Refine listing
Actions for selected content:
142352 results in Open Access
Regional variations in the demographic response to the arrival of rice farming in prehistoric Japan
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Creation of a Multi-Year Pediatric Candidemia Antibiogram in Georgia Identifies Changing Epidemiology and Resistance Trends
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s129-s130
-
- Article
-
- You have access
- Open access
- Export citation
Factors Associated with Inappropriate Urine Culture Orders in Hospitalized Patients with Indwelling Urinary Catheters
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s69
-
- Article
-
- You have access
- Open access
- Export citation
Perspectives and Awareness of Environmental Sustainability in the Infection Prevention and Control Community Nationally
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s25
-
- Article
-
- You have access
- Open access
- Export citation
Exposure to anti-refugee hate crimes and support for refugees in Germany
-
- Journal:
- Political Science Research and Methods / Volume 13 / Issue 3 / July 2025
- Published online by Cambridge University Press:
- 16 September 2024, pp. 755-764
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Clinical and Genomic Characteristics of Candida auris in Central Ohio: An Insight into Epidemiological Surveillance
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s93
-
- Article
-
- You have access
- Open access
- Export citation
Evaluation of a Sepsis Alert System at a Veterans Affairs Medical Center
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s61
-
- Article
-
- You have access
- Open access
- Export citation
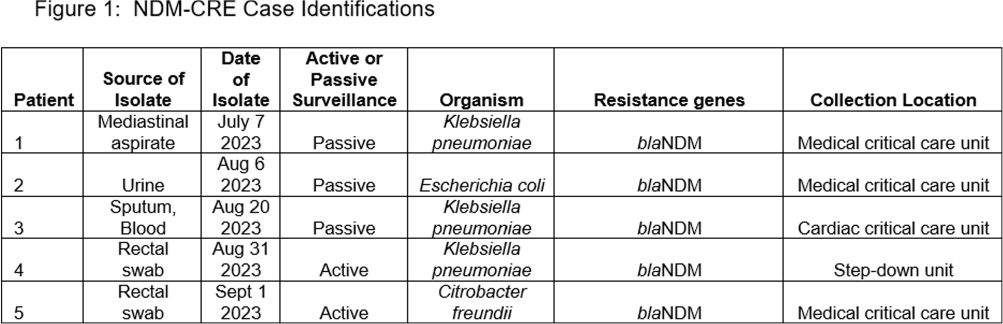
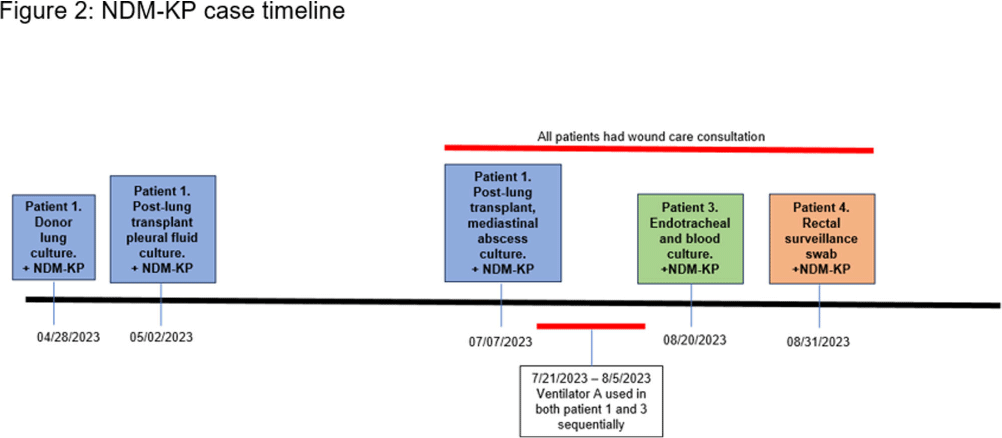
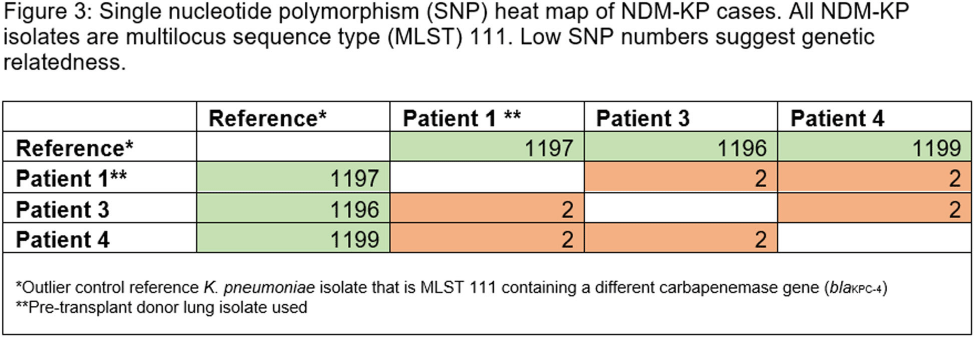
Investigation of a Donor-derived Carbapenamase-producing Carbapenem-resistant Enterobacterales Hospital Outbreak
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s126-s127
-
- Article
-
- You have access
- Open access
- Export citation
Morphological evaluation for the narrowest section of the patent ductus arteriosus in infants by CT: a crucial point for device closure
-
- Journal:
- Cardiology in the Young / Volume 34 / Issue 9 / September 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. 1983-1989
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Rapid Genomic Characterization of High-Risk, Antibiotic Resistant Pathogens Using Long-Read Sequencing to Identify Nosocomial
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s106
-
- Article
-
- You have access
- Open access
- Export citation
The distribution of non, nenny and non fait in Pre-Classical and Classical French
-
- Journal:
- Journal of French Language Studies / Volume 34 / Issue 3 / November 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. 431-456
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Multitask Neural Networks to Predict Antimicrobial Susceptibility Results of Escherichia coli Clinical Isolates
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s26-s27
-
- Article
-
- You have access
- Open access
- Export citation
Case validation of bloodstream infections with an antibiotic-resistant organism
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s156
-
- Article
-
- You have access
- Open access
- Export citation
Bad Habits that Stick: An Investigation into Adhesive Medical Tape Use Practices and Beliefs
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s29
-
- Article
-
- You have access
- Open access
- Export citation
Patient First Strategies for Reducing Inequities in HAI Prevention
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s76-s77
-
- Article
-
- You have access
- Open access
- Export citation
Inpatient Hospice Impact on Blood Culture Practices Near the Time of Death, Tertiary Center, Northern California, 2019–2023
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s85-s86
-
- Article
-
- You have access
- Open access
- Export citation
Estimating public opinion from surveys: the impact of including a “don't know” response option in policy preference questions
-
- Journal:
- Political Science Research and Methods / Volume 13 / Issue 3 / July 2025
- Published online by Cambridge University Press:
- 16 September 2024, pp. 663-679
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Methylome-wide association study of multidimensional resilience
-
- Journal:
- Development and Psychopathology / Volume 37 / Issue 4 / October 2025
- Published online by Cambridge University Press:
- 16 September 2024, pp. 1730-1741
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Impact of Discontinuing Contact Precautions for Multidrug-resistant Gram-negative Enterobacteriaceae in a Large Health System
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s139-s140
-
- Article
-
- You have access
- Open access
- Export citation
Analyzing the Relationship Between Socioeconomic Deprivation and Outpatient Medicare Part D Fluroquinolone Claims in Texas
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s75-s76
-
- Article
-
- You have access
- Open access
- Export citation