- AD
-
Alzheimer's disease
- ALA
-
α-linolenic acid
ApoE
ApoE is a prominent constituent of plasma and brain lipoproteins mediating cellular cholesterol uptake by interaction with cell surface receptors including LDL-receptor, LDL-receptor-related proteins and VLDL-receptor( Reference Innerarity and Mahley 1 , Reference Strickland, Gonias and Argraves 2 ). ApoE also binds to cell surface located glycosaminoglycans such as heparin sulphate proteoglycans to facilitate lipoprotein uptake( Reference Mahley and Ji 3 ). In addition to regulation of extra-hepatic cholesterol metabolism apoE is centrally involved in chylomicron clearance through uptake of remnants by the liver( Reference Hui, Innerarity and Mahley 4 ). The apoE protein is produced in various tissues with particular high concentrations in liver, brain, kidney, lymphocytes and adipose tissue. Beyond its known function in lipid and cholesterol metabolism apoE is believed to modulate many aspects of ageing in brain and artery walls( Reference Finch 5 ).
ApoE allelic variation
The human apoE gene is polymorphic with two major SNP (rs429358C>T, rs7412C>T) in the coding region of exon 4. The two nucleotide exchanges are revealed at the protein level as amino acid substitution (Arg→Cys) at positions 112 and 158 of the mature apoE protein( Reference Weisgraber, Rall and Mahley 6 ). There are three major protein isoforms (E2, E3 and E4) arising from the three possible genetic variants ε2, ε3 and ε4. Although other mammals express apoE, allelic variation was only found in human subjects. Sequence analysis revealed that primate apoE is identical to human apoE ε4 at the sites coding for Arg at positions 112 and 158( Reference Hanlon and Rubinsztein 7 ). Therefore, apoE ε4 is considered as the ancestral human allele that, after the human and primate lineages split, was modified by single successive mutations breeding the ε3 and ε2 alleles( Reference Fullerton, Clark and Weiss 8 ). As a result of combination of the allelic variants three homozygous (ε2/ε2, ε3/ε3 and ε4/ε4) and three heterozygous (ε2/ε3, ε2/ε4, ε3/ε4) genotypes emerge with varying frequency throughout human populations. ApoE ε4 is always the minor allele when compared with apoE ε3, whereas apoE ε2 is least common and even absent in particular aborigine populations( Reference Corbo and Scacchi 9 ).
Geographic distribution of apoE genotypes
Distribution of the three major apoE alleles varies worldwide (Table 1); however, the ε3 variant is most abundant in all human populations and ranges between 0·968 in Indians and 0·356 in Papuans( Reference Singh, Singh and Mastana 10 ). Highest apoE ε4 frequencies are found in Central Africa (including Pygmies (0·407)( Reference Corbo and Scacchi 9 ) and Tutsi (0·385)( Reference Singh, Singh and Mastana 10 )); Oceania (including Papuans (0·368) and Australian Aborigines (0·260)) and in Saami people (0·310)( Reference Corbo and Scacchi 9 ). Particularly low apoE ε4 frequencies are found in Mediterranean and several Asian populations (<0·10). The ε2 allele is rare or absent in Inuits, South Americans, Siberians and Mongolians, but relatively frequent in sub-Saharans, Malaysian and Papuans( Reference Singh, Singh and Mastana 10 ). At the continental level, allele frequencies of apoE ε3 and ε4 are inversely correlated in Europe, Africa and North America. In Asian and Oceanian populations, both ε2 and ε4 frequencies rise, when ε3 is less abundant. Of particular importance is the non-random north-to-south gradient of ε4 and ε3 alleles in Europe as shown in Fig. 1. The frequency of apoE ε4 increases with increasing latitude, whereas the ε3 allele frequency is negatively correlated with latitude. The occurrence of apoE ε2 is independent of the European latitude. The significant pattern in latitudinal apoE ε4 and ε3 allele distribution is also found in North but not in South America and Asia( Reference Singh, Singh and Mastana 10 ). Although in China existence of a south-to-north gradient in ε4 frequency was reported( Reference Hu, Qin and Jing 11 ). A more recent study additionally modelled a curvilinear relationship where worldwide ε4 allele frequencies first decrease with distance from the equator and then increase again at absolute latitudes higher than 35°. Importantly the population variation in apoE ε4 frequency was suggested to be shaped by natural selection and not due to underlying population structure( Reference Eisenberg, Kuzawa and Hayes 12 ).
Correlation of (a) apoE ε4, (b) apoE ε3 and (c) apoE ε2 allele frequencies in Europe with respective latitudes. Data on allele frequency and latitude were adapted from Singh et al. ( Reference Singh, Singh and Mastana 10 ) and Rodrigues et al. ( Reference Rodrigues, Fonseca and Matias 13 ). Linear regression of apoE allele frequency and latitude was calculated applying Pearson's correlation analysis and is given as regression coefficient R with corresponding P-values.

Allelic variation of apoE at the transcript and protein level and ranges of worldwide allele frequencies
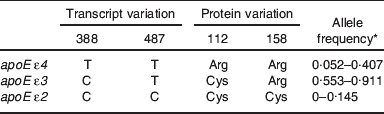
* Data from Corbo and Scacchi( Reference Corbo and Scacchi 9 ).
Generally apoE ε4 is more present in people either with dark skin pigmentation or living in regions with low insolation, while lower presence is found in people with moderate melanin pigmentation but exposed to relatively high solar irradiation. Therefore, the capability of better enduring low UV concentrations may be an advantage of the ancestral apoE ε4 compared with the new ε3 genotype. The evolution of apoE ε3 about 200 000 years ago( Reference Fullerton, Clark and Weiss 8 ) was accompanied by the establishment of more agricultural communities, as distinguished from simple hunters and gatherers, and subsequent emigration of the modern Homo sapiens from Africa( Reference Nei and Roychoudhury 14 ). However, it is uncertain precisely when and why apoE ε3 began to expand in frequency and supersede the ancestral ε4. Furthermore, emergence of the recent ε2 allele cannot be dated exactly yet. Due to the fact that apoE ε2 is absent in people coming from north Asia settled in Arctic regions and America 40 000–10 000 years ago, ε2 may likely first have emerged subsequent to this event( Reference Fullerton, Clark and Weiss 8 ).
ApoE protein isoforms
The mature apoE protein (34 kDa, 299 amino acids) comprises two structural helical domains, a bigger amino (N)-terminal (1–191) and a carboxyl (C)-terminal (216–299) region that are connected by a non-helical hinge region. The region responsible for receptor binding is determined in the N-terminal domain known to be rich in basic amino acids, whereas the region spanning residues 261–272 of the C-terminus determines lipoprotein and lipid-binding properties of apoE( Reference Dong and Weisgraber 15 ). In the apoE4 isoform, the positive charge of Arg112 facilitates a domain interaction within the protein determined by a salt bridge formation between residues Arg61 and Glu255. The Arg61–Glu255 salt bridge is not present in apoE3 and apoE2 as Arg is substituted by Cys at position 112 (Fig. 2). Due to the domain interaction (Arg61–Glu255), the C-terminal domain is organised differentially in apoE4 compared with apoE3 and apoE2 and therefore, lipoprotein-binding affinity is also altered( Reference Nguyen, Dhanasekaran and Nickel 16 ). ApoE4 prefers binding VLDL and intermediate density lipoproteins, while apoE3 and apoE2 display a preference for cholesterol-rich HDL particles( Reference Dong, Wilson and Wardell 17 ). In the apoE2 isoform, the mutation at position 158 (Arg→Cys) causes a salt bridge formation revealing conformational changes that affect its LDL-receptor-binding domain( Reference Wilson, Mau and Weisgraber 18 ). Interestingly, although primate apoE holds an Arg residue at position 112 (similar to apoE4), there is a Thr at position 61 (instead of Arg in human apoE) preventing the interaction with the C-terminal domain. Therefore, primate apoE is in terms of function more related to human apoE3 than apoE4.
Schematic protein structures of apoE4, apoE3 and apoE2 (adapted from Ye et al. ( Reference Ye, Huang and Mullendorff 19 )) showing amino acid residues that distinguish between the isoforms. Arg112 facilitates bridge formation (Arg61–Glu255) leading to domain interaction in the apoE4 isoform. Mutation at position 158 in apoE2 (Arg→Cys) changes domain charge from positive to negative (oval marking) and interferes with receptor binding. Additional oval marking of the C-terminal region that is responsible for apoE protein structure, self-association and ability to bind lipids and lipoprotein particles differentially organised in apoE4 and apoE3.

ApoE genotype and disease risk
Beyond genotype-dependent effects on blood lipids, which will be reviewed in the following section, the apoE polymorphism is associated with age-related chronic as well as infectious diseases. Risk of CVD is dramatically increased in apoE ε4 carriers with 40% increased incidence as compared with the ε3 genotype( Reference Song, Stampfer and Liu 20 ). This has been attributed to modestly elevated LDL-cholesterol in the ε4 genotype, although mechanisms underlying apoE ε4-CVD-risk associations may be more complex. Several lines of evidence suggest that apoE ε4 potentiates adverse effects of CVD-related risk factors such as smoking and physical inactivity( Reference Gustavsson, Mehlig and Leander 21 , Reference Humphries, Talmud and Hawe 22 ). In Alzheimer's disease (AD) association of apoE ε4 and disease prevalence is even more striking. Presence and number of apoE ε4 alleles increased AD risk (OR of 3·2 (ε4/ε3) and 14·9 (ε4/ε4) relative to ε3/ε3)( Reference Farrer, Cupples and Haines 23 ) with each additional ε4 allele shifting disease onset to younger age( Reference Corder, Saunders and Strittmatter 24 ). Poor neuronal repair, increased amyloid plaque burden and higher susceptibility towards oxidative insults have been suggested to underlie the positive association of apoE ε4 and AD development( Reference Arendt, Schindler and Bruckner 25 – Reference Huebbe, Jofre-Monseny and Boesch-Saadatmandi 27 ). In contrast apoE ε2 appears to be protective compared with ε3 both in CVD and AD( Reference Gustavsson, Mehlig and Leander 21 , Reference Farrer, Cupples and Haines 23 ).
There is increasing body of evidence that apoE may modulate susceptibility to viral infections in an isoform-dependent manner (extensively reviewed in Kuhlmann et al. ( Reference Kuhlmann, Minihane and Huebbe 28 )). ApoE4 increases fusion rate and cell entry of the HIV resulting in faster disease progression relative to apoE3, though the risk of acquiring HIV infection is independent of the apoE isoform( Reference Burt, Agan and Marconi 29 ). Risk of herpes labialis and development of herpes simplex-associated AD is potentiated in apoE ε4 carriers( Reference Itzhaki, Lin and Shang 30 , Reference Itzhaki and Wozniak 31 ). In contrast apoE4 protects against hepatitis C-induced liver damage and increases virus clearance attenuating chronic infection risk compared with apoE3( Reference Price, Bassendine and Norris 32 , Reference Wozniak, Itzhaki and Faragher 33 ). Although data are scarce, it was suggested that apoE4 may also reduce heavy burden of early childhood diarrhoea and improve disease outcome in children in the first 2 years( Reference Oria, Patrick and Oria 34 ).
Overall the apoE ε4 genotype is associated with increased morbidity and mortality in the elderly and the allele frequency is significantly declining from 85 years of age( Reference Corder, Lannfelt and Viitanen 35 , Reference Schachter, Faure-Delanef and Guenot 36 ). The influence of the ε4 allele on mortality is even increasing in advanced age (92–103 years)( Reference Jacobsen, Martinussen and Christiansen 37 ). Adverse effects of apoE4 may be attributed to altered lipid metabolism, but may also be mediated by differences in biomarkers of oxidative stress, inflammation and nuclear factor (erythroid-derived 2)-like 2-signalling.
Metabolic and molecular mechanisms of apoE isoforms
Lipid metabolism
Prospective cohort studies and human intervention studies have shown that the apoE polymorphism has a substantial effect on plasma lipids and lipoproteins (Table 2). Specifically, the apoE phenotypes have been associated with the variability of plasma total cholesterol concentrations and contribute to 4–12% of the variability of LDL-cholesterol concentrations in several populations( Reference Kolovou and Anagnostopoulou 38 ). In addition, a recent comprehensive meta-analysis demonstrated approximately linear relationships of apoE genotypes (when ordered ε2/ε2, ε2/ε3, ε2/ε4, ε3/ε3, ε3/ε4 and ε4/ε4) with LDL-cholesterol concentrations and with CHD risk( Reference Bennet, Di Angelantonio and Ye 39 ). The LDL-cholesterol concentrations were approximately 30% lower in people with ε2/ε2 than with ε4/ε4 genotypes, a difference comparable with that produced by ‘statin’ therapy.
The impact of apoE isoform on serum/plasma concentrations of lipids and lipoproteins in human subjects

Abbreviations: EPIC, European Prospective Investigation into Cancer and Nutrition; HDL-C, HDL-cholesterol; HRT, hormonal replacement therapy; HRT+, postmenopausal women HRT users; HRT−, postmenopausal women HRT non-users; LDL-C, LDL-cholesterol.
The impact of the different apoE isoforms on blood concentrations of lipids and lipoproteins has been explained by several mechanisms including (i) receptor-binding affinities of the different apoE-containing lipoproteins, (ii) dietary fat clearance, (iii) differences in the clearance of LDL apoB, and (iv) differences in the efficiency of intestinal cholesterol absorption (for review, see( Reference Kolovou and Anagnostopoulou 38 , Reference Hauser, Narayanaswami and Ryan 50 )).
In addition to the lower blood concentration of LDL-cholesterol (discussed earlier), the ε2 allele is associated with lower blood concentrations of apoB and increased concentrations of TAG and apoE when compared with the ε3 allele( Reference Hagberg, Wilund and Ferrell 51 ). Similarly, apoE ε2/ε2 and ε2/ε3 are associated with lower concentrations of LDL-cholesterol when compared with ε3/ε3 ( Reference Assmann, Schmitz and Menzel 52 ). The increased concentrations of TAG and apoE are consistent with an impaired clearance of remnant particles( Reference Havel, Chao and Windler 53 ). The metabolic explanation for the reduced LDL-cholesterol concentrations is less clear. Individuals with the ε2/ε2 genotype can develop a type III hyperlipoproteinemia. This is characterised by an accumulation of remnants of TAG-rich lipoprotein particles in plasma. It has been associated with several genetic abnormalities affecting lipoprotein metabolism including hepatic lipase deficiency and defects in the lipoprotein remnant receptor( Reference Assmann, Schmitz and Menzel 52 , Reference Davignon, Gregg and Sing 54 ). However, it should be noted that although the apoE ε2/ε2 genotype is present in about 1% of the general population, less than 5% of individuals with ε2/ε2 develop a type III hyperlipoproteinemia. Several secondary factors may thus promote type III hyperlipoproteinemia in individuals with ε2/ε2 genotype such as a hormonal disturbance (e.g. hypothyroidism, oestrogen withdrawal and pregnancy), environmental factors (e.g. positive energy balance leading to obesity) or changes associated with increasing age( Reference Assmann, Schmitz and Menzel 52 , Reference Davignon, Gregg and Sing 54 ).
Higher LDL-cholesterol, low TAG and apoE concentrations typically occur in individuals with ε4/ε3 and ε4/ε4 genotypes compared with ε3/ε3 individuals( Reference Davignon, Gregg and Sing 54 ). The low TAG is consistent with the fact that carriers of the ε4 allele clear circulating chylomicron remnants into the liver more rapidly than ε3/ε3 individuals and twice as fast as ε3/ε2 individuals( Reference Weintraub, Eisenberg and Breslow 55 ). ApoE is not a constituent of LDL particles, but it seems to have an indirect influence on LDL-cholesterol concentrations. In the fasting state, most plasma apoE resides in HDL particles. After intake of dietary fat, apoE shifts from HDL to postprandial particles. In ε4/ε3 individuals, VLDL and HDL are enriched in apoE protein( Reference Steinmetz, Jakobs and Motzny 56 ). ApoE4 preferentially associated with VLDL is removed from the circulation more rapidly than apoE3( Reference Gregg, Zech and Schaefer 57 ). Accordingly, it is supposed that individuals with the ε4 allele may more efficiently and rapidly deliver dietary fat to the liver. Faster hepatic clearance of dietary fat in apoE ε4/ε3 subjects could cause the down-regulation of LDL-receptors and an increase in plasma LDL-cholesterol( Reference Weintraub, Eisenberg and Breslow 55 ).
Oxidative stress, antioxidant defence and chronic inflammation
The first evidence that apoE may protect against oxidative stress was found in apoE-deficient mice with increased susceptibility of plasma lipoproteins to in vitro oxidation compared with wild-type mice( Reference Hayek, Oiknine and Brook 58 ). Miyata and Smith( Reference Miyata and Smith 59 ) then postulated that antioxidative activity of apoE would be isoform dependent and that E4 was least and E2 most effective. The authors suggested different metal-binding capacities of the individual apoE isoforms that were possibly involved in the observed antioxidant effects. The presence of Arg112 rather than the absence of any cysteinyl groups in the protein appears to contribute to the increased oxidative susceptibility (due to altered protein stability) of apoE4 compared with apoE3 and its associated lipoproteins( Reference Kashiwagi, Nakamura and Arai 60 , Reference Su, Wen and Chou 61 ). Neuronal cells cultured in apoE4 conditioned medium were more susceptible towards oxidative stress-induced cytotoxicity than in apoE3 conditioned medium( Reference Huebbe, Jofre-Monseny and Boesch-Saadatmandi 27 ). Furthermore, innate immune cells produce higher levels of reactive oxygen or nitrogen species in the presence of apoE4 than apoE3( Reference Brown, Wright and Colton 62 – Reference Jofre-Monseny, Pascual-Teresa and Plonka 64 ). Biomarkers of oxidative stress are elevated in apoE ε4 carriers notably in subjects suffering from AD or CVD( Reference Dietrich, Hu and Block 44 ,65–Reference Talmud, Stephens and Hawe 68 ). Expression of anti-atherogenic paraoxonase 1, which inhibits and reverses LDL-oxidation, is also lower in apoE4 than apoE3-targeted replacement mice( Reference Boesch-Saadatmandi, Egert and Schrader 69 ). Recent evidence suggests that the apoE ε4 genotype is associated with lower expression of the antioxidant enzyme heme oxygenase 1 and other nuclear factor (erythroid-derived 2)-like 2 target genes( Reference Graeser, Boesch-Saadatmandi and Lippmann 70 ). Although data are sometimes conflicting nuclear factor (erythroid-derived 2)-like 2 may play a role in preventing atherosclerosis( Reference Levonen, Inkala and Heikura 71 ). In summary, modulation of oxidative stress and antioxidant defence mechanisms may be a relevant physiological function of apoE, which is implemented in an isoform-dependent manner.
A number of studies have been conducted investigating the role of apoE in inflammatory processes mostly in models of neurodegeneration. Indeed chronic inflammation is associated with neurodegenerative disorders such as AD( Reference Akiyama, Barger and Barnum 72 ). ApoE has been shown to modulate inflammatory response in either direction, pro- and anti-inflammatory( Reference Guo, LaDu and Van Eldik 73 ). However, expression of pro-inflammatory markers such as cytokines and NO in stimulated microglia and macrophages was higher in the presence of apoE4 than apoE3( Reference Jofre-Monseny, Loboda and Wagner 74 – Reference Vitek, Brown and Colton 76 ). Higher pro-inflammatory response in apoE4 may be mediated by increased and prolonged activation of the redox-sensitive transcription factor NF-κB( Reference Jofre-Monseny, Loboda and Wagner 74 , Reference Ophir, Amariglio and Jacob-Hirsch 75 ). Chronic inflammation in the brain coincides with amyloid plaque pathology and both are more pronounced in apoE4 than apoE3 transgenic mice( Reference Belinson and Michaelson 77 ). ApoE ε4 is significantly associated with higher serum amyloid P (acute phase protein) in mice( Reference Boesch-Saadatmandi, Niering and Minihane 78 ) suggesting an elevated level of chronic low grade inflammation which may contribute to the increased chronic disease risk of ε4 as compared with non-ε4 carriers (Table 3).
The impact of apoE isoform on biomarkers of inflammation in human subjects

AD, Alzheimer's disease; CRP, C-reactive protein; CSF, cerebrospinal fluid; ICAM-1, intercellular adhesion molecule 1; MCFS, macrophage colony-stimulating factor; MCP-1, monocyte chemoattractant protein; MMP: matrix metalloproteinase; pSS, primary Sjögren's syndrome; SAA, serum amyloid A; TIMP-1, tissue inhibitor of metalloproteinase-1; VCAM-1, vascular cell adhesion molecule-1.
* Trend (0·05<P<0·1).
Responsiveness of the apoE genotype to dietary factors
Vitamin E
Vitamin E comprises eight different tocopherols and tocotrienols, α-tocopherol being biologically the most important vitamer (herein after referred to as vitamin E). Dietary vitamin E is postprandially delivered to the plasma via chylomicrons released from enterocytes or via VLDL following hepatic secretion. Under basal conditions, vitamin E is mainly associated with LDL particles with a constant flux existing between the different lipoprotein classes. Since apoE polymorphism affects concentration and clearance of plasma lipoproteins, it is conceivable that vitamin E metabolism is also impacted by the apoE genotype. Although a few studies found no difference in plasma vitamin E levels between apoE genotypes under baseline conditions( Reference Fernandes, Proenca and Nogueira 100 , Reference Gomez-Coronado, Entrala and Alvarez 101 ), a biokinetic approach using stable isotopes observed higher newly absorbed vitamin E levels among ε4 as compared with ε3 carriers( Reference Lodge, Hall and Jeanes 102 ). Extra-hepatic vitamin E concentration is lower in apoE4- than apoE3-targeted replacement mice, which is most likely due to the lower expression of LDL-receptor and related receptor classes mediating vitamin E uptake( Reference Huebbe, Jofre-Monseny and Rimbach 103 ). Furthermore, degradation of vitamin E may be increased in the apoE ε4 genotype contributing to lower tissue retention and therefore possibly lower vitamin E status in peripheral tissues( Reference Huebbe, Lodge and Rimbach 104 ).
Vitamin D
Unlike vitamin E the impact of the apoE polymorphism on vitamin D status is more pronounced. We recently provided first experimental and epidemiological evidence suggesting the apoE ε4 genotype is associated with higher circulating vitamin D levels( Reference Huebbe, Nebel and Siegert 46 ). In targeted gene replacement, mice expressing human apoE4 serum 25-hydroxy-vitamin D concentration was significantly higher compared with apoE3- and apoE2-expressing mice. The observed higher serum concentration may be a result of increased intestinal absorption of dietary vitamin D as the mRNA level encoding for the key enzyme in bile acid production was higher in apoE4 than E3 and E2 mice. Elevated renal reabsorption of vitamin D from primary urine may also contribute to better vitamin D status as loss of vitamin D due to renal excretion would be reduced in the apoE ε4 genotype. Furthermore, a higher femoral Ca concentration was evident in apoE4 v. apoE3 mice accompanied by relatively higher dietary Ca absorption. Supportive of a better vitamin D and Ca status, apoE4 mice showed increased renal Ca excretion and lower mRNA levels of renal Ca absorption genes that would have been induced upon hypocalcaemia (Table 4). These data illustrate that apoE4 compared with apoE3 mice have a better vitamin D and Ca status while dietary supply of both nutrients was similar in the apoE genotype groups( Reference Huebbe, Nebel and Siegert 46 ). In addition, circulating vitamin D was assessed in two independent human samples from northern Germany. Serum concentration of 25-hydroxy-vitamin D was significantly higher in subjects carrying ε4 as compared with non-ε4 carriers. Mean 25-hydroxy-vitamin D concentration of both samples was <50 nmol/l suggesting a mild vitamin D deficiency throughout study participants. Mild vitamin D deficiency is relatively common in people inhabiting the same geographic latitude and has been reported before( Reference Zittermann 105 , Reference Hintzpeter, Mensink and Thierfelder 106 ). Our data suggest apoE ε4 as a modulator of vitamin D and Ca status in apoE transgenic mice and in a population with insufficient vitamin D supply, which in the light of evolutionary aspects may explain non-random geographic distribution of apoE alleles (see subsection ‘Geographic distribution of apoE genotypes’).
Effects of the apoE isoform on parameters of vitamin D and Ca status evident in apoE4 compared with apoE3 targeted gene replacement mice( Reference Huebbe, Nebel and Siegert 46 )
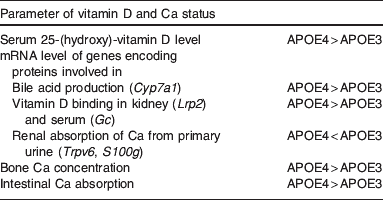
Cyp7a1, cholesterol-7-α-hydroxylase; Lrp2, LDL-receptor-related protein 2; Gc, vitamin D binding protein; Trpv6, Ca transport protein 1; S100g, calbindin D9K.
Flavonoids
Flavonoids are a large group of secondary plant metabolites with >6000 distinct flavonoids identified to date( Reference Erdman, Balentine and Arab 107 ). Epidemiological studies, together with data from animal models and some clinical trials, suggest a role of dietary flavonoids in the prevention of CVD and other age-related chronic diseases( Reference Graf, Milbury and Blumberg 108 – Reference Hooper, Kroon and Rimm 110 ). The flavonol quercetin exhibits a wide range of physiological effects such as inhibition of LDL-oxidation, lowering of arterial blood pressure and platelet aggregation, and improvement of endothelial function as shown in animal models and in human subjects( Reference Loke, Proudfoot and McKinley 111 – Reference Perez-Vizcaino, Duarte and Jimenez 117 ). Furthermore, cell culture and animal studies indicate a potent anti-inflammatory activity of quercetin( Reference Rivera, Moron and Sanchez 116 , Reference Rivera, Moron and Sanchez 118 – Reference Boesch-Saadatmandi, Loboda and Wagner 120 ).
Current scientific evidence from human and animal studies indicates that the apoE genotype may be an important determinant of the responsiveness to dietary quercetin. We have recently found that overweight and obese patients with metabolic syndrome traits carrying the apoE ε3 are highly responsive towards the blood pressure lowering effects of dietary quercetin supplementation, whereas apoE ε4 carriers, by large, do not benefit( Reference Egert, Boesch-Saadatmandi and Wolffram 45 ) (Table 5). We hypothesised that quercetin supplementation may have resulted in higher endothelial NO levels in apoE ε3 v. apoE ε4 carriers due to potential differences in the cellular redox and inflammatory states between the two genotypes. In addition, we found apoE genotype-specific effects of quercetin on fasting serum concentrations of HDL-cholesterol and apoA1 and on the ratio of LDL:HDL-cholesterol. Quercetin significantly decreased serum HDL-cholesterol and ApoA1 in apoE ε4 allele carriers but not in homozygous ε3/ε3. Moreover, our recent findings in apoE3- and apoE4-targeted gene replacement mice indicated that apoE3 animals were more responsive to the TNFα-lowering properties of dietary quercetin supplementation compared with apoE4 animals( Reference Boesch-Saadatmandi, Wolffram and Minihane 119 ). Therefore, the apoE genotype may in part explain the large heterogeneity of studies regarding potential health effects of flavonoids in human subjects where cohorts are not genotyped for apoE polymorphisms.
ApoE isoform and responsiveness to flavonoid manipulation in human subjects – evidence from randomised controlled intervention studies

Chol, cholesterol; CRP, C-reactive protein; DBP, diastolic blood pressure; HDL-C, HDL-cholesterol; LDL-C, LDL-cholesterol; ox-LDL, oxidized LDL; PAI-1, plasminogen activator inhibitor type 1; SBP, systolic blood pressure; sE-selectin, soluble endothelial-selectin; sICAM, soluble intracellular adhesion molecule; sVCAM, soluble vascular cell adhesion molecule; urinary 8-iso-PGF2α, 8-epimer of PG F2α.
Plant and marine n-3 fatty acids
A large body of epidemiological data and evidence from randomised controlled human trials has demonstrated the cardioprotective effects of the marine n-3 fatty acids EPA and DHA( Reference He 123 – Reference Saravanan, Davidson and Schmidt 125 ). For example, EPA and DHA have been shown to improve dyslipidaemia, to lower blood pressure and heart rate, to reduce inflammation, and to improve vascular function( Reference Saravanan, Davidson and Schmidt 125 – Reference Egert and Stehle 127 ). An alternative (n-3) PUFA is plant-derived α-linolenic acid (ALA), which in stable-isotope studies in human subjects was shown to be desaturated and elongated to long-chain (n-3) PUFA( Reference Burdge and Calder 128 ). ALA may also protect against CHD. However, the data concerning the protective role of ALA are less definitive than that for the long-chain n-3 PUFA, EPA and DHA( Reference Geleijnse, de Goede and Brouwer 129 ).
Recent evidence suggests that the apoE genotype may predict the lipid and lipoprotein response to n-3 fatty acid interventions (Table 6). Minihane et al.( Reference Minihane, Khan and Leigh-Firbank 47 ) examined the effect of apoE polymorphism and fish oil supplementation on volunteers with an atherogenic lipoprotein phenotype, which is characterised by moderate hypertriacylglycerolaemia, low concentrations of HDL-cholesterol and a predominance of small dense LDL3 particles. Fish oil (3 g/d EPA and DHA) was found to lower fasting and postprandial TAG responses, with a tendency towards greater responsiveness in individuals with the apoE ε2 allele. In the group as a whole there was a non-significant 7% rise in LDL-cholesterol following fish oil supplementation. However, in the subgroups based on apoE genotype, the greatest responsiveness was observed in the apoE ε4 carriers, with a more atherogenic shift in the plasma lipid profile, including a 7·4% (non-significant) decrease in HDL-cholesterol and 3·5% increase in total cholesterol, with a 16% increase in LDL-cholesterol. On the other hand, there was also a 26% reduction in the percentage of small dense LDL in this subgroup. This study demonstrated for the first time that the apoE genotype may in part determine the blood lipid response to fish oil intervention, and that the LDL-cholesterol increases may be largely evident in apoE ε4 carriers.
ApoE isoform and responsiveness to n-3 fatty acid manipulation in human subjects – evidence from randomised controlled intervention studies

ALA, α-linolenic acid; Chol, total cholesterol; CRP, C-reactive protein; HDL-C, HDL-cholesterol; LDL-C, LDL-cholesterol; LPL, lipoprotein lipase; MCSF, macrophage colony stimulating factor; ox-LDL, oxidized LDL; SAA, serum amyloid A; VLDL-C, VLDL-cholesterol.
In a subsequent trial using a prospectively genotyped cohort of metabolically healthy participants, Caslake et al. ( Reference Caslake, Miles and Kofler 130 ) systematically investigated the effect of apoE polymorphism, sex and age on lipid responses to modest fish oil supplementation (0·7 or 1·8 g/d EPA and DHA). In contrast with the previously described data( Reference Minihane, Khan and Leigh-Firbank 47 ), there was no significant effect of apoE genotype on LDL-cholesterol. It was speculated that the effect of apoE genotype on LDL-cholesterol response may be dose dependent( Reference Caslake, Miles and Kofler 130 ). In addition, there was no significant effect of apoE genotype on the responsiveness to TAG lowering by EPA and DHA. However, there was a trend towards greater responsiveness in carriers of the ε4 allele: a significant sex×genotype×treatment interaction was seen, and 15 and 23% reductions in TAG were evident in male ε4 carriers, respectively. It was speculated that the selective affinity of the E4 protein isoform for VLDL, in contrast with the E2 and E3 isoforms, which have a preference for the more lipid-poor large HDL protein, may explain the apparently greater TAG lowering in apoE4 subjects( Reference Caslake, Miles and Kofler 130 ). A recent study of the same research group systematically examined the individual impact of EPA- v. DHA-rich oils fed separately on plasma lipids in ε3/ε3 v. ε3/ε4 normolipidaemic males( Reference Olano-Martin, Anil and Caslake 131 ). In the ε3/ε4 group, within-treatment group analysis showed that DHA treatment, but not EPA, resulted in a significant increase in LDL-cholesterol, with a non-significant decrease in the ε3/ε3 group. As this proatherogenic shift may negate the cardioprotective actions of DHA, it was suggested that EPA-rich oils may be a more suitable therapy for apoE4 subjects( Reference Olano-Martin, Anil and Caslake 131 ).
The gene–nutrient interaction between apoE polymorphism and ALA and their subsequent effect on lipid metabolism and further CVD biomarkers has not been extensively studied until now. There is only one, uncontrolled study (no control group) in dyslipidaemic patients indicating that ALA may have beneficial effects on biomarkers of inflammation in carriers of the apoE ε3/ε3 and apoE ε3/ε4 genotypes, but not in carriers of the apoE ε2 allele( Reference Paschos, Yiannakouris and Rallidis 93 ). Owing to the limited number of ε2 (7%) and ε4 allele carriers (10%) these results need confirmation in larger, well-controlled and well-powered studies in prospectively genotyped participants.
Conclusions
In the present review, we have considered current data on apoE polymorphism. We have summarized metabolic impacts of the apoE isoforms and their responsiveness to dietary factors possibly underlying the geographic distribution and varying disease risk of the major apoE isoforms. The emergence and successful distribution of apoE ε3 have been put down to decreased susceptibility to AD and CVD in later life compared with apoE ε4; however, it could also be a result of varying responsiveness to dietary factors already present in younger life. The beneficial effects of quercetin and n-3 fatty acids were observed in individuals carrying the ε3 allele, but not in ε4 carriers indicating apoE3 a more flexible and responsive phenotype than apoE4. On the other hand, due to better clearance of dietary fat and reduced LDL uptake, the apoE ε4 genotype is associated with better vitamin D status and may provide protection against several infectious diseases. This could help to understand why the frequency of the ε4 allele follows a distinct pattern of geographic distribution and is enriched in particular regions (e.g. Northern Europe with insufficient UV-exposure in autumn and winter). Taken together, the apoE genotype appears to be an important determinant of individual responsiveness to dietary factors; however, large prospectively genotyped cohorts are required to confirm present data and to assess the clinical relevance of apoE isoform-dependent effects.
Acknowledgements
This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors. S.E., G.R. and P.H. declare no conflicts of interest. S.E., G.R. and P.H. conducted the literature research and wrote the manuscript. All authors read and approved the final manuscript.









