Introduction
Dipylidium caninum is a widespread cestode of carnivores occurring in both domestic and sylvatic settings. Its definitive hosts are canines and felines, while arthropods such as fleas and lice serve as intermediate hosts (Beugnet et al. Reference Beugnet, Labuschagne, Fourie, Jacques, Farkas, Cozma, Halos, Hellmann, Knaus and Rehbein2014). Definitive hosts become infected when they ingest fleas or lice containing cysticercoids, after which the parasites develop into the adult stage and establish in the small intestine. Eggs or gravid proglottids of the adult worms are shed in the feces and disseminated into the environment. Arthropod intermediate hosts acquire the infection by ingesting D. caninum eggs from the contaminated environment, where the larvae hatch and transform into cysticercoids within the body cavity. Humans may occasionally become infected by accidentally swallowing an infected arthropod (Rousseau et al. Reference Rousseau, Castro, Novo and Maia2022). In household environments, domestic dogs and cats are the principal hosts of this cestode, whereas various wild carnivores maintain the sylvatic cycle (East et al. Reference East, Kurze, Wilhelm, Benhaiem and Hofer2013; Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018).
Dipylidiasis is a cosmopolitan and common tapeworm infection of dogs and cats. Since the parasite is widespread in companion animals, human cases have been reported worldwide, although they remain relatively rare and usually cause only mild clinical problems (Rousseau et al. Reference Rousseau, Castro, Novo and Maia2022). Most infections occur in children living in resource-poor settings, where hygiene standards and veterinary care are inadequate, and close contact between humans and dogs or cats is frequent (Sahin et al. Reference Sahin, Köz, Atambay, Kayabas, Piskin and Unal2015; Xaplanteri et al. Reference Xaplanteri, Gkentzi, Stamouli, Kolonitsiou, Anastassiou, Marangos and Spiliopoulou2017; Bronstein et al. Reference Bronstein, Fedyanina, Maximova, Lukashev and Sergeev2020; Meena et al. Reference Meena, Singh, Pradeep Kumar, Gupta and Gupta2020; Benitez-Bolivar et al. Reference Benitez-Bolivar, Rondón, Ortiz, Díaz-Díaz, León, Riveros, Molina and González2022; Rousseau et al. Reference Rousseau, Castro, Novo and Maia2022). In both animals and humans, D. caninum infection often remains unnoticed because it is typically asymptomatic or produces non-specific clinical signs that may be mistaken for other helminth infections. Diagnosis relies on the morphological identification of eggs or proglottids, which requires experienced personnel, or on molecular tools that are too sophisticated or expensive to apply in many endemic areas. Consequently, information on its distribution and prevalence is scarce in most countries.
The taxonomy of D. caninum has evolved gradually, from its first description by Linnaeus in 1758 as Taenia canina, to its transfer to the newly erected genus Dipylidium by Leuckart in 1863 based on morphological features, and to its current refinement through genetic characterization. Like many parasites, D. caninum is experiencing a subspecific differentiation with the advent of molecular genetic data, revealing considerable genetic diversity that is not reflected by morphology (Beugnet et al. Reference Beugnet, Labuschagne, Fourie, Jacques, Farkas, Cozma, Halos, Hellmann, Knaus and Rehbein2014, Reference Beugnet, Labuschagne, Vos, Crafford and Fourie2018; Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018; Jesudoss Chelladurai et al. Reference Jesudoss Chelladurai, Abraham, Quintana, Ritchie and Smith2023). Two lineages, namely, the feline and canine genotypes, have been demonstrated to be in circulation. Genetic analysis showed these 2 genotypes to differ not only in their genetic makeup but also in their host preferences for cats or dogs, respectively, thereby supporting the proposal for species delimitation (Beugnet et al. Reference Beugnet, Labuschagne, Vos, Crafford and Fourie2018; Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018). However, the current volume of global molecular data is low and biased towards Europe and North America, while data from other important endemic regions, such as sub-Saharan Africa, the Middle East and Asia, remain limited or lacking. If the presumed ‘Dipylidium species complex’ is to be appraised, molecular genetic data will be required from all endemic regions. In the present study, we contribute to filling one of these gaps by reporting the molecular genetics of D. caninum collected from dogs in northern Ghana and demonstrating how the Ghanaian isolates fit into the global D. caninum phylogeny based on partial fragments of the mitochondrial cox1 and nad1 genes.
Materials and methods
Study area
The study was conducted in the Bolgatanga Municipality of the Upper East (UE) Region of Ghana. Bolgatanga town is the centre of economic activities of the UE Region, with a market that features various merchandise, agricultural produce and live animals, including livestock, poultry and dogs (Laar et al. Reference Laar, Abdulai, Agyen, Bortei, Abugri, Aovare, Jerela, Larbi, Adams and Abugri2020). The city is predominantly inhabited by the Frafra ethnic group, among whom dog meat is commonly consumed (Tasiame et al. Reference Tasiame, El-Duah, Johnson, Owiredu, Bleicker, Veith, Schneider, Emikpe, Folitse, Burimuah, Akyereko, Drosten and Corman2022).
Screening of animals and recovery of helminths
Dog slaughter points and meat-processing enterprises were visited between December 2022 and January 2023 to identify facilities consenting to participate in the study. The research objective – isolation and characterization of intestinal worms from dogs – was explained to the owners of all enterprises visited. Of 5 dog slaughter and meat-processing enterprises approached, 2 were selected based on the owners’ willingness to provide intestines from slaughtered dogs free of charge for parasitological screening. Material was strictly obtained only from dogs slaughtered traditionally – no animals were killed for the purpose of this study. Fresh intestines were obtained immediately after slaughter on the days of visit and were opened. Helminths visible to the naked eye were isolated, washed in normal saline and preserved in 75% ethanol. The remaining intestinal contents were then scraped into sampling bottles for subsequent laboratory examination. In the laboratory, the collected material was sieved, and the filtrates were examined microscopically for smaller helminths (data not reported here).
Helminth identification and characterization
Almost all isolates were identified as cestodes, and species determination was performed by analysing the mitochondrial cytochrome c oxidase subunit 1 (cox1) and NADH dehydrogenase subunit 1 (nad1) genes. DNA was extracted from worm tissues using the alkaline lysing method (Addy et al. Reference Addy, Alakonya, Wamae, Magambo, Mbae, Mulinge, Zeyhle, Wassermann, Kern and Romig2012). Briefly, a piece of the posterior proglottids of individual worms with intact scolex was excised and finely chopped, transferred into 30 µL 0.02 M NaOH and lysed at 99 °C for 30 min. The lysate was centrifuged for 5 min at 3000 g, and the supernatant was used as a DNA template for polymerase chain reaction (PCR). The following oligonucleotide primer pairs were used to amplify partial fragments of the cox1 gene (∼420 bp): 5′-TTT TTT GGG CAT CCT GA GGT TTA T-3′ and 5′-TAA AGA AAG AAC ATA ATG AAA ATG-3′ (Bowles et al. Reference Bowles, Blair and McManus1992), and for the nad1 gene (400 bp): 5′-AAA ACG TTG GGT TTG CGT CTC-3′ and 5′-CAC ATA AAA CAA GCC TCA-3′. The PCR was carried out in a 25 µL reaction mixture consisting of 12.5 μL Red Taq 2× Master Mix (VWR International Ltd, UK), 0.4 μM of each primer and 2 µL crude worm lysate as template DNA. The amplifications were performed under the following thermal conditions: initial denaturation at 95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s and elongation at 72 °C for 40 s, and final elongation at 72 °C for 5 min. Amplicons were visualized on 1.5 % agarose gel stained with ethidium bromide.
DNA sequencing and analysis
The PCR amplicons were sequenced using the reverse primers at inqaba biotec Ltd (Pretoria, South Africa). The raw sequences were viewed, edited and aligned using GENtle software (Manske M., 2003, University of Cologne, Germany). Species identification was performed by comparing the obtained sequences with reference data deposited in the NCBI GenBank using the Basic Alignment Search Tool.
Newly generated sequences, together with previously published ones, were used to construct haplotype networks based on the TCS model (Clement et al. Reference Clement, Snell, Walker, Posada and Crandall2002) in the PopART software (http://popart.otago.ac.nz). Bayesian analysis of cox1 sequences was performed with the BEAST X package (v. 10.5.0; BEAST Development Team, Geneva, Switzerland, 2025) and 10 000 000 iterations (sampling every 1000th), using the GTR + I + G substitution model and otherwise default parameter settings in BEAUTi (Suchard et al. Reference Suchard, Lemey, Baele, Ayres, Drummond and Rambaut2018). Trees were calculated with burn-in set to 10% of states in TreeAnnotator and visualized with FigTree (v. 1.4.4; Rambaut A., 2018, University of Edinburgh, Scotland). All DNA sequences generated in the present study were deposited in the GenBank database under the accession numbers: PX765912–PX765915 (Cox1 Hp01–4) and PX766076–PX766082 (nad1 Hp01–Hp07).
Results
Identification of cestodes
In total, 16 dog intestines were examined during the study period. Cestode infections were detected in 11 intestines (68.75%). At the species level, D. caninum was identified in 5 dogs (31.25%), Taenia hydatigena in 10 dogs (52.50%) and T. multiceps in 2 dogs (12.50%). Mixed cestode infections were observed in 6 dogs (37.50%), including co-infections with T. hydatigena and D. caninum in 4 dogs, T. hydatigena and T. multiceps in 1 dog, and all 3 cestode species in 1 dog. Roundworms (Toxocara canis) were seen in the intestines of 2 dogs, one of which was concurrently infected with T. multiceps.
Molecular variations within D. caninum
A total of 44 individual D. caninum isolates, taken from all 5 infected dogs, were subjected to molecular analysis. Of these, 4 specimens were assigned to the feline genotype and 40 to the canine genotype. Partial fragments of both the cox1 (373 bp) and nad1 (376 bp) genes were successfully sequenced in 24 isolates. In the remaining 20 isolates, sequence data could be obtained for only one of the 2 genes, with cox1 sequences generated from 16 and nad1 sequences from 4 isolates. The DNA sequences obtained in the present study showed between 98% and 100% similarity to previously deposited D. caninum sequences in the NCBI GenBank database.
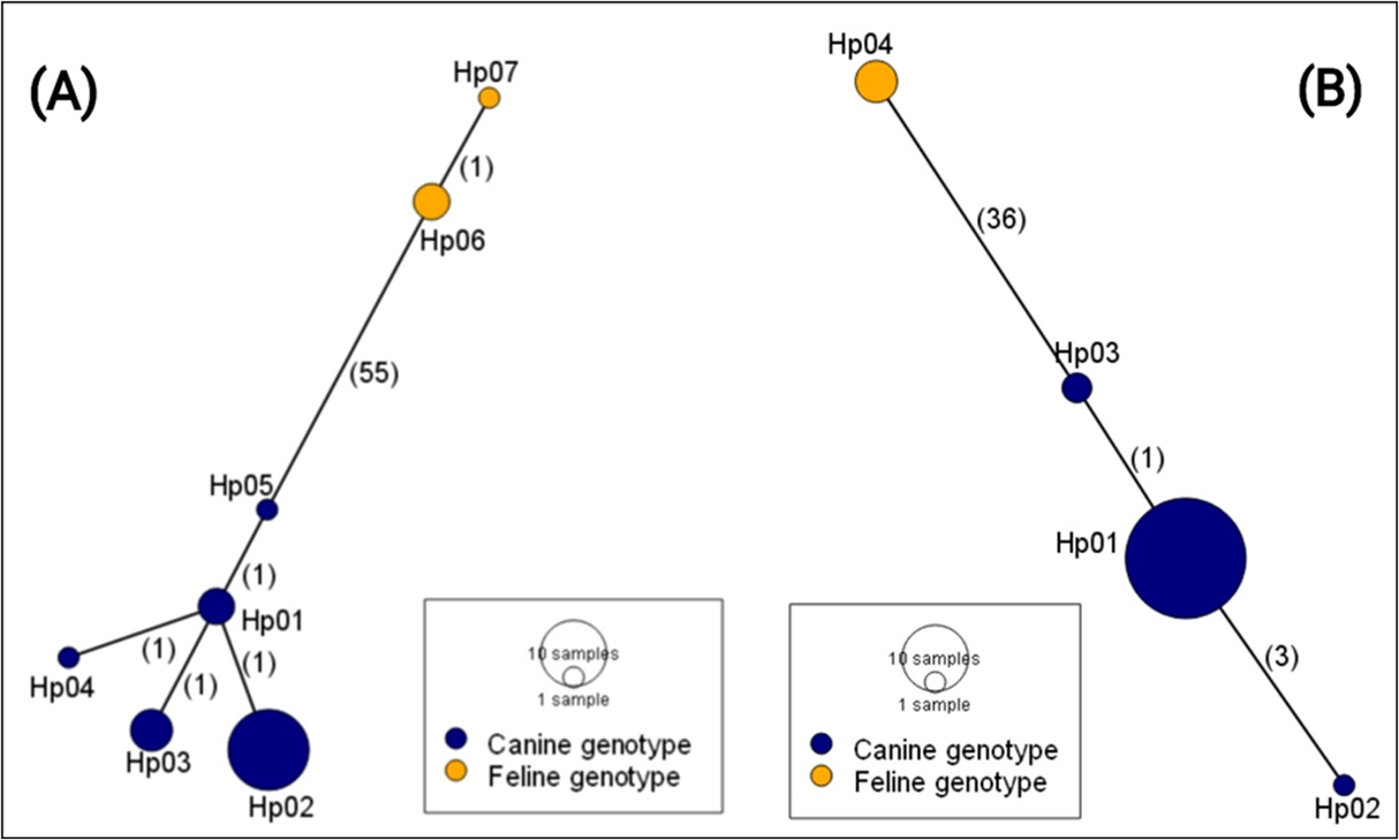
The haplotype analysis shows a linear network comprising 7 and 4 haplotypes based on the nad1 and cox1 sequences, respectively (Fig. 1). These microvariants occurred at different frequencies within the dataset, with haplotype Hp01 being the most dominant in the cox1 sequence set, and Hp02 in the nad1 set. The haplotype networks further revealed 2 major groups, each clearly separated by large mutational steps. In the cox1 dataset, 36 mutational steps separate Hp01–03 from Hp04, and 55 mutational steps were observed between Hp01–05 and Hp06–07 in the nad1 dataset. These steps clearly divide the canine and feline genotypes. Infected dogs usually harboured single haplotypes of the parasite, except in 2 cases where 5 (nad1) and 4 (cox1) haplotypes were identified in one dog in which both the canine and feline genotypes existed. The second dog had 2 haplotypes of nad1 (canine genotype).
Haplotype network of D. caninum isolates from Ghana based on the partial nad1 (A) (376 bp) and cox1 (B) (373 bp). Numbers in parentheses indicate the mutational steps between haplotypes.

Global haplotype network and phylogenetic tree of D. caninum
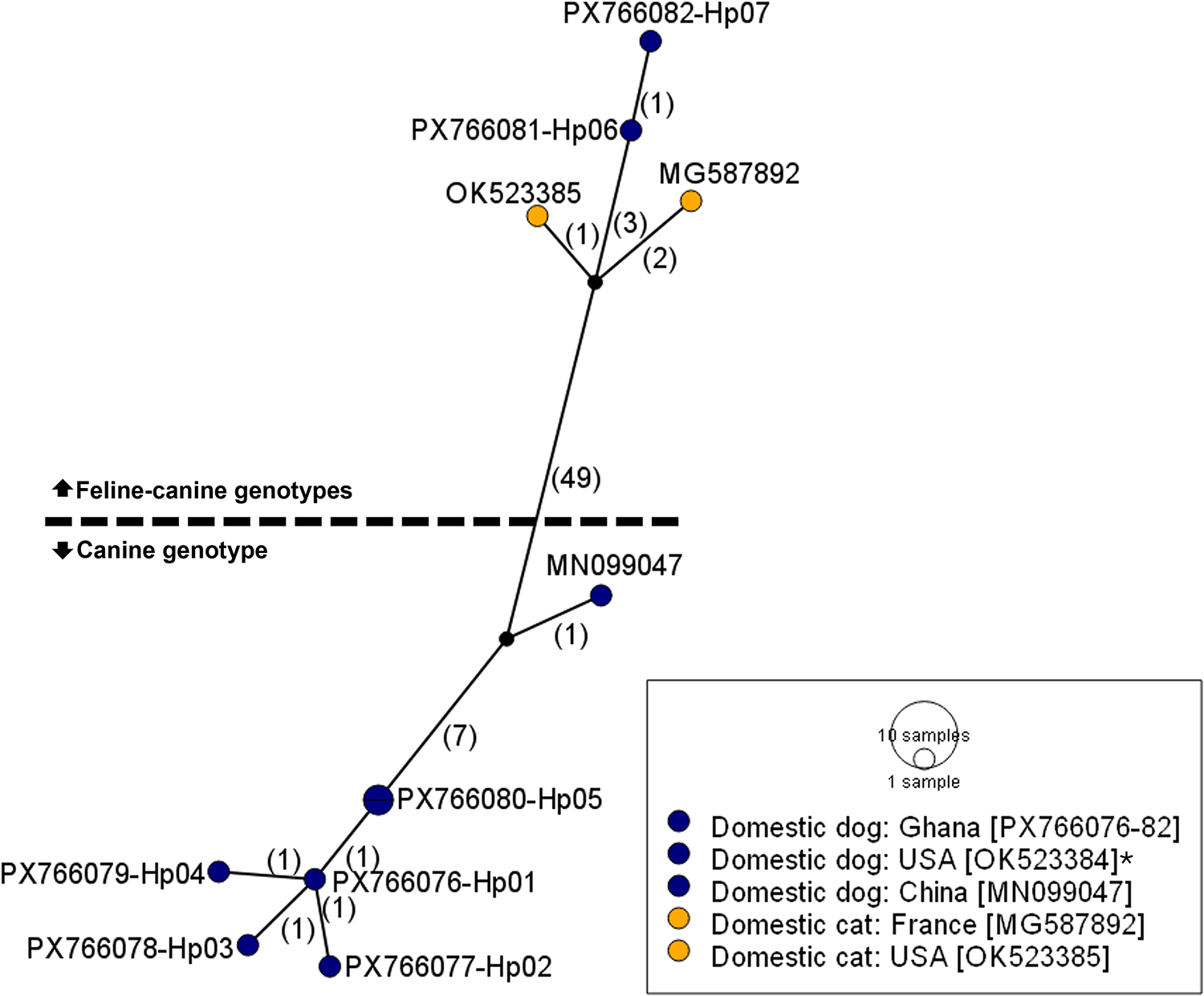
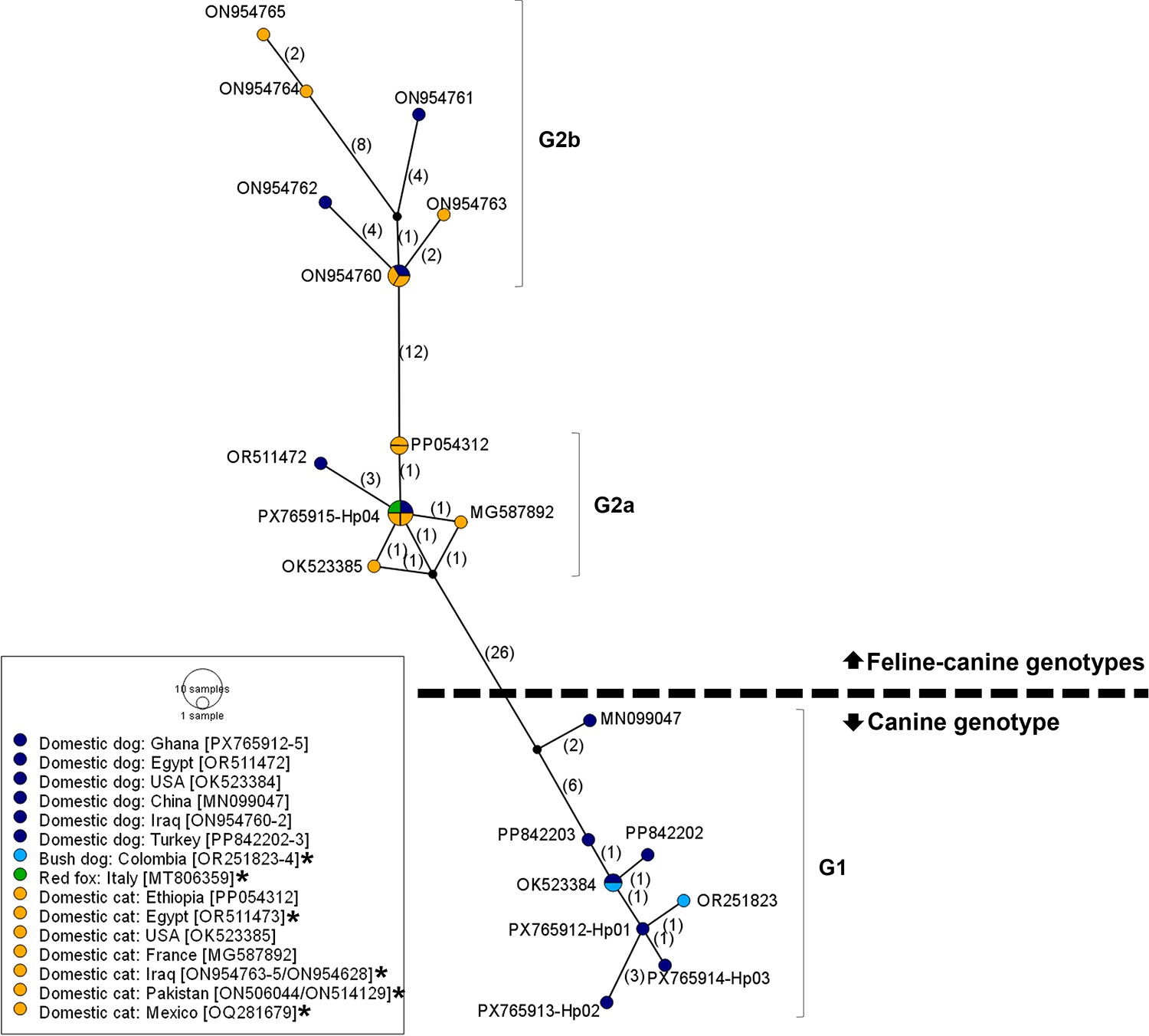
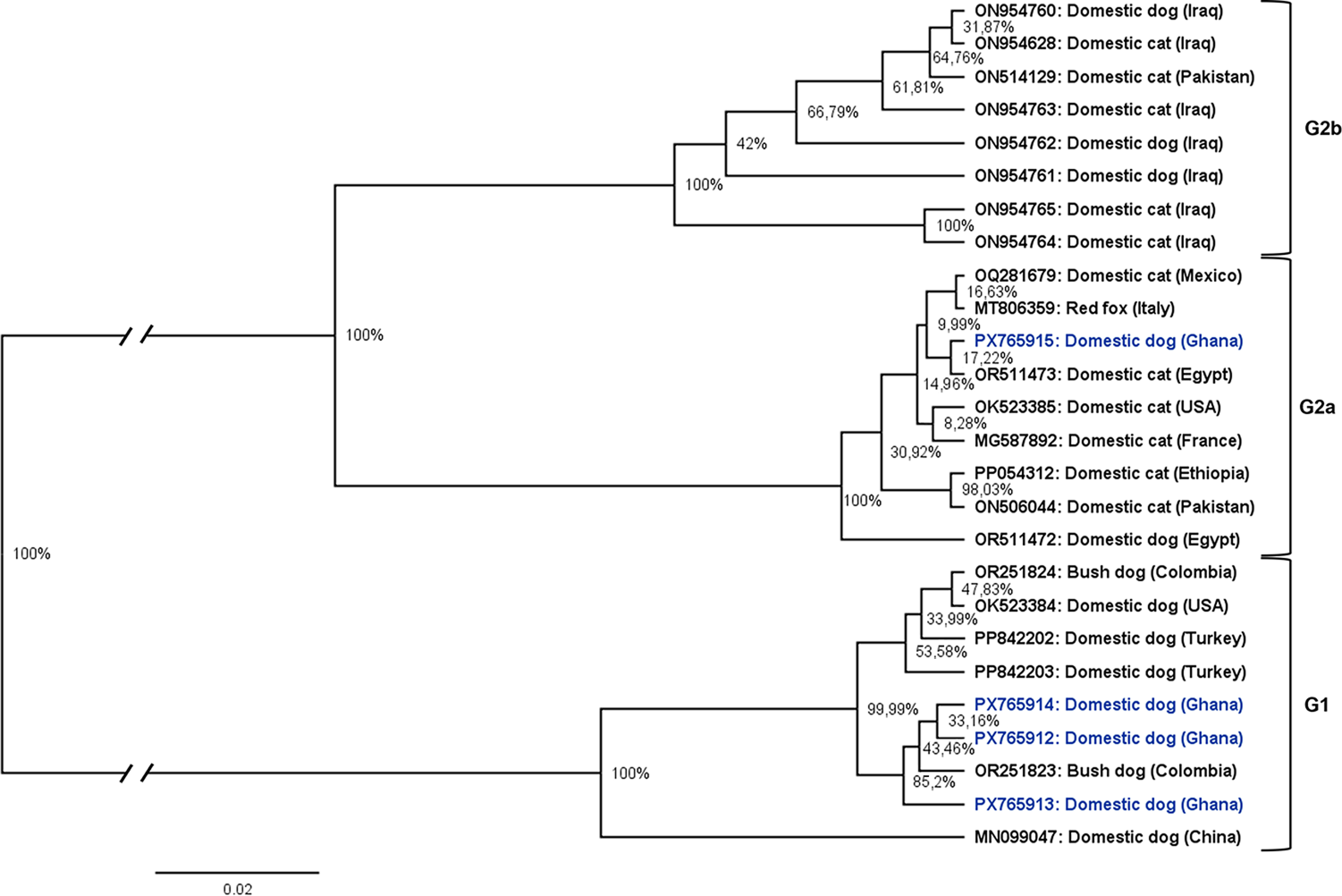
In a global comparison, the Ghanaian isolates clustered closely with previously deposited sequences from other endemic regions (Figs. 2 and 3). The 2 haplogroups identified among the Ghanaian isolates corresponded to the D. caninum canine and feline genotypes, respectively. Notably, within the cox1 sequences, the feline cluster appeared to be further divided into 2 subgroups (Fig. 3). Due to the limited nad1 reference sequences available for global comparison, only the cox1 data were used in the phylogenetic tree analysis. The genetic topology observed on the cox1 haplotype network was also reflected in the phylogenetic tree, which showed strong node support (posterior probability) (Fig. 4). We designated the observed clusters of global genetic variants into 3 genotypes: G1 representing the canine genotype, and G2a and G2b denoting the 2 haplogroups seen in the feline–canine admixture genotypes. Genotype G1 comprises haplotypes from Ghana (Hp01–03), Colombia, the USA, Turkey and China, all of which were isolated from canids. The genotype G2a consists of haplotypes from Africa (Ghana, Ethiopia, Egypt), America (Mexico, USA), Europe (Italy, France) and the Middle East (Pakistan), originating from domestic cats and dogs except the isolate from Italy, where the haplotype was identified in a red fox. The second feline–canine G2b includes haplotypes identified in domestic cats and dogs from the Middle East (Iraq and Pakistan) only.
Haplotype network of global D. caninum isolates based on the partial nad1 (336 bp). Numbers in parentheses indicate the mutational steps between haplotypes. *The sequence OK523384 is identical to PX766080-Hp05. Colour of haplotypes indicates host type: yellow, domestic cat and dark blue, domestic dog.

Haplotype network of global D. caninum isolates based on the partial cox1 (345 bp). Numbers in parentheses indicate the mutational steps between haplotypes. Colour of haplotypes indicates host type. *Sequences identical to those shown in the figure: OR251824 = OK523384; OR511473/OQ281679/MT806359 = PX765915-Hp04; ON506044 = PP054312; and ON954628/ON514129 = ON954760.

Phylogenetic tree constructed by Bayesian inference showing the phylogenetic position of Ghanaian D. caninum isolates within the global dataset, based on partial mitochondrial cox1 gene fragment (345 bp). G1 – canine genotype, G2a and G2b – feline–canine genotypes. Posterior probabilities are displayed on nodes as percentages. Basal branches were shortened by 0.06.

Discussion
The domestic dog is raised in homes and on farms for companionship, security and hunting purposes, but in certain regions and ethnicities, also for consumption. Globally, dog meat consumption is unpopular, but the practice is widespread, eaten as a cultural delicacy or for religious purposes. In sub-Saharan Africa, dog meat consumption is known among ethnic tribes in some countries such as Nigeria, Ghana and Namibia (Ekanem et al. Reference Ekanem, Eyong, Philip-Ephraim, Eyong, Adams and Asindi2013; Laar et al. Reference Laar, Abdulai, Agyen, Bortei, Abugri, Aovare, Jerela, Larbi, Adams and Abugri2020; Tasiame et al. Reference Tasiame, El-Duah, Johnson, Owiredu, Bleicker, Veith, Schneider, Emikpe, Folitse, Burimuah, Akyereko, Drosten and Corman2022; Tenzin et al. Reference Tenzin, Hikufe, Hedimbi, Athingo, Shikongo, Shuro, Iipinge, Herman, Naunyango, Haufiku, Peter, Hango, Gottlieb, Shoombe, Denzin, Busch, Lohr, Letshwenyo, Torres, Freuling, Müller and Shilongo2024). In northern Ghana, consumption of dog meat is common among the Frafra and Dagaaba tribal groups, where dogs are slaughtered at the household and communal levels during festivities or at local enterprises for sale, howbeit, without veterinary supervision (Laar et al. Reference Laar, Abdulai, Agyen, Bortei, Abugri, Aovare, Jerela, Larbi, Adams and Abugri2020; Tasiame et al. Reference Tasiame, El-Duah, Johnson, Owiredu, Bleicker, Veith, Schneider, Emikpe, Folitse, Burimuah, Akyereko, Drosten and Corman2022). Slaughter and consumption of dogs are identified as risk factors for parasitic, bacterial and viral zoonosis transmission to humans (Tasiame et al. Reference Tasiame, El-Duah, Johnson, Owiredu, Bleicker, Veith, Schneider, Emikpe, Folitse, Burimuah, Akyereko, Drosten and Corman2022). The present study reports the intestinal helminth infections, with a particular focus on D. caninum, in commercially slaughtered dogs from Bolgatanga, UE Region of Ghana.
The cestode D. caninum is a common helminth parasite of dogs and cats worldwide and can occasionally infect humans. Growing molecular evidence is revealing previously unrecognized cryptic variants, suggesting that D. caninum may represent a species complex. Based on nuclear markers and mitochondrial genome data from isolates in Europe, South Africa and the USA, 2 distinct genotypes, a canine and a feline genotype, were initially identified (Beugnet et al. Reference Beugnet, Labuschagne, Vos, Crafford and Fourie2018; Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018), leading to proposals for species delimitation (Jesudoss Chelladurai et al. Reference Jesudoss Chelladurai, Abraham, Quintana, Ritchie and Smith2023). Although these proposals are well supported, data from many endemic regions remain scarce, which currently limits comprehensive assessments of the parasite’s global genetic variability. The present study contributes to filling this gap by characterizing D. caninum isolates from dogs in Ghana (West Africa) and determining their phylogenetic position within the global dataset based on partial sequences of the mitochondrial nad1 and cox1 genes.
Essentially, the D. caninum isolates from Ghana included both the canine and feline–canine genotypes (Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018; Rousseau et al. Reference Rousseau, Castro, Novo and Maia2022) in their nad1 and cox1 sequences, with only minor variations. The majority (90%) of the isolates could be assigned to the canine genotype, which has previously been identified exclusively in canine hosts, suggesting a strong canine host preference, in contrast to the feline–canine associated genotypes which affect dogs and cats (Marshallf, Reference Marshallf1967; Beugnet et al. Reference Beugnet, Labuschagne, Fourie, Jacques, Farkas, Cozma, Halos, Hellmann, Knaus and Rehbein2014, Reference Beugnet, Labuschagne, Vos, Crafford and Fourie2018; Low et al. Reference Low, Prakash, Tan, Sofian-Azirun, Anwar, Vinnie-Siow and AbuBakar2017; Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018).
This study provides the first molecular data on D. caninum from Ghana and, more broadly, West Africa, although it represents only a snapshot of the situation. In the present study, worm samples were collected from dogs sourced from the animal market (Laar et al. Reference Laar, Abdulai, Agyen, Bortei, Abugri, Aovare, Jerela, Larbi, Adams and Abugri2020). These dogs were slaughtered under informal unregulated conditions, similar to household slaughter practice, handled with bare hands, and their waste was disposed of directly into the environment. Such practices may result in extensive environmental contamination with parasite eggs, frequent transmission to flea and louse intermediate hosts and consequently increase the risk of human infection (Marshallf, Reference Marshallf1967; Beugnet et al. Reference Beugnet, Labuschagne, Fourie, Jacques, Farkas, Cozma, Halos, Hellmann, Knaus and Rehbein2014, Reference Beugnet, Labuschagne, Vos, Crafford and Fourie2018; Labuschagne et al. Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018; Azarm et al. Reference Azarm, Dalimi, Pirestani and Mohammadiha2022; Rousseau et al. Reference Rousseau, Castro, Novo and Maia2022). Generally, the dynamics of zoonotic transmission in communities where companion animals are consumed, including those in the UE Region of northern Ghana (Laar et al. Reference Laar, Abdulai, Agyen, Bortei, Abugri, Aovare, Jerela, Larbi, Adams and Abugri2020), are poorly documented or entirely unknown. However, such data are required for informing local public health policies and guiding interventions aimed at reducing infection risks.
Beyond the local implications, these data also contribute to the global understanding of D. caninum diversity. The Ghanaian D. caninum sequences reported here clustered with isolates from other endemic regions for which sequence data were available, showing no distinct African clustering. Further, the data clearly support previous proposals to delimit the canine and feline genotypes as separate species. The 2 genotypes differed by approximately 16% in nad1 and about 10% in cox1, further underlining their deep genetic divergence, although only a partial fragment of the mitochondrial markers were used in the present study. Such levels of divergence are consistent with species-level differentiation in other cestode taxa. The current global cox1 dataset reveals a more complex picture than previously reported, with evidence of multiple feline-associated genotypes. In the analysis by Labuschagne et al. (Reference Labuschagne, Beugnet, Rehbein, Guillot, Fourie and Crafford2018) and Jesudoss Chelladurai et al. (Reference Jesudoss Chelladurai, Abraham, Quintana, Ritchie and Smith2023), no isolates from the Middle East were included. However, recently deposited sequences from Iraq and Pakistan now form a distinct clade, separate from the previously recognized feline–canine lineage. The robust genetic topology observed in the present analysis strongly suggests that the genetic diversity of D. caninum may be more complex than the 2-genotype model proposed so far. While we retain the host–origin-based classification, we designated the widely distributed canine genotype as G1 and the 2 feline–canine associated genotypes as G2a (detected globally) and G2b (identified in Iraq and Pakistan). We believe that this classification will provide clarity in future discussions of D. caninum genetic diversity as more data from other endemic regions become available. Obtaining complete mitochondrial genomes and important nuclear markers from a broader range of localities in the Middle East and Asia would provide valuable insights into the evolutionary structure and global diversity of this parasite.
Acknowledgements
The authors wish to express their profound gratitude to the dog meat sellers for accepting to participate and cooperating in the study.
Author contributions
FA coordinated sample collection, supervised laboratory analysis and drafted the manuscript; EA collected and analysed the samples, curated data and contributed to the writing of the manuscript; EKN and GAB collected samples and curated data; SRD, KCP, BW, MM, ED, FYL and AAWS analysed the samples; SD analysed the data and reviewed the manuscript; and TR and MW conceived and coordinated the study and reviewed manuscript drafts.
Financial support
This study was funded by the Deutsche Forschungsgemeinschaft – DFG, Germany (grant number: RO3753/9-1) within the project, Transmission Ecology and Risk Factors of Cystic Echinococcosis in sub-Saharan Africa.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
Not applicable.