chemicals can also damage DNA, resulting in strand breaks (Hiraku, Reference Hiraku2025). One possibility is inducing DNA damage in amoebae which could lead to enhanced transgene integration. In support of this, introducing DSBs into the genome in other organisms including yeast and mammals results in a significant increase in random integration (Zelensky et al., Reference Zelensky, Schoonakker, Brandsma, Tijsterman, van Gent, Essers and Kanaar2020) as well as gene targeting events (Rudin et al., Reference Rudin, Sugarman and Haber1989; Rouet et al., Reference Rouet, Smih and Jasin1994; Choulika et al., Reference Choulika, Perrin, Dujon and Nicolas1995; Smih et al., Reference Smih, Rouet, Romanienko and Jasin1995).
Proteins involved in DSB repair that may be required for transgenesis
Given the importance of DSB repair pathways in promoting integration of transgenes, it is possible that proteins involved in DNA repair could be important for the generation of stable transgenes in the pFLA. Here, we describe several of the known key players in DSB repair and consider differences found in pFLA that could explain the refractory nature of the amoebae to transgenesis.
The MRN complex
The MRE11-RAD50-NBS1 protein complex (the MRN complex) is one of the first repair complexes that localizes to DSBs. This complex has been proposed to have a structural role in tethering the broken chromosomes together while modulating DNA damage signalling (van den Bosch et al., Reference van den Bosch, Bree and Lowndes2003). The carboxy-terminal interacting protein (CtIP) has also been identified to play a critical role in promoting HR-mediated DSB repair through its interaction with the MRN complex. CtIP binds to BRCA1 as well, forming the MRN-CtIP-BRCA1 complex, which is important for end-resection to initiate HR (Wang et al., Reference Wang, Shao, Shi, Hwang, Truong, Berns, Chen and Wu2012).
The pFLA have a homolog for the MRE11 protein, with sequence identity ranging from 38% to 44% (Table 2). Additionally, a CtIP homolog, which may promote the MRE11 end resection of DSB, is present in all three amoebae (Ranjha et al., Reference Ranjha, Howard and Cejka2018). However, it is less clear if other protein components of the MRN complex, including RAD50 and NBS1, are present. In addition, there is no obvious BRCA1 homolog among the pFLA. The absence of BRCA homologs likely does not reflect a deficiency in homology-driven repair (HDR), as yeast lack BRCA homologs yet have very high rates of HDR (Guaragnella et al., Reference Guaragnella, Palermo, Galli, Moro, Mazzoni and Giannattasio2014).
pFLA orthologs of human and yeast HDR proteins

a Proteins identified with lowest E-value after query using human orthologs against AmoebaDB (https://amoebadb.org/amoeba/app/) or NCBI (for yeast). E-values are in parentheses.
b While having greatest %identity to RAD51, this N. fowleri gene had the lowest E-value for multiple Rad51 paralogs.
RAD51
In mitotic cells, RAD51 is the central enzyme that carries out HR. RAD51 is an ATP-dependent DNA-binding protein that forms a nucleoprotein filament on ssDNA after end resection. The RAD51 filament searches the genome for a homologous sequence and promotes strand exchange to use the intact sequence as a template for repair (Harris et al., Reference Harris, Rabellino and Khanna2018). RAD51 interacts with several partners that promote HR in cells. For example, BRCA2 contains eight conserved ∼35 amino acid motifs called BRC repeats that can all independently interact with RAD51 to promote its nucleation on ssDNA and others stimulating the growth of the nucleofilament. RAD51 nucleoprotein filament assembly is also stimulated by RAD54 (Ranjha et al., Reference Ranjha, Howard and Cejka2018), which stimulates RAD51 DNA exchange activity. Additionally, RAD54 binds to Holliday junctions and drives branch migration while helping to stimulate DNA cleavage activity (Mazin et al., Reference Mazin, Mazina, Bugreev and Rossi2010).
The RAD51 paralogs, including RAD51B, RAD51C, RAD51D, XRCC2 and XRCC3, share limited sequence homology to RAD51 beyond their well-conserved ATP-binding domains. The paralogs form two major multi-protein complexes: the BCDX2 complex, consisting of RAD51B/C/D/XRCC2, and the CX3 complex, consisting of RAD51C/XRCC3. These paralog complexes have a critical role in regulating the structure and stability of RAD51 ssDNA filaments. The increased stability of the nucleofilament favours DNA repair by HR as well as increases the efficiency of the homology search. These roles are critical, as deletion of any of the paralogs inhibits the localization of RAD51 to radiation-induced DSBs and leads to a reduction in sister chromatid exchange (Harris et al., Reference Harris, Rabellino and Khanna2018).
RAD51 is an essential gene that is required for mitotic HR-mediated DSB repair. Therefore, it is not surprising that the pFLA harbour-related homologs, with N. fowleri and A. castellanii having closely related protein homologs (62% and 83% identical, respectively) (Table 2). The pFLA also harbour potential homologs of the human RAD51 paralog family (RAD51B/C/D/XRCC2/3), although modest protein identity (25–40%) emphasizes the need for experimental confirmation of these proteins. Last, the pFLA species likely harbour BRCA2 homologs, but the limited identity (27–30%) of these proteins calls for caution in assigning these functions without experimental support.
What do similarities and differences in DNA repair suggest regarding transgenesis approaches in the pFLA?
DNA repair pathways in the pFLA may be the limiting factor in our efforts to generate transgenic pathogens. The absence of genetic tools makes understanding repair in these organisms challenging. However, numerous well-characterized inhibitors target NHEJ and HR from other organisms and could be candidates to test against the homologous targets in amoebae (Ray et al., Reference Ray, Vartak and Raghavan2020; Groelly et al., Reference Groelly, Fawkes, Dagg, Blackford and Tarsounas2023). These would enable the dissection of various repair mechanisms and could prove useful for manipulating pathways that limit transgene integration (e.g., by being too high-fidelity).
Gene editing
Gene editing, here described to alter genes in a site-directed manner through the addition, elimination, or modification of nucleotides, generates DNA breaks as part of the process. Editing occurs when NHEJ/MMEJ incorrectly repair DSBs leading to addition or deletion of nucleotides resulting in frameshift mutations. HR-mediated repair can be used to generate precise nucleotide changes (e.g., base substitutions) by providing a donor template with the desired modifications. Efforts to edit genomes in other systems have led to the development of a diverse set of reagents and approaches to generate a site-specific DSB to promote gene editing, including those that utilize transcription activator-like effector nucleases (TALENs) or CRISPR-Cas9.
TALEN proteins were originally developed from transcription activator-like effectors (TALEs) identified in the bacterial plant pathogen Xanthomonas (Boch et al., Reference Boch, Scholze, Schornack, Landgraf, Hahn, Kay, Lahaye, Nickstadt and Bonas2009). The TALE proteins were found to bind DNA and alter host gene transcription, much like a transcription factor. Generation of chimeric proteins bearing the DNA-binding domain of TALE with the catalytic domain of the restriction endonuclease Fok1 yielded TALENS. The Fok1 endonuclease acts as a dimer requiring two TALENS to be engineered per target site, limiting its utility for gene editing. This fusion protein introduces double-stranded DNA breaks in target genes, which use NHEJ/MMEJ or HR repair (Kim et al., Reference Kim, Skowron and Szybalski1996; Porteus and Carroll, Reference Porteus and Carroll2005).
CRISPR-Cas9 has been developed into a powerful and versatile genome editing tool. Originally identified in bacteria, the system has been found to play an important role in bacterial adaptive immunity. Short regions of DNA separated by palindromic repeats, Clustered Regulatory Interspaced Short Palindromic Repeats (CRISPR), were found to be homologous to phages and plasmids (Bolotin et al., Reference Bolotin, Quinquis, Sorokin and Ehrlich2005; Mojica et al., Reference Mojica, Diez-Villasenor, Garcia-Martinez and Soria2005; Pourcel et al., Reference Pourcel, Salvignol and Vergnaud2005). Association of CRISPR RNA (crRNA) with Cas9, a nuclease encoded nearby the CRISPR region, yielded a target-specific ribonucleoprotein complex capable of preventing phage infection (Garneau et al., Reference Garneau, Dupuis, Villion, Romero, Barrangou, Boyaval, Fremaux, Horvath, Magadan and Moineau2010; Horvath and Barrangou, Reference Horvath and Barrangou2010).
CRISPR/Cas9 cleavage specificity is directed by complementarity between the target DNA and the Cas9-associated guide RNA, allowing generation of ribonucleoprotein complexes for programmed cleavage of specific DNA sites (Jinek et al., Reference Jinek, Chylinski, Fonfara, Hauer, Doudna and Charpentier2012; Ran et al., Reference Ran, Hsu, Wright, Agarwala, Scott and Zhang2013). The system has been simplified through an improved understanding of the components required for high fidelity editing, now only requiring the Cas9 nuclease and a single guide RNA that serves as a scaffold for interacting with the nuclease while also providing sequence specificity for the cleavage. This has made the approach a flexible and user-friendly tool for gene editing (Gaj et al., Reference Gaj, Sirk, Shui and Liu2016).
What are the challenges and solutions to using gene editing in the pFLA?
Gene editing will likely prove to be a useful tool for developing transgenic pFLA, as CRISPR/Cas9 has been found useful for gene manipulation in many eukaryotic systems. Direct delivery of exogenous recombinant purified Cas9 complexed with in vitro synthesized guide RNAs or transfection of plasmids harbouring genes for the enzyme and guide RNA may be a potential means of delivery of the activity into cells. Donor repair DNA templates could be included or delivered independently. In either case, cargo delivery approaches will need to be developed and refined first before this tool can be thoroughly assessed.
RNAi and the pFLA
RNA interference (RNAi) is another useful tool for the manipulation of gene expression in many systems. RNAi is triggered by introducing dsRNA into cells, leading to the degradation of cognate mRNAs bearing the same sequence as the dsRNA (Fire et al., Reference Fire, Xu, Montgomery, Kostas, Driver and Mello1998). DsRNAs are processed to ∼20 bp small interfering RNAs (siRNAs) that are associated with the RNA-induced silencing complex (RISC), leading to homology-dependent degradation of the target mRNA during RNAi (Hamilton and Baulcombe, Reference Hamilton and Baulcombe1999; Piatek and Werner, Reference Piatek and Werner2014). RNAi has advantages over other approaches, including offering the opportunity to study the impact of the loss of essential genes on cells. Unlike knockouts, RNAi attenuates gene expression through reduction of mRNA and the rate of lethality is impacted by the penetrance of the silencing and the half-life of the mRNA. This makes RNAi a powerful approach for exploring gene function on a genome-wide scale (Hannon, Reference Hannon2002; Morris et al., Reference Morris, Wang, Drew and Englund2002; Wilson et al., Reference Wilson, Kwok, Porter, Payne, McElroy, Ohle, Greenhill, Blahna, Yamamoto, Jean, Mizgerd and Kotton2013).
Key proteins involved in the RNAi pathway
Dicer
Dicer is the RNase III responsible for cleavage of dsRNAs into siRNAs, activating the RISC, which occurs in the first stages of RNAi. It is unclear if the pFLA harbour homologs of this protein as predicted proteins with the best E-values (thereby being unlikely to be chance events but rather reflective of a potentially shared ancestry) have low identity (24% and 26%, for N. fowleri and A. castellanii proteins, respectively) and differ from the human enzyme in size; however, both have a predicted RNase III catalytic domain, which is similar to that found in Dicer (Table 3). Adding to the complexity of this comparison, humans harbor a related RNase III, Drosha, which cleaves endogenous RNAs that have hairpin structures into microRNAs (miRNAs) that regulate gene expression. The same A. castellanii predicted protein had the best E-value for both Dicer and Drosha, raising the possibility of a function in either (or both) pathways.
pFLA orthologs of human RNAi proteins

a Proteins identified with lowest E-value after query using human orthologs against AmoebaDB (https://amoebadb.org/amoeba/app/) or NCBI (for human). E-values are in parentheses.
b ACA1_291000 was also the only potential homolog identified for Dicer in A. castellanii, with a p-value of 5E-12 and 26% identity to the human protein.
c While humans have four argonaute RISC components, the pFLA have a single predicted ortholog.
Argonaute
In humans, Argonaute is composed of four related proteins. These proteins are a component of the RISC, serving to cleave the target mRNA strand that is complementary to the siRNA. The pFLA harbour single potential homologs to the four proteins (based on E-value, with N. fowleri having the best score), though the identity to the human proteins is low (Table 3).
MOV10
MOV10 is an RNA helicase component of the RISC that likely plays a role in resolving secondary structures that can be components of RNA. This helicase has additional activities in development and in antiviral responses (Nawaz et al., Reference Nawaz, Shilikbay, Skariah and Ceman2022). N. fowleri and A. castellanii harbour potential homologs that share 62% identity with each other.
C3PO
The RNAi regulator C3PO is a complex of a Mg2+-dependent endoribonuclease that removes the siRNA passenger strand cleavage products, which promotes the action of the RNAi machinery (Liu et al., Reference Liu, Ye, Jiang, Liang, Chen, Peng, Kinch, Grishin and Liu2009). Both N. fowleri and A. castellanii harbour possible homologs that are similarly sized to the human enzyme, although identity of the proteins was modest (36% and 39%).
Notably, homologs of several key putative RNAi proteins were not identified in the pFLA genomes. For example, DGCR8 (DiGeorge syndrome critical region gene 8, also called Pasha), a microprocessor complex subunit and binding partner of Drosha that participates in the repair of UV photoproducts, was not identified (Calses et al., Reference Calses, Dhillon, Tucker, Chi, Huang, Kawasumi, Nghiem, Wang, Clurman, Jacquemont, Gafken, Sugasawa, Saijo and Taniguchi2017). While this could reflect the absence of miRNA pathways in the pFLA, the identification of conserved miRNAs in A. castellanii suggests the amoebae may use mechanisms distinct from other organisms to generate regulatory small RNAs (Edelbroek et al., Reference Edelbroek, Kjellin, Biryukova, Liao, Lundberg, Noegel, Eichinger, Friedlander and Soderbom2024).
The pFLA also lack clear homologs of the RISC loading complex transactivation response RNA-binding protein, TRBP2, and the protein kinase RNA activator (PACT). Both proteins are part of the dsRNA-binding components of RISC, with TRBP2 also participating in the loading of AGO2 to the complex while PACT functions to regulate processing of pre-siRNA substrates. Both proteins also participate in the PKR.
Outlook for using RNAi in the pFLA
RNAi is not a universal eukaryotic activity, with some organisms lacking the ability to silence gene expression. These include the model species Saccharomyces cerevisiae, which lacks key proteins like Dicer and Argonaute. The pFLA have homologs to those proteins, suggesting they may have an intact RNAi pathway. The lack of clear understanding on optimized delivery of the dsRNA triggers required for RNAi (by transfection or similar) creates challenges in exploiting the pathway for manipulating gene expression. The giant virus Catovirus naegleriensis may provide clues on how to transfect the amoebae. This virus can infect and kill bacteria-fed Naegleria but does not infect amoebae adapted to axenic conditions. This suggests that the virus requires phagocytosis for entry into the host – a path we have attempted to exploit by feeding N. fowleri bacteria that express dsRNAs without success (McKeon and Morris, unpublished).
Additional considerations
This review has focused on cellular responses to transfection that could be impacting nucleic acid uptake. There are additional hurdles that need to be addressed before routine transfection approaches are resolved. First, it is unclear what nucleic acid elements are required for transgene expression. Regulatory sequences like promoters will likely be key components of a transgene expression system. These regulatory sequences, in combination with useful selectable markers, will also be important for development of heritable transgene expression tools. Other regulatory components, including polyA addition sequences and terminators, may be required for this approach to work.
Conclusions
The pFLA are responsible for infections that are difficult to treat and typically have poor outcomes. Efforts to develop new therapeutics for the treatment of these infections have been hampered by a lack of understanding of the basic cell biology of the organisms. As a result, amoebae-specific treatment options are not available. Instead, repurposed agents have been identified that can be anti-amebic in vitro, but these have yet to be proven effective in the treatment of disease.
Resolving approaches for genetic manipulation of the pathogens would open the door to new treatment avenues providing a clearer understanding of amoebae gene function. This would have a 2-fold impact, enabling the selection of targets that are amoebae-specific and the assessment of their essentiality to the organism. Unfortunately, the challenges associated with transfection of the pFLA have limited the use of reverse and forward genetics to study gene function, validate drug target choices and create reporter cell lines.
The reasons for these challenges are likely multifaceted. First, the developmental stage of the amoebae may influence transfection efficiency. For example, transfection of N. gruberi flagellates has been reported (Faktorova et al., Reference Faktorova, Nisbet, Fernandez Robledo, Casacuberta, Sudek, Allen, Ares, Areste, Balestreri, Barbrook, Beardslee, Bender, Booth, Bouget, Bowler, Breglia, Brownlee, Burger, Cerutti, Cesaroni, Chiurillo, Clemente, Coles, Collier, Cooney, Coyne, Docampo, Dupont, Edgcomb, Einarsson, Elustondo, Federici, Freire-Beneitez, Freyria, Fukuda, Garcia, Girguis, Gomaa, Gornik, Guo, Hampl, Hanawa, Haro-Contreras, Hehenberger, Highfield, Hirakawa, Hopes, Howe, Hu, Ibanez, Irwin, Ishii, Janowicz, Jones, Kachale, Fujimura-Kamada, Kaur, Kaye, Kazana, Keeling, King, Klobutcher, Lander, Lassadi, Li, Lin, Lozano, Luan, Maruyama, Matute, Miceli, Minagawa, Moosburner, Najle, Nanjappa, Nimmo, Noble, Novak Vanclova, Nowacki, Nunez, Pain, Piersanti, Pucciarelli, Pyrih, Rest, Rius, Robertson, Ruaud, Ruiz-Trillo, Sigg, Silver, Slamovits, Jason Smith, Sprecher, Stern, Swart, Tsaousis, Tsypin, Turkewitz, Turnsek, Valach, Verge, von Dassow, von der Haar, Waller, Wang, Wen, Wheeler, Woods, Zhang, Mock, Worden and Lukes2020), although this approach has not been successfully adopted to N. fowleri (Palmentiero and Morris, unpublished observation). The cell cycle stage may also be critical for success, although this has not yet been explored in N. fowleri. In N. gruberi, CERE abundance increases during encystment, possibly in preparation for future excystment when the environment is more nutrient-rich (Nguyen, Reference Nguyen2024). This increase in copy number may impact transfection efficiency of exogenous DNA by depleting biosynthetic intermediates needed for the replication and repair of the transgenic episome. Second, successful transgenesis may require cells to be actively phagocytosing food/prey, rather than using pinocytosis associated with axenic growth to acquire nutrients. Supporting this supposition, viral uptake is enhanced in phagocytosing amoeba.
Other potential inhibitors of transfection include cellular nucleases. In humans, DNase II, an acidic lysosomal endonuclease, can impact transfection efficiency of foreign DNAs (Howell et al., Reference Howell, Krieser, Eastman and Barry2003). While the amoebae lack a clear homolog to this enzyme (by sequence comparison), other endonucleases may be at work impacting transfection, including those used to break down prey nucleic acids. Alternatively, these could include the intron-encoded homing endonucleases (Elde et al., Reference Elde, Haugen, Willassen and Johansen1999, Reference Elde, Willassen and Johansen2000), though their restrictive sequence specificity suggests this is unlikely.
This review explores various cellular pathways known to influence transfection in a variety of systems, using sequence gazing to assess the likelihood of the presence of homologs in the pFLA. The lack of sequence homologs for various cellular processes potentially related to transgenesis cannot rule out the possibility that other unrecognized proteins provide the activity. To fully validate potential candidates, heterologous expression in tractable systems, such as yeast, followed by careful assessment of function, could be used. Nevertheless, multiple missing conserved components raise the possibility that a pathway may be diminished or absent in the pathogens.
Author contributions
All authors contributed to the writing, editing, and conceptualization of this manuscript. Caroline Palmentiero and Colm Roster contributed equally to this document.
Financial support
Work from the JCM laboratory was partially supported by the National Institutes of Health (R21 AI175463, R21 AI171217, and a NIH Center for Biomedical Excellence (COBRE) grant under award number P20GM146584). JA was partially supported by a Seifert Scholars Summer internship award and a Creative Inquiry award.
Competing interests
None.
Ethical standards
None.