Introduction
It is universally recognized that there is a global biodiversity crisis (Mace et al., Reference Mace, Masundire and Baille2005) and, despite some uncertainties, contemporary rates of species extinction are estimated to be at least three orders of magnitude above the background extinction rate (Mace et al., Reference Mace, Masundire and Baille2005). However, reduction of biodiversity is even higher within species, affecting population size (Loh et al., Reference Loh, Green, Ricketts, Lamoreux, Jenkins, Kapos and Randers2005) and reducing species distributions, with the resultant loss of local, genetically adapted populations (Ceballos & Ehrlich, Reference Ceballos and Ehrlich2002).
There is considerable debate over how to allocate limited resources to conservation initiatives below the species level. Several authors (Erwin, Reference Erwin1991; Moritz, Reference Moritz1999) have suggested that conservation goals should be to conserve ecological and evolutionary processes rather than to preserve specific phenotypic variants, the products of those processes. From this perspective, one prominent idea has been the concept of the evolutionary significant unit. Evolutionary significant units are populations that merit separate management and are a conservation priority (Fraser & Bernatchez, Reference Fraser and Bernatchez2001). Frequently, genetic differentiation indicating a long history of reproductive isolation of two populations has been considered sufficient to recognize these units (Moritz, Reference Moritz1994). However, Crandall et al. (Reference Crandall, Bibinda-Emonds, Mace and Wayne2000) suggest that some kind of ecological differentiation should be added to genetic distance to recognize populations that could be managed as different units.
Lack of regulation could be facilitating the mixing of two clearly different evolutionary significant units of the European rabbit Oryctolagus cuniculus in Spain because of deliberate translocations to facilitate hunting and, in some cases, conservation goals. The European rabbit is a keystone species in the Mediterranean ecosystem (Delibes-Mateos et al., Reference Delibes-Mateos, Redpath, Angulo, Ferreras and Villafuerte2007), a biodiversity hotspot (Myers et al., Reference Myers, Mittermeier, Mittermeier, da Fonseca and Kent2000). The rabbit is native to the Iberian Peninsula in south-western Europe, and two divergent genetic lineages (usually called A and B) can be recognized by analyses of mitochondrial DNA (mtDNA) variation (Branco et al., Reference Branco, Ferrand and >Monnerot2000, Reference Branco, Monnerot, Ferrand and Templeton2002) and the study of nuclear genes (Geraldes et al., Reference Geraldes, Ferrand and Nachman2006). Lineages A and B differentiated approximately two million years ago in two different Quaternary refugia in southern Spain (Branco et al., Reference Branco, Ferrand and >Monnerot2000). At present, lineage A is found in the south-west of the Iberian Peninsula and lineage B in other areas, including north-east Spain (Fig. 1) and everywhere else in the world where rabbits have been introduced (Branco et al., Reference Branco, Ferrand and >Monnerot2000, Reference Branco, Monnerot, Ferrand and Templeton2002). During the last postglacial period rabbits have come into secondary contact in central Iberia (Fig. 1; Branco et al., Reference Branco, Ferrand and >Monnerot2000, Reference Branco, Monnerot, Ferrand and Templeton2002). Besides genetic differentiation, lineages A and B can be associated with the traditional subspecies O. c. algirus, and O. c. cuniculus, respectively. Differences between these subspecies include body size, sexual maturation and litter size (R. Villafuerte, unpubl. data). Therefore, these two rabbit lineages meet all the criteria to warrant separate management.
Distribution area of European rabbit lineages A and B in the Iberian Peninsula and the area of overlap between them (100 km wide strip, as delimited by Branco et al., Reference Branco, Monnerot, Ferrand and Templeton2002), and the locations in which 60 interviews were carried out in 1993 and 2002. Translocations in the decade prior to the 2002 interviews (see text for details) are indicated by arrows showing the origin and destination of the released rabbits. Inset indicates location of main figure in Europe.
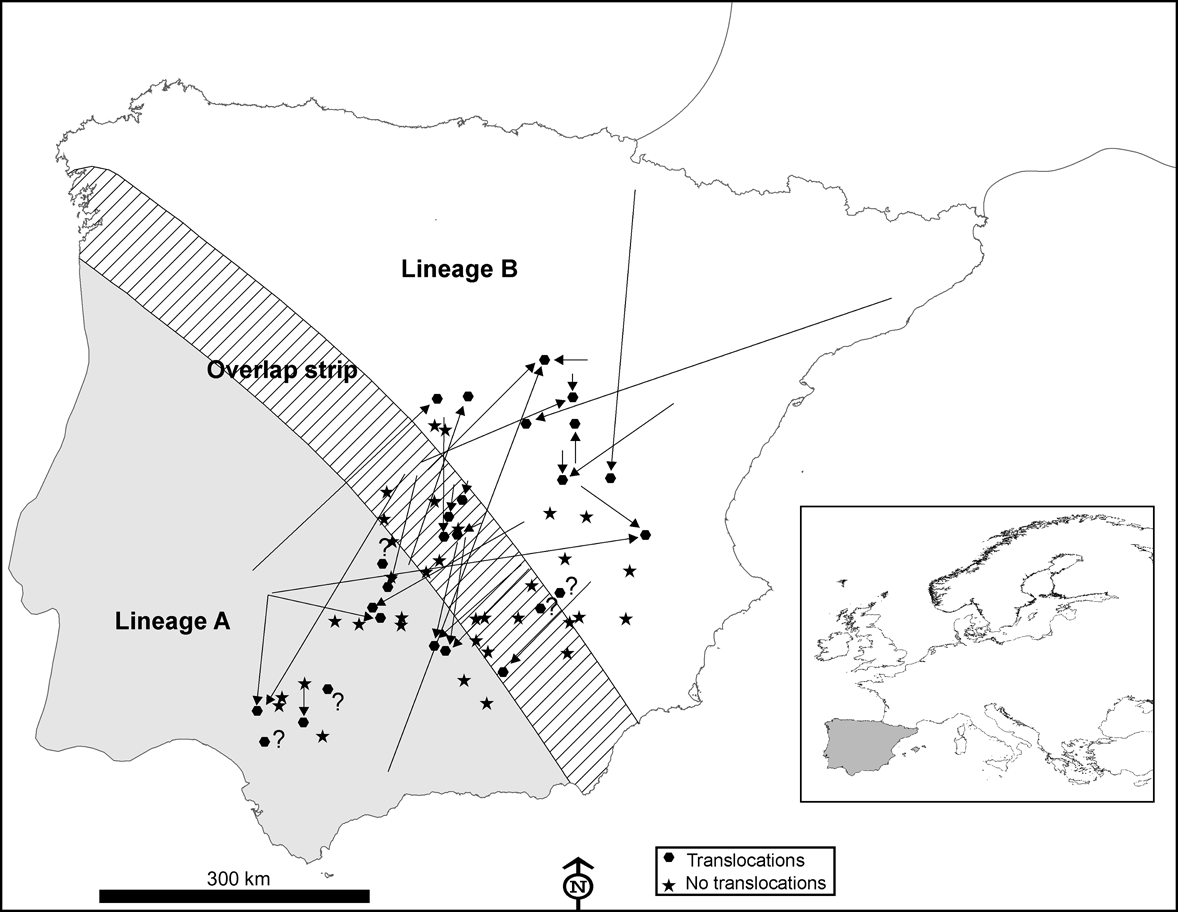
Wild rabbits were formerly abundant in Spain, supporting many predators and intensive sport hunting (Delibes-Mateos et al., Reference Delibes-Mateos, Redpath, Angulo, Ferreras and Villafuerte2007). However, rabbit populations have undergone a sharp decline since the 1950s (Virgós et al., Reference Virgós, Cabezas-Díaz and Lozano2007). As a consequence, many efforts, including translocations for restocking, are being made in Spain to increase local population numbers for both hunting and conservation purposes (Calvete et al., Reference Calvete, Villafuerte, Lucientes and Osácar1997).
Although European (Habitats Directive 92/43/CEE) and Spanish laws (4/1989 national law and various regional laws) have the intention of controlling the introduction of non-native lineages (Galvez, Reference Galvez2004), no tool exists in Spain to develop and implement these laws. For instance, to capture and release rabbits in Spain a governmental permit and a rabbit transit permit (granted by an authorized veterinary official) are required. Both permits are granted after some considerations regarding diseases, whereas no questions are asked on the origin of released rabbits or their genetic lineage.
Our aim here is to analyse the frequency and importance of rabbit translocations in central-southern Spain and their potential risk for the conservation of the two rabbit lineages. Our broad objectives are: (1) to evaluate whether translocations are currently increasing, as well as to examine the frequency of these practices in central-southern Spain, (2) to analyse the goals of restocking (for conservation or hunting priorities), (3) to estimate the numbers of wild rabbits involved in translocations, (4) to gain the opinions of hunters and conservationists about the effectiveness of translocations, (5) to evaluate the legality of this management practice, (6) to determine the origin of the individual rabbits released in each locality to estimate the risk of mixture of lineages, and (7) to estimate directly, by analysis of mtDNA, the current level of mixture of both lineages. Finally, we provide recommendations, for instances where translocations have to be employed, to avoid current or future problems in rabbit conservation.
Methods
Restocking information
We performed 60 interviews with local hunting and/or conservation managers to gather information concerning game management activities in central-southern Spain during 1993, and again during 2002 (Fig. 1). Interviews were carried out in 60 locations that had previously been used in a more comprehensive survey performed in 1993 when over 300 localities were surveyed throughout the entire country (Villafuerte et al., Reference Villafuerte, Viñuela and Blanco1998). However, because of logistic constraints, on the second occasion we repeated only the 60 surveys in central-southern Spain (Fig. 1). Approximately 90% of the interviews were carried out in face-to-face meetings and the rest were conducted by fax, post or e-mail. Interviews were performed in the same locations in 1993 and 2002, and in many cases the people interviewed were the same on both occasions. Landowners have changed in only 6.6% of the 60 locations (Delibes-Mateos, unplubl. data).
In the 1993 interviews we only asked questions about translocations during the previous 10 years. In 2002, however, we included new questions concerning management practices. To evaluate whether translocations were increasing we asked whether they had been employed during the decade prior to the survey. To estimate the frequency of translocations we rated the intensity of this management activity from 1 to 4: 1, never; 2, rare (once in the last decade); 3, sporadic; 4, frequent (almost annually). We also asked whether rabbits were translocated for conservation or hunting purposes, what the total number of rabbits released per year at each locality was, and whether the translocations were successful. To evaluate the legality of this practice we asked if the required permit had been applied for or not. Finally, we asked about the origin of the released rabbits to estimate the potential risk of mixing lineages.
Collection of rabbit samples
During the 2003-2004 and 2004-2005 hunting seasons we sampled a total of 750 hunted rabbits from 43 different locations across the Iberian Peninsula (Fig. 2). Where possible, a portion of ear from at least 20 shot rabbits was collected at each locality. Samples were preserved in 100% ethanol; mtDNA variation was analysed using restriction fragment length polymorphisms (RFLPs) to distinguish between lineages A and B (Branco et al., Reference Branco, Ferrand and >Monnerot2000). In this method, digestion with a restriction enzyme (AluI) of a 1120-bp fragment comprising most of the cytochrome b gene resolved 8 different genetic profiles (Branco et al., Reference Branco, Ferrand and >Monnerot2000). Profiles A, B and C correspond to lineage B, whereas the other five profiles correspond to lineage A (Branco et al., Reference Branco, Ferrand and >Monnerot2000).
Number of hunted rabbits sampled (of a total of 750) for mtDNA variation in 43 locations across the Iberian Peninsula in the 2003-2004 and 2004-2005 hunting seasons. The size of the symbols is proportional to the number of rabbits sampled in each locality (small, 4-11; medium, 10-20; large, >20). Numbers correspond to the ID of each locality in Table 1. Inset indicates location of main figure in Europe.
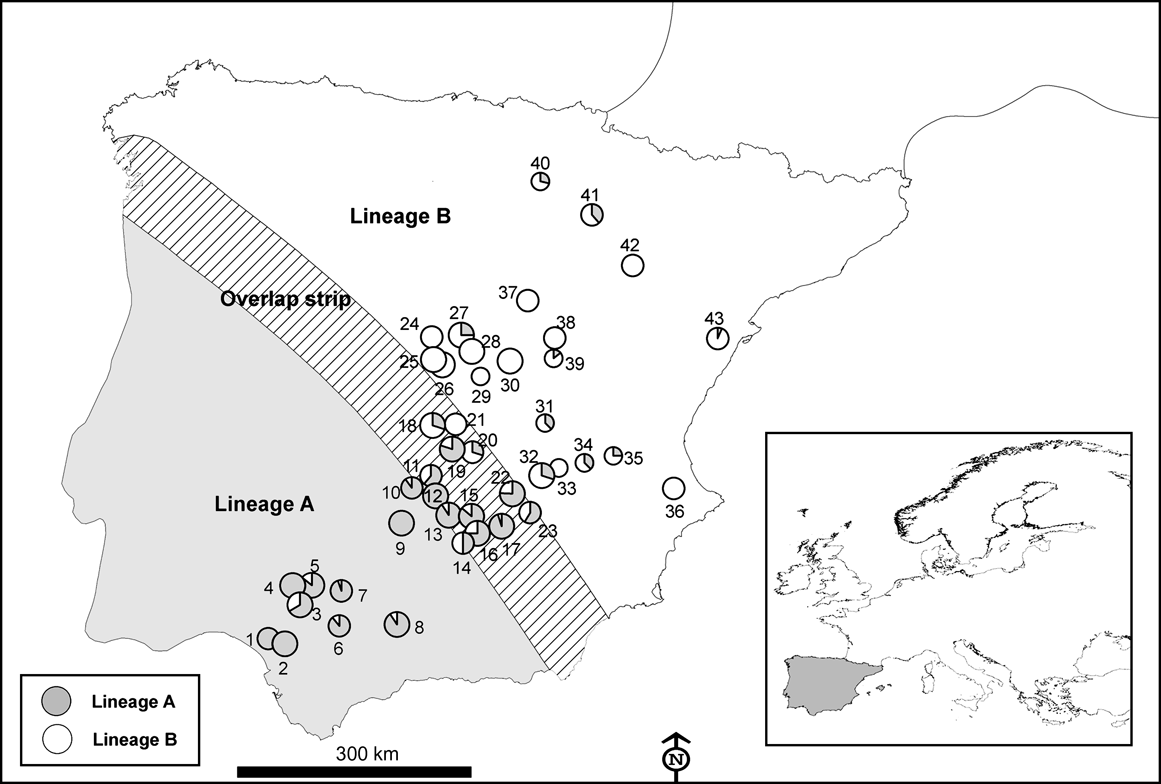
Results
According to the interviews carried out in 1993, rabbits were translocated at least once during the prior decade in 25% (n = 15) of the 60 localities studied; this increased to 43% during the decade before the 2002 interviews (n = 26; Fig. 1). In 51% of the localities surveyed in the 2 decades from 1983 to 2002 rabbits were translocated at least once (n = 31). Results from the 2002 interviews also indicated that: (1) translocations occurred almost annually in six (23%) localities, sporadically in 14 (54%) localities and rarely (once in the last decade) in six (23%) localities; (2) rabbits were released with the purpose of preserving threatened predators in only two (8%) of the 26 localities where translocations were carried out, the majority being for hunting purposes (24 cases, 92%); (3) a mean of 320 ± SE 65.22 rabbits were released in each translocation (range = 30-1,000); (4) only nine (35%) of the 26 managers who had liberated rabbits at least once in the decade prior to 2002 regarded this practice as a successful game management strategy; (5) nearly half of those making translocations (12 of 26, 46%) admitted that they did not apply for the required permit to capture alive, transport and release wild rabbits.
In 10 of 26 (38%) instances of translocations a mixing of different rabbit lineages may have occurred. Individuals from the naturally occurring area of lineage A had been released in areas populated by rabbits of lineage B, or vice versa (Fig. 1). We recorded one extreme case of a translocation of rabbits captured from a donor population 670 km away (Fig. 1).
Using mtDNA analysis 401 (53.5%) of the 750 sampled rabbits were found to correspond to lineage A and 349 (46.5%) to lineage B (Table 1). As we expected from the results of the interviews, some rabbits corresponding to lineage B appeared within the natural distribution of lineage A, and vice versa (Table 1; Fig. 2). For example, we found individuals belonging to lineage B in five of nine populations sampled in the lineage A distribution area described by Branco et al. (Reference Branco, Ferrand and >Monnerot2000, Reference Branco, Monnerot, Ferrand and Templeton2002). Similarly, rabbits belonging to lineage A were found in nine of 20 populations studied in the lineage B distribution area. Rabbits belonging to lineages A and B were found in the 13 populations studied in the area of overlap between the two lineages (a 100 km wide strip, as delimited by Branco et al., Reference Branco, Monnerot, Ferrand and Templeton2002).
Percentage of sampled rabbits corresponding to each lineage in the survey locations.
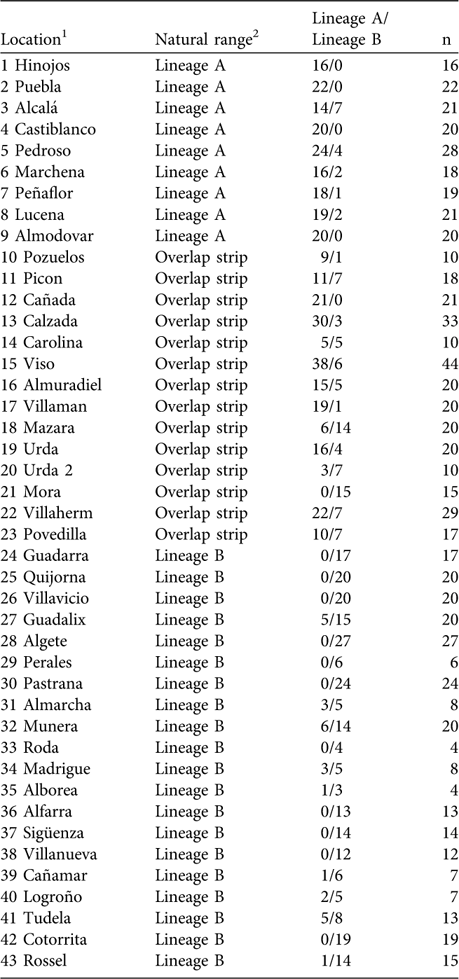
1 Corresponds to numbers plotted in Fig. 2
2 According to the distribution of rabbit lineages described in Branco et al. (Reference Branco, Monnerot, Ferrand and Templeton2002); see also Figs 1 & 2
Discussion
Although it was already known that rabbit translocation is a common management activity in Spain (Calvete et al., Reference Calvete, Villafuerte, Lucientes and Osácar1997) no estimation of the scale or consequences of the translocations was available. Our results indicate that rabbit restocking occurred at least once in over half of the surveyed localities in the 2 decades from 1983 to 2002. Furthermore, even though 65% of gamekeepers consider the translocations to be unsuccessful the use of this management practice increased from 1983 to 2002. Large numbers of rabbits are involved in each translocation. The increased frequency of translocations may be related to the need to increase rabbit numbers following the spread of viral haemorrhagic disease across Spain from 1988 onwards (Calvete et al., Reference Calvete, Villafuerte, Lucientes and Osácar1997).
Our results show that during 1993-2002 the main purpose of rabbit releases in central-southern Spain was the restocking of hunting reserves (92%), with only a small number of translocations carried out to supply prey for the conservation of threatened predators. This is in concordance with the observations of Griffith et al. (Reference Griffith, Scott, Carpenter and Reed1989) from Australia, Canada, New Zealand and the USA: 90% of introduction and translocation programmes of birds and mammals were made to enhance game populations, with only 10% made primarily for conservation of threatened species. In addition, our findings also show that most hunting and conservation managers do not persist with rabbit translocations over time. Thus, rabbit translocations occur sporadically or rarely (e.g. just once in the last decade) in most of the surveyed localities. This finding is not surprising, as rabbit translocations are not usually successful (Calvete et al., Reference Calvete, Villafuerte, Lucientes and Osácar1997), and involve a large economic investment.
Almost half of those managers who carried out rabbit translocations in Spain did not apply for the required permit and, when permission was given, usually no consideration was given to the origin of the animals concerned and the risk of mixing different evolutionary units. Consequently, rabbits belonging to one lineage seem to have been freely introduced within the distribution area of the other lineage (Fig. 1) and we found that the area where both lineages now occur is wider than that previously observed (Fig. 2; Branco et al., Reference Branco, Ferrand and >Monnerot2000, Reference Branco, Monnerot, Ferrand and Templeton2002). It is possible that the zone in which both lineages coexist has broadened naturally but the occurrence of lineage A rabbits in several localities within the distribution area of the other lineage, and vice versa, is almost certainly a result of translocations.
The mixture of different evolutionary significant units (sensu Fraser & Bernatchez, Reference Fraser and Bernatchez2001) has been frequently discouraged (Morales et al., Reference Morales, Andau, Supriatna, Zainuddin and Melnick1997) and, wherever possible, should be avoided. In the case of rabbits, unnatural contact between lineages A and B may have serious consequences for both conservation and hunting. Recent studies have demonstrated the existence of hybrids within the area where the lineages naturally overlap (Geraldes et al., Reference Geraldes, Ferrand and Nachman2006). Therefore, hybridization could also be expected in areas where rabbits of both lineages are mixed as a consequence of translocations. The effects of such hybridization on rabbit populations remain unknown but in the long-term could result in the loss of approximately 2 million years of genetic differentiation and underlying adaptation. In this sense, the loss of genetic variability between populations is contrary to the aim of maintaining evolutionary processes (Moritz, Reference Moritz1999). We can only speculate about the possible effects on rabbit populations of the translocations of individuals without consideration of their evolutionary history. For example, releasing rabbits of one lineage outside their range could result in introducing strains of myxoma virus or rabbit haemorrhagic disease virus that are non-virulent to them but virulent for the individuals of the native lineage. This would be particularly so in areas where rabbits have adapted to locally prevalent strains of pathogens (Parer et al., Reference Parer, Sobey, Conolly and Morton1994), aggravating the effects of rabbit diseases. Also, as the two rabbit lineages occur in regions of contrasting habitats, fitness of resident populations could be reduced because of introgression of poorly adapted gene complexes (Storfer, Reference Storfer1999).
From the point of view of biodiversity conservation rabbit translocations should be avoided wherever possible (IUCN, 1987). Nevertheless, in some instances translocations are required, and recommendations can be made on the basis of our findings. Firstly, regional authorities in Spain should inform hunting associations of the existence of both rabbit lineages and the importance of avoiding mixing through translocation. Secondly, only rabbits with the same genetic lineage as that of the receiving population should be released. If DNA analysis is not possible, the donor locality should be selected based on the proximity to the receiving population, and certainly within the known geographical limits (Fig. 2). The regional authorities could recommend potential donor populations for each region according to known rabbit abundance and genetic characteristics. Finally, the appropriate authorities should better regulate several aspects of rabbit translocations to ensure that rabbits of one lineage are not released within the distribution area of the other (excluding the narrow zone of overlap in lineage distribution). In addition, using existing regulations, the appropriate authorities should be more cautious when either hunters or conservationists apply for permits to translocate rabbits and, wherever possible, reduce the widespread practice of non-authorized translocations.
Acknowledgements
MDM was supported by an I3P grant (CSIC). Funding was provided by the projects REN-2001-0448/GLO and CGL-2005-02340/BOS. We are indebted to Drs M. Delibes, B. Cooke and J.C. Avise and two anonymous referees for helpful comments on previous drafts of the manuscript. We thank M. Branco, A. Centeno, J. Castillo and R. Estrada for their help with the genetic analyses. Special thanks are due to J. Blanco-Aguiar, G. Calabuig, J. Castillo, P. Castro, A. Delibes, A. Finque, A. Linares, S. Luna, L. Minguez, A. Pérez, M. Reglero, O. Rodríguez and C. Rouco for their field assistance.
Biographical sketches
Miguel Delibes-Mateos's interests include the study of the conflicts between nature conservation and management of small game species in the Iberian Peninsula. Esther Ramírez has diverse interests in population genetics, especially of small game species in the Iberian Peninsula. Pablo Ferreras's research focuses on the ecology of carnivores and predator-prey relationships in game management and conservation. Rafael Villafuerte works on ecological effects of management practices carried out to enhance prey populations for hunting and conservationist purposes.




